1. Introduction
Phospholipids (PL), specifically glycerophospholipids, are amphiphilic molecules and key components of cell membranes [
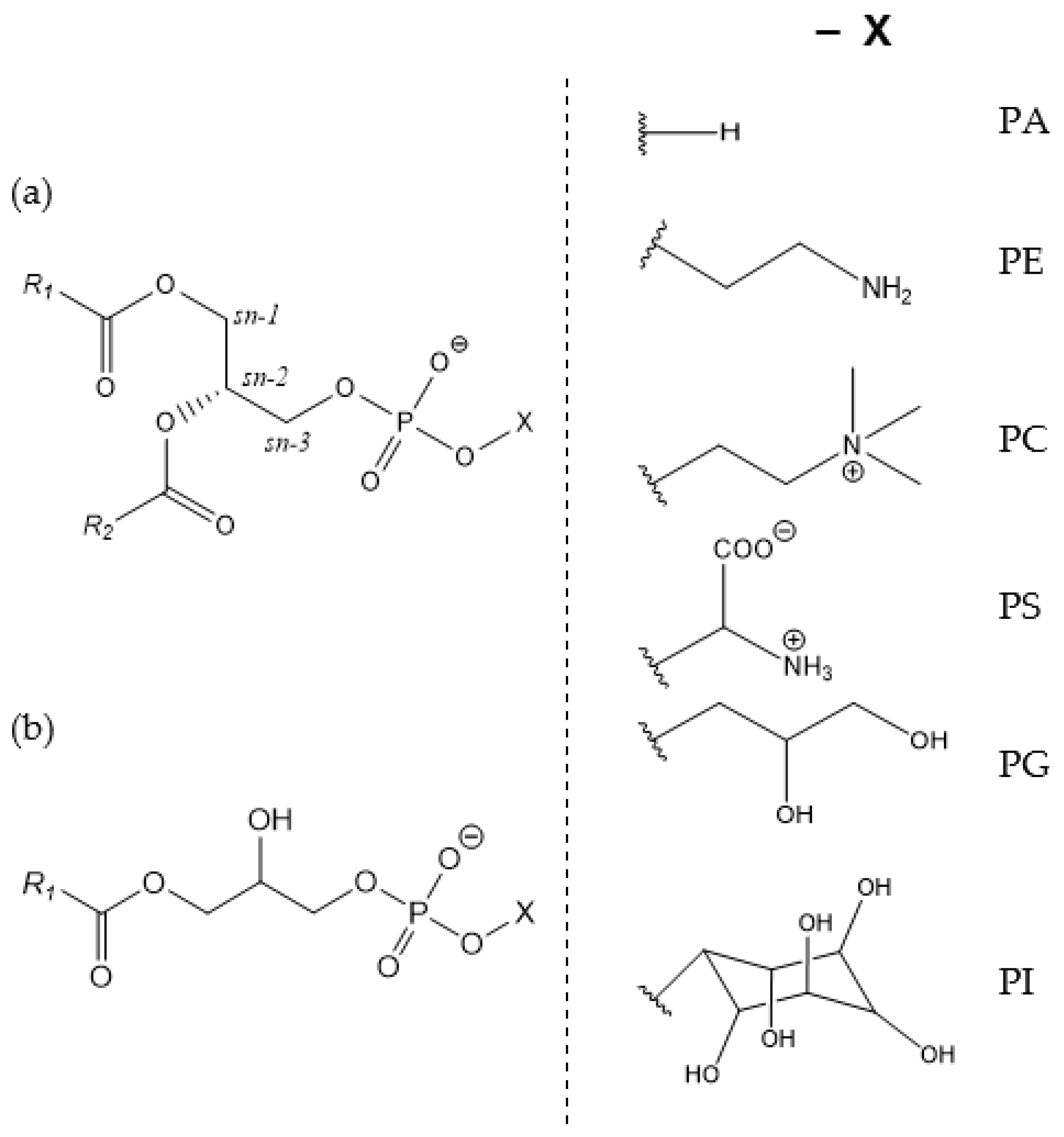
1]. They consist of a hydrophilic phosphate head and two hydrophobic fatty acid tails linked by a glycerol backbone (
Figure 1). In PL, fatty acids of varying lengths and degrees of saturation (R1 and R2) are esterified to the
sn-1 and
sn-2 positions of the glycerol. At the
sn-3 position, a phosphate group is esterified and modified with an alcohol—together forming the polar head group (-X). Depending on the type of alcohol, different classes of PL are distinguished: phosphatidic acid (PA), phosphatidylethanolamine (PE), phosphatidylcholine (PC), phosphatidylserine (PS), phosphatidylglycerol (PG), and phosphatidylinositol (PI). If one acyl chain is missing, as shown in
Figure 1b, the resulting molecules are classified as lysophospholipids (LPL) [
2,
3,
4].
The amphiphilic nature of PL enables their spontaneous self-assembly in aqueous environments into various structures, including micelles, bilayers, and liposomes. The type of self-assembled structure formed is determined by the interplay between the polar head group and the hydrophobic fatty acid chains, particularly their length and degree of unsaturation. Even subtle differences in molecular structure can significantly affect the aggregation behavior, stability, and functional performance of PL-based systems [
5,
6]. These structure-dependent properties directly determine the suitability of PL for use as encapsulating and emulsifying agents, which is widely exploited in the pharmaceutical [
2,
7], cosmetic [
8], and food industries [
9,
10] to improve the solubility, stability, and bioavailability of active compounds.
In pharmaceutical applications, synthetic PL with precisely defined composition is preferred to ensure consistency and controlled drug delivery [
10]. In contrast, the food and cosmetic industries primarily rely on lecithin [
9], a natural mixture of different PL, fatty acids, triglycerides, glycolipids, and sterols [
4]. The PL composition of lecithin varies depending on its source (egg yolk, soybean, milk, sunflower, etc.) [
9], influencing its suitability for specific formulations such as micelles, liposomes, and other assemblies [
11,
12,
13,
14]. Therefore, accurate identification of individual PL in lecithin is essential for predicting its functional properties or quality control of the industrial process.
Numerous analytical techniques have been applied, differing primarily in the amount of structural information they provide [
15]. One of the most widely used methods for determining total PL content in lecithin is phosphorus determination. Because all glycerophospholipids contain a phosphate group, quantifying phosphorus provides an indirect measure of total PL content [
16,
17]. This method is commonly used for routine quality control, as it is relatively simple, robust, and suitable for industrial applications. However, phosphorus determination does not provide information on the distribution of individual PL classes or molecular species and therefore offers limited insight into the functional properties of lecithin.
To obtain class-level information, thin-layer chromatography (TLC) [
18,
19] and normal-phase high-performance liquid chromatography (NP-HPLC) [
20,
21,
22] are frequently used. These techniques separate PL primarily based on the polarity of their head groups and enable differentiation between major PL classes. When coupled with detectors such as evaporative light scattering detection (ELSD) [
23,
24,
25] or ultraviolet (UV) detection [
26,
27,
28], NP-HPLC allows determination of PL classes and is therefore widely used for quality control and compositional comparison of lecithin samples. However, both TLC and NP-HPLC generally do not resolve individual molecular species within the same PL class and thus fail to capture the structural diversity arising from differences in fatty acid chain length and degree of unsaturation.
For applications requiring deeper insight into PL composition, such as studies of emulsification behavior, oxidative stability, or structure–function relationships, information at the level of individual molecular species is essential. In this context, reversed-phase high-performance liquid chromatography (RP-HPLC) is particularly advantageous [
21,
29]. The RP-HPLC separates PL based on hydrophobic interactions, which are determined mainly by the length and degree of unsaturation of the fatty acid chains. As a result, PL within the same class but differing in acyl chain composition can be effectively resolved. This makes RP-HPLC a powerful tool for investigating molecular-level variability in lecithin that cannot be accessed by class-based analytical methods. The RP-HPLC is often coupled with mass spectrometry (MS), which provides molecular mass information for PL species [
30,
31,
32].
However, a major challenge in PL identification using MS is that many PL species share the same molecular ion because of their isomeric structures. Consequently, most analytical methods rely heavily on reference standards to ensure accurate identification. However, obtaining a complete set of PL standards is both costly and impractical, limiting the feasibility of comprehensive PL analysis. Therefore, there is a growing need for an alternative approach that enables PL identification without full dependence on commercial standards. Developing a method that accurately determines PL composition based on MS data would enhance analytical efficiency, reduce costs, and make PL analysis more accessible for research and industrial applications.
The main aim of this research work was to develop an analytical method for identifying PL in lecithin samples without using standards, to facilitate further study of the structural dependence of lecithin composition on emulsification properties and oxidation processes in developed food supplement formulations. This study is the first in a series; however, due to the complexity of the field, it is presented in a separate paper. The aim of this work is to provide proof-of-concept rather than to systematically analyze a large variety of lecithins.
2. Materials and Methods
2.1. Chemicals
Acetonitrile (LC-MS grade) and methanol (HPLC grade) were purchased from J. T. Baker (Phillipsburg, NJ, USA), while formic acid (99%, HPLC grade) was obtained from Carlo Erba (Cornaredo, Italy). The deionized water was purified using an Adrona (Riga, Latvia) purification system (18.2 MΩcm at 25 °C) prior to use.
Phospholipid standards (18:0–18:2) PC (1-octadecanoyl-2-(9Z,12Z-octadecadienoyl)-sn-glycero-3-phosphocholine), (18:1) LPE (1-(9Z-octadecenoyl)-sn-glycero-3-phosphoethanolamine), (18:1) PC (1,2-di-(6Z-petroselenoyl)-sn-glycero-3-phosphatidylcholine) and (18:1) LPC (1-(9Z-octadecenoyl)-sn-glycero-3-phosphocholine) were purchased from Avanti Research (Alabaster, AL, USA). The lecithin sample PC30 was purchased from Lecico GmbH (Hamburg, Germany).
2.2. Preparation of Standards and Samples Solutions
The concentrated sample solution for method development was prepared by dissolving lecithin PC30 in water at a concentration of 50 mg/mL, then diluting it to working concentrations of 5.0, 7.5, 10.0, 12.5, and 15.0 mg/mL for method development and validation.
The standard solutions for qualification confirmation were prepared by dissolving (18:0 18:2) PC, (18:1) LPE, (18:1) PC, and (18:1) LPC in ethanol at a concentration of 0.5 mg/mL.
2.3. Instrumentation
A Thermo Scientific Dionex UltiMate HPLC system, equipped with a quaternary pump (LPG-3400SD), an autosampler (WPS-3000 SplitLoopRS), a column thermostat (TCC-3000), and a variable wavelength detector (VWD 3100), was used for chromatographic separation. The HPLC system was coupled to an Exactive Plus Orbitrap tandem mass spectrometer (ThermoFisher Scientific, Waltham, MA, USA) via an electrospray ionization interface. Chromeleon 6.8, Chromeleon 7.2, Tune 2.9, and XCalibur 3.0 (ThermoFisher Scientific, Waltham, MA, USA), Microsoft Excel (Microsoft Office 365, Redmond, WA, USA), and ChemDraw 22.2.0 (Revvity, Waltham, MA, USA) were used for instrument settings and data processing.
2.4. HPLC and MS Measurements
In the procedure developed, we used HPLC coupled with high-resolution mass spectrometry (HRMS). HRMS provides high mass accuracy and resolution, enabling precise determination of molecular and fragment ions, which is essential for distinguishing structurally similar PL species. Given the complex nature of lecithin, which contains a mixture of different PL classes and molecular species, HPLC was used to separate individual analytes before MS analysis. The PL molecular ions were detected in full scan mode, while fragmentation spectra for structural characterization were obtained in MS-MS mode. In-house software was developed to propose individual candidates based on MS spectra and is available upon request.
Analysis of samples was performed using a reverse-phase separation column, Waters XBridge Protein BEH C4, with dimensions of 2.1 × 150 mm and a particle size of 1.7 µm. The flow rate was set to 0.4 mL/min, the injection volume to 1 µL, and the column oven temperature to 25 °C. UV detection was performed at a wavelength of 203 nm. The mobile phase consisted of 0.1% (
v/
v) formic acid in water (A) and acetonitrile (B). The gradient elution used to achieve separation is presented in
Table 1.
The MS analysis was performed on an Exactive Plus Orbitrap MS (Thermo Scientific, Bremen, Germany) with an electrospray ionization (ESI) source. A high-resolution mass spectrum was acquired in full scan mode over a mass range of m/z 67–1000 at a resolution of 70,000. MS/MS data were obtained at a resolution of 70,000. Mass spectrometric detection was performed in positive ion mode. The ion source parameters were as follows: sheath gas and auxiliary gas were nitrogen (purity ≥ 99.99%) with flow rates of 4 L/min and 15 L/min, respectively; capillary temperature was 380 °C; probe heater temperature was 380 °C. The collision energy was set to 40 eV.
2.5. Software
To support the identification of PL in lecithin samples based on LC–HRMS data, an in-house software tool was developed using the Python 3.8 (PSF, Wilmington, DE, USA) programming language. The code consists of 53 lines. The software was designed to generate a list of theoretically possible PL candidates corresponding to experimentally observed molecular ions detected in Full MS mode. An example of the software output is presented on
Figure S1 in Supplementary Materials. For this purpose, a structural framework of glycerophospholipids was implemented, consisting of a glycerol backbone, a phosphate-containing polar head group, and one or two fatty acid chains. The algorithm systematically combined predefined polar head groups with fatty acid chains of varying length and degree of unsaturation to calculate the exact monoisotopic masses of potential PL species. Experimentally detected m/z values were compared with the calculated masses within a defined mass tolerance (±0.01 Da), and all matching structures were retained as possible candidates. The mass tolerance (±0.01 Da) was selected to reflect the expected mass accuracy of the Orbitrap instrument under the applied acquisition settings and to provide a robust screening window while minimizing false-positive candidate matches. Candidate generation was performed assuming protonated molecular ions [M+H]
+ in ESI+ mode. Other common adducts (e.g., [M+Na]
+, [M+NH
4]
+) were not included in the current implementation in order to keep candidate lists compact and because protonated species dominated under the applied mobile-phase conditions. However, the future versions of the software will include these options as well.
3. Results and Discussion
3.1. Optimization of Separation
The aim of this study was to develop a method primarily for the identification and possible quantification of PL in lecithin samples to enable further studies on the emulsification properties and stability of developed formulations. To achieve this, a chromatographic separation method needed to be developed to ensure that individual PL could be efficiently resolved prior to mass spectrometric (MS) analysis. As the goal was to identify individual PL, reverse phase chromatography was used to achieve separation based on the composition of the hydrophobic tail. This method allows differentiation of PL by chain length and degree of saturation. In contrast, normal phase chromatography primarily separates PL based on head group polarity, resulting in type-based separation without differentiation of individual species. Several separation columns with stationary C18, C8, and C4 phases were tested. Columns with C18 and C8 stationary phases, although commonly used for lipid analysis, exhibited significant limitations in this study. Both columns caused excessive retention of hydrophobic analytes, requiring very high proportions of the organic phase for elution. With the C18 column, most analytes eluted only at extremely high concentrations of the organic phase, indicating overly strong interactions with the stationary phase. The C8 column provided slightly shorter retention times due to its shorter alkyl chains, but some analytes still eluted only at high concentrations of the organic phase, indicating that retention remained problematic. Such strong retention can result in incomplete elution of analytes and reduced separation efficiency. Additionally, both columns exhibited broad peaks with significant tailing, reducing the overall resolution of the separation. The best selectivity and peak symmetry were achieved using the Waters XBridge Protein BEH C4 (2.1 × 150 mm) separation column with a particle size of 1.7 µm. The details of the chromatographic separation are described above in the Materials and Methods section.
Figure 2 shows the chromatogram of lecithin PC30 (Lecico) obtained using optimized separation conditions.
Once chromatographic separation was achieved, the method was validated on six selected peaks shown in
Figure 2, with retention times (t
r) listed in
Table 2. The precision of the chromatographic method, reported as RSD of peak areas, was estimated by measuring repeatability on six replicate injections at a lecithin concentration of 10 mg/mL. The RSD values of retention times for the selected peaks ranged from 0.31% to 1.70% (
Table 2), indicating good precision given the complexity of the analytical method. Five-point calibration curves were prepared with lecithin solutions in the concentration range of 5.0–15.0 mg/mL to evaluate linearity. Linearity was assessed for the six selected chromatographic peaks by plotting peak areas against sample concentration. The calibration curves for all six peaks showed strong linearity within the tested concentration range, with correlation coefficients (R
2) between 0.9985 and 0.9992. The R
2 values for each selected peak are summarized in
Table 2. The optimized chromatographic method provides sufficient separation of the major analytes in the lecithin sample, which is crucial for accurate identification. It should be noted that the validation (linearity and repeatability) refers to UV peak areas of the selected chromatographic peaks in the lecithin matrix and primarily demonstrates method performance for chromatographic separation and signal reproducibility. It does not constitute quantitative validation for individually confirmed PL species, which would require species-specific calibration with appropriate standards.
3.2. Identification of Phospholipids by LC-MS
After optimizing and validating the chromatographic separation, the lecithin sample was further analyzed using a coupled LC-MS technique. The LC was used to separate the analytes, and the MS was used to identify them in a two-stage procedure. In the first phase of the identification process, a total ion current (TIC) chromatogram of the lecithin sample was recorded in full scan mode. By examining the mass spectra associated with each chromatographic peak, was obtained the exact m/z values corresponding to the molecular ions of the analytes. These exact m/z values generated possible candidates, and further selection was accomplished using MS/MS information.
To facilitate the identification of PL candidates, a program was developed in Python. The tool was designed for two main purposes: (i) to assess whether a given molecular ion could correspond to a PL, and (ii) if so, to generate a list of all theoretically possible PL that could correspond to the observed m/z value. The tool compares experimental m/z values with the theoretical masses of all possible PL species, considering variations in headgroup, chain length, and degree of saturation. For diacyl phospholipids, the program pairs a headgroup with two fatty acid chains, while for LPL, only a single chain is considered. If the calculated exact mass is within ±0.01 Da of the experimental value, the PL structure is listed as a candidate. It is important to note that an m/z value may correspond to multiple PL structures due to the presence of structural isomers, such as those differing in fatty acid chain length, degree of saturation, or positional arrangement. If no matches are found, it is assumed that the peak originates from another compound present in the lecithin matrix. The output consists of a list of all PL candidates that could correspond to a particular peak, or no result if none are found.
The TIC chromatogram of the lecithin PC30 sample is shown in
Figure 3. For improved clarity and visualization, the TIC chromatogram was divided into two segments:
Figure 3a displays the TIC chromatogram from 7 to 18 min, while
Figure 3b covers 18 to 29 min. Peaks assigned to PL based on m/z matching are labeled, while unlabeled peaks correspond to other components in the lecithin.
Based on Python-supported analysis, the first 12 peaks were identified as LPL: lysophosphatidylcholine (LPC) or lysophosphatidylethanolamine (LPE), while the remaining assigned peaks corresponded to phosphatidylcholine (PC) or phosphatidylethanolamine (PE). This indicates that only two types of PL were detected in the lecithin sample. The observed retention behavior is consistent with expectations, as LPL elute earlier due to their lower hydrophobicity. Because they contain only one fatty acid chain, they interact less strongly with the non-polar stationary phase and therefore have shorter retention times compared to diacyl phospholipids.
It is noteworthy that two molecular ions were detected for the same peak at peaks 9 and 14, indicating the co-elution of different PL species. Co-elution was assessed by inspecting extracted-ion chromatograms for both ions. The extracted-ion chromatograms showed overlapping retention profiles with maxima at the same retention time, supporting chromatographic co-elution. For the LPL (the first 12 peaks), one or two possible structures corresponding to LPC or LPE were assigned to each peak. In contrast, for the remaining peaks corresponding to PC and PE, up to 53 different structural candidates were identified for a single peak due to the high variability of possible fatty acid compositions. After generating the candidate list for each chromatographic peak, the next step was to determine which specific PL corresponded to each peak by analyzing the presence of characteristic fragment ions using MS/MS analysis.
The MS/MS fragmentation analysis was performed in positive electrospray ionization mode (ESI+) at a collision energy of 40 eV. The proposed fragmentation patterns for PC, PE, LPC, and LPE species were adopted from the study by Pi et al. [
33], who examined the fragmentation patterns of five types of PL to identify characteristic fragment ions for each PL class. The theoretical fragment ions for each PL candidate were first calculated based on known fragmentation pathways (
Table 3). These theoretical values were then compared with the experimental MS/MS spectra. If a PL candidate had at least four fragment ions that matched those in the experimental spectrum, and their intensity was at least ten times higher than spectral noise level, it was confirmed as the correct assignment for that peak.
Table 4 summarizes the identified PL species that correspond to each chromatographic peak in the TIC chromatogram.
The results in
Table 4 show that certain LPL peaks, although occurring at different retention times, were identified as the same LPL species. This indicates the presence of isomeric forms that elute separately due to differences in structural configuration. For example, (18:2) LPC was detected at three different retention times, which can be attributed to geometric isomerism, as its two double bonds allow for up to four geometric isomers. However, (18:1) LPC, which contains only one double bond, was observed at four different retention times, which cannot be explained by cis–trans isomerism alone. A plausible explanation is acyl migration, a process described by Subbaiah and Sugasini [
34], in which the acyl chain shifts from the
sn-1 to the
sn-2 position. This type of isomerization, combined with geometric variations, is likely the cause of the large number of (18:1) LPC peaks.
Up to this point, all identifications have been made solely by matching molecular and fragment ion data, without using reference substances. However, to further evaluate the reliability of this approach, we performed experimental verification with defined PL standards.
3.3. Confirmation of Identification with Standards
From the list of PL previously identified in the lecithin sample, four were selected for which pure standards were commercially available. These included (18:1) LPE, (18:1) LPC, (18:1) PC, and (18:0–18:2) PC, covering both LPL and diacyl phospholipids, as well as two different PL types, allowing validation across structural subclasses and headgroup types.
Each of these standards were analyzed by LC-MS under the same conditions as the lecithin sample to verify the accuracy of peak assignment. For each standard, a TIC chromatogram was recorded in full scan mode and overlaid with the TIC chromatogram of the lecithin PC30 sample. The retention time of each standard was compared with that of the corresponding peak in the lecithin sample, which had previously been assigned to the same PL. As shown in
Figure 4, peaks 9, 10, 20, and 21 in the lecithin TIC chromatogram, which were assigned in
Table 4 to (18:1) LPE, (18:1) LPC, (18:0–18:2) PC, and (18:1) PC, respectively, exhibit identical retention times as the corresponding standards. In addition, the MS/MS fragmentation pattern of the peak in the lecithin sample was compared with that of the standard to confirm the match.
In all four cases, the retention times of the standards exactly matched those of the corresponding peaks in the lecithin TIC chromatogram. The MS/MS spectra also showed characteristic fragment ions that matched those of the respective standard compounds. This provided strong evidence that the previous peak assignments were correct.
It should be emphasized; however, that although the method reliably identifies molecular species, it does not allow differentiation between positional or stereoisomers that may still be present in the sample. These isomeric forms may differ in the position of the fatty acid chains on the glycerol backbone (sn-1 vs. sn-2) or in the configuration of the double bonds (cis vs. trans), both of which affect chromatographic behavior and elution profiles. Although shifts in retention times may indicate the presence of isomeric species, the current approach does not allow a definitive structural distinction. This limitation is particularly evident in the case of (18:1) LPC, where multiple isomeric peaks were observed in the lecithin sample, each eluting at a specific retention time. In contrast, the commercially available (18:1) LPC standard corresponds to a single defined isomer. Its comparison allowed confirmation of one peak in the sample, but not the full range of isomeric variants likely to be present.
4. Conclusions
This study presents a robust LC–HRMS-based analytical strategy for identifying PL in lecithin without relying on comprehensive sets of reference standards. By combining optimized reversed-phase chromatographic separation on a C4 column with high-resolution mass spectrometry and tandem MS, individual PL species were successfully resolved and structurally characterized within a complex lecithin matrix. The developed chromatographic method showed excellent linearity (R2 = 0.9985–0.9992) and good repeatability (RSD = 0.31–1.70%) based on peak area evaluation, ensuring reliable separation and reproducible detection of major PL components.
A key strength of the proposed approach is the integration of a Python-based in-house software tool that systematically generates, and screens theoretically possible PL candidates based on accurate mass data. Combined with class-specific MS/MS fragmentation analysis, this workflow enabled confident identification of both lysophospholipids and diacyl phospholipids, even in the presence of numerous isomeric species. The validity of the identification strategy was further confirmed by targeted analysis of selected commercially available standards, demonstrating excellent agreement in both retention behavior and fragmentation patterns.
While the method does not allow unambiguous differentiation of positional or geometric isomers, it provides a practical, cost-effective, and broadly applicable solution for molecular-level PL profiling when reference standards are limited or unavailable. Thus, the developed methodology represents a valuable analytical platform for future quantitation of individual PL based on a quantitative structure–property relationship approach. The final analytical procedure will facilitate studies investigating structure–function relationships, emulsification behavior, and oxidative stability of lecithin-based formulations.