HER2 Heterogeneity Is Associated with Poor Survival in HER2-Positive Breast Cancer
Abstract
1. Introduction
2. Results
2.1. Mouse Model
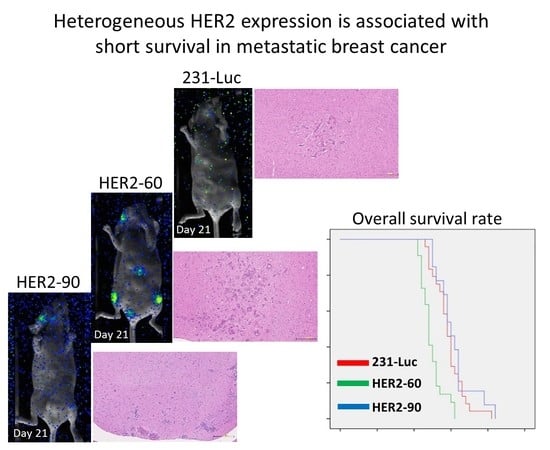
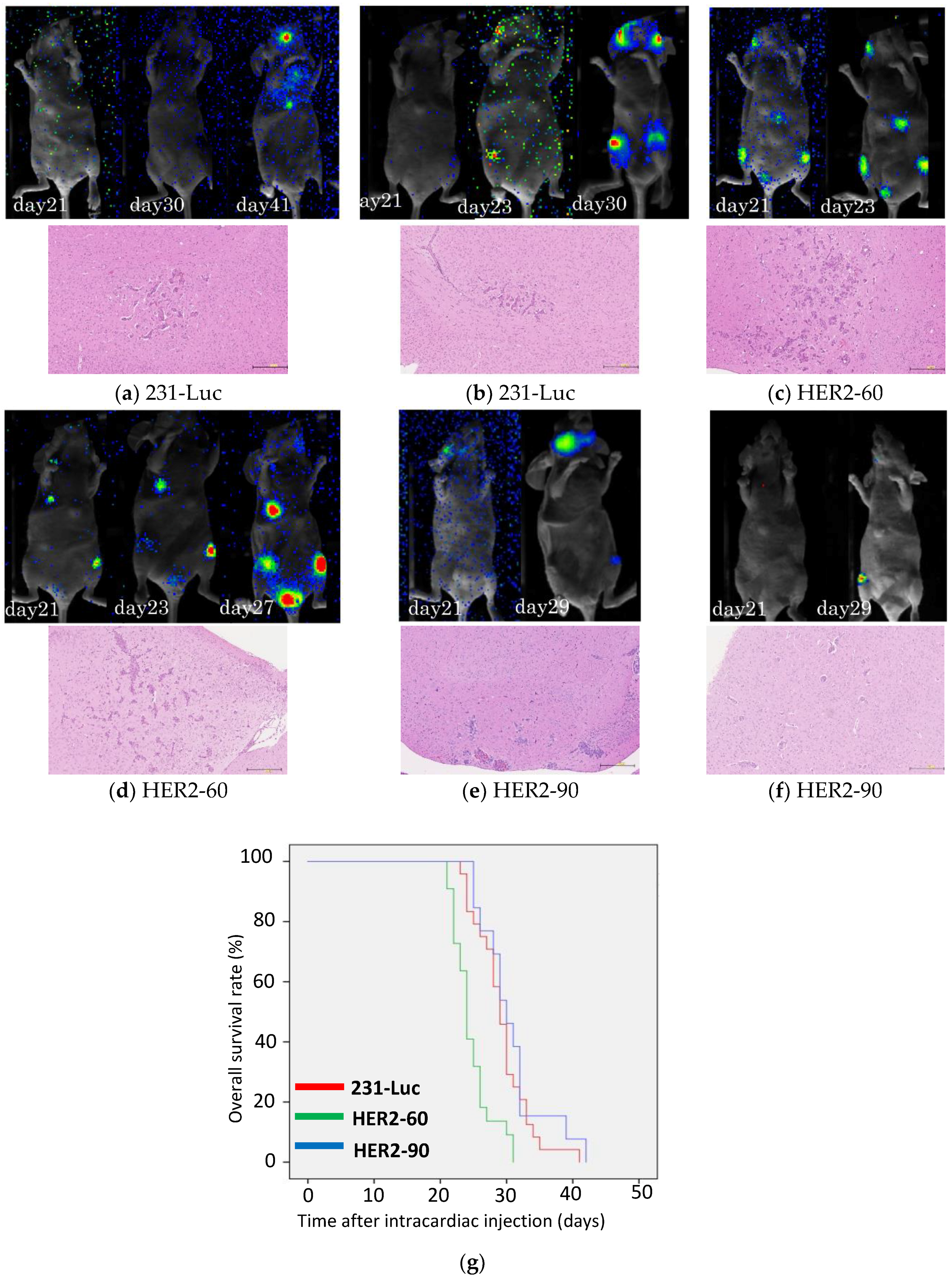
2.1.1. Heterogeneous HER2 Expression Is Associated with Short Survival in Metastatic Breast Cancer
2.1.2. HER2-60 Brain Metastasis Showed Low Expression of Caspase-1 on RNA Sequence Analysis
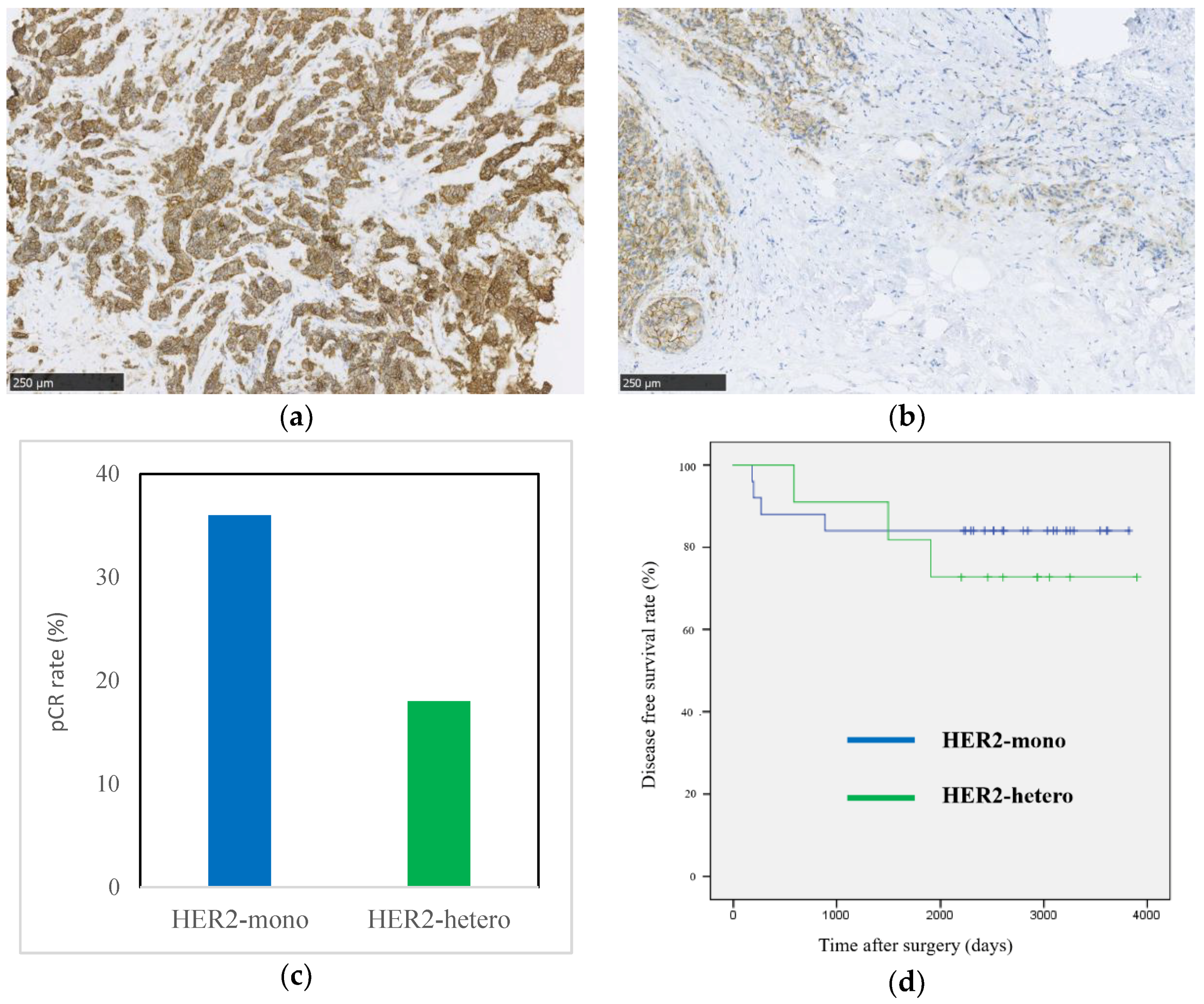
2.2. Clinical Study
3. Discussion
4. Materials and Methods
4.1. Basic Study
4.1.1. Cell Culture and Establishment of HER2-60 and HER2-90 Cell Lines
4.1.2. Brain Metastasis Model
4.1.3. Bioluminescence Imaging
4.1.4. Selecting Samples for RNA Sequence Analysis
4.1.5. RNA Sequence
4.1.6. Real Time RT-PCR
4.2. Clinical Study
4.2.1. Definition of HER2 Genetic Heterogeneity
4.2.2. Classification of HER2-positive Breast Cancer with or without HER2 Heterogeneity by IHC Staining
4.3. Survival Analysis
4.4. Ethics Statement
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| HER2 | human epidermal growth factor receptor 2 |
| CEP17 | centromere on chromosome 17 |
| NLRs | nucleotide oligomerization domain (NOD)-like receptors |
| AIM2 | absent in melanoma |
| NLRP3 | NLR family pyrin domain-containing protein 3 |
| NLRC4 | NLR family caspase activation and recruitment domain (CARD) domain-containing protein 4 |
| PYCARD | apoptosis-associated speck-like protein containing a CARD |
| IHC | immunohistochemistry |
| DFS | disease free survival |
| ASCO/CAP | American society of clinical oncology and College of American pathologist |
| OS | overall survival |
References
- Gerlinger, M.; Rowan, A.J.; Horswell, S.; Larkin, J.; Stewart, A.; Tarpey, P.; Varela, I. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 2012, 366, 883–892. [Google Scholar] [CrossRef] [PubMed]
- Shackleton, M.; Quintana, E.; Fearon, E.R.; Morrison, S.J. Heterogeneity in cancer: Cancer stem cells versus clonal evolution. Cell 2009, 138, 822–829. [Google Scholar] [CrossRef] [PubMed]
- Huang, S. Non-genetic heterogeneity of cells in development: More than just noise. Development 2009, 136, 3853–3862. [Google Scholar] [CrossRef] [PubMed]
- Shah, S.P.; Morin, R.D.; Khattra, J.; Prentice, L.; Pugh, T.; Burleigh, A.; Delaney, A.; Gelmon, K.; Guliany, R.; Senz, J.; et al. Mutational evolution in a lobular breast tumour profiled at single nucleotide resolution. Nature 2009, 461, 809–813. [Google Scholar] [CrossRef] [PubMed]
- Park, S.Y.; Gonen, M.; Kim, H.J.; Michor, F.; Polyak, K. Cellular and genetic diversity in the progression of in situ human breast carcinomas to an invasive phenotype. J. Clin. Investig. 2010, 120, 636–644. [Google Scholar] [CrossRef] [PubMed]
- Seol, H.; Lee, H.J.; Choi, Y.; Lee, H.E.; Kim, Y.J.; Kim, J.H.; Kang, E.; Kim, S.W.; Park, S.Y. Intratumoral heterogeneity of HER2 gene amplification in breast cancer: Its clinicopathological significance. Mod. Pathol. 2012, 25, 938–948. [Google Scholar] [CrossRef] [PubMed]
- Glockner, S.; Buurman, H.; Kleeburger, W.; Lehmann, U.; Kreipe, H. Marked intratumoral heterogeneity of c-myc and CyclinD1 but not c-erbB2 amplification in breast cancer. Lab. Investig. 2002, 82, 1419–1426. [Google Scholar] [CrossRef] [PubMed]
- Bartlett, A.I.; Starcyznski, J.; Robson, T.; MacLellan, A.; Campbell, F.M.; Van de Velde, C.J.; Hasenburg, A.; Markopoulos, C.; Seynaeve, C.; Rea, D.; et al. Heterogeneous HER2 gene amplification. Impact on patient outcome and a clinical relevant definition. Am. J. Clin. Pathol. 2011, 136, 266–274. [Google Scholar] [CrossRef] [PubMed]
- Hosonaga, M.; Arima, Y.; Sugihara, E.; Kohno, N.; Saya, H. Expression of CD24 is associated with HER2 expression and supports HER2-Akt signaling in HER2-positive breast cancer. Cancer Sci. 2014, 105, 779–787. [Google Scholar] [CrossRef] [PubMed]
- Sato, R.; Nakano, T.; Hosonaga, M.; Sampetrean, O.; Harigai, R.; Sasaki, T.; Koya, I.; Okano, H.; Kudoh, J.; Saya, H.; et al. RNA sequencing analysis reveals interactions between breast cancer or melanoma cells and the tissue microenvironment during brain metastasis. BioMed. Res. 2017, 2017. [Google Scholar] [CrossRef] [PubMed]
- Brennan, M.A.; Cookson, B.T. Salmonella induces macrophages death by caspase-1-dependent necrosis. Mol. Microbiol. 2000, 38, 31–40. [Google Scholar] [CrossRef] [PubMed]
- Fantuzzi, G.; Dinarello, C.A. Interleukin-18 and interleukin-1β: Two cytokines substrates for ICE (caspase-1). J. Clin. Immunol. 1999, 19, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Walsh, J.G.; Muruve, D.A.; Power, C. Inflammasomes in the CNS. Nat. Rev. Neurosci. 2014, 15, 84–97. [Google Scholar] [CrossRef] [PubMed]
- Vance, G.H.; Barry, T.S.; Bloom, K.J.; Fitzgibbons, P.L.; Hicks, D.G.; Jenkins, R.B.; Persons, D.L.; Tubbs, R.R.; Hammond, M.E.H. Genetic heterogeneity in HER2 testing in breast cancer. Arch. Pathol. Lab. Med. 2009, 133, 611–612. [Google Scholar] [PubMed]
- Brunelli, M.; Manfrin, E.; Martignoni, G.; Miller, K.; Remo, A.; Reghellin, D.; Bersani, S.; Gobbo, S.; Eccher, A.; Chilosi, M.; et al. Genotypic intratumoral heterogeneity in breast carcinoma with HER2/neu amplification: Evaluation according to ASCO/CAP criteria. Am. J. Clin. Pathol. 2009, 131, 678–682. [Google Scholar] [CrossRef] [PubMed]
- Yoon, H.H.; Shi, Q.; Sukov, W.R.; Lewis, M.A.; Sattler, C.A.; Wiktor, A.E.; Wu, T.T.; Diasio, R.B.; Jenkins, R.B.; Sinicrope, F.A. Adverse prognostic impact of intratumor heterogeneous HER2 gene amplification in patients with esophageal adenocarcinoma. J. Clin. Oncol. 2012, 30, 3932–3938. [Google Scholar] [CrossRef] [PubMed]
- Carey, L.A.; Berry, D.A.; Cirrincione, C.T.; Barry, W.T.; Pitcher, B.N. Molecular heterogeneity and response to human epidermal growth factor receptor 2 targeted GALGB 40601, a randomized phase III trial of paclitaxel plus trastuzumab with or without lapatinib. J. Clin. Oncol. 2016, 34, 542–549. [Google Scholar] [CrossRef] [PubMed]
- Winter, R.N.; Kramer, A.; Borkowski, A.; Kyprianou, N. Loss of caspase-1 and caspase-3 protein expression in human prostate cancer. Cancer Res. 2001, 61, 122–132. [Google Scholar]
- Hu, B.; Elinav, E.; Huber, S.; Booth, C.J.; Strowig, T.; Jin, C.; Eisenbarth, S.C.; Flavell, R.A. Inflammation-induced tumorigenesis in the colon is regulated by caspase-1 and NLRC4. Proc. Natl. Acad. Sci. USA 2010, 107, 21635–21640. [Google Scholar] [CrossRef] [PubMed]
- Jee, C.D.; Lee, H.S.; Bae, S.I.; Yang, H.K.; Lee, Y.M.; Rho, M.S.; Kim, W.H. Loss of caspase-1 gene expression in human gastric carcinomas and cell lines. Int. J. Oncol. 2005, 26, 1265–1271. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Guo, Y. Expression of caspase-1 in breast cancer tissues and its effects on cell proliferation, apoptosis and invasion. Oncol. Lett. 2018, 15, 6431–6435. [Google Scholar] [CrossRef] [PubMed]
- Ramos, H.J.; Lanteri, M.C.; Blahnik, G.; Negash, A.; Suthar, M.S.; Brassil, M.M.; Sodhi, K.; Treuting, P.M.; Busch, M.P.; Norris, P.J. IL-1β signaling promotes CNS-intrinsic immune control of West Nile virus infection. PLoS Pathog. 2012, 8, e1003039. [Google Scholar] [CrossRef] [PubMed]
- Jamilloux, Y.; Pierini, R.; Querenet, M.; Juruj, C.; Fauchais, A.L.; Jauberteau, M.O.; Jarraud, S.; Lina, G.; Etienne, J.; Roy, C.R. Inflammasome activation restricts Legionella pneumophila replication in primary microglial cells through flagellin detection. Glia 2013, 61, 539–549. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef] [PubMed]
- Komura, D.; Isagawa, T.; Kishi, K.; Suzuki, R.; Sato, R.; Tanaka, M.; Katoh, H.; Yamamoto, S.; Tatsuno, K.; Fukayama, M.; et al. CASTIN: A system for comprehensive analysis of cancer-stromal interactome. BMC Genom. 2016, 17, 899. [Google Scholar] [CrossRef] [PubMed]
- Curtis, C.; Shah, S.P.; Chin, S.F.; Turashvili, G.; Rueda, O.M.; Dunning, M.J.; Speed, D.; Lynch, A.G.; Samarajiwa, S.; Yuan, Y.; et al. The genomic and transcriptomic architecture of 2000 breast tumours reveals novel subgroups. Nature 2012, 486, 346–352. [Google Scholar] [CrossRef] [PubMed]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio Cancer Genomics Portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012, 2, 401–404. [PubMed]
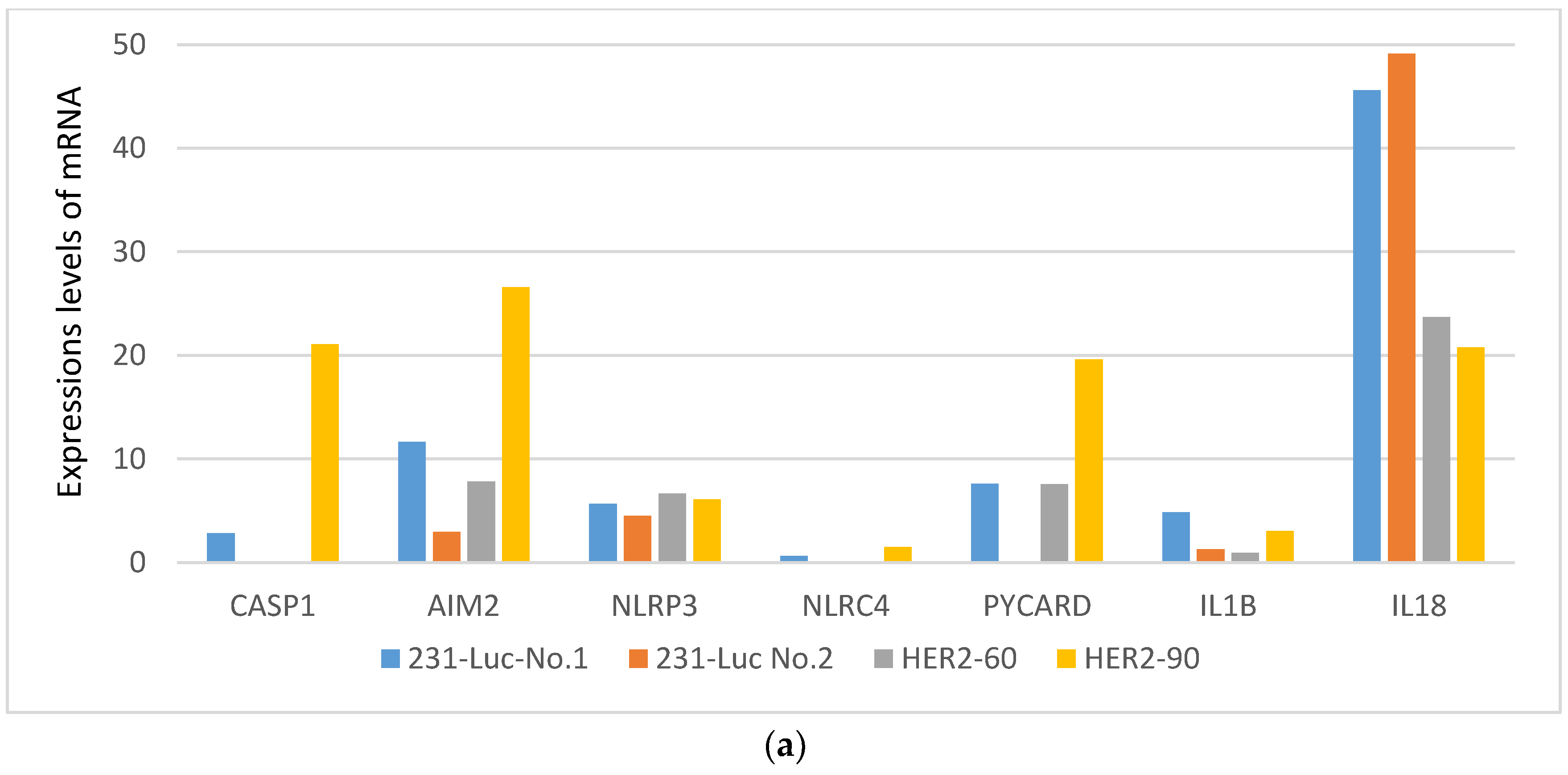
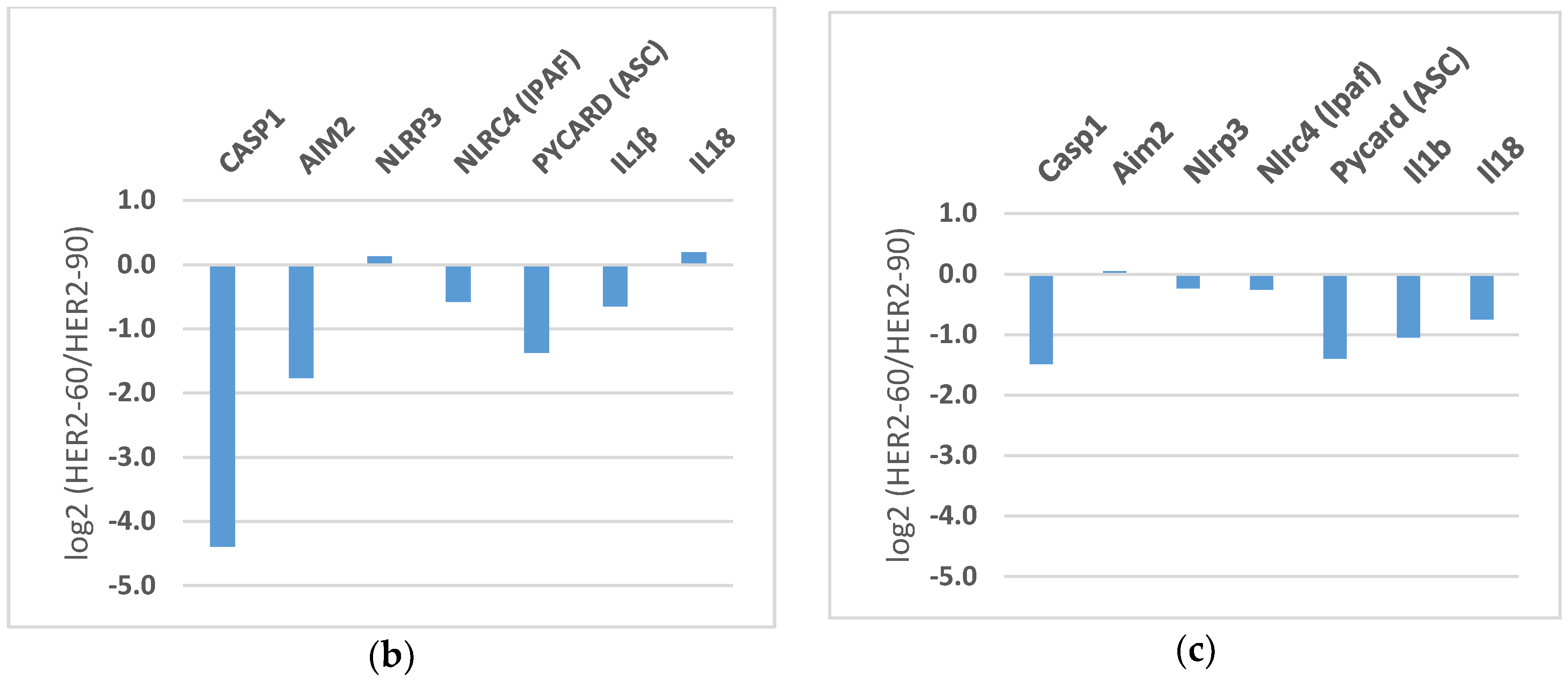
| Gene Symbol | 231-Luc No. 1 | 231-Luc No. 2 | HER2-60 | HER2-90 |
|---|---|---|---|---|
| CASP1 | 2.83 | 0 | 0 | 21.07 |
| AIM2 | 11.62 | 2.96 | 7.80 | 26.59 |
| NLRP3 | 5.65 | 4.51 | 6.67 | 6.10 |
| NLRC4 | 0.61 | 0 | 0 | 1.49 |
| PYCARD | 7.59 | 0 | 7.56 | 19.61 |
| IL1B | 4.85 | 1.28 | 0.95 | 3.06 |
| IL18 | 45.61 | 49.12 | 23.71 | 20.75 |
| Primer | Forward | Reverse |
|---|---|---|
| mouse Gapdh | 5-ACTAACATCAAATGGGGTGAGGCC-3 | 5-GGATGCATTGCTGACAATCTTGAGTGA-3 |
| human GAPDH | 5-CAAAATCAAGTGGGGCGATGCTGGC-3 | 5-GGCATTGCTGATGATCTTGAGGCT-3 |
| CASP1 | 5-TCCCTAGAAGAAGCTCAAAGGATATG-3 | 5-CGTGTGCGGCTTGACTTG-3 |
| AIM2 | 5-CAGAAATGATGTCGCAAAGCA-3 | 5-TCAGTACCATAACTGGCAAACAG-3 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hosonaga, M.; Arima, Y.; Sampetrean, O.; Komura, D.; Koya, I.; Sasaki, T.; Sato, E.; Okano, H.; Kudoh, J.; Ishikawa, S.; et al. HER2 Heterogeneity Is Associated with Poor Survival in HER2-Positive Breast Cancer. Int. J. Mol. Sci. 2018, 19, 2158. https://doi.org/10.3390/ijms19082158
Hosonaga M, Arima Y, Sampetrean O, Komura D, Koya I, Sasaki T, Sato E, Okano H, Kudoh J, Ishikawa S, et al. HER2 Heterogeneity Is Associated with Poor Survival in HER2-Positive Breast Cancer. International Journal of Molecular Sciences. 2018; 19(8):2158. https://doi.org/10.3390/ijms19082158
Chicago/Turabian StyleHosonaga, Mari, Yoshimi Arima, Oltea Sampetrean, Daisuke Komura, Ikuko Koya, Takashi Sasaki, Eiichi Sato, Hideyuki Okano, Jun Kudoh, Shumpei Ishikawa, and et al. 2018. "HER2 Heterogeneity Is Associated with Poor Survival in HER2-Positive Breast Cancer" International Journal of Molecular Sciences 19, no. 8: 2158. https://doi.org/10.3390/ijms19082158
APA StyleHosonaga, M., Arima, Y., Sampetrean, O., Komura, D., Koya, I., Sasaki, T., Sato, E., Okano, H., Kudoh, J., Ishikawa, S., Saya, H., & Ishikawa, T. (2018). HER2 Heterogeneity Is Associated with Poor Survival in HER2-Positive Breast Cancer. International Journal of Molecular Sciences, 19(8), 2158. https://doi.org/10.3390/ijms19082158