Evolution Analysis of the Fasciclin-Like Arabinogalactan Proteins in Plants Shows Variable Fasciclin-AGP Domain Constitutions
Abstract
1. Introduction
2. Results and Discussions
2.1. Identification of the FLA Family in Plants
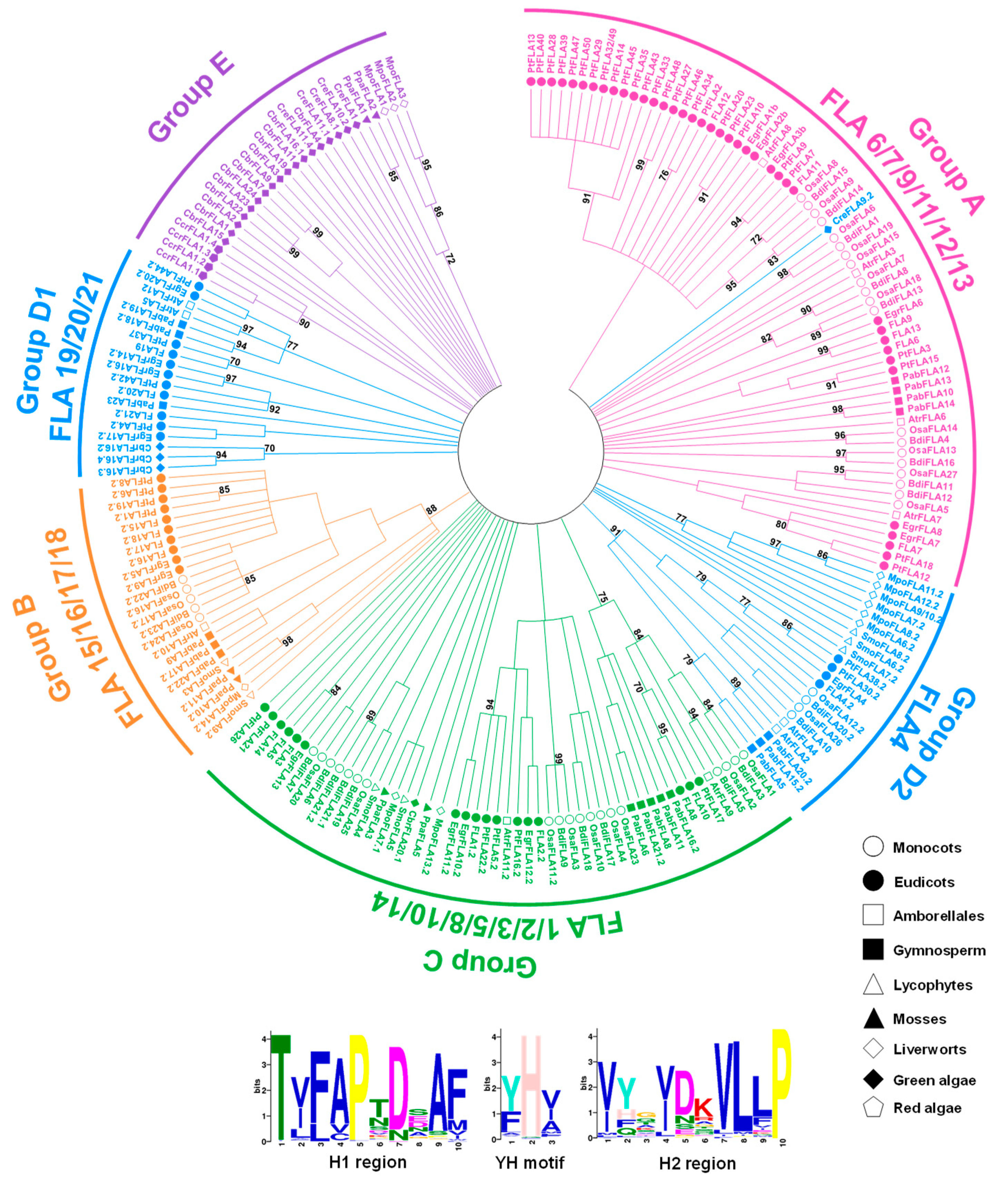
2.2. Phylogenetic Analysis and Classification of FLAs
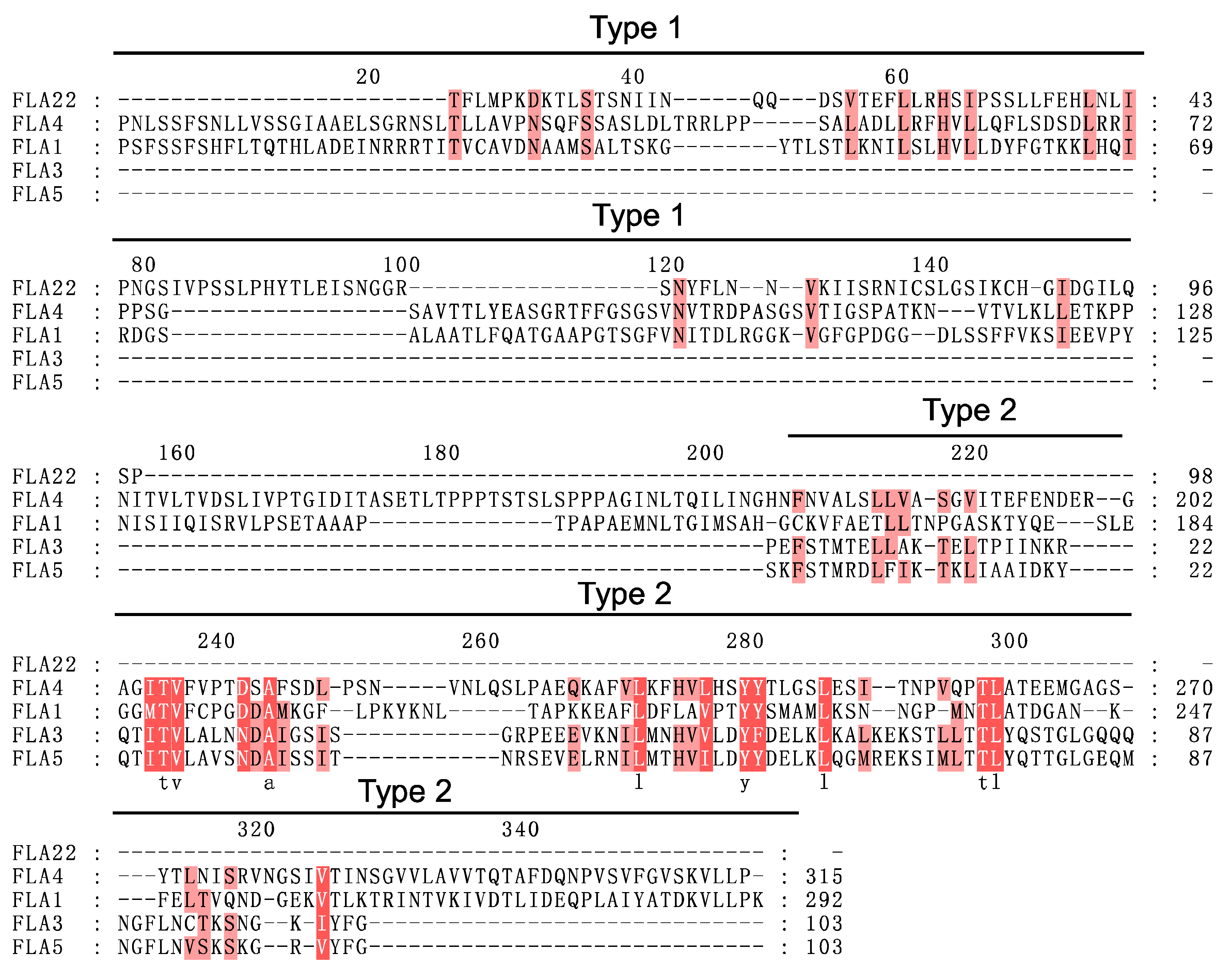
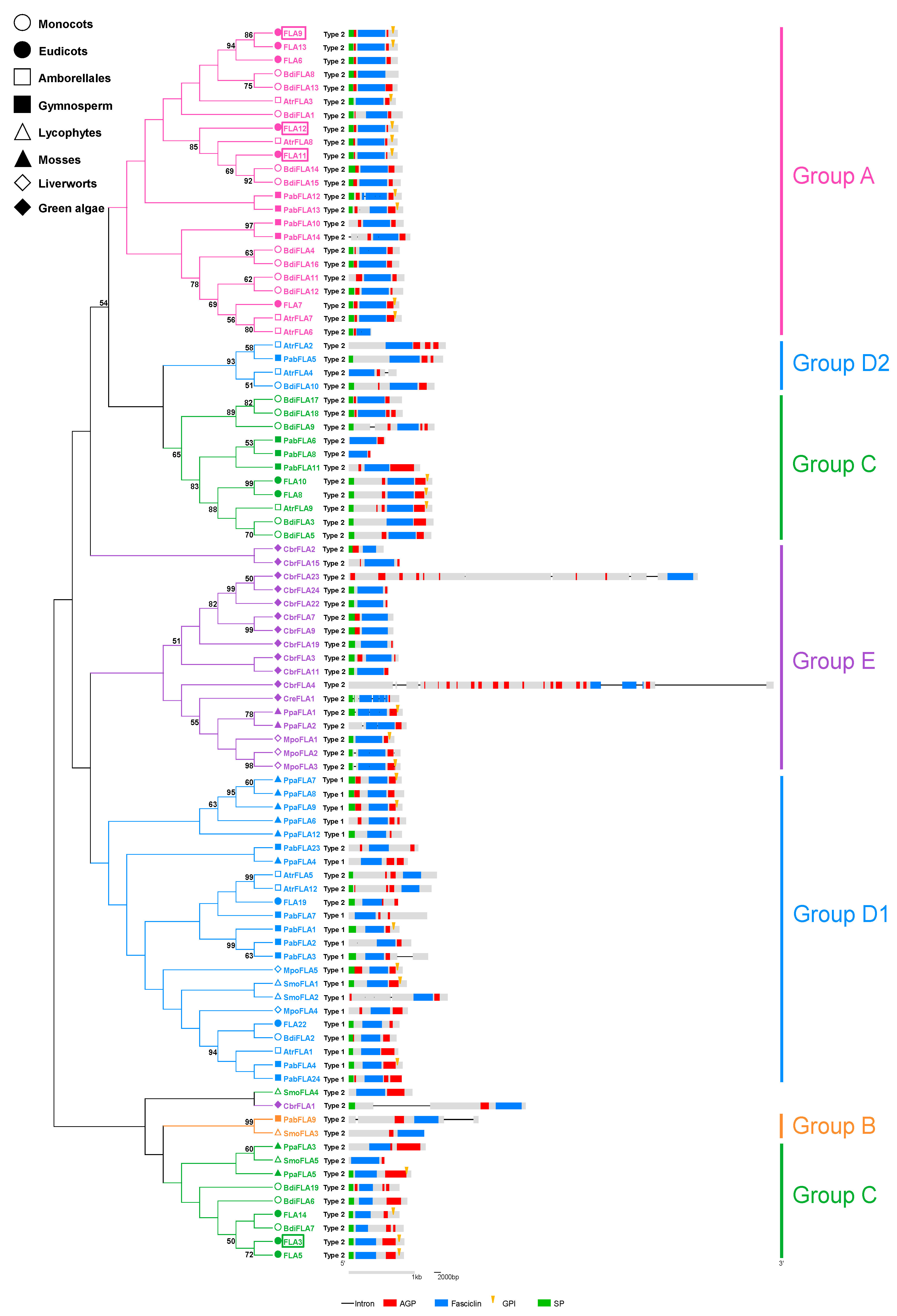
2.3. Structural and Evolutionary Analysis of FLAs
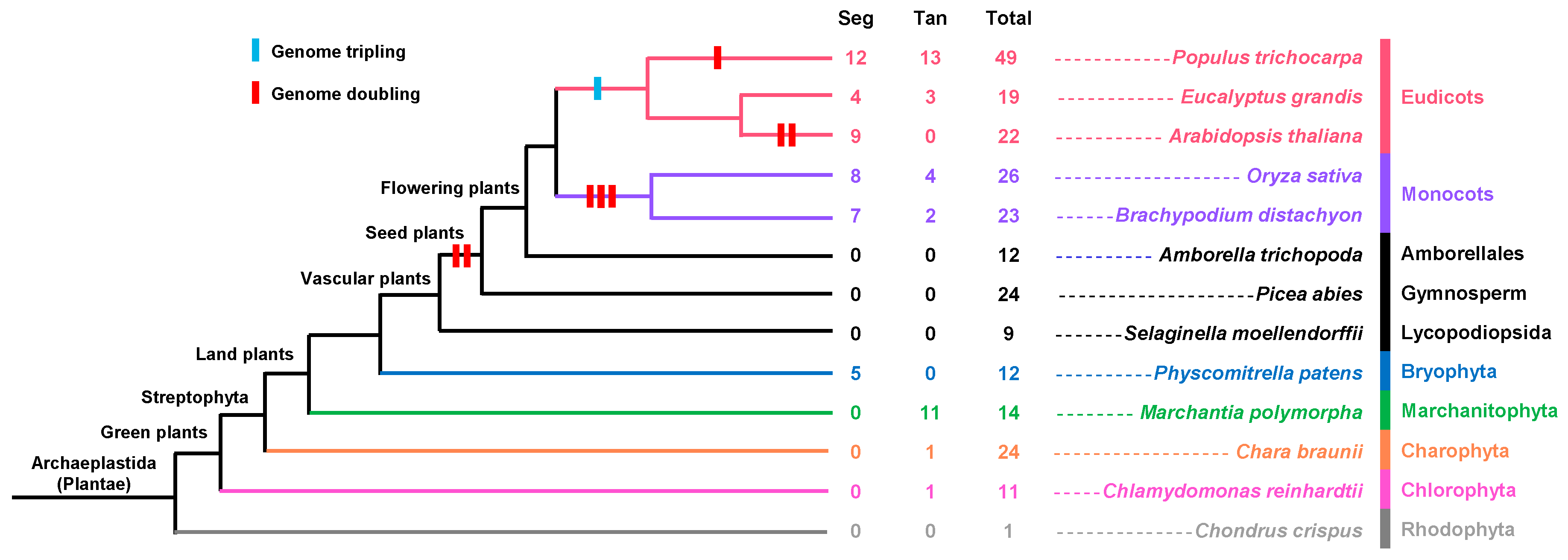
2.4. Analysis of FLA Duplication Patterns during the Process of Evolution
3. Materials and Methods
3.1. Bioinformatics Identification of FLAs
3.2. Multiple Sequence Alignment and Phylogenetic Analysis
3.3. Motif Prediction
3.4. Gene Duplication and Molecular Evolution
4. Conclusions
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
Abbreviations
| FLA | Fasciclin-like arabinogalactan protein |
| AGP | Arabinogalactan protein |
| GPI | Glycosylphosphatidylinositol |
| PAST% | The percentage of Pro, Ala, Ser, and Thr residues in a protein amino-acid sequence |
| Ccr | Chondrus crispus |
| Cre | Chlamydomonas reinhardtii |
| Mpo | Marchantia polymorpha |
| Smo | Selaginella moellendorffii |
| Pab | Picea abies |
| Atr | Amborella trichopoda |
| Egr | Eucalyptus grandis |
| Pt | Populus trichocarpa |
| Bdi | Brachypodium distachyon |
| Os | Oryza sativa |
| Ka | Nonsynonymous substitution rate |
| Ks | Synonymous substitution rate |
| PGDD | Plant Genome Duplication Database |
| NCBI | National Center for Biotechnology Information |
| ConGenIE | Conifer Genome Integrative Explorer |
| HMM | Hidden Markov Model |
| BLASTP | Protein Basic Local Alignment Search Tool |
| MYN | Modified Yang-Nielsen Algorithm |
| MCSCAN | Multiple Collinearity Scan |
| MEME | Multiple Expectation maximization for Motif Elicitation |
| LG | Le_Gascuel_2008 model |
| G | Gamma distribution |
| I | Evolutionarily invariable |
| GSDS | Gene Structure Display Server |
| OrcAE | Online Resource for Community Annotation of Eukaryotes |
References
- Nothnagel, E.A. Proteoglycans and related components in plant cells. Int. Rev. Cytol. 1997, 174, 195–291. [Google Scholar] [PubMed]
- Chivasa, S.; Ndimba, B.K.; Simon, W.J.; Robertson, D.; Yu, X.L.; Knox, J.P.; Bolwell, P.; Slabas, A.R. Proteomic analysis of the Arabidopsis thaliana cell wall. Electrophoresis 2002, 23, 1754–1765. [Google Scholar] [CrossRef]
- Johnson, K.L.; Cassin, A.M.; Lonsdale, A.; Wong, G.K.-S.; Soltis, D.E.; Miles, N.W.; Melkonian, M.; Melkonian, B.; Deyholos, M.K.; Leebens-Mack, J.; et al. Insights into the Evolution of Hydroxyproline-rich Glycoproteins from 1000 plant Transcriptomes. Plant Physiol. 2017, 174, 904–921. [Google Scholar] [CrossRef]
- Jamet, E.; Albenne, C.; Boudart, G.; Irshad, M.; Canut, H.; Pont-Lezica, R. Recent advances in plant cell wall proteomics. Proteomics 2008, 8, 893–908. [Google Scholar] [CrossRef] [PubMed]
- Nothnagel, E.A.; Bacic, A.; Clarke, A.E. Cell and Developmental Biology of Arabinogalactan-Proteins; Springer Science Business Media: New York, NY, USA, 2000. [Google Scholar]
- Showalter, A.M. Arabinogalactan-proteins: Structure, expression and function. Cell. Mol. Life Sci. 2001, 58, 1399–1417. [Google Scholar] [CrossRef]
- Ma, Y.; Yan, C.; Li, H.; Wu, W.; Liu, Y.; Wang, Y.; Chen, Q.; Ma, H. Bioinformatics Prediction and Evolution Analysis of Arabinogalactan Proteins in the Plant Kingdom. Front. Plant Sci. 2017, 8, 66. [Google Scholar] [CrossRef]
- Ellis, M.; Egelund, J.; Schultz, C.J.; Bacic, A. Arabinogalactan-proteins: Key Regulators at the Cell Surface? Plant Physiol. 2010, 153, 403–419. [Google Scholar] [CrossRef]
- Shpak, E.; Barbar, E.; Leykam, J.F.; Kieliszewski, M.J. Contiguous Hydroxyproline Residues Direct Hydroxyproline Arabinosylation in Nicotiana tabacum. J. Biol. Chem. 2001, 276, 11272–11278. [Google Scholar] [CrossRef] [PubMed]
- Showalter, A.M.; Basu, D. Extensin and Arabinogalactan-Protein Biosynthesis: Glycosyltransferases, Research Challenges, and Biosensors. Front. Plant Sci 2016, 7, 814. [Google Scholar] [CrossRef] [PubMed]
- Johnson, K.L.; Jones, B.J.; Bacic, A.; Schultz, C.J. The fasciclin-like arabinogalactan proteins of Arabidopsis. A multigene family of putative cell adhesion molecules. Plant Physiol. 2003, 133, 1911–1925. [Google Scholar] [CrossRef] [PubMed]
- Ma, H.; Zhao, J. Genome-wide identification, classification, and expression analysis of the arabinogalactan protein gene family in rice (Oryza sativa L.). J. Exp. Bot. 2010, 61, 2647–2668. [Google Scholar] [CrossRef]
- MacMillan, C.P.; Taylor, L.; Bi, Y.; Southerton, S.G.; Evans, R.; Spokevicius, A. The fasciclin-like arabinogalactan protein family of Eucalyptus grandis contains members that impact wood biology and biomechanics. New Phytol. 2015, 206, 1314–1327. [Google Scholar] [CrossRef] [PubMed]
- Mashiguchi, K.; Asami, T.; Suzuki, Y. Genome-wide identification, structure and expression studies, and mutant collection of 22 early nodulin-like protein genes in Arabidopsis. Biosci. Biotechnol. Biochem. 2009, 73, 2452–2459. [Google Scholar] [CrossRef]
- Ma, H.; Zhao, H.; Liu, Z.; Zhao, J. The phytocyanin gene family in rice (Oryza sativa L.): genome-wide identification, classification and transcriptional analysis. PLoS ONE 2011, 6, e25184. [Google Scholar] [CrossRef]
- Motose, H.; Sugiyama, M.; Fukuda, H. A proteoglycan mediates inductive interaction during plant vascular development. Nature 2004, 429, 873–878. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, Y.; Motose, H.; Iwamoto, K.; Fukuda, H. Expression and genome-wide analysis of the xylogen-type gene family. Plant Cell Physiol. 2011, 52, 1095–1106. [Google Scholar] [CrossRef] [PubMed]
- Seifert, G.J. Fascinating Fasciclins: A Surprisingly Widespread Family of Proteins that Mediate Interactions between the Cell Exterior and the Cell Surface. Int. J. Mol. Sci. 2018, 19, 1628. [Google Scholar] [CrossRef]
- Bastiani, M.J.; Harrelson, A.L.; Snow, P.M.; Goodman, C.S. Expression of fasciclin I and II glycoproteins on subsets of axon pathways during neuronal development in the grasshopper. Cell 1987, 48, 745–755. [Google Scholar] [CrossRef]
- Elkins, T.; Zinn, K.; McAllister, L.; Hoffmann, F.M.; Goodman, C.S. Genetic analysis of a Drosophila neural cell adhesion molecule: interaction of fasciclin I and Abelson tyrosine kinase mutations. Cell 1990, 60, 565–575. [Google Scholar] [CrossRef]
- Schultz, C.J.; Rumsewicz, M.P.; Johnson, K.L.; Jones, B.J.; Gaspar, Y.M.; Bacic, A. Using Genomic Resources to Guide Research Directions. The Arabinogalactan Protein Gene Family as a Test Case. Plant Physiol. 2002, 129, 1448–1463. [Google Scholar] [CrossRef]
- Faik, A.; Abouzouhair, J.; Sarhan, F. Putative fasciclin-like arabinogalactan-proteins (FLA) in wheat (Triticum aestivum) and rice (Oryza sativa): identification and bioinformatic analyses. Mol. Genet. Genom. 2006, 276, 478–494. [Google Scholar] [CrossRef]
- Lafarguette, F.; Leplé, J.C.; Déjardin, A.; Laurans, F.; Costa, G.; Lesage-Descauses, M.C.; Pilate, G. Poplar genes encoding fasciclin-like arabinogalactan proteins are highly expressed in tension wood. New Phyto. 2004, 164, 107–121. [Google Scholar] [CrossRef]
- Showalter, A.M.; Keppler, B.D.; Liu, X.; Lichtenberg, J.; Welch, L.R. Bioinformatic Identification and Analysis of Hydroxyproline-Rich Glycoproteins in Populus trichocarpa. Bmc Plant Biol. 2016, 16, 229. [Google Scholar] [CrossRef] [PubMed]
- Dahiya, P.; Findlay, K.; Roberts, K.; McCann, M.C. A fasciclin-domain containing gene, ZeFLA11, is expressed exclusively in xylem elements that have reticulate wall thickenings in the stem vascular system of Zinnia elegans cv Envy. Planta 2006, 223, 1281–1291. [Google Scholar] [CrossRef] [PubMed]
- Huang, G.Q.; Xu, W.L.; Gong, S.Y.; Li, B.; Wang, X.L.; Xu, D.; Li, X.B. Characterization of 19 novel cotton FLA genes and their expression profiling in fiber development and in response to phytohormones and salt stress. Physiol. Plant 2008, 134, 348–359. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Shi, R.; Wang, X.; Pan, Y.; Li, Z.; Yang, X.; Zhang, G.; Ma, Z. Characterization and expression analysis of a fiber differentially expressed Fasciclin-like arabinogalactan protein gene in sea island cotton fibers. PLoS ONE 2013, 8, e70185. [Google Scholar] [CrossRef]
- Jun, L.; Xiaoming, W. Genome-wide identification, classification and expression analysis of genes encoding putative fasciclin-like arabinogalactan proteins in Chinese cabbage (Brassica rapa L.). Mol. Biol. Rep. 2012, 39, 10541–10555. [Google Scholar] [CrossRef]
- Guerriero, G.; Mangeot-Peter, L.; Legay, S.; Behr, M.; Lutts, S.; Siddiqui, K.S.; Hausman, J.F. Identification of fasciclin-like arabinogalactan proteins in textile hemp (Cannabis sativa L.): in silico analyses and gene expression patterns in different tissues. Bmc Genom. 2017, 18, 741. [Google Scholar] [CrossRef]
- MacMillan, C.P.; Mansfield, S.D.; Stachurski, Z.H.; Evans, R.; Southerton, S.G. Fasciclin-like arabinogalactan proteins: specialization for stem biomechanics and cell wall architecture in Arabidopsis and Eucalyptus. Plant J. 2010, 62, 689–703. [Google Scholar] [CrossRef] [PubMed]
- Collen, J.; Porcel, B.; Carre, W.; Ball, S.G.; Chaparro, C.; Tonon, T.; Barbeyron, T.; Michel, G.; Noel, B.; Valentin, K.; et al. Genome structure and metabolic features in the red seaweed Chondrus crispus shed light on evolution of the Archaeplastida. Proc. Natl. Acad. Sci. USA 2013, 110, 5247–5252. [Google Scholar] [CrossRef] [PubMed]
- Bowman, J.L.; Kohchi, T.; Yamato, K.T.; Jenkins, J.; Shu, S.; Ishizaki, K.; Yamaoka, S.; Nishihama, R.; Nakamura, Y.; Berger, F.; et al. Insights into Land Plant Evolution Garnered from the Marchantia polymorpha Genome. Cell 2017, 171, 287–304. [Google Scholar] [CrossRef] [PubMed]
- Lang, D.; Ullrich, K.K.; Murat, F.; Fuchs, J.; Jenkins, J.; Haas, F.B.; Piednoel, M.; Gundlach, H.; Van Bel, M.; Meyberg, R.; et al. The Physcomitrella patens chromosome-scale assembly reveals moss genome structure and evolution. Plant J. 2018, 93, 515–533. [Google Scholar] [CrossRef] [PubMed]
- Banks, J.A.; Nishiyama, T.; Hasebe, M.; Bowman, J.L.; Gribskov, M.; dePamphilis, C.; Albert, V.A.; Aono, N.; Aoyama, T.; Ambrose, B.A.; et al. The Selaginella Genome Identifies Genetic Changes Associated with the Evolution of Vascular Plants. Science 2011, 332, 960–963. [Google Scholar] [CrossRef] [PubMed]
- Nystedt, B.; Street, N.R.; Wetterbom, A.; Zuccolo, A.; Lin, Y.-C.; Scofield, D.G.; Vezzi, F.; Delhomme, N.; Giacomello, S.; Alexeyenko, A.; et al. The Norway spruce genome sequence and conifer genome evolution. Nature 2013, 497, 579–584. [Google Scholar] [CrossRef] [PubMed]
- Ambrella Genome Project. The Amborella Genome and the Evolution of Flowering Plants. Science 2013, 342, 1241089. [Google Scholar] [CrossRef] [PubMed]
- The International Brachypodium Initiative. Genome sequencing and analysis of the model grass Brachypodium distachyon. Nature 2010, 463, 763–768. [Google Scholar] [CrossRef]
- Ouyang, S.; Zhu, W.; Hamilton, J.; Lin, H.; Campbell, M.; Childs, K.; Thibaud-Nissen, F.; Malek, R.L.; Lee, Y.; Zheng, L.; et al. The TIGR rice Genome Annotation Resource: improvements and new features. Nucleic Acids Res. 2007, 35, D883–D887. [Google Scholar] [CrossRef]
- Merchant, S.S.; Prochnik, S.E.; Vallon, O.; Harris, E.H.; Karpowicz, S.J.; Witman, G.B.; Terry, A.; Salamov, A.; Fritz-Laylin, L.K.; Marechal-Drouard, L.; et al. The Chlamydomonas Genome Reveals the Evolution of Key Animal and Plant Functions. Science 2007, 318, 245–250. [Google Scholar] [CrossRef] [PubMed]
- Lamesch, P.; Berardini, T.Z.; Li, D.; Swarbreck, D.; Wilks, C.; Sasidharan, R.; Muller, R.; Dreher, K.; Alexander, D.L.; Garcia-Hernandez, M.; et al. The Arabidopsis Information Resource (TAIR): improved gene annotation and new tools. Nucleic Acids Res. 2012, 40, D1202–D1210. [Google Scholar] [CrossRef]
- Myburg, A.A.; Grattapaglia, D.; Tuskan, G.A.; Hellsten, U.; Hayes, R.D.; Grimwood, J.; Jenkins, J.; Lindquist, E.; Tice, H.; Bauer, D.; et al. The genome of Eucalyptus grandis. Nature 2014, 510, 356–362. [Google Scholar] [CrossRef]
- Tuskan, G.A.; Difazio, S.; Jansson, S.; Bohlmann, J.; Grigoriev, I.; Hellsten, U.; Putnam, N.; Ralph, S.; Rombauts, S.; Salamov, A.; et al. The Genome of Black Cottonwood, Populus trichocarpa (Torr. & Gray). Science 2006, 313, 1596–1604. [Google Scholar]
- Nishiyama, T.; Sakayama, H.; de Vries, J.; Buschmann, H.; Saint-Marcoux, D.; Ullrich, K.K.; Haas, F.B.; Vanderstraeten, L.; Becker, D.; Lang, D.; et al. The Chara Genome: Secondary Complexity and Implications for Plant Terrestrialization. Cell 2018, 174, 448–464. [Google Scholar] [CrossRef]
- Hu, B.; Jin, J.; Guo, A.-Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: architecture and applications. Bmc Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef] [PubMed]
- Sievers, F.; Wilm, A.; Dineen, D.; Gibson, T.J.; Karplus, K.; Li, W.; Lopez, R.; McWilliam, H.; Remmert, M.; Soding, J.; et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 2011, 7, 539. [Google Scholar] [CrossRef]
- Sievers, F.; Higgins, D.G. Clustal Omega for making accurate alignments of many protein sequences. Protein Sci. 2018, 27, 135–145. [Google Scholar] [CrossRef]
- Thompson, J.D.; Plewniak, F.; Poch, O. A comprehensive comparison of multiple sequence alignment programs. Nucleic Acids Res. 1999, 27, 2682–2690. [Google Scholar] [CrossRef] [PubMed]
- Hall, B.G. Phylogenetic Trees Made Easy: A How-To Manual, 4th ed.; Oxford University Press: Cary, NC, USA, 2011. [Google Scholar]
- Ogden, T.H.; Rosenberg, M.S. Multiple Sequence Alignment Accuracy and Phylogenetic Inference. Syst. Biol. 2006, 55, 314–328. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Le, S.Q.; Gascuel, O. An Improved General Amino Acid Replacement Matrix. Mol. Biol. Evol. 2008, 25, 1307–1320. [Google Scholar] [CrossRef]
- Bailey, T.L.; Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 1994, 2, 28–36. [Google Scholar] [PubMed]
- Klepikova, A.V.; Logacheva, M.D.; Dmitriev, S.E.; Penin, A.A. RNA-seq analysis of an apical meristem time series reveals a critical point in Arabidopsis thaliana flower initiation. Bmc Genom. 2015, 16, 466. [Google Scholar] [CrossRef] [PubMed]
- Klepikova, A.V.; Kasianov, A.S.; Gerasimov, E.S.; Logacheva, M.D.; Penin, A.A. A high resolution map of the Arabidopsis thaliana developmental transcriptome based on RNA-seq profiling. Plant J. 2016, 88, 1058–1070. [Google Scholar] [CrossRef]
- Li, J.; Yu, M.; Geng, L.L.; Zhao, J. The fasciclin-like arabinogalactan protein gene, FLA3, is involved in microspore development of Arabidopsis. Plant J. 2010, 64, 482–497. [Google Scholar] [CrossRef] [PubMed]
- Basu, D.; Tian, L.; Debrosse, T.; Poirier, E.; Emch, K.; Herock, H.; Travers, A.; Showalter, A.M. Glycosylation of a Fasciclin-Like Arabinogalactan-Protein (SOS5) Mediates Root Growth and Seed Mucilage Adherence Via a Cell Wall Receptor-Like Kinase (FEI1/FEI2) Pathway in Arabidopsis. PLoS ONE 2016, 11, e0145092. [Google Scholar] [CrossRef]
- Johnson, K.L.; Kibble, N.A.; Bacic, A.; Schultz, C.J. A Fasciclin-Like Arabinogalactan-Protein (FLA) Mutant of Arabidopsis thaliana, FLA1, Shows Defects in Shoot Regeneration. PLoS ONE 2011, 6, e25154. [Google Scholar] [CrossRef]
- Cagnola, J.I.; Dumont de Chassart, G.J.; Ibarra, S.E.; Chimenti, C.; Ricardi, M.M.; Delzer, B.; Ghiglione, H.; Zhu, T.; Otegui, M.E.; Estevez, J.M.; et al. Reduced expression of selected FASCICLIN-LIKE ARABINOGALACTAN PROTEIN genes associates with the abortion of kernels in field crops of Zea mays (maize) and of Arabidopsis seeds. Plant Cell Env. 2018, 41, 661–674. [Google Scholar] [CrossRef] [PubMed]
- Turupcu, A.; Almohamed, W.; Oostenbrink, C.; Seifert, G.J. A speculation on the tandem fasciclin 1 repeat of FLA4 proteins in angiosperms. Plant Signal. Behav. 2018, 13, e1507403. [Google Scholar] [CrossRef]
- Xue, H.; Veit, C.; Abas, L.; Tryfona, T.; Maresch, D.; Ricardi, M.M.; Estevez, J.M.; Strasser, R.; Seifert, G.J. Arabidopsis thaliana FLA4 functions as a glycan-stabilized soluble factor via its carboxy-proximal fasciclin 1 domain. Plant J. 2017, 91, 613–630. [Google Scholar] [CrossRef]
- Moore, R.C.; Purugganan, M.D. The early stages of duplicate gene evolution. Proc. Natl. Acad. Sci. USA 2003, 100, 15682–15687. [Google Scholar] [CrossRef] [PubMed]
- Cao, J.; Li, X. Identification and phylogenetic analysis of late embryogenesis abundant proteins family in tomato (Solanum lycopersicum). Planta 2015, 241, 757–772. [Google Scholar] [CrossRef]
- Kong, H.; Landherr, L.L.; Frohlich, M.W.; Leebens-Mack, J.; Ma, H.; dePamphilis, C.W. Patterns of gene duplication in the plant SKP1 gene family in angiosperms: evidence for multiple mechanisms of rapid gene birth. Plant J. 2007, 50, 873–885. [Google Scholar] [CrossRef] [PubMed]
- Cannon, S.B.; Mitra, A.; Baumgarten, A.; Young, N.D.; May, G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. Bmc Plant Biol. 2004, 4, 10. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Wu, N.; Song, W.; Yin, G.; Qin, Y.; Yan, Y.; Hu, Y. Soybean (Glycine max) expansin gene superfamily origins: segmental and tandem duplication events followed by divergent selection among subfamilies. Bmc Plant Biol. 2014, 14, 93. [Google Scholar] [CrossRef] [PubMed]
- Cao, J.; Li, X.; Lv, Y.; Ding, L. Comparative analysis of the phytocyanin gene family in 10 plant species: A focus on Zea mays. Front. Plant Sci. 2015, 6, 515. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Mistry, J.; Mitchell, A.L.; Potter, S.C.; Punta, M.; Qureshi, M.; Sangrador-Vegas, A.; et al. The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 2016, 44, D279–D285. [Google Scholar] [CrossRef]
- Eddy, S.R. Accelerated profile HMM searches. Plos Comput. Biol. 2011, 7, e1002195. [Google Scholar] [CrossRef]
- Petersen, T.N.; Brunak, S.; von Heijne, G.; Nielsen, H. SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat. Methods 2011, 8, 785–786. [Google Scholar] [CrossRef] [PubMed]
- Eisenhaber, B.; Wildpaner, M.; Schultz, C.J.; Borner, G.H.; Dupree, P.; Eisenhaber, F. Glycosylphosphatidylinositol Lipid Anchoring of Plant Proteins. Sensitive Prediction from Sequence- and Genome-wide Studies for Arabidopsis and Rice. Plant Physiol. 2003, 133, 1691–1701. [Google Scholar] [CrossRef] [PubMed]
- Nicholas, K.B.; Nicholas, H.B., Jr.; Deerfield, D.W., II. GeneDoc: Analysis and visualization of genetic variation. Embnew 1997, 4, 1–4. [Google Scholar]
- Nei, M.; Kumar, S. Molecular Evolution and Phylogenetics; Oxford University Press: New York, NY, USA, 2000. [Google Scholar]
- Lee, T.H.; Tang, H.; Wang, X.; Paterson, A.H. PGDD: a database of gene and genome duplication in plants. Nucleic Acids Res. 2013, 41, D1152–D1158. [Google Scholar] [CrossRef]
- Tang, H.; Bowers, J.E.; Wang, X.; Ming, R.; Alam, M.; Paterson, A.H. Synteny and Collinearity in Plant Genomes. Science 2008, 320, 486–488. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.; Wang, X.; Bowers, J.E.; Ming, R.; Alam, M.; Paterson, A.H. Unraveling ancient hexaploidy through multiply-aligned angiosperm gene maps. Genome Res. 2008, 18, 1944–1954. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.; Bowers, J.E.; Wang, X.; Paterson, A.H. Angiosperm genome comparisons reveal early polyploidy in the monocot lineage. Proc. Natl. Acad. Sci. USA 2010, 107, 472–477. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Zhang, Y.; Zhang, Z.; Zhu, J.; Yu, J. KaKs_Calculator 2.0: A Toolkit Incorporating Gamma-Series Methods and Sliding Window Strategies. Genom. Proteom. Bioinform. 2010, 8, 77–80. [Google Scholar] [CrossRef]


| Lineage | Organism | Genome Size (Mb) | No. of Predicted Genes | No. of FLA Genes | Reference |
|---|---|---|---|---|---|
| Red algae | Chondrus crispus | 104.98 | 9843 | 1 | This study |
| Green algae | Chlamydomonas reinhardtii | 120.405 | 14,488 | 11 | This study |
| Chara braunii | 1751.21 | 35,424 | 24 | This study | |
| Liverworts | Marchantia polymorpha | 215.739 | 19,287 | 14 | This study |
| Mosses | Physcomitrella patens | 472.081 | 23,733 | 12 | This study |
| Lycophytes | Selaginella moellendorffii | 212.315 | 34,782 | 9 | This study |
| Gymnosperm | Picea abies | 19,600 | 28,354 | 24 | This study |
| Amborellales | Amborella trichopoda | 706.495 | 19,354 | 12 | This study |
| Eudicots | Arabidopsis thaliana | 119.148 | 38,093 | 22 | Schultz et al. [21] |
| Eucalyptus grandis | 691.43 | 45,226 | 19 | MacMillan et al. [13] | |
| Populus trichocarpa | 434.29 | 37,197 | 49 | Showalter et al. [24] | |
| Monocots | Brachypodium distachyon | 218.345 | 34,310 | 23 | This study |
| Oryza sativa | 374.423 | 33,185 | 26 | Ma and Zhao [12] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
He, J.; Zhao, H.; Cheng, Z.; Ke, Y.; Liu, J.; Ma, H. Evolution Analysis of the Fasciclin-Like Arabinogalactan Proteins in Plants Shows Variable Fasciclin-AGP Domain Constitutions. Int. J. Mol. Sci. 2019, 20, 1945. https://doi.org/10.3390/ijms20081945
He J, Zhao H, Cheng Z, Ke Y, Liu J, Ma H. Evolution Analysis of the Fasciclin-Like Arabinogalactan Proteins in Plants Shows Variable Fasciclin-AGP Domain Constitutions. International Journal of Molecular Sciences. 2019; 20(8):1945. https://doi.org/10.3390/ijms20081945
Chicago/Turabian StyleHe, Jiadai, Hua Zhao, Zhilu Cheng, Yuwei Ke, Jiaxi Liu, and Haoli Ma. 2019. "Evolution Analysis of the Fasciclin-Like Arabinogalactan Proteins in Plants Shows Variable Fasciclin-AGP Domain Constitutions" International Journal of Molecular Sciences 20, no. 8: 1945. https://doi.org/10.3390/ijms20081945
APA StyleHe, J., Zhao, H., Cheng, Z., Ke, Y., Liu, J., & Ma, H. (2019). Evolution Analysis of the Fasciclin-Like Arabinogalactan Proteins in Plants Shows Variable Fasciclin-AGP Domain Constitutions. International Journal of Molecular Sciences, 20(8), 1945. https://doi.org/10.3390/ijms20081945





