2.1. Graph Definition
An unattributed graph is a set of unlabeled vertices and links. is the set of vertices. is the adjacency matrix. If there is a link between and , then . Otherwise, .
A graph is an (induced) subgraph of graph , if and only if and . For an unattributed graph, a subgraph is also called as a pattern of a graph.
A (sub)graph is isomorphic to , if there exists as least one bijective function so that . We denote it as .
A graph alignment
is defined by a set of several subgraphs
and a special order of the nodes
in each subgraph [
15]; this joint order is denoted by
. Therefore
means that the two graphs
and
are aligned.
2.2. Kernel Method
Kernel method is an important concept in machine learning. Through a mapping
from the input space
into the reproducing kernel Hilbert space
(RKHS), the kernel function is defined as:
where
and
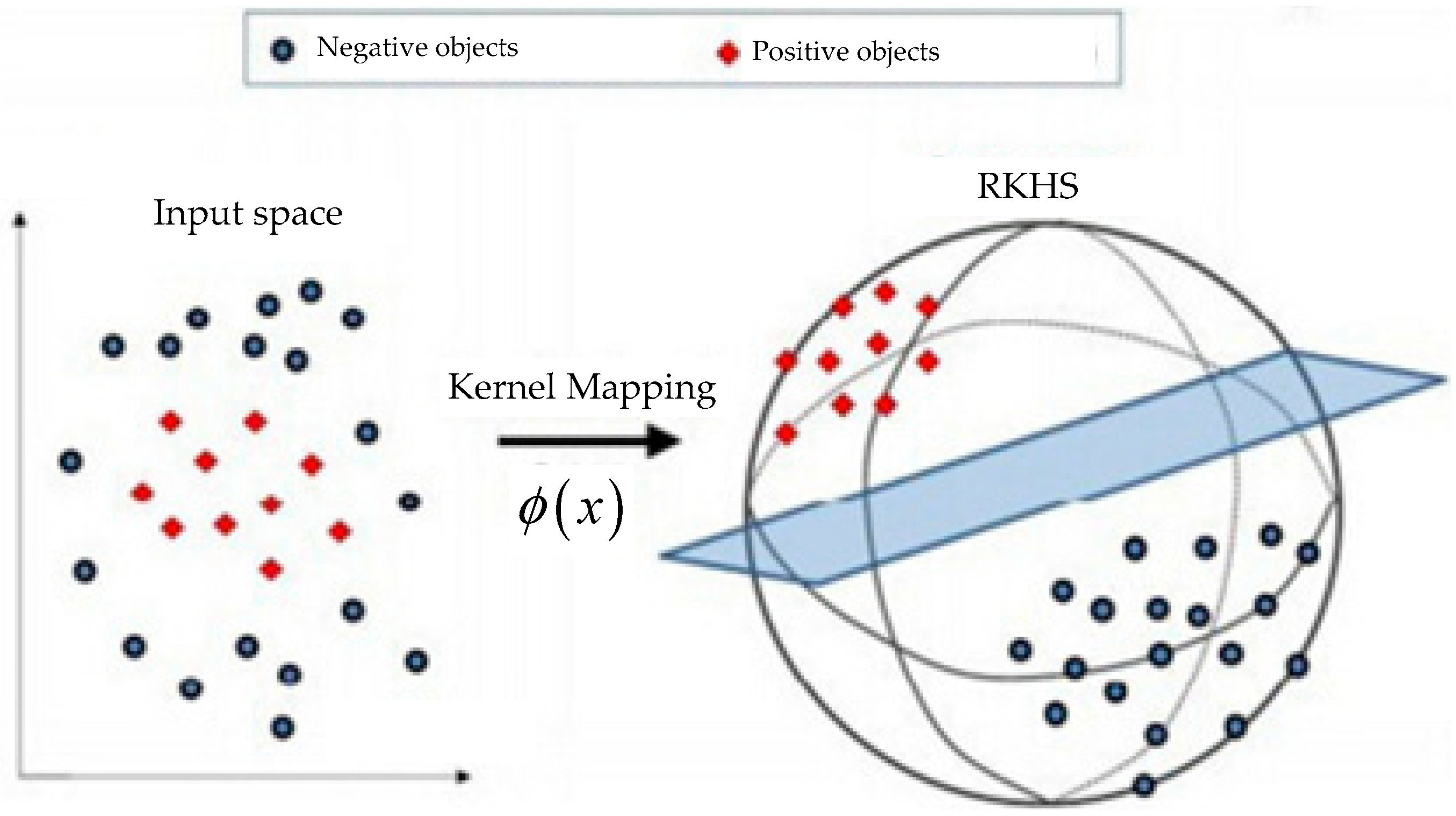
is the inner product operation in RKHS. An illustration of kernel method is shown in
Figure 1. The most obvious advantage of graph kernel is that a problem which cannot be linearly separated is changed into a linear separable problem through a kernel mapping.
Specifically, we call the method as graph kernel if the input space is the graph space. From Equation (1), it is clear that via a feature extraction method , graphs can be mapped into vectors in a RKHS, and the inner product of two vectors can represent the graph similarity.
Usually, the graph similarity can be computed directly, without the explicitly definition of the feature extraction method
. In the real world, it is quite difficult to compute an explicit embedding representation of structured data. Therefore, compared to topology descriptors and other graph comparison methods, graph kernel could achieve a smart and accurate similarity measurement because of the implicit feature extraction. In the field of machine learning, it has been witnessed that graph kernels can bridge the gap between graph-based data and a large group of kernel-based machine learning algorithms including support vector machines (SVM), kernel regression, and kernel principle component analysis (kPCA) [
9].
According to the Mercer’s theorem [
16], a valid graph kernel must be symmetric and positive semi-definite (p.s.d.):
Symmetric. Obviously, for two graphs and , .
p.s.d. For a dataset with graphs, any finite sequences of graphs and any choices of arbitrary real numbers , we have , i.e., all the eigenvalues of the kernel matrix for the dataset are nonnegative.
Note that although the learning ability of valid graph kernels has been proven, there is no theoretical research which demonstrates that invalid graph kernel could not support the learning algorithms. Actually, there exist discussions of how similarities (i.e., non-p.s.d. kernels) can support learning [
17,
18]. Therefore, some graph kernels are not proved to be p.s.d. in the literatures.
2.3. Kernel Groups
In the literature, the existing graph kernels are usually designed based on the distinct topological analysis of graph structure. According to five different dimensions of kernel design details, the whole family of graph kernels can be categorized as follows:
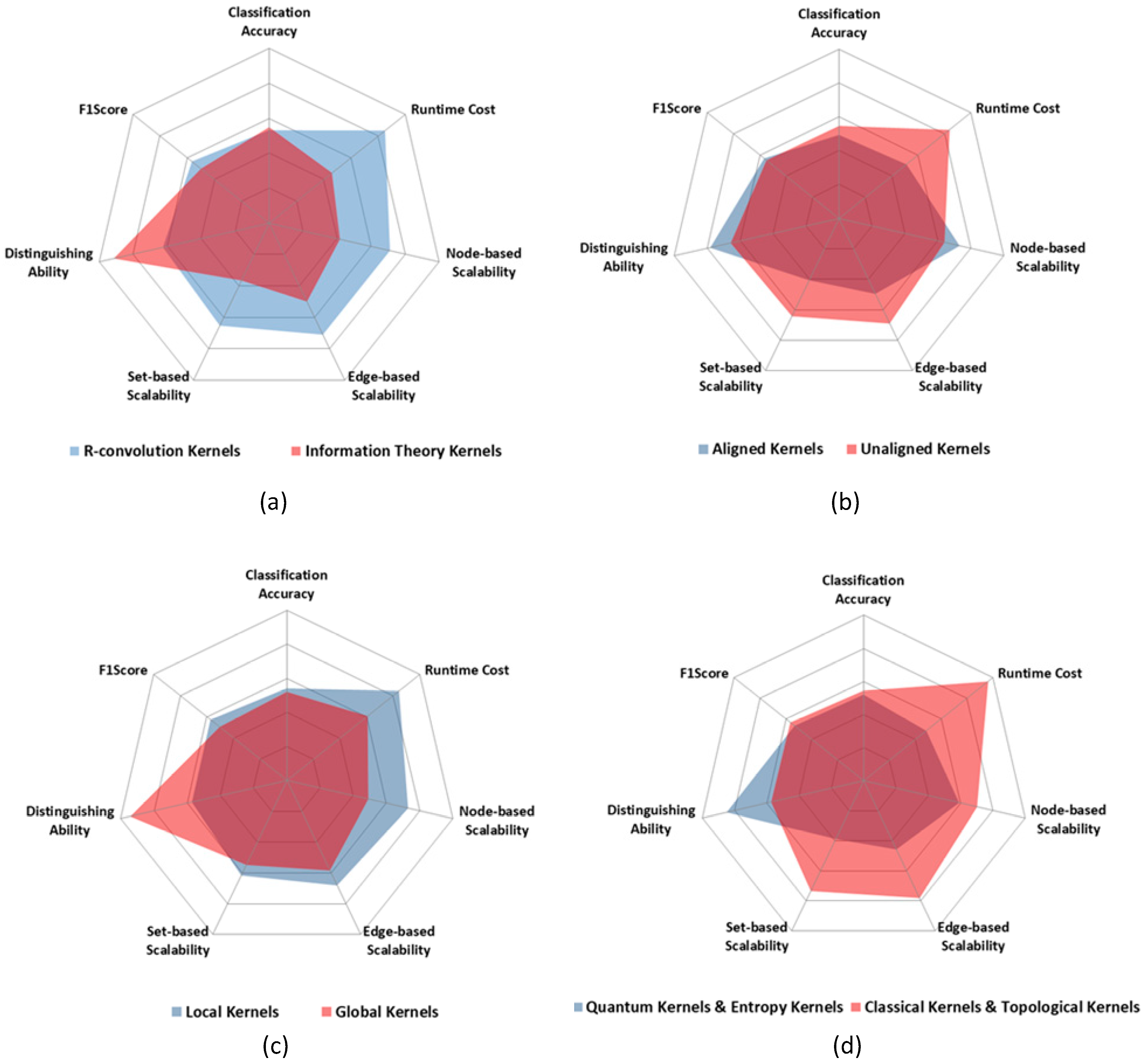
1. Framework: R-convolution kernels vs. Information theoretic kernels
In 1999, Haussler proposed R-convolution kernels [
19], which discover the relationship between graph similarity and the appearances of same or similar substructures of two graphs. As the formal definition in [
20], for two graphs
and
,
and
are sub-graph sets of
and
respectively. A standard R-convolution kernel
for these two graphs is defined as:
where
denotes a Dirac kernel shown as:
where
indicates the substructure
is isomorphic (or approximately isomorphic) to
.
So far, most existing kernels [
9,
10,
21,
22,
23] belong to this group. Intuitively, two similar graphs should have many common substructures. However, the main drawback of the R-convolution kernels is that they neglect the relative locations of substructures. This is because R-convolution kernels cannot establish reliable structural correspondences among substructures [
21]. Meanwhile, the teeter-totter problem and the complexity issue of the graph decomposition challenge the development of R-convolution kernels [
6].
Recently, Bai et al. utilized information theory methods to compute the probability distribution diffusion of two graphs as the similarity measurement [
20,
24,
25,
26,
27,
28]. By mapping all the data points of the input space into a fitted distribution in a parametric family S, a kernel for the distributions can be defined. This group of kernels for the data points in terms of distributions can be automatically induced in the original input space. Therefore, this framework provides us an alternative way of defining kernels that maps graphs into a statistical manifold. In real-world applications, these kernels outperform the linear kernels using SVM classifiers. Some of them create a bridge between kernel methods and information theory, and thus have an information theoretic interpretation. These methods are called information theoretic kernels. However, its high computational complexity of the information entropy is the bottleneck of this group.
2. Graph Pre-processing: Aligned Kernels vs. Unaligned Kernels
Graph alignment can locate the mapping relationship between the node sequences of two graphs. It is a pre-processing procedure for the original graphs. Common substructures will be pinned in the same position in these two graphs after the alignment. Through assigning parts of one object to parts of the other, the most similar parts of the two objects can be found out. Finding such a bijection is known as the assignment problem and well-studied in combinatorial optimization. This approach has been successfully applied to graph comparison, e.g., in general graph matching as well as kernel-based classification. In contrast to convolution kernels, assignments establish structural correspondences, thereby alleviating the problem of diagonal dominance at the same time. And then it can achieve an accurate similarity computation without false positive. The research on optimal assignment kernels was reported in [
29], in which each pair of structures is aligned before comparison. However, the similarities derived in this way are not necessarily positive semi-definite (p.s.d.) and thus do not give rise to valid kernels, severely limiting their use in kernel methods [
30]. Kriege et al. discussed the condition for guaranteeing an optimal assignment kernel to be positive semi-definite [
31].
The performance of the aligned kernel depends on the alignment accuracy. During graph alignment, every node/subgraph in a graph could only map to one node/subgraph in another graph, which will lead to information loss. Therefore, unaligned kernels are more common in the literature.
3. Matching Pattern: Global kernels vs. Local kernels
A novel choice of matching pattern is usually the core of designing a new graph kernel. The similarity computation of a kernel mainly depends on the exploration of the matching patterns.
Most existing graph kernels belong to the group of local kernels, which focus on local patterns of data structure, such as walks [
22], paths [
23], sub-trees [
21] and limit-sized sub-graphs [
10]. For example, the random walk kernel will embed graph into a feature space, in which the feature vector consists of walk numbers with different lengths.
A few studies changed to explore the global characteristics of graphs such as the Lovász number [
32] and the whole probability distribution [
11]. The kernels based on these methods are called global kernels. In this kernel group, all the members directly obtain the similarity among graphs based on the whole structure information. Therefore, the graph decomposition is not necessary.
In the most recent research, some hybrid patterns were utilized to design a kernel [
33]. Because graph matching is a special case of sub-graph matching, the kernels based on graph matching are allocated in the group of local pattern in this paper.
4. Computing Models: Quantum kernels vs. Classical kernels
Quantum computation differs from its classical counterparts. It can store, process and transport information using the peculiar properties of quantum mechanics such as the superposition and the entangled state. These properties result in an exponential increase of the dimensionality of the state-space, which is the basis of the quantum speedup.
As the quantum counterpart of random walk, quantum walk [
34] becomes a novel computing model to analyze graph structures. Because of the amazing properties of quantum parallel and quantum interference, the quantum amplitude of quantum walks on a graph could represent more complex structural information. For kernels based on the discrete-time edge-based quantum walk (DTQW) [
12,
35] or the concrete-time node-based quantum walk (CTQW) [
11,
21], all of them are called quantum kernels, and the others are called classical kernels. In fact, quantum walk is the only method used to design the quantum kernels in literature.
Here we use DTQW as an instance. The discrete-time quantum walk is the quantum counterpart of the classical random walk. We denote the state space of DTQW as
, which is the edge set of a graph. And a general state of the quantum walk is:
where the quantum amplitudes
are complex. Using the Grover diffusion matrix, the entries of the transition matrix
is shown as follows:
where
shows the quantum amplitude for the transition
,
means the vertex degree and
is the Kronecker delta, i.e.,
if
, otherwise
.
Different from random walk where the probability propagates, what propagates during quantum walk is the quantum amplitude. Therefore, the quantum interference will happen between two crossing walks. In this paper, we only consider quantum computing as a novel computation model and evaluate the quantum kernels by running on a classical computer to simulate quantum walk.
5. Methodologies: Topological kernels vs. Entropy kernels
Graph and sub-graph matching is the key procedure for all the kernels. For most of the existing kernels, match mapping is computed via the topological analysis. In this kernel group, graph isomorphism or sub-graph isomorphism is the main method. However, the pairwise matching will cost a lot of time for the kernel computation. Therefore, adding some toy pattern constraints (e.g., edge, fixed-length walk, triangles) or constructing the product graph are the common methods to locate the matching. The product graph is usually an auxiliary structure to locate common sub-graphs, which is constructed by two graphs [
22]. However, the product graph will be large if the two graphs are big, which may still lead to unacceptable complexity.
In information theory, mutual information represents the diffusion of two probability distributions. Utilizing the mutual information entropy of the probability distributions of substructures is a novel trend to search the similar substructures. Therefore these methods are called entropy kernels in this paper.
Here 15 popular graph kernels are chosen and shown in
Table 1. Every kernel can be grouped according to the above five dimensions. From the groups of these kernels, we find that:
Most of these kernels are R-convolution, unaligned and local-pattern kernels.
All the entropy kernels here utilize quantum walk model to compute the probability distribution.
All of the information theory kernels here belong to the group of entropy kernels. Meanwhile, some R-convolution kernels which detect the similar substructure via the computation of information entropy also belong to this group.