1. Introduction
Many key insights and discoveries in the life sciences have been derived from atomic-scale 3D structures of proteins. Thanks to steady improvements in experimental structure determination methods, the PDB, or Protein Data Bank [
1], which stores these structures recently went past 125,000 entries—a landmark in our understanding of the molecular processes of life. This still lags far behind the growth of protein sequence information, with less than 0.1% of UniProt [
2] proteins linked to a PDB [
1] structure. However, the understanding that evolution conserves structure more than sequence has led to the large-scale computation of structural model efforts [
3]. Previously, we contributed to Aquaria [
4], a source built upon a systematic all-against-all comparison of Swiss-Prot [
2] and PDB sequences, resulting in a large number of template models [
4]; this provides a depth of sequence-to-structure information currently not available from other resources to the visible proteome. However, there is a dark side of the proteome which we were the first to categorize [
5], i.e., regions of protein sequences, or whole sequences, still remain inaccessible to either experimental structure determination or modelling approaches, such as Aquaria and others. This knowledge was updated and is kept in the independent Dark Proteome Database (DPD) [
6] as a synonym for the structurally unknown part of the protein sequence universe, i.e., full sequences and/or regions of sequences for which the structure is currently undetermined [
5,
6]. The intrinsically disordered proteins (IDP) and/or regions (IDR), and more recently also dark proteome [
7] are intrinsically unstructured proteins and/or regions, where their structure determination by conventional methods such as X-ray, nuclear magnetic resonance (NMR) crystallography and electron microscopy (EM) are arduous, due to the failure of homology methods, but most important due to their restless nature. Besides the fact that both definitions are focused on the structurally undetermined protein sequences, the first definition is broader and wider while the later narrows it to a small part of the former, i.e., only to the intrinsically unstructured regions. Nevertheless, both research paths stress the importance of studying the dark proteome, since it is still largely incomprehensible.
The present study serves as an introduction for two new features that were added to the Dark Proteome Database: (i) The availability of the ‘Features’ (FT) field of Swiss-Prot to deepen the knowledge of dark and non-dark proteins that allow the characterization of life domains and model organisms to get more thorough analysis; and (ii) the availability of an autonomy value per protein, allowing the analysis of the level of autonomy of dark and non-dark proteomes of those organisms as a whole.
Therefore, the main goals of this work are: (i) To detail even more the dark proteins present in the four domains of life, model organisms, and Homo Sapiens and (ii) to map how much of darkness is still present in these model organisms, since they are so important for mankind, i.e., to how much we do and do not know about a certain organism just by looking at the amount of darkness that it holds? Due to the number of existing organisms, we had to select the most important ‘model’ organisms (for us humans) while the ones remaining are work in progress. This choice is not innocent, since sequences originated from non-model organisms are known by the lack of sequence annotation, without being true orphan sequences. In the DPD [
6] we started with information from Homo Sapiens because it is of direct concern to us, but Earth is the home of millions of organisms and only approximately two handfuls of them were adopted as “model” for biological experiments, concerning food, infirmities, diseases, and threats. In an ideal world, we would have access to all the knowledge concerning every single organism that inhabits this planet. Since, we are not in the ideal world, these models were selected according to our present reality. In short, studying all of them in depth with today’s technology is almost impossible because experimentation is complex, time consuming, and very expensive.
Keeping our reality in mind, we have to remember that some of the most valuable methods in biological research are invasive or require an organism’s death. For all the previous reasons and more, much of this work is impossible or unethical to perform on humans and in certain organisms. As a solution, biologists have selected some model organisms to be used as testers. The following list contains a simple plant, a worm, a bacterium or prokaryotic, a simple eukaryotic, and a complex eukaryotic or mammal.
Finally, we will analyze the human organism (Homo Sapiens) in much more detail, including the genes from where dark proteins came from (‘dark’ genes) and tissues where dark proteins are expressed (‘dark’ tissues).
Arabidopsis thaliana (Plant)
Arabidopsis thaliana is considered a weed, also known as mouse-ear cress and is the most widely used plant as a model organism. One reason Arabidopsis makes a good model is because it undergoes the same exact processes of growth, flowering, and reproduction as most complex plants, taking only about one month and a half to grow completely producing a huge quantity of seeds in the process. Another reason is due to the fact that Arabidopsis has one of the smallest genomes in the plant kingdom with only 135 mega base pairs and five diploid chromosomes. It is the first plant with a completely sequenced genome.
Caenorhabditis Elegans (Worm)
Caenorhabditis Elegans is a soil worm or nematode and is considered a model for multicellular organisms. The reason why it is such a good model is related to the fact that it shares a common ancestor (“urbilaterian ancestor”) with humans that lived 500 million years ago, therefore sharing most of the genes that govern most modern organismal development and disease, such as the human and nematode. This is extremely important because many genetic and developmental experiments are impossible in humans (technically and ethically). Some evasive procedures associated with experiments are extremely time-consuming or cost-intensive. C. Elegans presents itself as a reliable alternative because of its short generation time (four days), and due to its complete anatomy also being known. The adult hermaphrodite has exactly 959 cells, while the adult male has exactly 1031 cells and both are transparent. This allows researchers to relate behaviors to particular cells, to trace the effects of genetic mutations and gain relevant insights into the mechanisms of development and ageing. Therefore, due to the evolutionary conservation of gene function, C. Elegans is the ideal model organism to trace basic genetic mechanisms of human development and disease, such as cancer and neurodegenerative diseases. C. Elegans was the first multicellular eukaryotic organism to have its whole genome sequenced.
Escherichia Coli (Bacteria)
The Escherichia Coli (E. Coli) can be found in the intestine of warm blood organisms. The reasons why this bacterium is such a good model are firstly, because it is a simple organism, and secondly that it can be cultured and grown easily and inexpensively in a laboratory. However, the main reason why it is heavily tested and trialed even though a bacterium, is related to the fact that its basic biochemical mechanisms are common to the human organism. Another important aspect is that E. Coli was the reason for first understanding the transcription factors that activate and deactivate genes in the presence of a virus. From that point on, E. Coli was used as a host in genetic engineering and especially in health, producing several types of proteins, encoding them with the majority of human genes that are applied as medicinal drugs. There are several variants of E. Coli, most of them harmless. However, some of them can be lethal and are responsible for product recall due to food contamination or food poisoning. This bacterium was the first prokaryotic model organism to have its genome sequenced (K12 strain) and has a single circular chromosome with 4.6 million base pairs.
Saccharomyces cerevisiae (Yeast)
Saccharomyces cerevisiae is recognized as the key factor in brewing for centuries. The main reasons for it being considered a model organism are related to its easy culture, fast growing process, and inexpensive production in a laboratory. Being a eukaryote, it share the same complex internal cell structure of plants or animals without the high percentage of junk DNA present in more complex eukaryotes facilitating research. Since S. cerevisiae is biochemically very similar to the human organism, many studies about the molecular processes involved in cell cycle, meiosis, recombination, DNA reparation, ageing, and other fundamental areas of biology were possible. S. cerevisiae was the first eukaryotic genome to be completely sequenced. It is composed of 12 chromosomes containing approximately 12.2 million base pairs.
Mus musculus (Mouse)
The Mus Musculus (Mouse) is the most famous model organism because is the mostly used mammal in medicine and biology scientific communities. The main reason why is such a good model is that it is a mammal and, therefore, has organs and development processes that are very similar to the human organism; next, they are easily reproductible, grow fast, and very easy to maintain and manipulate in a laboratory; finally, mice suffer from most of the diseases and calamities that affect mankind. Therefore, mice have an extremely important role in the development of new pharmaceuticals for humans. The mouse genome consists of 40 chromosomes with 2.63 billion amino acids.
In our previous work [
5], we analyzed the four domains of life where the conclusions were: The dark proteome is mostly not disordered, mostly not compositionally biased, mostly not transmembrane, but more important and unexpectedly, it is mostly “Unknown Unknowns” [
5]. The dark protein portion of “Unknown Unknowns” in eukaryota is almost 50%. It is composed of ordered, globular, and low compositional bias proteins. In the case of bacteria this percentage is over 50% and in case of archaea reached almost 70%. Finally, in viruses this percentage reached almost 75% [
5]. There were several questions raised at that time, that are still valid today, such as: Could we detail more dark proteins location and environment? Could we detail even more its functions?
2. Materials and Methods
Dataset: The set of protein sequences selected for this work were prevenient from Swiss-Prot release of July 2016 [
6]. The protein structures were extracted from PDB on July 2016. Predictions from PSSH2 [
4], PMP [
8] and Predict Protein [
9] are versions from July 2016. Finally, Protein-Protein Interaction (PPI) information is prevenient from STRING [
10], also from July 2016. The Swiss-Prot dataset is composed of 550.116 proteins and divided in four kingdoms: 19.370 protein sequences from archaea, 332.327 from bacteria, 181.814 from eukaryota and 16.605 from viruses. The number of proteins sequences for each model organism are: 14.349 protein sequences for Arabidopsis, 3.652 for C. Elegans, 669 for E. Coli, 42 for S. cerevisiae, 16.747 for Mouse and finally 20.209 for Human.
Mapping Darkness: For each Swiss-Prot protein, each residue was categorized as “non-dark” if it met either one of the following criteria: If the residue was aligned onto the “ATOM” record of any PDB entry [
1] in the corresponding Aquaria matching structures entry (criterion A); or if the residue was aligned onto a PDB entry in the corresponding UniProt entry (criterion B). All other residues were categorized as “dark.” We then calculated a “darkness” score (
D) as defined in Reference [
5]. If
D = 0 this means it is PDB or a white protein, otherwise, if
D = 1, this means it is a dark protein. If 0 <
D < 1 means it is a grey protein with grey regions containing dark regions [
5].
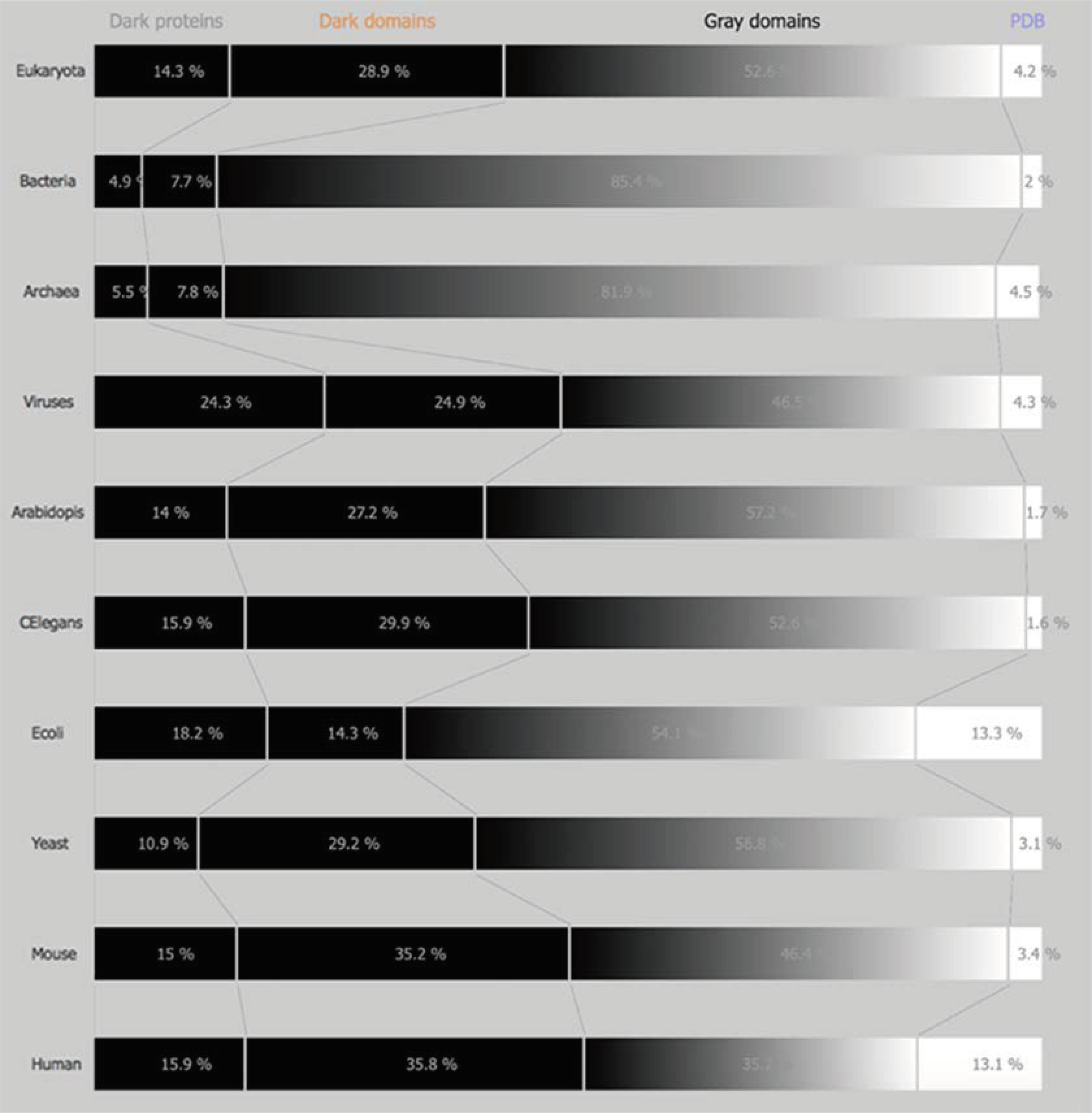
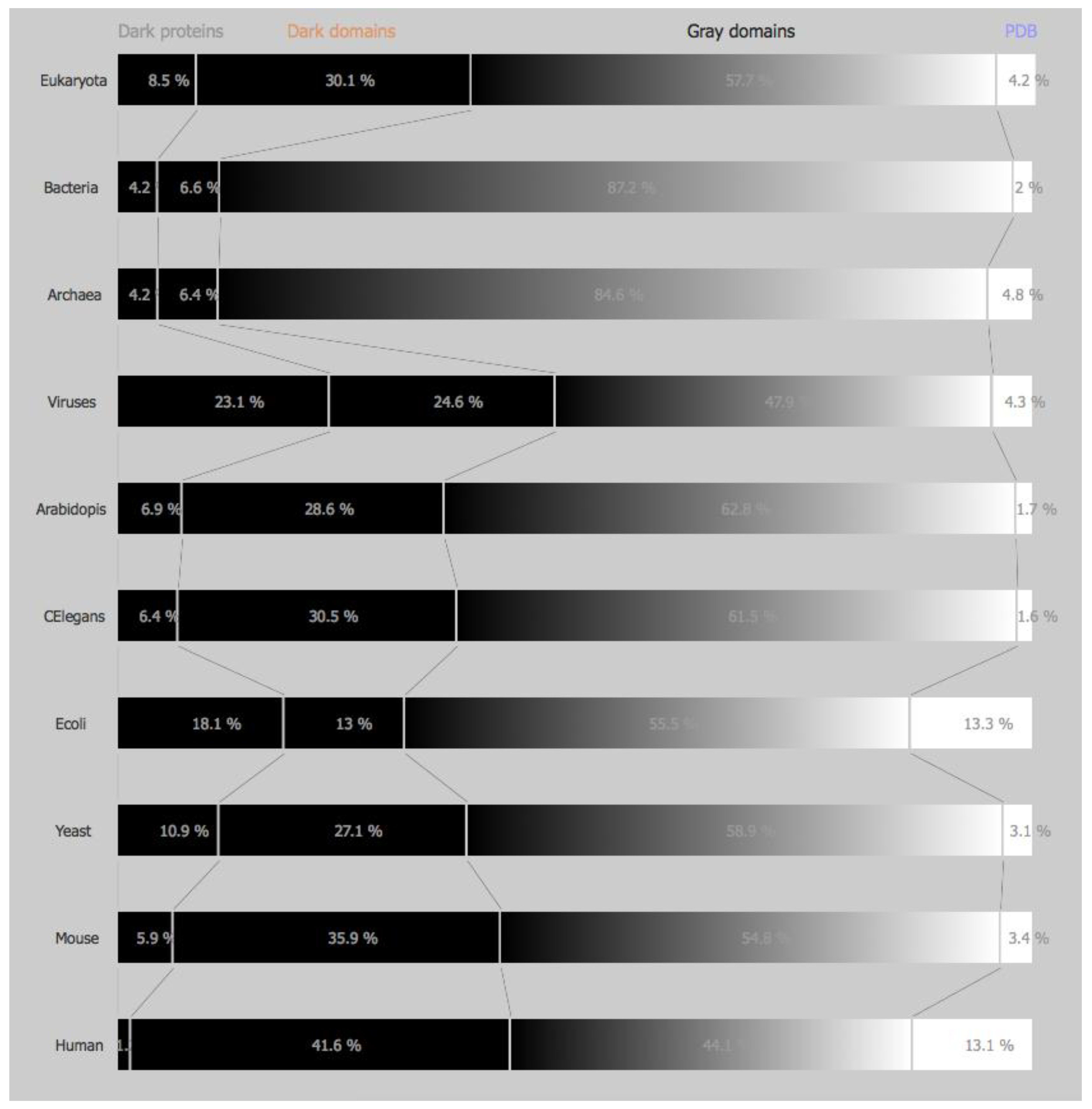
Dark and non-Dark Percentages: The percentages displayed for “dark” proteins, “dark” regions, grey regions, and PDB regions present in the above sets (domains of life and model organisms) consist first in obtaining both “dark” and PDB proteins in the sets mentioned above. Next, “dark” as well as, “non-dark” regions are mapped, subtracting the “dark” proteins from the former, and subtracting the PDB proteins from the later, obtaining the cardinality of “dark” regions and “non-dark” regions. If we divide the above cardinalities by the total amount of dark and non-dark regions, we obtain the percentages presented in
Figure 1 and
Figure 2.
Annotation Enrichment: The functional analysis compares annotations between dark and non-dark proteins in a reliable manner, by the application of annotation enrichment to the ‘Description’ (DE) field, which were now extended with the ‘Features’ (FT) field of the Swiss-Prot proteins through Fisher exact tests [
11,
12] followed by the Benjamini–Hochberg false discovery correction [
13] with α, the fraction of false positives was considered acceptable, set to 1%, and accepted only annotations with an adjusted
p value of ≤ 1%, calculated via:
where
p is from Fisher’s test,
n is the total of number of annotations in the set, and
k is the rank of the largest
p value that satisfies the false discovery criteria as in Reference [
6]. This approach was then repeatedly applied to compare dark and non-dark proteins across various sets of organisms.
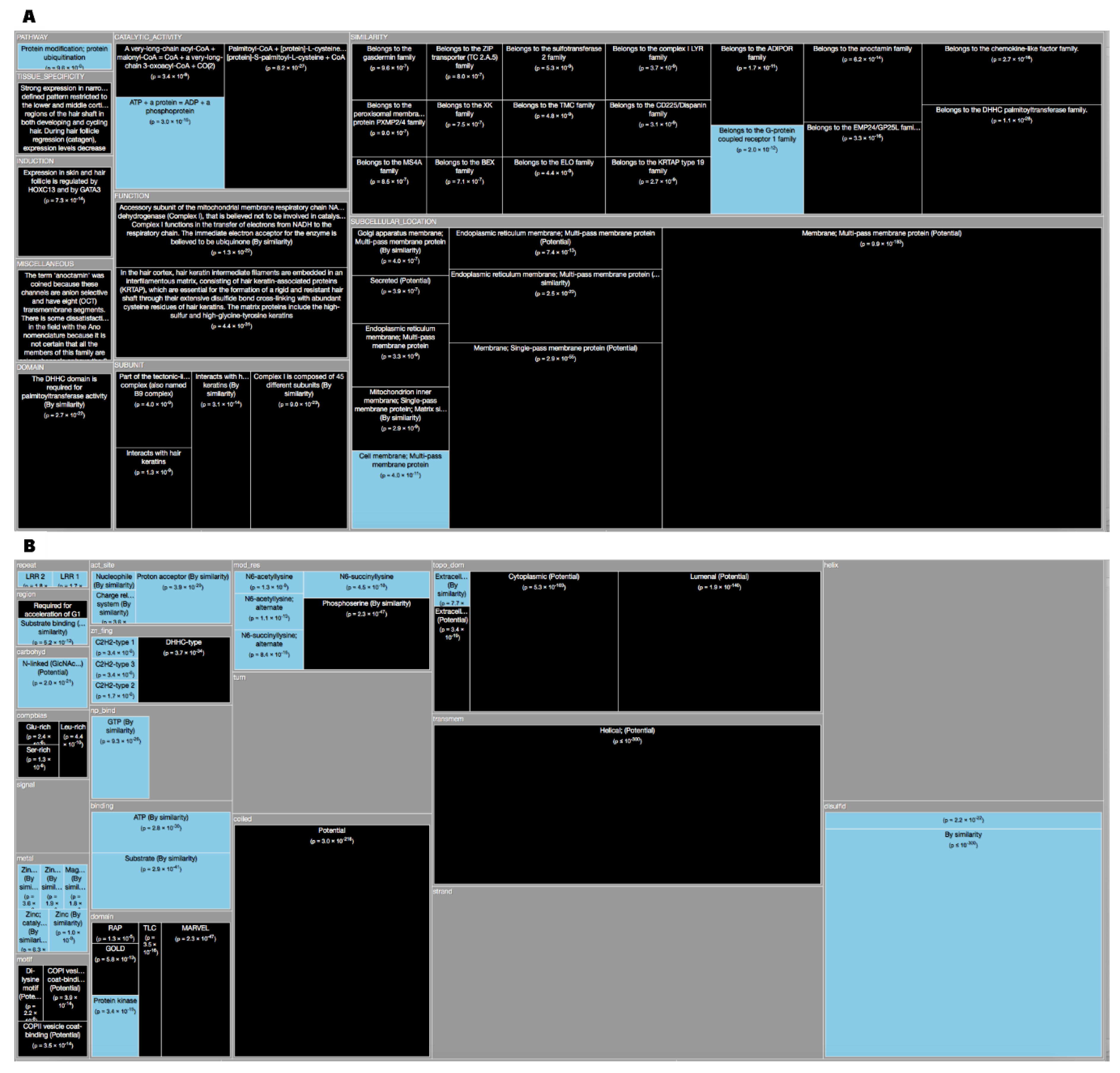
Tree Maps. From the ‘Description’ enrichment analysis results, we selected 21 (of 25) subcategories judged to be most significative and visualized them using a tree map [
14]. For the ‘Features’ enrichment analysis results we selected 36 (of 39) subcategories. The removed subcategories included those with relatively few results—or results with relatively high adjusted
p values—as well as subcategories such as “Similarity,” which only give information about groups of very similar proteins and the specific functions they perform; although interesting, these specific annotations do not reveal more general properties of dark proteins. In
Figure 3,
Figure 4,
Figure 5,
Figure 6,
Figure 7,
Figure 8 and
Figure 9 the results were displayed using the D3 zoomable tree map library (bost.ocks.org/mike/treemap); some annotation terms have also been reworded to improve readability.
Mapping Autonomy per protein. For every human protein we evaluated its autonomy, i.e., using STRING [
10] we counted how many others it interacts with. The STRING scheme classifies its functional link confidence into three different scores [
15]: Low (< 400), medium (400 < score < 700) and high (> 700) confidence scores measuring the confidence in the pair-wise functional interactions of the networks produced. Even assuming that sequence data is accurate, computational tools can introduce noise when generation sequence similarity data occurs. Taking this noise into account, it is suggested to set a cut-off score above which an interaction is highly probable. In terms of a functional classification accuracy, what matters is a high confidence score of 700 or higher [
16], however, low and medium confidence were done for comparison purposes (results not shown). Therefore, for each Swiss-Prot protein, we categorized its autonomy as:
where
m(N) indicates the number of matches that occur for a link score of
N. This means, if a protein has
m(0) equals to zero matches, then the protein is fully autonomous because at the lowest quality cut-off score no interactions occur between it and other proteins. On the other hand, if at the highest cut-off score there still exist interactions with other proteins (i.e.,
m(900) is not zero) then it can be concluded that the protein is completely non-autonomous.
Dark Genes. For each chromosome in Homo Sapiens, we then constructed a list of dark proteins sorted by the position of the central nucleotide of the corresponding gene, determined using UCSC assembly hg19 [
17]. In some cases, due to gene duplication, multiple copies of the same dark protein were annotated as arising from multiple genes in the same chromosome; in such cases, we considered only the first occurrence, and removed all other copies from the list. For each chromosome, we then calculated the longest run of dark proteins, and assigned a
p value by calculating how many times a run with the same number of dark proteins or more occurred by chance in 1,000 random re-orderings of proteins along the chromosome. Note, that the cluster results are very conservative where the chance of a false positive is 1/1000 on a per-chromosome basis; thus, there are probably more such ‘dark’ gene clusters.
Dark Tissues. Finally, we have used ProteomicsDB [
18] that contains mass-spectrometry data from protein expression measurements from 16,857 liquid chromatography tandem-mass-spectrometry (LC-MS/MS) experiments involving human tissues, cell lines, body fluids including data from PTM studies, and affinity purifications. To obtain the normalized intensity values for each protein from ProteomicsDB, the protein expression API was used. These values measure the relative abundance of peptides of each protein in a specific sample in a logarithmic scale. As we did not find any mass-spectrometry data for 1,391 dark proteins and 2,762 non-dark proteins, we considered these empty entries as 0.
4. Discussion
Our previous work [
5] didn’t point out solutions, but it opened a new field to be explored. This study is complementary to [
6] through the delivery of new information concerning darkness present in Homo Sapiens and in model organisms related with it pharmacologically. It can be concluded that the amount of dark proteome present in all of them is still high, whereas in higher eukaryotes like mouse and human, it is around 50%. The results presented above are consistent with previous works [
6,
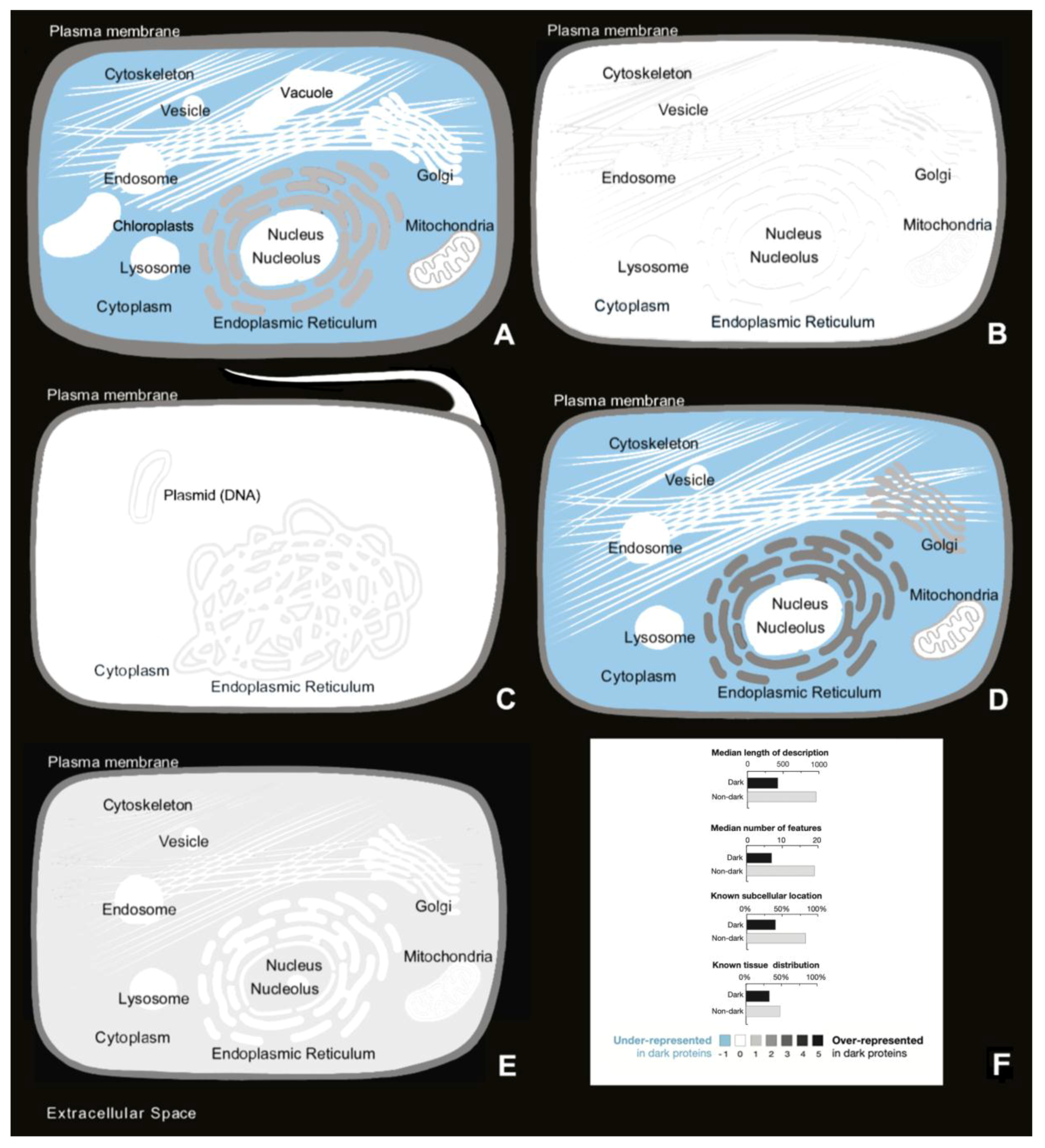
20] since Arabidopsis dark proteins are mainly located in extracellular space, cellular membrane, and endoplasmic reticulum membranes. C. Elegans dark proteins are present again in cell membrane where they are secreted. In the E. Coli case, it is reconfirmed that they are present in inner and outer membrane. Finally, in higher eukaryotic organisms like mouse, we observed that dark proteins are located in endoplasmic reticulum and in mitochondrion membrane, which is consistent with the previous results that state that dark proteins are mostly over-represented in specific secretory tissues and exterior environments, being also related to cancer endogenous retroviral proteins in the human organism [
6]. Therefore, it was shown that dark proteins are not uniformly distributed throughout the different areas of the cell in organisms, where their presence is more common in some regions than in others. There are a lot of them in membranes, cell membranes or associated with transmembrane regions and cleavage, but they are less common in cytoplasm, where many globular proteins perform their activity.
Concerning functions, the results confirmed that dark proteins perform a wide spectrum of functions depending on the organism in question, being more focused in simpler organisms and wider in higher organisms [
6]. Again, it was found that a vast amount of them is programmed to live outside the cell, where many are associated with secretion (through secretory glands and ducts) or with extracellular areas in tissues, being an indicator that they possibly are designed for being defensive agents against external threats such as bacteria and/or virus. But we also observed that some of these dark proteins are subject to post-translational modifications, therefore being chemically modified after translation be applied.
Concerning autonomy, up to now there is no comprehensive map of all relevant functionally for PPI’s in simple or complex organisms. The existence of this map is of crucial importance to understand cellular behavior. Several databases started to flourish helping in the construction of this global protein interactions map. Some databases are dedicated to register interaction experiments such as physical binding detection among proteins [
24,
25,
26,
27]; others are centered on specific model organisms [
28,
29] However, there are two difficulties: The first is the “tsunami” of genome and proteome sequencing information that must be processed putting the above map in standby; The second difficulty is in the way proteins interact i.e., they also interact through indirect associations such as shared pathways which are not registered in interaction databases, but instead are registered in pathway databases [
30,
31,
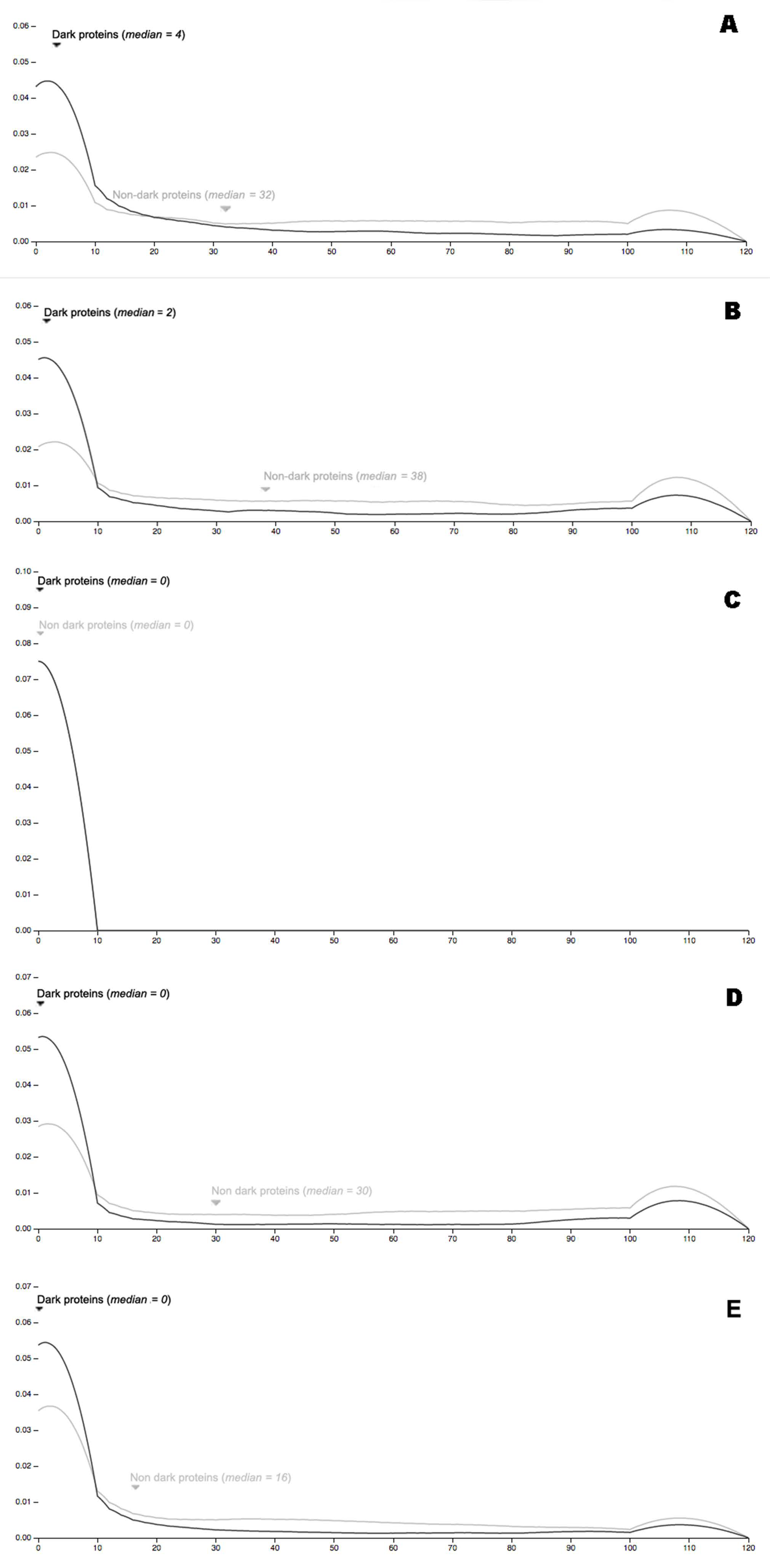
32] This is our contribution to the above map, especially to its dark side. The results show clear evidence that—independently of the organism evaluated—dark proteins have significantly fewer interactions with other proteins, in comparison with non-dark proteins. In general, we can conclude that dark proteins are more independent and autonomous than non-dark proteins. Therefore, the DPD is a map for the dark proteome at the present time where the model organisms described are already available together with its functional analysis, augmenting the knowledge about them, where we have work in progress for all the remaining organisms.
A point that we want to bring to discussion is the difference between intrinsically disordered proteins (IDP’s) and their relationship with the Dark Proteome (DP) as we coined it in 2015 [
5], where we concluded that the Dark Proteome is mostly not disordered using the predictor IUPred [
33]. The need to use a predictor was because only 62 proteins of Swiss-Prot (data from 2014) existed with ´disordered´ annotations from a total of 546,000 proteins. In our subsequent work [
6] (data from 2016), the same 62 proteins remained but now among a set of 550,116 Swiss-Prot proteins. For this work, considering ‘disordered’ annotations for the Human organism we would have (data from 2016 [
6]) only one protein to work with: Q8WYP5 (the same for 2014 data [
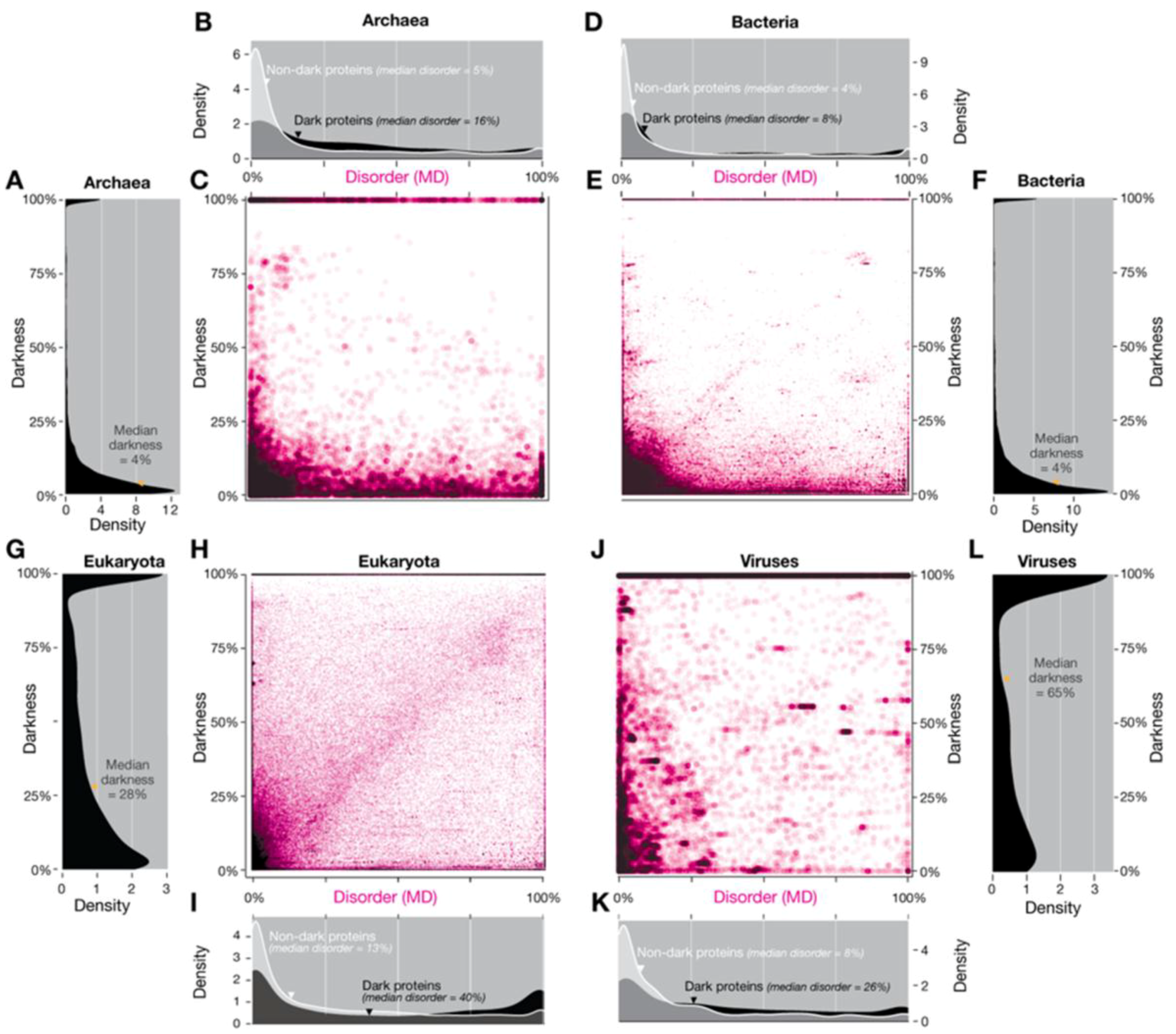
5]). Hence, we make the hypothesis: If we use another predictor do we get a different result? The answer is yes, but no. The disorder values shown in Figure 2 and Figure S3 of Perdigão et al. [
5] were calculated using IUPred [
33] because it is one of the most widely used methods for predicting disorder. Residues were defined as disordered if they had an IUPred score ≥0.5 [
5]. In this study, we also calculated a second set of disorder values using MD (META-Disorder) [
34], a machine-learning method that calculates a consensus disorder from several orthogonal methods. Re-plotting the density and scatterplots Figure 2 and Figure S3 of Reference [
5] using MD disorder gave a similar overall pattern, although some differences were apparent (
Figure 12). MD includes as one of its input methods DISOPRED2 [
35], which is one of several available methods that are optimized to predict residues missing from PDB structures. For a small fraction of proteins there were not MD predictions; to balance the comparisons, these proteins were removed from the density and scatterplots in Figure 2 and Figure S3 of Reference [
5]—thus reducing the number of proteins to 175.646 in archaea, 18.999 in bacteria, 326.945 in eukaryotes and 16.316 in viruses, respectively.
Intrinsic disorder in proteins is a complex and poorly understood phenomenon, in addition to IUPred, many other prediction methods have been developed focusing on a range of different aspects of disorder [
36]. It would certainly be of interest to compare darkness with disorder predictions from a range of methods, however the output from these algorithms is difficult to decode due to the lack of metrics or references to compare with [
37]. The DPD wants to help in a near future with the introduction of three new disorder predictors applied in the Swiss-Prot universe.