Polysialic Acid Modulates the Binding of External Lactoferrin in Neutrophil Extracellular Traps
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Isolation of Bovine Neutrophils
2.3. Polysialylated Fluorescent Bead Accumulation in NETs
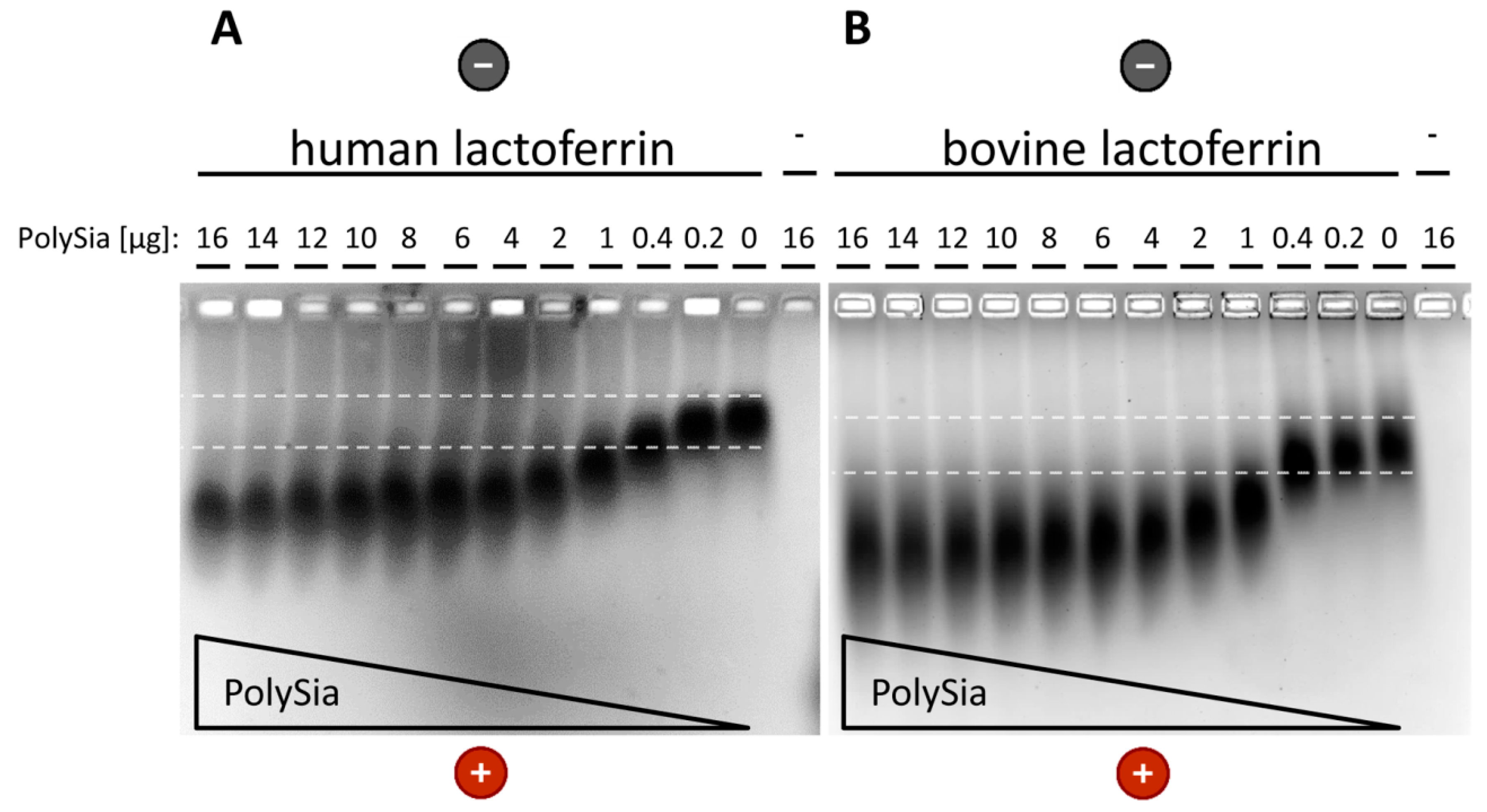
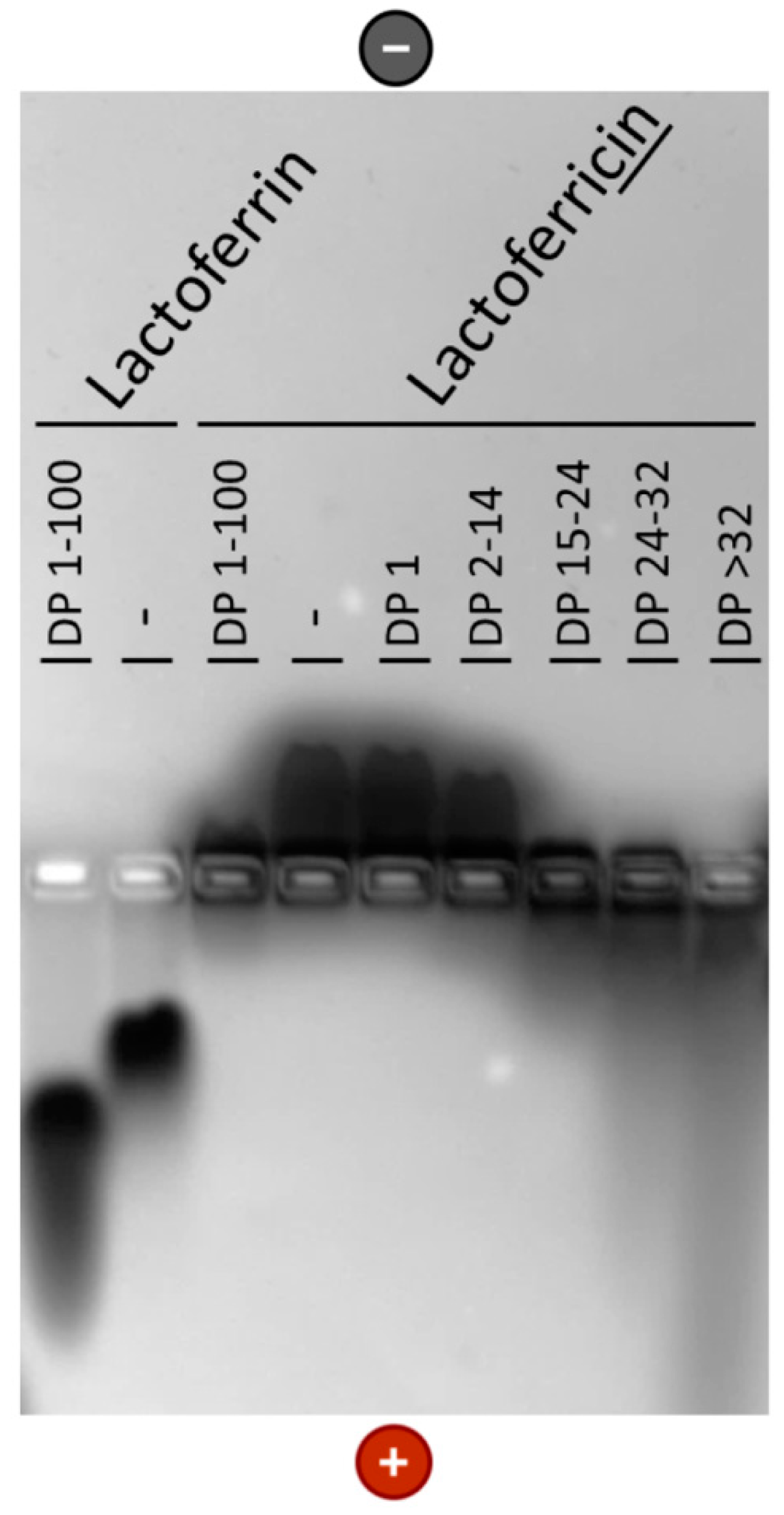
2.4. Native Gel Electrophoresis
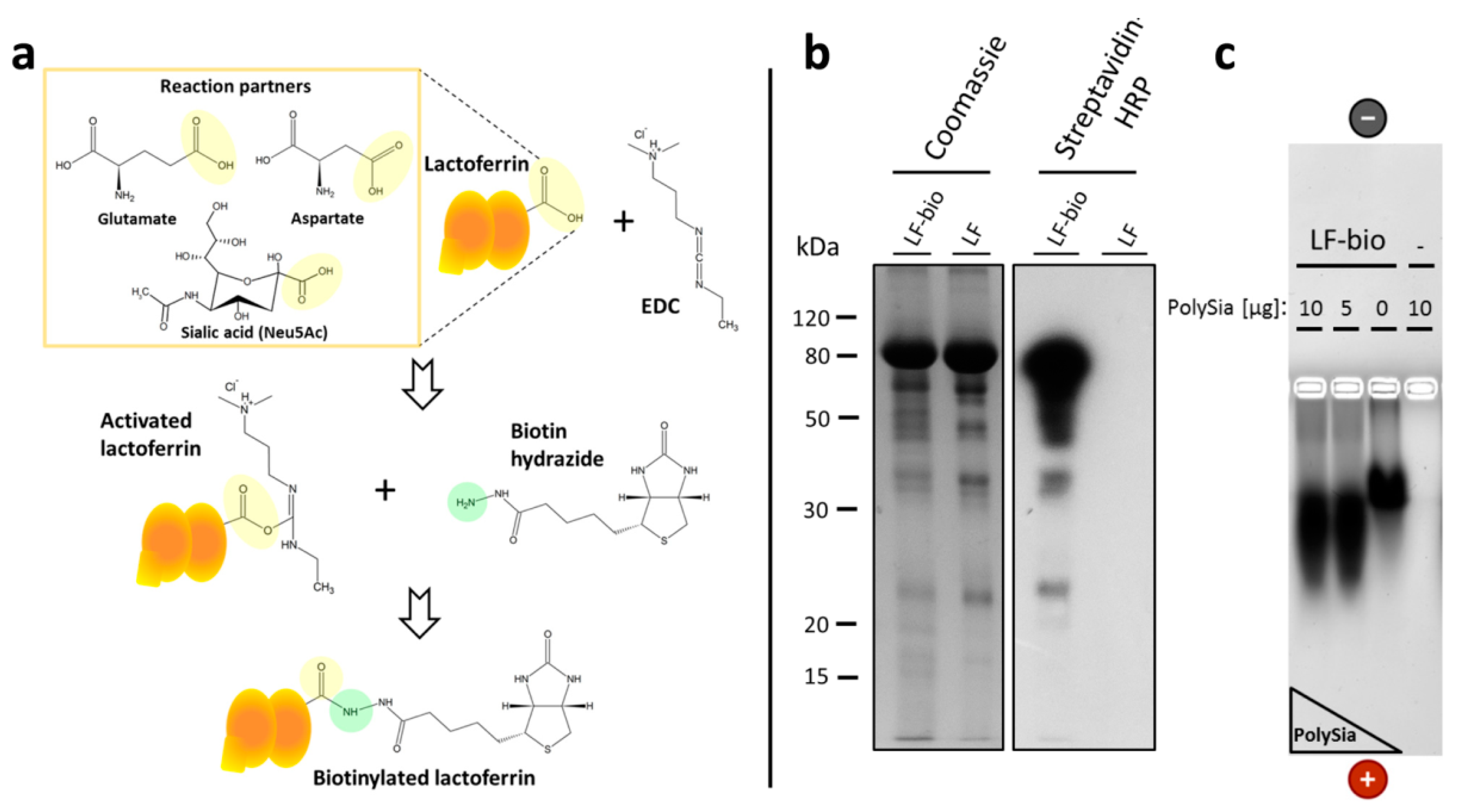
2.5. Biotinylation of Lactoferrin
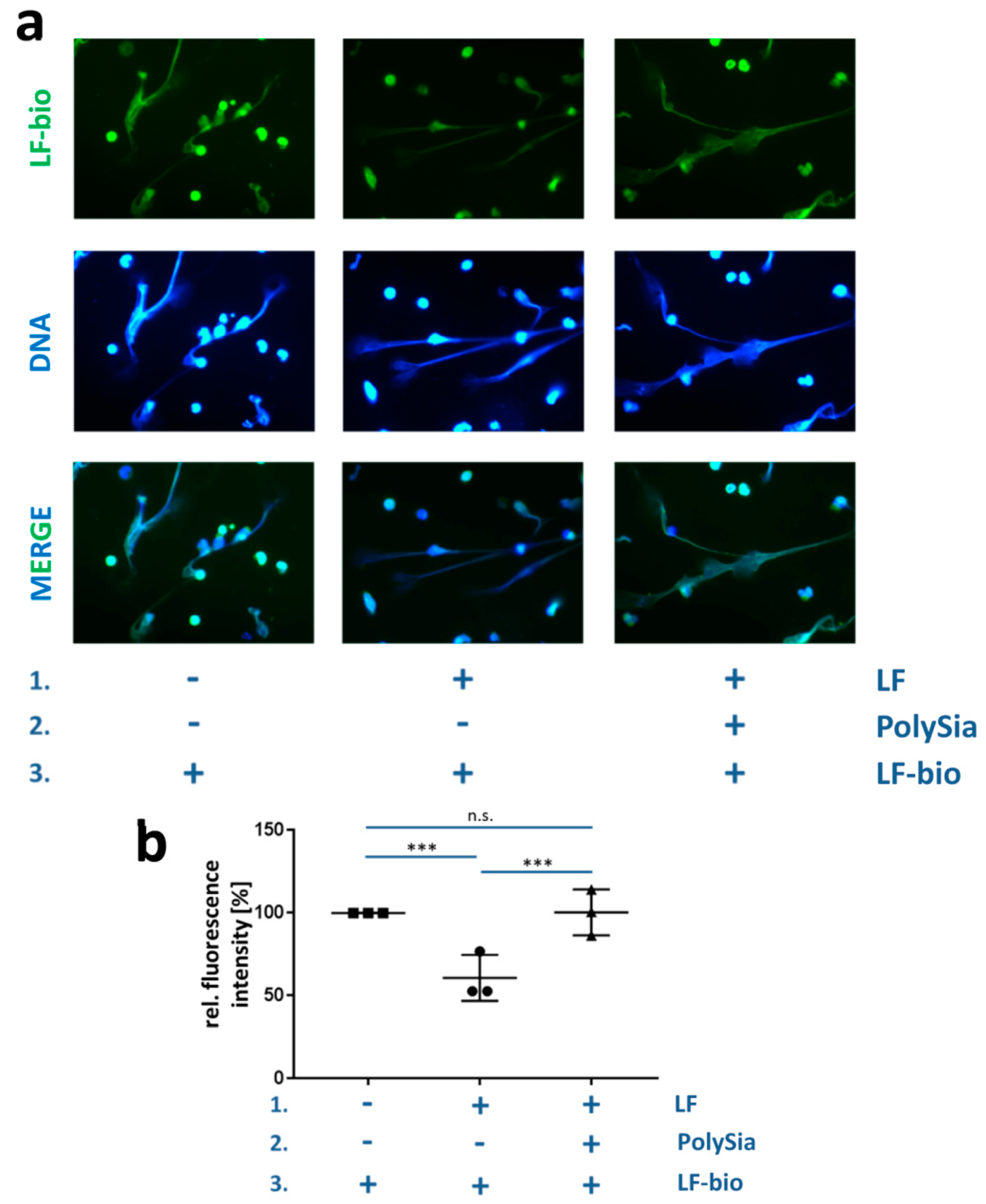
2.6. Enrichment of Lactoferrin-Biotin in Bovine NET Fibers
2.7. Statistical Analyses
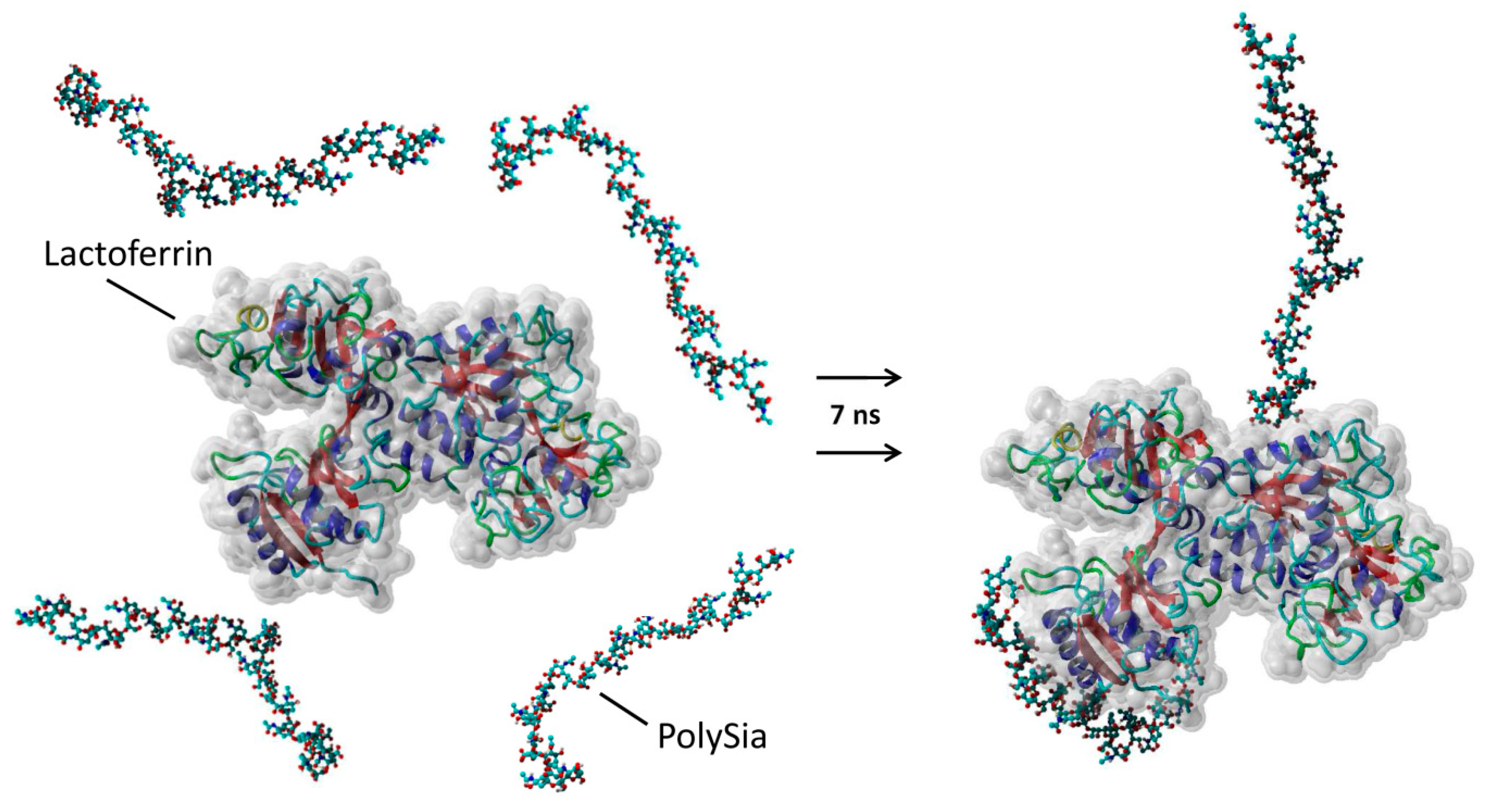
2.8. 3D Simulations
3. Results and Discussion
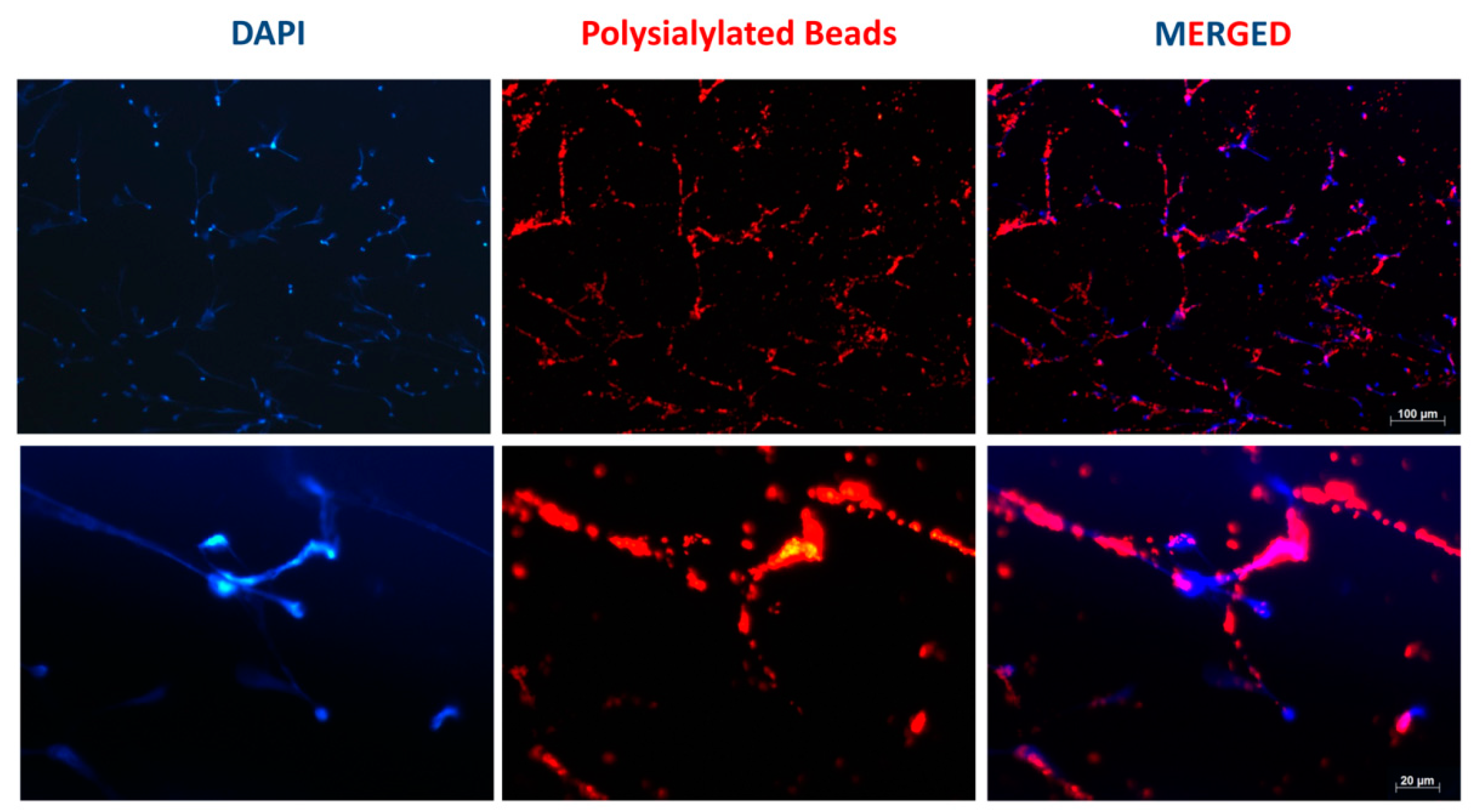
3.1. Accumulation of PolySia Coupled to Red Fluorescent Particles in NET Fibers
3.2. Interaction of PolySia with LFcin Depends on Its DP
3.3. PolySia Supports the Enrichment of External Lactoferrin into NET Fibers
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Gray, R.D.; McCullagh, B.N.; McCray, P.B. Nets and cf lung disease: Current status and future prospects. Antibiotics 2015, 4, 62–75. [Google Scholar] [CrossRef] [PubMed]
- Fournier, B.M.; Parkos, C.A. The role of neutrophils during intestinal inflammation. Mucosal Immunol. 2012, 5, 354–366. [Google Scholar] [CrossRef]
- Phillipson, M.; Kubes, P. The neutrophil in vascular inflammation. Nat. Med. 2011, 17, 1381–1390. [Google Scholar] [CrossRef]
- Brinkmann, V.; Reichard, U.; Goosmann, C.; Fauler, B.; Uhlemann, Y.; Weiss, D.S.; Weinrauch, Y.; Zychlinsky, A. Neutrophil extracellular traps kill bacteria. Science 2004, 303, 1532–1535. [Google Scholar] [CrossRef]
- Urban, C.F.; Ermert, D.; Schmid, M.; Abu-Abed, U.; Goosmann, C.; Nacken, W.; Brinkmann, V.; Jungblut, P.R.; Zychlinsky, A. Neutrophil extracellular traps contain calprotectin, a cytosolic protein complex involved in host defense against candida albicans. PLoS Pathog. 2009, 5, e1000639. [Google Scholar] [CrossRef] [PubMed]
- Brinkmann, V.; Zychlinsky, A. Beneficial suicide: Why neutrophils die to make nets. Nat. Rev. Microbiol. 2007, 5, 577–582. [Google Scholar] [CrossRef]
- Saffarzadeh, M.; Juenemann, C.; Queisser, M.A.; Lochnit, G.; Barreto, G.; Galuska, S.P.; Lohmeyer, J.; Preissner, K.T. Neutrophil extracellular traps directly induce epithelial and endothelial cell death: A predominant role of histones. PLoS ONE 2012, 7, e32366. [Google Scholar] [CrossRef]
- Xu, J.; Zhang, X.; Pelayo, R.; Monestier, M.; Ammollo, C.T.; Semeraro, F.; Taylor, F.B.; Esmon, N.L.; Lupu, F.; Esmon, C.T. Extracellular histones are major mediators of death in sepsis. Nat. Med. 2009, 15, 1318–1321. [Google Scholar] [CrossRef]
- Kawasaki, H.; Iwamuro, S. Potential roles of histones in host defense as antimicrobial agents. Infect. Disord. Drug Targets 2008, 8, 195–205. [Google Scholar] [CrossRef]
- Hoeksema, M.; van Eijk, M.; Haagsman, H.P.; Hartshorn, K.L. Histones as mediators of host defense, inflammation and thrombosis. Future Microbiol. 2016, 11, 441–453. [Google Scholar] [CrossRef] [PubMed]
- Varju, I.; Longstaff, C.; Szabo, L.; Farkas, A.Z.; Varga-Szabo, V.J.; Tanka-Salamon, A.; Machovich, R.; Kolev, K. DNA, histones and neutrophil extracellular traps exert anti-fibrinolytic effects in a plasma environment. Thromb. Haemost. 2015, 113, 1289–1298. [Google Scholar] [CrossRef]
- Fuchs, T.A.; Bhandari, A.A.; Wagner, D.D. Histones induce rapid and profound thrombocytopenia in mice. Blood 2011, 118, 3708–3714. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Qi, H.; Kan, K.; Quan, C.; Xie, H.; Guo, X.; Zhang, L. Neutrophil extracellular traps promote hypercoagulability in patients with sepsis. Shock 2017, 47, 132–139. [Google Scholar] [CrossRef] [PubMed]
- Wright, T.K.; Gibson, P.G.; Simpson, J.L.; McDonald, V.M.; Wood, L.G.; Baines, K.J. Neutrophil extracellular traps are associated with inflammation in chronic airway disease. Respirology 2016, 21, 467–475. [Google Scholar] [CrossRef] [PubMed]
- Sorensen, O.E.; Borregaard, N. Neutrophil extracellular traps—The dark side of neutrophils. J. Clin. Investig. 2016, 126, 1612–1620. [Google Scholar] [CrossRef] [PubMed]
- Porto, B.N.; Stein, R.T. Neutrophil extracellular traps in pulmonary diseases: Too much of a good thing? Front. Immunol. 2016, 7, 311. [Google Scholar] [CrossRef]
- Nakazawa, D.; Kumar, S.; Desai, J.; Anders, H.J. Neutrophil extracellular traps in tissue pathology. Histol. Histopathol. 2016, 32, 203–2013. [Google Scholar]
- Hasler, P.; Giaglis, S.; Hahn, S. Neutrophil extracellular traps in health and disease. Swiss Med. Wkl. 2016, 146, w14352. [Google Scholar] [CrossRef] [PubMed]
- Boeltz, S.; Amini, P.; Anders, H.-J.; Andrade, F.; Bilyy, R.; Chatfield, S.; Cichon, I.; Clancy, D.M.; Desai, J.; Dumych, T.; et al. To net or not to net:Current opinions and state of the science regarding the formation of neutrophil extracellular traps. Cell Death Differ. 2019, 26, 395–408. [Google Scholar] [CrossRef]
- Zlatina, K.; Saftenberger, M.; Kuhnle, A.; Galuska, C.E.; Gartner, U.; Rebl, A.; Oster, M.; Vernunft, A.; Galuska, S.P. Polysialic acid in human plasma can compensate the cytotoxicity of histones. Int. J. Mol. Sci. 2018, 19, 1679. [Google Scholar] [CrossRef]
- Zlatina, K.; Lutteke, T.; Galuska, S.P. Individual impact of distinct polysialic acid chain lengths on the cytotoxicity of histone h1, h2a, h2b, h3 and h4. Polymers 2017, 9, 720. [Google Scholar] [CrossRef]
- Galuska, S.P.; Galuska, C.E.; Tharmalingam, T.; Zlatina, K.; Prem, G.; Husejnov, F.C.O.; Rudd, P.M.; Vann, W.F.; Reid, C.; Vionnet, J.; et al. In vitro generation of polysialylated cervical mucins by bacterial polysialyltransferases to counteract cytotoxicity of extracellular histones. FEBS J. 2017, 284, 1688–1699. [Google Scholar] [CrossRef] [PubMed]
- Galuska, C.E.; Dambon, J.A.; Kuhnle, A.; Bornhofft, K.F.; Prem, G.; Zlatina, K.; Lutteke, T.; Galuska, S.P. Artificial polysialic acid chains as sialidase-resistant molecular-anchors to accumulate particles on neutrophil extracellular traps. Front. Immunol. 2017, 8, 1229. [Google Scholar] [CrossRef]
- Ulm, C.; Saffarzadeh, M.; Mahavadi, P.; Müller, S.; Prem, G.; Saboor, F.; Simon, P.; Middendorff, R.; Geyer, H.; Henneke, I.; et al. Soluble polysialylated ncam: A novel player of the innate immune system in the lung. Cell. Mol. Life Sci. 2013, 70, 3695–3708. [Google Scholar] [CrossRef]
- Mishra, B.; von der Ohe, M.; Schulze, C.; Bian, S.; Makhina, T.; Loers, G.; Kleene, R.; Schachner, M. Functional role of the interaction between polysialic acid and extracellular histone h1. J. Neurosci. 2010, 30, 12400–12413. [Google Scholar] [CrossRef] [PubMed]
- Galuska, C.E.; Lutteke, T.; Galuska, S.P. Is polysialylated ncam not only a regulator during brain development but also during the formation of other organs? Biology 2017, 6, 27. [Google Scholar] [CrossRef]
- Finne, J. Occurrence of unique polysialosyl carbohydrate units in glycoproteins of developing brain. J. Biol. Chem. 1982, 257, 1966–1970. [Google Scholar]
- Galuska, S.P.; Geyer, R.; Gerardy-Schahn, R.; Mühlenhoff, M.; Geyer, H. Enzyme-dependent variations in the polysialylation of the neural cell adhesion molecule (ncam) in vivo. J. Biol. Chem. 2008, 283, 17–28. [Google Scholar] [CrossRef] [PubMed]
- Galuska, S.P.; Oltmann-Norden, I.; Geyer, H.; Weinhold, B.; Kuchelmeister, K.; Hildebrandt, H.; Gerardy-Schahn, R.; Geyer, R.; Mühlenhoff, M. Polysialic acid profiles of mice expressing variant allelic combinations of the polysialyltransferases st8siaii and st8siaiv. J. Biol. Chem. 2006, 281, 31605–31615. [Google Scholar] [CrossRef]
- Simon, P.; Bäumner, S.; Busch, O.; Röhrich, R.; Kaese, M.; Richterich, P.; Wehrend, A.; Müller, K.; Gerardy-Schahn, R.; Mühlenhoff, M.; et al. Polysialic acid is present in mammalian semen as a post-translational modification of the neural cell adhesion molecule ncam and the polysialyltransferase st8siaii. J. Biol. Chem. 2013, 288, 18825–18833. [Google Scholar] [CrossRef]
- Yabe, U.; Sato, C.; Matsuda, T.; Kitajima, K. Polysialic acid in human milk. Cd36 is a new member of mammalian polysialic acid-containing glycoprotein. J. Biol. Chem. 2003, 278, 13875–13880. [Google Scholar] [CrossRef] [PubMed]
- Kühnle, A.; Veelken, R.; Galuska, C.E.; Saftenberger, M.; Verleih, M.; Schuppe, H.C.; Rudloff, S.; Kunz, C.; Galuska, S.P. Polysialic acid interacts with lactoferrin and supports its activity to inhibit the release of neutrophil extracellular traps. Carbohydr. Polym. 2019, 208, 32–41. [Google Scholar] [CrossRef] [PubMed]
- Arnljots, K.; Sorensen, O.; Lollike, K.; Borregaard, N. Timing, targeting and sorting of azurophil granule proteins in human myeloid cells. Leukemia 1998, 12, 1789–1795. [Google Scholar] [CrossRef] [PubMed]
- Nazir, S.; Nasir, M.; Yasmeen, A.; Usman, S. Review study on lactoferrin: A multifunctional protein. Sky J. Food Sci. 2017, 6, 014–020. [Google Scholar]
- Vogel, H.J. Lactoferrin, a bird’s eye view. Biochem. Cell Biol. 2012, 90, 233–244. [Google Scholar] [CrossRef] [PubMed]
- Legrand, D. Overview of lactoferrin as a natural immune modulator. J. Pediatr. 2016, 173, S10–S15. [Google Scholar] [CrossRef] [PubMed]
- Inoue, M.; Yamada, J.; Kitamura, N.; Shimazaki, K.; Andren, A.; Yamashita, T. Immunohistochemical localization of lactoferrin in bovine exocrine glands. Tissue Cell 1993, 25, 791–797. [Google Scholar] [CrossRef]
- Steijns, J.M.; van Hooijdonk, A.C.M. Occurrence, structure, biochemical properties and technological characteristics of lactoferrin. Br. J. Nutr. 2000, 84, S11–S17. [Google Scholar] [CrossRef]
- Levay, P.F.; Viljoen, M. Lactoferrin—A general-review. Haematologica 1995, 80, 252–267. [Google Scholar] [PubMed]
- Bullen, J.J.; Rogers, H.J.; Leigh, L. Iron-binding proteins in milk and resistance to escherichia-coli infection in infants. BMJ Br. Med. J. 1972, 1, 69–75. [Google Scholar] [CrossRef]
- Sherman, M.P.; Adamkin, D.H.; Radmacher, P.G.; Sherman, J.; Niklas, V. Protective proteins in mammalian milks: Lactoferrin steps forward. NeoReviews 2012, 13, e293–e301. [Google Scholar] [CrossRef]
- Okubo, K.; Kamiya, M.; Urano, Y.; Nishi, H.; Herter, J.M.; Mayadas, T.; Hirohama, D.; Suzuki, K.; Kawakami, H.; Tanaka, M.; et al. Lactoferrin suppresses neutrophil extracellular traps release in inflammation. EBioMedicine 2016, 10, 204–215. [Google Scholar] [CrossRef] [PubMed]
- Schuberth, H.-J.; Krueger, C.; Hendricks, A.; Bimczok, D.; Leibold, W. Superantigen-dependent accelerated death of bovine neutrophilic granulocytes in vitro is mediated by blood mononuclear cells. Immunobiology 2000, 202, 493–507. [Google Scholar] [CrossRef]
- Kim, R.; Yokota, H.; Kim, S.H. Electrophoresis of proteins and protein-protein complexes in a native agarose gel. Anal. Biochem. 2000, 282, 147–149. [Google Scholar] [CrossRef] [PubMed]
- Hensley, K. Detection of protein carbonyls by means of biotin hydrazide-streptavidin affinity methods. Methods Mol. Biol. 2009, 536, 457–462. [Google Scholar] [PubMed]
- Kang, C.D.; Cao, C.; Lee, J.; Choi, I.S.; Kim, B.W.; Sim, S.J. Surface plasmon resonance-based inhibition assay for real-time detection of cryptosporidium parvum oocyst. Water Res. 2008, 42, 1693–1699. [Google Scholar] [CrossRef] [PubMed]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef] [PubMed]
- Duan, Y.; Wu, C.; Chowdhury, S.; Lee, M.C.; Xiong, G.; Zhang, W.; Yang, R.; Cieplak, P.; Luo, R.; Lee, T.; et al. A point-charge force field for molecular mechanics simulations of proteins based on condensed-phase quantum mechanical calculations. J. Comput. Chem. 2003, 24, 1999–2012. [Google Scholar] [CrossRef]
- Waterhouse, A.M.; Procter, J.B.; Martin, D.M.; Clamp, M.; Barton, G.J. Jalview version 2—A multiple sequence alignment editor and analysis workbench. Bioinformatics 2009, 25, 1189–1191. [Google Scholar] [CrossRef] [PubMed]
- Van Berkel, P.H.; Geerts, M.E.; van Veen, H.A.; Mericskay, M.; de Boer, H.A.; Nuijens, J.H. N-terminal stretch arg2, arg3, arg4 and arg5 of human lactoferrin is essential for binding to heparin, bacterial lipopolysaccharide, human lysozyme and DNA. Biochem. J. 1997, 328, 145–151. [Google Scholar] [CrossRef]
- Bellamy, W.; Takase, M.; Yamauchi, K.; Wakabayashi, H.; Kawase, K.; Tomita, M. Identification of the bactericidal domain of lactoferrin. Biochim. Biophys. Acta 1992, 1121, 130–136. [Google Scholar] [CrossRef]
- ElassRochard, E.; Roseanu, A.; Legrand, D.; Trif, M.; Salmon, V.; Motas, C.; Montreuil, J.; Spik, G. Lactoferrin-lipopolysaccharide interaction: Involvement of the 28-34 loop region of human lactoferrin in the high-affinity binding to escherichia coli 055b5 lipopolysaccharide. Biochem. J. 1995, 312, 839–845. [Google Scholar] [CrossRef]
- Wang, B.; Timilsena, Y.P.; Blanch, E.; Adhikari, B. Lactoferrin: Structure, function, denaturation and digestion. Crit. Rev. Food Sci. Nutr. 2017, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Lonnerdal, B. Nutritional roles of lactoferrin. Curr. Opin. Clin. Nutr. Metab. Care 2009, 12, 293–297. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kühnle, A.; Lütteke, T.; Bornhöfft, K.F.; Galuska, S.P. Polysialic Acid Modulates the Binding of External Lactoferrin in Neutrophil Extracellular Traps. Biology 2019, 8, 20. https://doi.org/10.3390/biology8020020
Kühnle A, Lütteke T, Bornhöfft KF, Galuska SP. Polysialic Acid Modulates the Binding of External Lactoferrin in Neutrophil Extracellular Traps. Biology. 2019; 8(2):20. https://doi.org/10.3390/biology8020020
Chicago/Turabian StyleKühnle, Andrea, Thomas Lütteke, Kim F. Bornhöfft, and Sebastian P. Galuska. 2019. "Polysialic Acid Modulates the Binding of External Lactoferrin in Neutrophil Extracellular Traps" Biology 8, no. 2: 20. https://doi.org/10.3390/biology8020020
APA StyleKühnle, A., Lütteke, T., Bornhöfft, K. F., & Galuska, S. P. (2019). Polysialic Acid Modulates the Binding of External Lactoferrin in Neutrophil Extracellular Traps. Biology, 8(2), 20. https://doi.org/10.3390/biology8020020




