Optimized Extraction by Response Surface Methodology Used for the Characterization and Quantification of Phenolic Compounds in Whole Red Grapes (Vitis vinifera)
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. Chemicals and Reagents
2.3. Polyphenol Extraction
2.4. Single-Factor Studies
2.5. Response Surface Design
2.6. Kinetic Study
2.7. Sequential Extractions
2.8. Application of the Method
2.9. Analysis of Response Variables
2.9.1. Total Phenolic Content
2.9.2. Total Anthocyanin Content
2.9.3. Total Flavanol Content
2.9.4. HPLC-DAD Analysis of Phenolic Compounds
2.9.5. HPLC-DAD Method Validation
2.10. Statistics
3. Results and Discussion
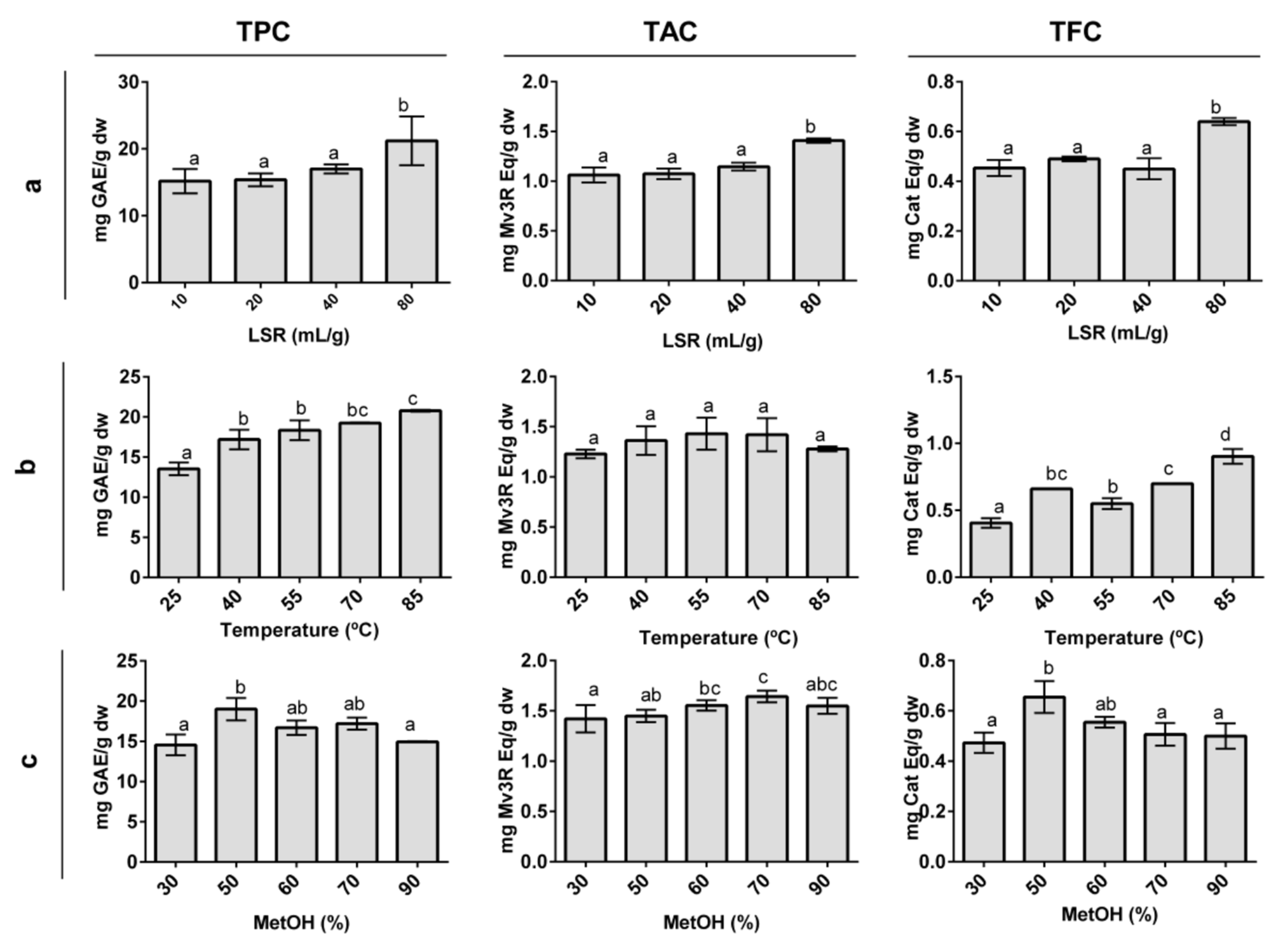
3.1. Single-Factor Studies
3.1.1. Effect of Liquid-to-Solid Ratio
3.1.2. Effect of Temperature
3.1.3. Effect of Methanol Proportion
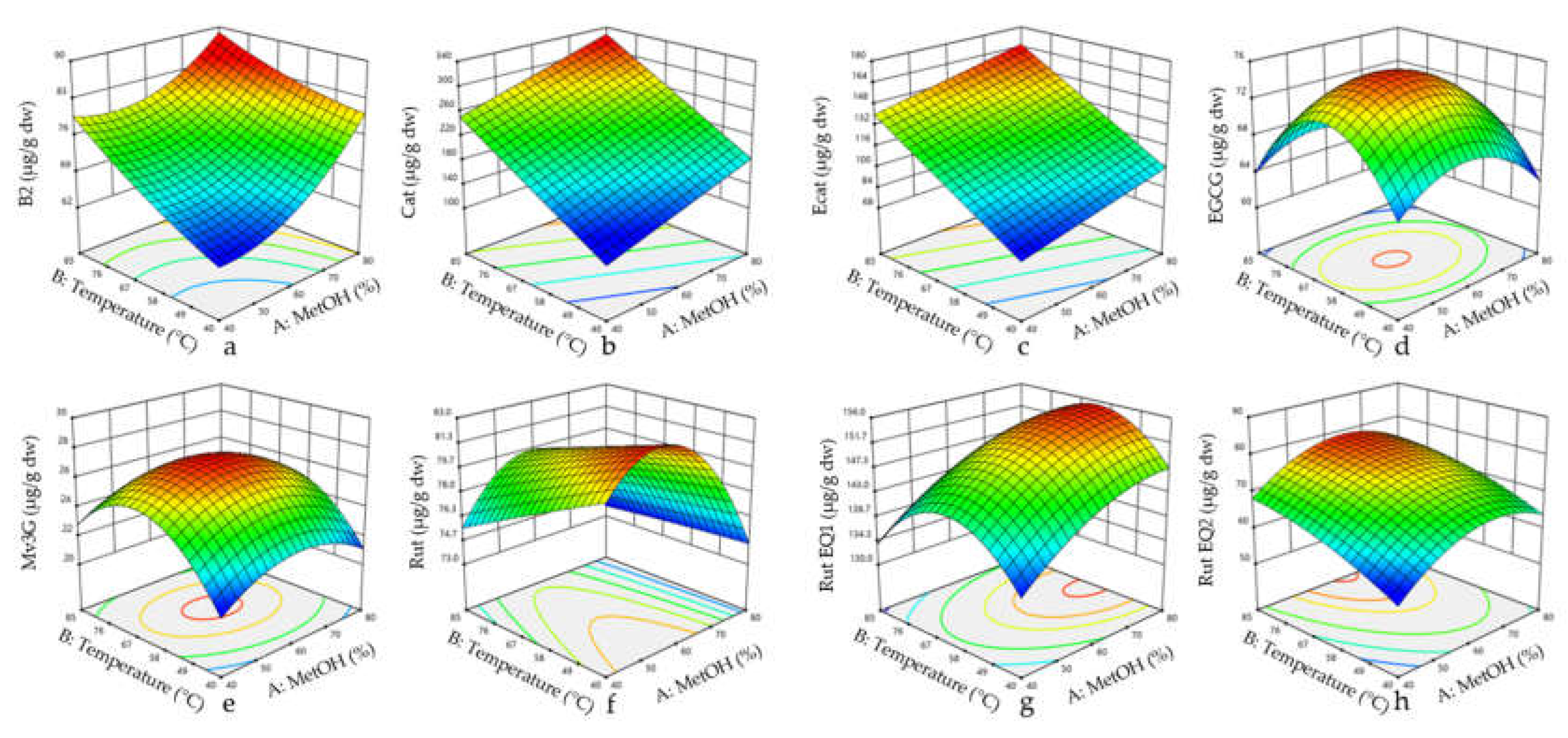
3.2. Surface Response Results
3.2.1. Fitting the Model
3.2.2. Combined Effect of Temperature and Methanol
3.2.3. Method Validation
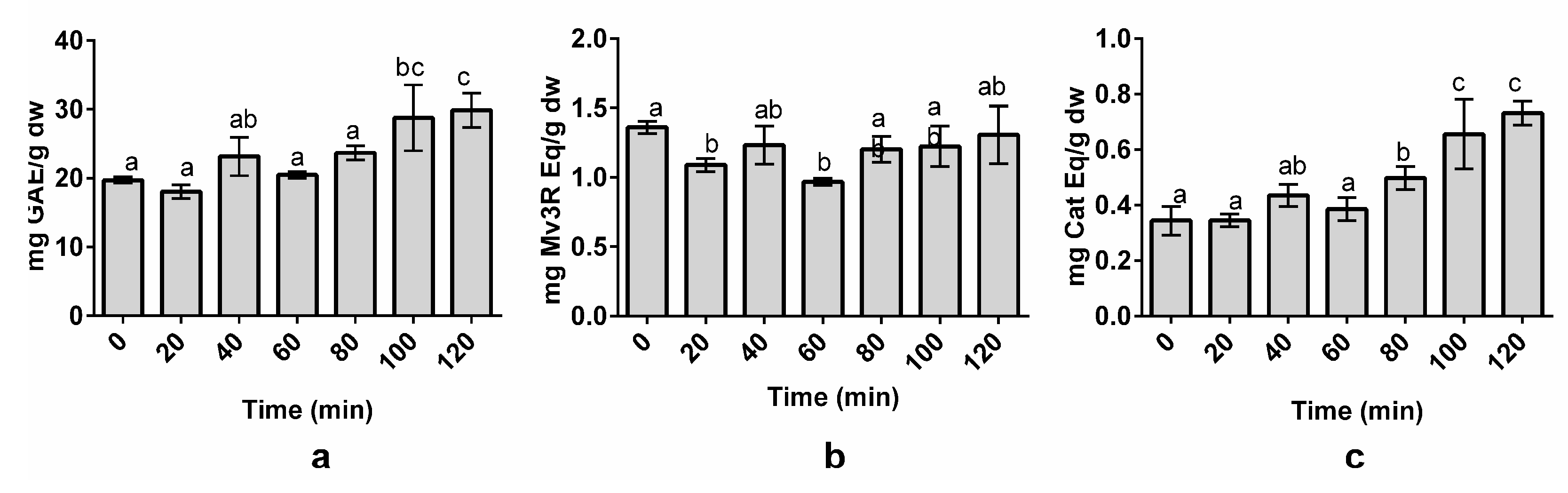
3.3. Effect of Time on Polyphenol Extraction
3.4. Multi-Step Extractions
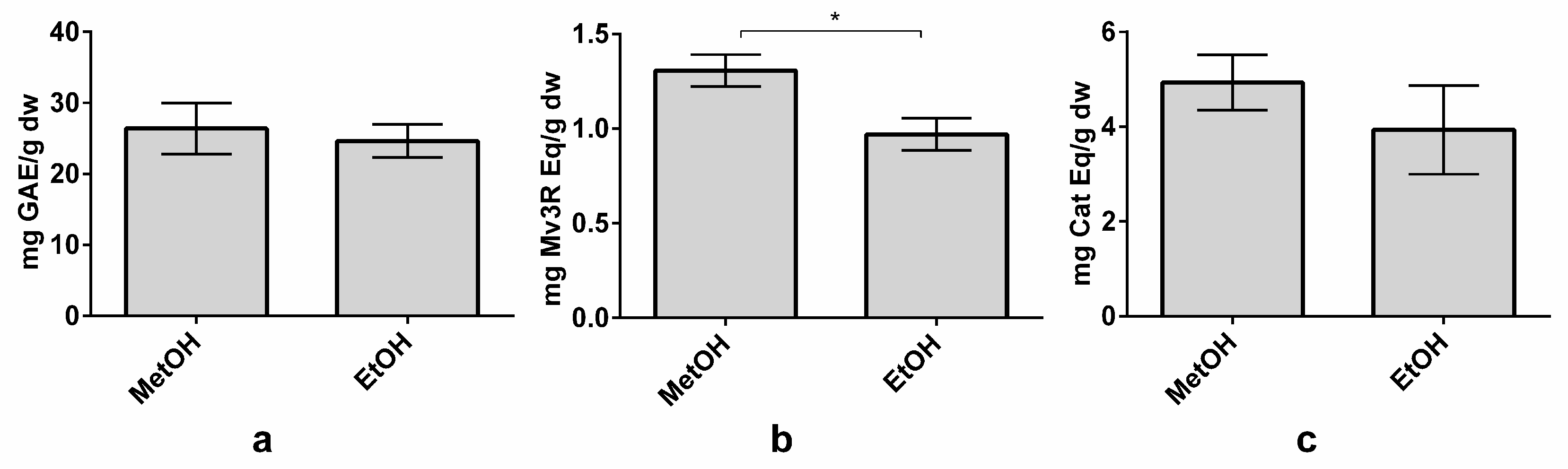
3.5. Methanol–Ethanol Comparison
3.6. Full Characterization of Whole Red Grapes
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Crozier, A.; Jaganath, I.B.; Clifford, M. Dietary phenolics: Chemistry, bioavailability and effects on health. Nat. Prod. Rep. 2009, 26, 1001–1043. [Google Scholar] [CrossRef] [PubMed]
- Garrido, J.; Borges, F. Wine and grape polyphenols—A chemical perspective. Food Res. Int. 2013, 54, 1844–1858. [Google Scholar] [CrossRef]
- Ammerer, D.I.K.; Laus, A.C.C.; Arle, R.E.C.; Chieber, A.N.S. Polyphenol Screening of Pomace from Red and White Grape Varieties (Vitis vinifera L.) by HPLC-DAD-MS/MS. J. Agric. Food Chem. 2004, 52, 27–30. [Google Scholar] [CrossRef]
- Te, U. Comparative Study of the Phenolic Composition of Seeds and Skins from Carménère and Cabernet Souvignon Grape Varieties (Vitis vinifera L.) during Ripening. J. Agric. Food Chem. 2010, 58, 3591–3599. [Google Scholar] [CrossRef]
- Di Lecce, G.; Arranz, S.; Jáuregui, O.; Tresserra-Rimbau, A.; Quifer-Rada, P.; Lamuela-Raventós, R.M. Phenolic profiling of the skin, pulp and seeds of Albariño grapes using hybrid quadrupole time-of-flight and triple-quadrupole mass spectrometry. Food Chem. 2014, 145, 874–882. [Google Scholar] [CrossRef]
- Farhadi, K.; Esmaeilzadeh, F.; Hatami, M.; Forough, M.; Molaie, R. Determination of phenolic compounds content and antioxidant activity in skin, pulp, seed, cane and leaf of five native grape cultivars in West Azerbaijan province, Iran. Food Chem. 2016, 199, 847–855. [Google Scholar] [CrossRef]
- Miller, S.; White, J.A.; Chowdhury, R.; Gales, D.N.; Tameru, B.; Tiwari, A.K.; Samuel, T. Effects of consumption of whole grape powder on basal NF-k B signaling and inflammatory cytokine secretion in a mouse model of inflammation. J. Nutr. Intermed. MeTable 2018, 11, 1–8. [Google Scholar] [CrossRef]
- Rodrigues, A.D.; Scheffel, T.B.; Scola, G.; Dos Santos, M.T.; Fank, B.; De Freitas, S.C.V.; Dani, C.; Vanderlinde, R.; Henriques, J.A.P.; Coitinho, A.S.; et al. Neuroprotective and anticonvulsant effects of organic and conventional purple grape juices on seizures in Wistar rats induced by pentylenetetrazole. Neurochem. Int. 2012, 60, 799–805. [Google Scholar] [CrossRef]
- Akçay, Y.D.; Yildirim, H.K.; Güvenç, U.; Sözmen, E.Y. The effects of consumption of organic and nonorganic red wine on low-density lipoprotein oxidation and antioxidant capacity in humans. Nutr. Res. 2004, 24, 541–554. [Google Scholar] [CrossRef]
- Bladé, C.; Aragonès, G.; Arola-Arnal, A.; Muguerza, B.; Bravo, F.I.; Salvadó, M.J.; Arola, L.; Suárez, M. Proanthocyanidins in health and disease. BioFactors 2016, 42, 5–12. [Google Scholar] [CrossRef]
- Wrolstad, R.E. Color and Pigment Analyses in Fruit Products; Agricultural Experiment Station, Oregon State University: Corvallis, OR, USA, 1993; pp. 1–17. [Google Scholar]
- Arti, B.; Rakesh, M.; Pushpa, L.; Guru, S.K.; Sandeep, A. Role of secondary metabolites and brassinosteroids in plant defense against environmental stresses. J. Plant Growth Regul. 2013, 32, 216–232. [Google Scholar] [CrossRef]
- Winkel-Shirley, B. Biosynthesis of flavonoids and effects of stress. Curr. Opin. Plant Biol. 2002, 5, 218–223. [Google Scholar] [CrossRef]
- Arbera, Ä.S. Varietal Differences among the Polyphenol Profiles of Seven Table Grape Cultivars Studied by LC−DAD−MS−MS. J. Agric. Food Chem. 2002, 50, 5691–5696. [Google Scholar] [CrossRef]
- Mulero, J.; Pardo, F.; Zafrilla, P. Antioxidant activity and phenolic composition of organic and conventional grapes and wines. J. Food Compos. Anal. 2010, 23, 569–574. [Google Scholar] [CrossRef]
- Alonso, R.; Berli, F.J.; Fontana, A.; Piccoli, P.; Bottini, R. Malbec grape (Vitis vinifera L.) responses to the environment: Berry phenolics as influenced by solar UV-B, water deficit and sprayed abscisic acid. Plant Physiol. Biochem. 2016, 109, 84–90. [Google Scholar] [CrossRef]
- Iacopini, P.; Baldi, M.; Storchi, P.; Sebastiani, L. Catechin, epicatechin, quercetin, rutin and resveratrol in red grape: Content, in vitro antioxidant activity and interactions. J. Food Compos. Anal. 2008, 21, 589–598. [Google Scholar] [CrossRef]
- Pompeu, D.R.; Silva, E.M.; Rogez, H. Optimisation of the solvent extraction of phenolic antioxidants from fruits of Euterpe oleracea using Response Surface Methodology. Bioresour. Technol. 2009, 100, 6076–6082. [Google Scholar] [CrossRef]
- Karacabey, E.; Mazza, G. Optimization of solid-liquid extraction of resveratrol and other phenolic compounds from milled grape canes (Vitis vinifera). J. Agric. Food Chem. 2008, 56, 6318–6325. [Google Scholar] [CrossRef]
- Karacabey, E.; Mazza, G. Optimisation of antioxidant activity of grape cane extracts using response surface methodology. Food Chem. 2010, 119, 343–348. [Google Scholar] [CrossRef]
- Pinelo, M.; Rubilar, M.; Jerez, M.; Sineiro, J.; Núñez, M.J. Effect of solvent, temperature, and solvent-to-solid ratio on the total phenolic content and antiradical activity of extracts from different components of grape pomace. J. Agric. Food Chem. 2005, 53, 2111–2117. [Google Scholar] [CrossRef]
- Amendola, D.; De Faveri, D.M.; Spigno, G. Grape marc phenolics: Extraction kinetics, quality and stability of extracts. J. Food Eng. 2010, 97, 384–392. [Google Scholar] [CrossRef]
- Sant’Anna, V.; Brandelli, A.; Marczak, L.D.F.; Tessaro, I.C. Kinetic modeling of total polyphenol extraction from grape marc and characterization of the extracts. Sep. Purif. Technol. 2012, 100, 82–87. [Google Scholar] [CrossRef]
- Casazza, A.A.; Aliakbarian, B.; Mantegna, S.; Cravotto, G.; Perego, P. Extraction of phenolics from Vitis vinifera wastes using non-conventional techniques. J. Food Eng. 2010, 100, 50–55. [Google Scholar] [CrossRef]
- Silva, E.M.; Rogez, H.; Larondelle, Y. Optimization of extraction of phenolics from Inga edulisleaves using response surface methodology. Sep. Purif. Technol. 2007, 55, 381–387. [Google Scholar] [CrossRef]
- Prasad, K.N.; Hassan, F.A.; Yang, B.; Kong, K.W.; Ramanan, R.N.; Azlan, A.; Ismail, A. Response surface optimisation for the extraction of phenolic compounds and antioxidant capacities of underutilised Mangifera pajang Kosterm. peels. Food Chem. 2011, 128, 1121–1127. [Google Scholar] [CrossRef]
- Yang, L.; Jiang, J.G.; Li, W.F.; Chen, J.; Wang, D.Y.; Zhu, L. Optimum extraction process of polyphenols from the bark of Phyllanthus emblica L. based on the response surface methodology. J. Sep. Sci. 2009, 32, 1437–1444. [Google Scholar] [CrossRef]
- Canals, R.; Llaudy, M.C.; Valls, J.; Canals, J.M.; Zamora, F. Influence of ethanol concentration on the extraction of color and phenolic compounds from the skin and seeds of tempranillo grapes at different stages of ripening. J. Agric. Food Chem. 2005, 53, 4019–4025. [Google Scholar] [CrossRef]
- Gómez-Alonso, S.; García-Romero, E.; Hermosín-Gutiérrez, I. HPLC analysis of diverse grape and wine phenolics using direct injection and multidetection by DAD and fluorescence. J. Food Compos. Anal. 2007, 20, 618–626. [Google Scholar] [CrossRef]
- Mané, C.; Souquet, J.M.; Ollé, D.; Verriés, C.; Véran, F.; Mazerolles, G.; Cheynier, V.; Fulcrand, H. Optimization of simultaneous flavanol, phenolic acid, and anthocyanin extraction from grapes using an experimental design: Application to the characterization of champagne grape varieties. J. Agric. Food Chem. 2007, 55, 7224–7233. [Google Scholar] [CrossRef]
- Teixeira, A.; Eiras-Dias, J.; Castellarin, S.D.; Gerós, H. Berry phenolics of grapevine under challenging environments. Int. J. Mol. Sci. 2013, 14, 18711–18739. [Google Scholar] [CrossRef]
- Martínez-Gil, A.M.; Gutiérrez-Gamboa, G.; Garde-Cerdán, T.; Pérez-Álvarez, E.P.; Moreno-Simunovic, Y. Characterization of phenolic composition in Carignan noir grapes (Vitis vinifera L.) from six wine-growing sites in Maule Valley, Chile. J. Sci. Food Agric. 2018, 98, 274–282. [Google Scholar] [CrossRef] [PubMed]
- Lingua, M.S.; Fabani, M.P.; Wunderlin, D.A.; Baroni, M.V. In vivo antioxidant activity of grape, pomace and wine from three red varieties grown in Argentina: Its relationship to phenolic profile. J. Funct. Foods 2016, 20, 332–345. [Google Scholar] [CrossRef]
- Clifford, M.N. Chlorogenic acids and other cinnamates—Nature, occurence and dietary burden. J. Sci. Food Agric. 1999, 79, 362–372. [Google Scholar] [CrossRef]
- Nenadis, N.; Boyle, S.; Bakalbassis, E.G.; Tsimidou, M. An experimental approach to structure-activity relationships of caffeic and dihydrocaffeic acids and related monophenols. JAOCS J. Am. Oil Chem. Soc. 2003, 80, 451–458. [Google Scholar] [CrossRef]
- Karvela, E.; Makris, D.P.; Kalogeropoulos, N.; Karathanos, V.T.; Kefalas, P. Factorial design optimisation of grape (Vitis vinifera) seed polyphenol extraction. Eur. Food Res. Technol. 2009, 229, 731–742. [Google Scholar] [CrossRef]
- Spigno, G.; De Faveri, D.M. Antioxidants from grape stalks and marc: Influence of extraction procedure on yield, purity and antioxidant power of the extracts. J. Food Eng. 2007, 78, 793–801. [Google Scholar] [CrossRef]
- Arnous, A.; Makris, D.P.; Kefalas, P. Correlation of pigment and flavanol content with antioxidant properties in selected aged regional wines from Greece. J. Food Compos. Anal. 2002, 15, 655–665. [Google Scholar] [CrossRef]
- Bucić-Kojić, A.; Planinić, M.; Tomas, S.; Bilić, M.; Velić, D. Study of solid-liquid extraction kinetics of total polyphenols from grape seeds. J. Food Eng. 2007, 81, 236–242. [Google Scholar] [CrossRef]
- Cacace, J.E.; Mazza, G. Mass transfer process during extraction of phenolic compounds from milled berries. J. Food Eng. 2003, 59, 379–389. [Google Scholar] [CrossRef]
- Galili, S.; Hovav, R. Determination of Polyphenols, Flavonoids, and Antioxidant Capacity in Dry Seeds. In Polyphenols in Plants: Isolation, Purification and Extract Preparation; Academic Press, 2014; pp. 305–323. [Google Scholar]
- Tabaraki, R.; Nateghi, A. Optimization of ultrasonic-assisted extraction of natural antioxidants from rice bran using response surface methodology. Ultrason. Sonochem. 2011, 18, 1279–1286. [Google Scholar] [CrossRef]
- Li, W.; Wang, Z.; Wang, Y.P.; Jiang, C.; Liu, Q.; Sun, Y.S.; Zheng, Y.N. Pressurised liquid extraction combining LC-DAD-ESI/MS analysis as an alternative method to extract three major flavones in Citrus reticulata “Chachi” (Guangchenpi). Food Chem. 2012, 130, 1044–1049. [Google Scholar] [CrossRef]
- Mukhopadhyay, S.; Luthria, D.L.; Robbins, R.J. Optimization of extraction process for phenolic acids from black cohosh (Cimicifuga racemosa) by pressurized liquid extraction. J. Sci. Food Agric. 2006, 86, 156–162. [Google Scholar] [CrossRef]
- Borges, G.D.S.C.; Vieira, F.G.K.; Copetti, C.; Gonzaga, L.V.; Fett, R. Optimization of the extraction of flavanols and anthocyanins from the fruit pulp of Euterpe edulis using the response surface methodology. Food Res. Int. 2011, 44, 708–715. [Google Scholar] [CrossRef]
- Wijngaard, H.H.; Brunton, N. The optimisation of solid-liquid extraction of antioxidants from apple pomace by response surface methodology. J. Food Eng. 2010, 96, 134–140. [Google Scholar] [CrossRef]
- Yılmaz, F.M.; Karaaslan, M.; Vardin, H. Optimization of extraction parameters on the isolation of phenolic compounds from sour cherry (Prunus cerasus L.) pomace. J. Food Sci. Technol. 2015, 52, 2851–2859. [Google Scholar] [CrossRef] [PubMed]
- Metivier, R.P.; Francis, F.J.; Clydesdale, F.M. Solvent Extraction of Anthocyanins From Wine Pomace. J. Food Sci. 1980, 45, 1099–1100. [Google Scholar] [CrossRef]
- Kushwaha, R.; Karanjekar, S. Standardization of ashwagandharishta formulation by TLC method. Int. J. ChemTech Res. 2011, 3, 1033–1036. [Google Scholar] [CrossRef]
- Chassy, A.W.; Bui, L.; Renaud, E.N.C.; Van Horn, M.; Mitchell, A.E. Three-year comparison of the content of antioxidant microconstituents and several quality characteristics in organic vs conv. J. Agric. Food Chem. 2006, 54, 8244–8252. [Google Scholar] [CrossRef]

| Run 1 | MetOH (%) | T (°C) | B2 | Cat | Ecat | EGCG | Rut | Rut EQ1 | Rut EQ2 | Chl | Chl EQ1 | pCou | Rvt | Cy3R | Mv3G | Mv3G EQ1 | Mv3G EQ2 | Mv3G EQ3 | Mv3G EQ4 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 40 | 40 | 60.66 | 136.93 | 74.27 | 63.95 | 81.08 | 133.00 | 56.47 | n.d. | 825.48 | n.d. | n.d. | 135.47 | 1159.52 | 104.39 | 37.57 | 20.26 | 181.29 |
| 2 | 80 | 40 | 79.63 | 177.73 | 100.23 | 62.89 | 75.20 | 146.23 | 65.66 | n.d. | 796.71 | n.d. | n.d. | 151.31 | 1253.41 | 119.17 | 43.96 | 21.19 | 216.30 |
| 3 | 40 | 85 | 79.82 | 230.51 | 124.36 | 63.03 | 74.90 | 135.20 | 67.32 | n.d. | 760.54 | n.d. | n.d. | 144.71 | 1194.49 | 111.79 | 33.17 | 22.99 | 188.68 |
| 4 | 80 | 85 | 90.47 | 363.22 | 180.12 | 60.74 | 73.44 | 146.81 | 73.75 | n.d. | 925.75 | n.d. | n.d. | 139.57 | 1209.93 | 113.83 | 36.78 | 22.74 | 201.51 |
| 5 | 40 | 65 | 72.29 | 168.54 | 128.96 | 73.44 | 78.05 | 143.62 | 63.99 | n.d. | 853.20 | n.d. | n.d. | 142.10 | 1284.35 | 133.39 | 32.91 | 25.37 | 210.41 |
| 6 | 80 | 65 | 83.00 | 251.54 | 124.24 | 69.12 | 73.76 | 153.93 | 70.50 | n.d. | 838.44 | n.d. | n.d. | 133.90 | 1163.91 | 113.00 | 36.41 | 24.52 | 200.34 |
| 7 | 60 | 40 | 68.95 | 158.76 | 78.99 | 69.09 | 81.25 | 147.06 | 63.77 | n.d. | 860.99 | n.d. | n.d. | 145.19 | 1176.84 | 119.36 | 36.24 | 22.90 | 198.69 |
| 8 | 60 | 85 | 78.02 | 350.76 | 146.46 | 67.14 | 80.61 | 138.68 | 80.25 | n.d. | 826.50 | n.d. | n.d. | 141.28 | 1276.78 | 117.86 | 42.36 | 23.53 | 214.59 |
| 9 | 60 | 65 | 70.79 | 208.79 | 128.10 | 72.67 | 81.34 | 150.34 | 75.91 | n.d. | 743.96 | n.d. | n.d. | 134.92 | 1175.77 | 120.11 | 38.09 | 26.86 | 191.47 |
| 10 | 60 | 65 | 74.60 | 191.36 | 122.34 | 73.67 | 82.88 | 151.35 | 76.61 | n.d. | 758.96 | n.d. | n.d. | 140.03 | 1244.41 | 136.07 | 40.21 | 28.83 | 205.79 |
| 11 | 60 | 65 | 72.44 | 221.15 | 129.73 | 76.89 | 78.79 | 156.21 | 77.25 | n.d. | 799.85 | n.d. | n.d. | 132.07 | 1271.07 | 117.57 | 34.31 | 26.69 | 216.11 |
| Model Parameters | Regression Coefficient | B2 | Cat | Ecat | EGCG | Mv3G Eq3 | Rut | Rut Eq1 | Rut Eq2 |
|---|---|---|---|---|---|---|---|---|---|
| Intercept | β0 | 66.132 | −98.331 | −10.753 | −20.457 | −23.042 | 55.243 | 33.532 | −45.193 |
| Linear | |||||||||
| MetOH | β1 | −0.742 * | 1.912 | 0.642 | 1.293 | 0.624 # | 1.228 * | 1.307 * | 2.593 * |
| T | β2 | 0.285 * | 3.217 * | 1.475 * | 1.914 | 0.973 | −0.193 # | 2.314 | 0.926 * |
| Interaction | |||||||||
| MetOH × T | β12 | −4.775 × 10−3 | - | - | −7.870 × 10−4 | −6.990 × 10−4 | 2.422 × 10−3 | −9.810 × 10−4 | −1.579 × 10−3 |
| Quadratic | |||||||||
| MetOH × MetOH | β11 | 1.151 × 10−2 * | - | - | −1.089 × 10−2 * | −4.842 × 10−3 * | −1.232 × 10−2 * | −7.934 × 10−3 | −1.924 × 10−2 * |
| T × T | B22 | 2.328 × 10−3 | - | - | −1.523 × 10−2 * | −7.156 × 10−3 * | −1.250 × 10−4 | −1.837 × 10−2 * | −4.548 × 10−3 |
| R2 | 0.961 | 0.847 | 0.874 | 0.9456 | 0.9283 | 0.8935 | 0.9037 | 0.99553 | |
| Adjusted R2 | 0.922 | 0.809 | 0.843 | 0.8912 | 0.8567 | 0.7871 | 0.8074 | 0.9105 | |
| p-value | 0.002 | <0.001 | <0.001 | 0.0035 | 0.007 | 0.018 | 0.014 | 0.002 | |
| F-value | 24.63 | 22.12 | 27.82 | 17.38 | 12.95 | 8.39 | 9.38 | 21.36 | |
| Lack of fit | 0.442 | 0.173 | 0.080 | 0.776 | 0.742 | 0.821 | 0.495 | 0.056 |
| Extraction Variables 1 | Parameter | Predicted | Experimental | ||||
|---|---|---|---|---|---|---|---|
| T (°C) | MetOH (%) | Desirability | |||||
| 72 | 65 | 0.686 | B2 | 76.70 | 77.73 | ± | 1.17 |
| Cat | 257.57 | 264.44 | ± | 17.64 | |||
| Ecat | 137.16 | 153.03 | ± | 17.03 | |||
| EGCG | 72.74 | 73.71 | ± | 3.25 | |||
| Cy3R | 136.56 | 144.59 | ± | 1.73 | |||
| Mv3G | 1233.23 | 1363.78 | ± | 7.49 | |||
| Mv3G Eq1 | 118.77 | 131.95 | ± | 8.61 | |||
| Mv3G Eq2 | 37.64 | 38.23 | ± | 1.45 | |||
| Mv3G Eq3 | 26.72 | 26.33 | ± | 1.98 | |||
| Mv3G Eq4 | 202.23 | 219.43 | ± | 6.19 | |||
| Rut | 79.80 | 82.75 | ± | 1.05 | |||
| Rut Eq1 | 151.80 | 151.53 | ± | 1.47 | |||
| Rut Eq2 | 77.80 | 76.14 | ± | 3.37 | |||
| Chl Eq1 | 828.07 | 888.50 | ± | 4.73 | |||
| CG | OG | PG | |
|---|---|---|---|
| TPC 1 | 23.98 ± 2.60 | 26.37 ± 3.60 | 17.18 ± 1.87 * |
| TAC 1 | 0.98 ± 0.11 | 1.31 ± 0.08* | 0.71 ± 0.05 * |
| TFC 1 | 4.42 ± 0.92 | 4.93 ± 0.58 | 3.68 ± 0.20 |
| Cat | 519.51 ± 179.55 | 382.30 ± 90.06 | 255.82 ± 68.72 |
| B2 | 139.29 ± 43.04 | 95.88 ± 10.09 | 39.15 ± 4.14 * |
| EGCG | N.D. | 99.47 ± 6.45 | N.D. |
| Ecat | 276.45 ± 98.22 | 162.42 ± 46.17 | 241.48 ± 14.81 |
| Chl EQ1 | 797.05 | 853.16 ± 57.44 | 176.31 ± 6.69 * |
| Chl | N.D. | N.D. | N.D. |
| Rvt | N.D. | N.D. | N.D. |
| pCou | N.D. | N.D. | N.D. |
| Mv3G EQ1 | 34.38 ± 11.23 | 133.81 ± 10.29 * | N.D. |
| Cy3R | 79.57 ± 8.81 | 159.83 ± 6.20 * | 161.31 ± 11.97 * |
| Mv3G | 1116.80 ± 67.21 | 1534.89 125.26 * | 887.14 ± 57.14 * |
| Mv3G EQ2 | 36.95 ± 10.36 | 53.36 ± 15.01 | N.D. |
| Mv3G EQ3 | 69.58 ± 1.50 | 79.52 ± 10.42 | N.D. |
| Mv3G EQ4 | 198.09 ± 28.08 | 199.30 ± 31.98 | 42.39 ± 5.22* |
| Rut | 157.72 ± 20.31 | 117.13 ± 13.26 * | 60.81 ± 11.6 * |
| Rut EQ1 | 318.48 ± 63.43 | 172.48 ± 13.57 * | 116.17 ± 19.60 * |
| Rut EQ2 | 111.03 ± 21.27 | 82.49 ± 4.01 | N.D. |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Iglesias-Carres, L.; Mas-Capdevila, A.; Sancho-Pardo, L.; Bravo, F.I.; Mulero, M.; Muguerza, B.; Arola-Arnal, A. Optimized Extraction by Response Surface Methodology Used for the Characterization and Quantification of Phenolic Compounds in Whole Red Grapes (Vitis vinifera). Nutrients 2018, 10, 1931. https://doi.org/10.3390/nu10121931
Iglesias-Carres L, Mas-Capdevila A, Sancho-Pardo L, Bravo FI, Mulero M, Muguerza B, Arola-Arnal A. Optimized Extraction by Response Surface Methodology Used for the Characterization and Quantification of Phenolic Compounds in Whole Red Grapes (Vitis vinifera). Nutrients. 2018; 10(12):1931. https://doi.org/10.3390/nu10121931
Chicago/Turabian StyleIglesias-Carres, Lisard, Anna Mas-Capdevila, Lucía Sancho-Pardo, Francisca Isabel Bravo, Miquel Mulero, Begoña Muguerza, and Anna Arola-Arnal. 2018. "Optimized Extraction by Response Surface Methodology Used for the Characterization and Quantification of Phenolic Compounds in Whole Red Grapes (Vitis vinifera)" Nutrients 10, no. 12: 1931. https://doi.org/10.3390/nu10121931
APA StyleIglesias-Carres, L., Mas-Capdevila, A., Sancho-Pardo, L., Bravo, F. I., Mulero, M., Muguerza, B., & Arola-Arnal, A. (2018). Optimized Extraction by Response Surface Methodology Used for the Characterization and Quantification of Phenolic Compounds in Whole Red Grapes (Vitis vinifera). Nutrients, 10(12), 1931. https://doi.org/10.3390/nu10121931








