The RsRlpA Effector Is a Protease Inhibitor Promoting Rhizoctonia solani Virulence through Suppression of the Hypersensitive Response
Abstract
1. Introduction
2. Results
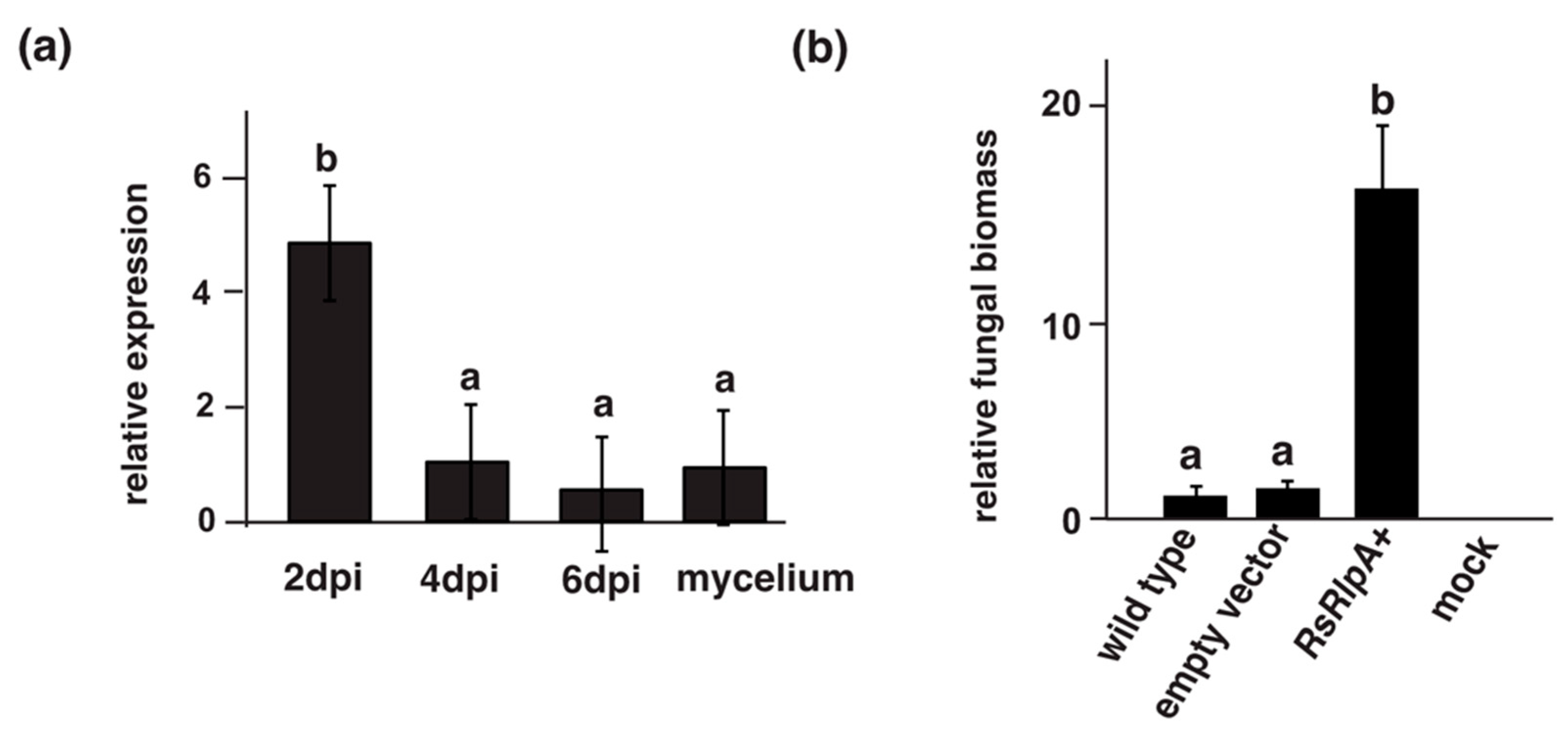
2.1. The RsRlpA Contains a DPBB Fold and Is Highly Induced upon Early Infection
2.2. Heterologous Expression of the RsRlpA Promotes Fungal Infection
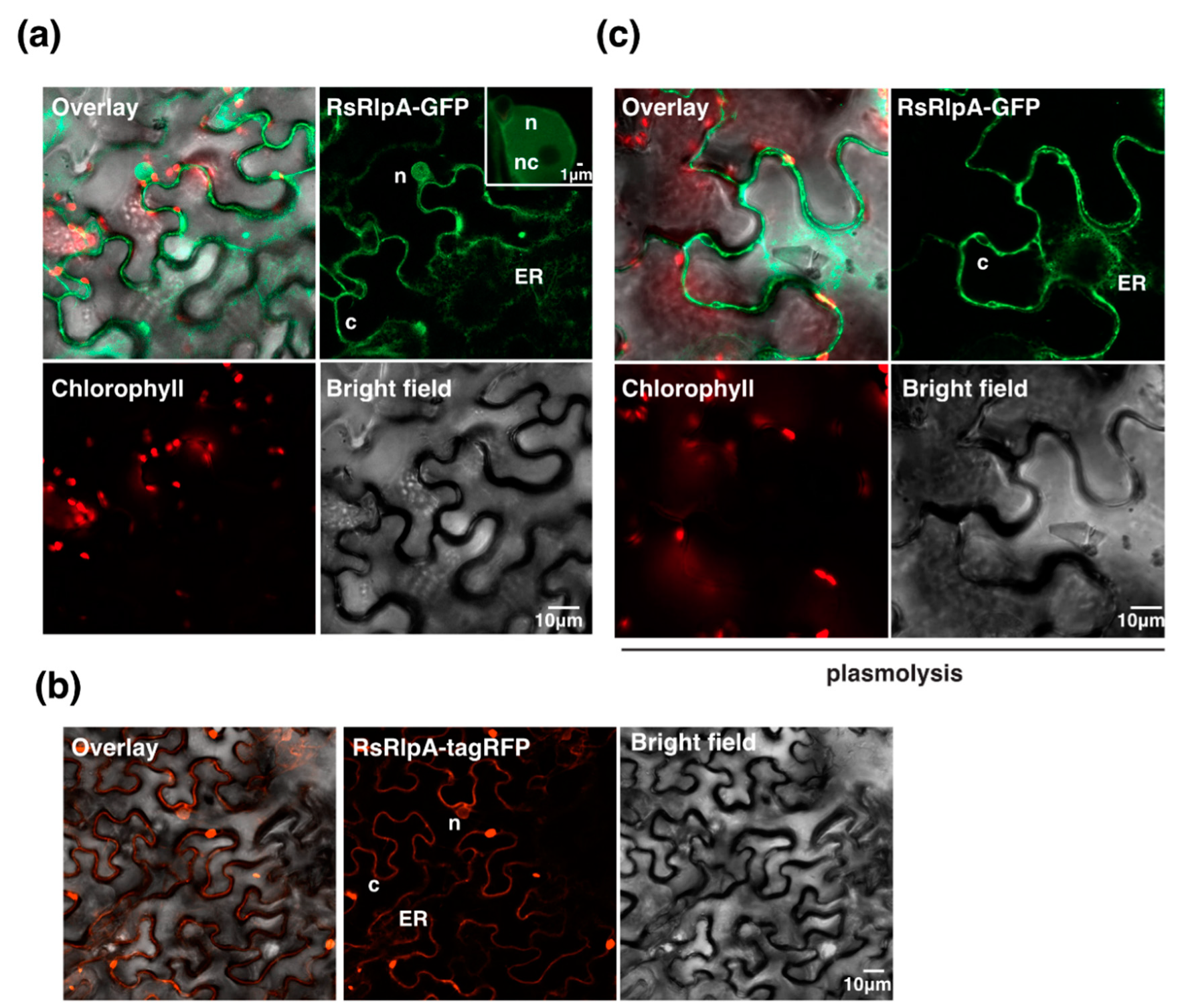
2.3. The RsRplA is Localized to the Cell Periphery and Suppresses Hypersensitive Response Induced by the Avr4/Cf4 Complex
2.4. The RsRlpA is an Active Protease Inhibitor Suppressing Oxidative Burst
2.5. The RsRplA Associates with a Plant Cathepsin
2.6. The Ser120 is Involved in RsRplA Ability to Suppress Cell Death
3. Discussion
4. Materials and Methods
4.1. Fungal Isolates and Growth Conditions
4.2. RNA Isolation and qRT-PCR Analysis
4.3. Construction of Cercospora Beticola Transgenic Strains and Infection Assays
4.4. Transgenic Arabidopsis Plants and Pathogenicity Assays
4.5. Effector Subcellular Localization and Hypersensitive Response Assays
4.6. Prediction of Protein 3D Structure, Production and Purification
4.7. Protease Inhibition and ROS Burst Suppression Assays
4.8. Pull-Down and Mass Spectrometry
4.9. Protein-Protein Interaction
4.10. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| AG | Anastomosis group |
| DAMP | Damage-associated molecular patterns |
| CAZy | Carbohydrate-Active enZYmes |
| DPBB | Double psi beta barrel |
| DPI | Days post inoculation |
| ETI | Effector-triggered immunity |
| HR | Hypersensitive response |
| PAMPs | Pathogen-associated molecular patterns |
| PLCP | Papain-like cysteine protease |
| PRR | Pattern recognition receptors |
| PTI | Pattern-triggered immunity |
| RlpA | Rare lipoprotein A |
| ROS | Reactive oxygen species |
References
- Chisholm, S.T.; Coaker, G.; Day, B.; Staskawicz, B.J. Host-microbe interactions: Shaping the evolution of the plant immune response. Cell 2006, 124, 803–814. [Google Scholar] [CrossRef] [PubMed]
- Jones, J.D.; Dangl, J.L. The plant immune system. Nature 2006, 444, 323–329. [Google Scholar] [CrossRef] [PubMed]
- Boller, T.; Felix, G. A renaissance of elicitors: Perception of microbe-associated molecular patterns and danger signals by pattern-recognition receptors. Annu. Rev. Plant Biol. 2009, 60, 379–406. [Google Scholar] [CrossRef] [PubMed]
- Giraldo, M.C.; Valent, B. Filamentous plant pathogen effectors in action. Nat. Rev. Microbiol. 2013, 11, 800–814. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Tyler, B.M.; Wang, Y. Defense and counter defense during plant-pathogenic oomycete infection. Annu. Rev. Microbiol. 2019, 8, 667–696. [Google Scholar] [CrossRef] [PubMed]
- Mukhopadhyay, A.N. Handbook on Diseases of Sugar Beet, 1st ed.; CRC Press: Boca Raton, FL, USA, 1987. [Google Scholar]
- Cubeta, M.A.; Vilgalys, R. Population Biology of the Rhizoctonia solani complex. Phytopathology 1997, 87, 480–484. [Google Scholar] [CrossRef]
- Gonzalez, M.; Pujol, M.; Metraux, J.P.; Gonzalez-Garcia, V.; Bolton, M.D.; Borras-Hidalgo, O. Tobacco leaf spot and root rot caused by Rhizoctonia solani Kuhn. Mol. Plant Pathol. 2011, 12, 209–216. [Google Scholar] [CrossRef]
- Sneh, B.; Burpee, L.; Ogoshi, A. Identification of Rhizoctonia Species; APS Press Inc.: St Paul, MI, USA, 1991. [Google Scholar]
- Bolton, M.D.; Panella, L.; Campbell, L.; Khan, M.F. Temperature, moisture, and fungicide effects in managing Rhizoctonia root and crown rot of sugar beet. Phytopathology 2010, 100, 689–697. [Google Scholar] [CrossRef]
- Zheng, A.; Lin, R.; Zhang, D.; Qin, P.; Xu, L.; Ai, P.; Ding, L.; Wang, Y.; Chen, Y.; Liu, Y.; et al. The evolution and pathogenic mechanisms of the rice sheath blight pathogen. Nat. Commun. 2013, 4, 1424. [Google Scholar] [CrossRef]
- Anderson, J.P.; Sperschneider, J.; Win, J.; Kidd, B.; Yoshida, K.; Hane, J.; Saunders, D.; Singh, K.B. Comparative secretome analysis of Rhizoctonia solani isolates with different host ranges reveals unique secretomes and cell death inducing effectors. Sci. Rep. 2017, 7, 10410. [Google Scholar] [CrossRef]
- Li, S.; Peng, X.; Wang, Y.; Hua, K.; Xing, F.; Zheng, Y.; Liu, W.; Sun, W.; Wei, S. The effector AGLIP1 in Rhizoctonia solani AG1 IA triggers cell death in plants and promotes disease development through inhibiting PAMP-triggered immunity in Arabidopsis thaliana. Front. Microbiol. 2019, 10, 2228. [Google Scholar] [CrossRef] [PubMed]
- Wei, M.; Wang, A.J.; Liu, Y.; Ma, L.; Niu, X.; Zheng, A. Identification of the novel effector RsIA_NP8 in Rhizoctonia solani AG1 IA that induces cell death and triggers defense responses in non-host plants. Front. Plant Sci. 2020, 11, 01115. [Google Scholar] [CrossRef] [PubMed]
- Wibberg, D.; Andersson, L.; Rupp, O.; Goesmann, A.; Pühler, A.; Varrelmann, M.; Dixelius, C.; Schlüter, A. Draft genome sequence of the sugar beet pathogen Rhizoctonia solani AG2-2IIIB strain BBA69670. J. Biotechnol. 2016, 222, 11–12. [Google Scholar] [CrossRef]
- Gerding, M.A.; Liu, B.; Bendezu, F.O.; Hale, C.A.; Bernhardt, T.G.; de Boer, P.A. Self-enhanced accumulation of FtsN at division sites and roles for other proteins with a SPOR domain (DamX, DedD, and RlpA) in Escherichia coli cell constriction. J. Bacteriol. 2009, 191, 7383–7401. [Google Scholar] [CrossRef]
- Joosten, M.H.; Vogelsang, R.; Cozijnsen, T.J.; Verberne, M.; de Wit, P.J. The biotrophic fungus Cladosporium fulvum circumvents Cf-4-mediated resistance by producing unstable AVR4 elicitors. Plant Cell 1997, 9, 367–379. [Google Scholar]
- van den Burg, H.A.; Harrison, S.J.; Joosten, M.H.; Vervoort, J.; de Wit, P.J. Cladosporium fulvum Avr4 protects fungal cell walls against hydrolysis by plant chitinases accumulating during infection. Mol. Plant Microbe Interact. 2006, 19, 1420–1430. [Google Scholar] [CrossRef]
- Stergiopoulos, I.; de Wit, P.J. Fungal effector proteins. Annu. Rev. Phytopathol. 2009, 47, 233–263. [Google Scholar] [CrossRef] [PubMed]
- Lo Presti, L.; Lanver, D.; Schweizer, G.; Tanaka, S.; Liang, L.; Tollot, M.; Zuccaro, A.; Reissmann, S.; Kahmann, R. Fungal effectors and plant susceptibility. Annu. Rev. Plant Biol. 2015, 66, 513–545. [Google Scholar] [CrossRef] [PubMed]
- Collmer, A.; Lindeberg, M.; Petnicki-Ocwieja, T.; Schneider, D.J.; Alfano, J.R. Genomic mining type III secretion system effectors in Pseudomonas syringae yields new picks for all TTSS prospectors. Trends Microbiol. 2002, 10, 462–469. [Google Scholar] [CrossRef]
- Martínez, M.; Cambra, I.; González-Melendi, P.; Santamaría, M.E.; Díaz, I. C1A cysteine-proteases and their inhibitors in plants. Physiol. Plant 2012, 145, 85–94. [Google Scholar] [CrossRef]
- Bárány, I.; Berenguer, E.; Solís, M.T.; Pérez-Pérez, Y.; Santamaría, M.E.; Crespo, J.L.; Risueño, M.C.; Díaz, I.; Testillano, P.S. Autophagy is activated and involved in cell death with participation of cathepsins during stress-induced microspore embryogenesis in barley. J. Exp. Bot. 2018, 69, 1387–1402. [Google Scholar] [PubMed]
- Tian, M.; Win, J.; Song, J.; van der Hoorn, R.; van der Knaap, E.; Kamoun, S. A Phytophthora infestans cystatin-like protein targets a novel tomato papain-like apoplastic protease. Plant Physiol. 2007, 143, 364–377. [Google Scholar] [PubMed]
- Rooney, H.C.; van’t Klooster, J.W.; van der Hoorn, R.A.; Joosten, M.H.; Jones, J.D.; de Wit, P.J. Cladosporium Avr2 inhibits tomato Rcr3 protease required for Cf-2-dependent disease resistance. Science 2005, 308, 1783–1786. [Google Scholar] [PubMed]
- Wibberg, D.; Andersson, L.; Tzelepis, G.; Rupp, O.; Blom, J.; Jelonek, L.; Pühler, A.; Fogelqvist, J.; Varrelmann, M.; Schlüter, A.; et al. Genome analysis of the sugar beet pathogen Rhizoctonia solani AG2-2IIIB revealed high numbers in secreted proteins and cell wall degrading enzymes. BMC Genom. 2016, 17, 245. [Google Scholar] [CrossRef]
- Castillo, R.M.; Mizuguchi, K.; Dhanaraj, V.; Albert, A.; Blundell, T.L.; Murzin, A.G. A six-stranded double-psi beta barrel is shared by several protein superfamilies. Structure 1999, 7, 227–236. [Google Scholar] [CrossRef]
- Quiroz-Castaneda, R.E.; Martinez-Anaya, C.; Cuervo-Soto, L.I.; Segovia, L.; Folch-Mallol, J.L. Loosenin, a novel protein with cellulose-disrupting activity from Bjerkandera adusta. Microb. Cell Fact. 2011, 10, 8. [Google Scholar] [CrossRef]
- Mosquera, G.; Giraldo, M.C.; Khang, C.H.; Coughlan, S.; Valent, B. Interaction transcriptome analysis identifies Magnaporthe oryzae BAS1-4 as biotrophy-associated secreted proteins in rice blast disease. Plant Cell 2009, 21, 1273–1290. [Google Scholar]
- Pellegrin, C.; Morin, E.; Martin, F.M.; Veneault-Fourrey, C. Comparative analysis of secretomes from ectomycorrhizal fungi with an emphasis on small-secreted proteins. Front. Microbiol. 2015, 6, 1278. [Google Scholar] [CrossRef]
- Ismail, I.A.; Able, A.J. Secretome analysis of virulent Pyrenophora teres f. teres isolates. Proteomics 2016, 16, 2625–2636. [Google Scholar]
- Arends, S.J.; Williams, K.; Scott, R.J.; Rolong, S.; Popham, D.L.; Weiss, D.S. Discovery and characterization of three new Escherichia coli septal ring proteins that contain a SPOR domain: DamX, DedD, and RlpA. J. Bacteriol. 2010, 192, 242–255. [Google Scholar] [CrossRef]
- Jorgenson, M.A.; Chen, Y.; Yahashiri, A.; Popham, D.L.; Weiss, D.S. The bacterial septal ring protein RlpA is a lytic transglycosylase that contributes to rod shape and daughter cell separation in Pseudomonas aeruginosa. Mol. Microbiol. 2014, 93, 113–128. [Google Scholar] [CrossRef]
- Kim, N.H.; Hwang, B.K. Pepper pathogenesis-related protein 4c is a plasma membrane-localized cysteine protease inhibitor that is required for plant cell death and defense signaling. Plant J. 2015, 81, 81–94. [Google Scholar] [CrossRef]
- Krüger, J.; Thomas, C.M.; Golstein, C.; Dixon, M.S.; Smoker, M.; Tang, S.; Mulder, L.; Jones, J.D. A tomato cysteine protease required for Cf-2-dependent disease resistance and suppression of autonecrosis. Science 2002, 296, 744–747. [Google Scholar] [CrossRef]
- Xu, Q.F.; Cheng, W.S.; Li, S.S.; Li, W.; Zhang, Z.X.; Xu, Y.P.; Zhou, X.P.; Cai, X.Z. Identification of genes required for Cf-dependent hypersensitive cell death by combined proteomic and RNA interfering analyses. J. Exp. Bot. 2012, 63, 2421–2435. [Google Scholar] [CrossRef] [PubMed]
- Misas-Villamil, J.C.; van der Hoorn, R.A.; Doehlemann, G. Papain-like cysteine proteases as hubs in plant immunity. New Phytol. 2016, 212, 902–907. [Google Scholar] [CrossRef]
- Pearce, G.; Yamaguchi, Y.; Barona, G.; Ryan, C.A. A subtilisin-like protein from soybean contains an embedded, cryptic signal that activates defense-related genes. Proc. Natl. Acad. Sci. USA 2010, 107, 14921–14925. [Google Scholar] [CrossRef] [PubMed]
- Bozkurt, T.O.; Schornack, S.; Win, J.; Shindo, T.; Ilyas, M.; Oliva, R.; Cano, L.M.; Jones, A.M.E.; Huitema, E.; van der Hoorn, R.A.L.; et al. Phytophthora infestans effector AVRblb2 prevents secretion of a plant immune protease at the haustorial interface. Proc. Natl. Acad. Sci. USA 2011, 108, 20832–20837. [Google Scholar] [CrossRef] [PubMed]
- Müeller, A.N.; Ziemann, S.; Treitschke, S.; Assmann, D.; Doehlemann, G. Compatibility in the Ustilago maydis-maize interaction requires inhibition of host cysteine proteases by the fungal effector Pit2. PLoS Pathog. 2013, 9, e1003177. [Google Scholar] [CrossRef]
- Clark, K.; Franco, J.Y.; Schwizer, S.; Pang, Z.; Hawara, E.; Liebrand, T.W.H.; Pagliaccia, D.; Zeng, L.; Gurung, F.B.; Wang, P.; et al. An effector from the Huanglongbing-associated pathogen targets citrus proteases. Nat. Commun. 2018, 9, 1718. [Google Scholar] [CrossRef]
- D’Silva, I.; Poirier, G.G.; Heath, M.C. Activation of cysteine proteases in cowpea plants during the hypersensitive response: A form of programmed cell death. Exp. Cell Res. 1998, 245, 389–399. [Google Scholar] [CrossRef]
- Lozano-Torres, J.L.; Wilbers, R.H.; Gawronski, P.; Boshoven, J.C.; Finkers-Tomczak, A.; Cordewener, J.H.; America, A.H.; Overmars, H.A.; Van’t Klooster, J.W.; Baranowski, L.; et al. Dual disease resistance mediated by the immune receptor Cf-2 in tomato requires a common virulence target of a fungus and a nematode. Proc. Natl. Acad. Sci. USA 2012, 109, 10119–10124. [Google Scholar] [CrossRef]
- Spanu, P.D.; Panastruga, R. Editorial: Biotrophic plant-microbe interactions. Front. Plant Sci. 2017, 8, 192. [Google Scholar] [CrossRef] [PubMed]
- Dölfors, F.; Holmquist, L.; Dixelius, C.; Tzelepis, G. A LysM effector protein from the basidiomycete Rhizoctonia solani contributes to virulence through suppression of chitin-triggered immunity. Mol. Genet. Genom. 2019, 294, 1211–1218. [Google Scholar] [CrossRef]
- Holmquist, L.; Dölfors, F.; Fogelqvist, J.; Cohn, J.; Kraft, T.; Dixelius, C. Major latex protein-like encoding genes contribute to Rhizoctonia solani defense responses in sugar beet. Mol. Genet. Genom. 2020, accepted. [Google Scholar] [CrossRef]
- Marcuzzo, L.L.; Haveroth, R.; Nascimento, A. Induction technique of sporulation in vitro of Cercospora beticola. Summa Phytopathol. 2015, 41, 74. [Google Scholar] [CrossRef][Green Version]
- Staats, M.; van Kan, J.A. Genome update of Botrytis cinerea strains B05.10 and T4. Eukaryot Cell 2012, 11, 1413–1414. [Google Scholar] [CrossRef] [PubMed]
- Tzelepis, G.; Melin, P.; Jensen-Funck, D.; Stenlid, J.; Karlsson, M. Functional analysis of glycoside hydrolase family 18 and 20 genes in Neurospora crassa. Fungal Genet. Biol. 2012, 49, 717–730. [Google Scholar] [CrossRef] [PubMed]
- Chamoun, R.; Samsatly, J.; Pakala, S.B.; Cubeta, M.A.; Jabaji, S. Suppression subtractive hybridization and comparative expression of a pore-forming toxin and glycosyl hydrolase genes in Rhizoctonia solani during potato sprout infection. Mol. Genet. Genom. 2015, 290, 877–900. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Crespo-Sempere, A.; Lopez-Perez, M.; Martinez-Culebras, P.V.; Gonzalez-Candelas, L. Development of a green fluorescent tagged strain of Aspergillus carbonarius to monitor fungal colonization in grapes. Int. J. Food Microbiol. 2011, 148, 135–140. [Google Scholar] [CrossRef]
- Utermark, J.; Karlovsky, P. Genetic transformation of filamentous fungi by Agrobacterium tumefaciens. Protocol Exchange. 2008. Available online: http://www.nature.com/protocolexchange/protocols/427 (accessed on 1 March 2017).
- Möller, E.M.; Bahnweg, G.; Sandermann, H.; Geiger, H.H. A simple and efficient protocol for isolation of high molecular weight DNA from filamentous fungi, fruit bodies, and infected plant tissues. Nucleic Acids Res. 1992, 20, 6115–6116. [Google Scholar] [PubMed]
- Davis, A.M.; Hall, A.; Millar, A.J.; Darrah, C.; Davis, S.J. Protocol: Streamlined sub-protocols for floral-dip transformation and selection of transformants in Arabidopsis thaliana. Plant Methods 2009, 5, 3. [Google Scholar] [PubMed]
- Li, X. Infiltration of Nicotiana benthamiana protocol for transient expression via Agrobacterium. Bio-Protocol 2011, 1, e95. [Google Scholar]
- Gimenez-Ibanez, S.; Hann, D.R.; Chang, J.H.; Segonzac, C.; Boller, T.; Rathjen, J.P. Differential suppression of Nicotiana benthamiana innate immune responses by transiently expressed Pseudomonas syringae Type III effectors. Front. Plant Sci. 2018, 9, 688. [Google Scholar]
- Zheng, L.; Baumann, U.; Reymond, J.L. An efficient one-step site-directed and site-saturation mutagenesis protocol. Nucleic Acids Res. 2004, 32, e115. [Google Scholar]
- Letunic, I.; Doerks, T.; Bork, P. SMART 6, Recent updates and new developments. Nucleic Acids Res 2009, 37, D229–D232. [Google Scholar] [PubMed]
- Jones, P.; Binns, D.; Chang, H.Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5, Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; de Beer, T.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar]
- Wang, S.; Li, W.; Liu, S.; Xu, J. RaptorX-Property: A web server for protein structure property prediction. Nucleic Acids Res. 2016, 44, W430–W435. [Google Scholar] [PubMed]
- Charova, S.N.; Gazi, A.D.; Mylonas, E.; Pozidis, C.; Sabarit, B.; Anagnostou, D.; Psatha, K.; Aivaliotis, M.; Beuzon, C.R.; Panopoulos, N.J.; et al. Migration of type III secretion system transcriptional regulators links gene expression to secretion. mBio 2018, 9, e01096-18. [Google Scholar]
- Gawlik, K.; Gutowicz, J. Inhibitory activity against papain, a CA1 cysteine peptidase, in Saccharomycetaceae. Microbiol. Res. 2008, 163, 545–555. [Google Scholar] [CrossRef] [PubMed]
- Albert, M.; Butenko, M.A.; Aalen, R.B.; Felix, G.; Wildhagen, M. Chemiluminescence detection of the oxidative burst in plant leaf pieces. Bioprotocol 2015, 5, e1423. [Google Scholar] [CrossRef]
- Moggridge, S.; Sorensen, P.H.; Morin, G.B.; Hughes, C.S. Extending the compatibility of the SP3 paramagnetic bead processing approach for proteomics. J. Proteome Res. 2018, 17, 1730–1740. [Google Scholar] [CrossRef]






Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Charova, S.N.; Dölfors, F.; Holmquist, L.; Moschou, P.N.; Dixelius, C.; Tzelepis, G. The RsRlpA Effector Is a Protease Inhibitor Promoting Rhizoctonia solani Virulence through Suppression of the Hypersensitive Response. Int. J. Mol. Sci. 2020, 21, 8070. https://doi.org/10.3390/ijms21218070
Charova SN, Dölfors F, Holmquist L, Moschou PN, Dixelius C, Tzelepis G. The RsRlpA Effector Is a Protease Inhibitor Promoting Rhizoctonia solani Virulence through Suppression of the Hypersensitive Response. International Journal of Molecular Sciences. 2020; 21(21):8070. https://doi.org/10.3390/ijms21218070
Chicago/Turabian StyleCharova, Spyridoula N., Fredrik Dölfors, Louise Holmquist, Panagiotis N. Moschou, Christina Dixelius, and Georgios Tzelepis. 2020. "The RsRlpA Effector Is a Protease Inhibitor Promoting Rhizoctonia solani Virulence through Suppression of the Hypersensitive Response" International Journal of Molecular Sciences 21, no. 21: 8070. https://doi.org/10.3390/ijms21218070
APA StyleCharova, S. N., Dölfors, F., Holmquist, L., Moschou, P. N., Dixelius, C., & Tzelepis, G. (2020). The RsRlpA Effector Is a Protease Inhibitor Promoting Rhizoctonia solani Virulence through Suppression of the Hypersensitive Response. International Journal of Molecular Sciences, 21(21), 8070. https://doi.org/10.3390/ijms21218070




