Development of Analytical Strategies for the Determination of Olive Fruit Bioactive Compounds Using UPLC-HRMS and HPLC-DAD. Chemical Characterization of Kolovi Lesvos Variety as a Case Study
Abstract
1. Introduction
2. Results and Discussion
2.1. Determination of Phenolic Compounds by LC-QTOF-MS
2.1.1. Sample Preparation Optimization
Sample Pretreatment
Selection of Extraction Solvent
Selection of Purification Step
2.1.2. Method Validation
2.1.3. Samples Results
Target Screening
Suspect Screening
2.2. Determination of Pigments, Tocopherols, Squalene by HPLC-DAD
2.2.1. Method Validation
2.2.2. Samples Results
Pigments, Tocopherols, Squalene
3. Materials and Methods
3.1. Chemicals and Standards
3.1.1. Chemicals and Standards for Phenolic Analysis
3.1.2. Chemicals and Standards for Pigment, Tocopherols and Squalene Analysis
3.2. Olive Fruit Samples
3.3. Phenolic Compounds Determination
3.3.1. Sample Preparation
Method Optimization
Final Sample Preparation Protocol
3.3.2. LC-HRMS Methodology
3.3.3. Method Validation
3.3.4. Screening Strategies
Target Screening
Suspect Screening
3.4. Pigment, Tocopherol and Squalene Determination
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Sample Availability
References
- Boskou, G. Antioxidant capacity and phenolic profile of table olives from the Greek market. Food Chem. 2010, 94, 925–934. [Google Scholar] [CrossRef]
- Brahmi, F.; Mechri, B.; Dhibi, M.; Hammami, M. Variations in phenolic compounds and antiradical scavenging activity of Olea europaea leaves and fruits extracts collected in two different seasons. Ind. Crops Prod. 2013, 49, 256–264. [Google Scholar] [CrossRef]
- Ghanbari, R.; Anwar, F.; Alkharfy, K.M.; Gilani, A.H.; Saari, N. Valuable nutrients and functional bioactives in different parts of olive (Olea europaea L.)—A review. Int. J. Mol. Sci. 2012, 13, 3291–3340. [Google Scholar] [CrossRef]
- Savarese, M.; Demarco, E.; Sacchi, R. Characterization of phenolic extracts from olives (Olea europaea cv. Pisciottana) by electrospray ionization mass spectrometry. Food Chem. 2007, 105, 761–770. [Google Scholar] [CrossRef]
- European Commission. COMMISSION REGULATION (EU) No 432/2012. Establishing a list of permitted health claims made on foods, other than those referring to the reduction of disease risk and to children’s development and health. Official J. Eur. Union 2012, 43, 281–320. [Google Scholar]
- Baiano, A.; Gambacorta, G.; Terracone, C.; Previtali, M.A.; La Notte, E. Characteristics of drupes, phenolic content and antioxidant capacity of Italian olive fruits. J. Food Lipids 2009, 16, 209–226. [Google Scholar] [CrossRef]
- Blekas, G.; Vassilakis, C.; Harizanis, C.; Tsimidou, M.; Boskou, D.G. Biophenols in table olives. J. Agric. Food Chem. 2002, 50, 3688–3692. [Google Scholar] [CrossRef] [PubMed]
- Crawford, L.M.; Holstege, D.M.; Wang, S.C. High-throughput extraction method for phenolic compounds in olive fruit (Olea europaea). J. Food Compost. Anal. 2018, 66, 136–144. [Google Scholar] [CrossRef]
- Durante, M.; Tufariello, M.; Tommasi, L.; Lenucci, M.S.; Bleve, G.; Mita, G. Evaluation of bioactive compounds in black table olives fermented with selected microbial starters. J. Sci. Food Agric. 2018, 98, 96–103. [Google Scholar] [CrossRef] [PubMed]
- Romero, C.; García, P.; Brenes, M.; García, A.; Garrido, A. Phenolic compounds in natural black Spanish olive varieties. Eur. Food Res. Technol. 2002, 215, 489–496. [Google Scholar] [CrossRef]
- Trapani, S.; Breschi, C.; Cecchi, L.; Guerrini, L.; Mulinacci, N.; Parenti, A.; Canuti, V.; Picchi, M.; Caruso, G.; Gucci, R.; et al. Indirect indices of oxidative damage to phenolic compounds for the implementation of olive paste malaxation optimization charts. J. Food Eng. 2017, 207, 24–34. [Google Scholar] [CrossRef]
- Peng, L.Q.; Li, Q.; Chang, Y.X.; An, M.; Yang, R.; Tan, Z.; Hao, J.; Cao, J.; Xu, J.J.; Hu, S.S. Determination of natural phenols in olive fruits by chitosan assisted matrix solid-phase dispersion microextraction and ultrahigh performance liquid chromatography with quadrupole time-of-flight tandem mass spectrometry. J. Chromatogr. A 2016, 1456, 68–76. [Google Scholar] [CrossRef]
- Olmo-Garcia, L.; Kessler, N.; Neuweger, H.; Wendt, K.; Olmo-Peinado, J.M.; Fernandez-Gutierrez, A.; Baessmann, C.; Carrasco-Pancorbo, A. Unravelling the distribution of secondary metabolites in Olea Europaea L.: Exhaustive characterization of eight olive-tree derived matrices by complementary platforms (LC-ESI/APCI-MS and GC-APCI-MS). Molecules 2018, 23, 2419. [Google Scholar] [CrossRef] [PubMed]
- Perez, A.G.; Leon, L.; Sanz, C.; de la Rosa, R. Fruit phenolic profiling: A new selection criterion in olive breeding programs. Front. Plant Sci. 2018, 9, 241. [Google Scholar] [CrossRef] [PubMed]
- Aprile, A.; Negro, C.; Sabella, E.; Luvisi, A.; Nicoli, F.; Nutricati, E.; Vergine, M.; Miceli, A.; Blando, F.; De Bellis, L. Antioxidant activity and anthocyanin contents in olives (cv Cellina di Nardo) during ripening and after fermentation. Antioxidants 2019, 8, 138. [Google Scholar] [CrossRef] [PubMed]
- Klen, T.J.; Wondra, A.G.; Vrhovsek, U.; Vodopivec, B.M. Phenolic profiling of olives and olive oil process-derived matrices using UPLC-DAD-ESI-QTOF-HRMS analysis. J. Agric. Food Chem. 2015, 63, 3859–3872. [Google Scholar] [CrossRef] [PubMed]
- Lucci, P.; Saurina, J.; Núñez, O. Trends in LC-MS and LC-HRMS analysis and characterization of polyphenols in food. TrAC Trends Anal. Chem. 2017, 88, 1–24. [Google Scholar] [CrossRef]
- Ben Othman, N.; Roblain, D.; Thonart, P.; Hamdi, M. Tunisian table olive phenolic compounds and their antioxidant capacity. J. Food Sci. 2008, 73, C235–C240. [Google Scholar] [CrossRef]
- Dagdelen, A.; Tumen, G.; Ozcan, M.M.; Dundar, E. Phenolics profiles of olive fruits (Olea europaea L.) and oils from Ayvalik, Domat and Gemlik varieties at different ripening stages. Food Chem. 2013, 136, 41–45. [Google Scholar] [CrossRef]
- Pereira, J.A.; Pereira, A.P.; Ferreira, I.C.; Valentao, P.; Andrade, P.B.; Seabra, R.; Estevinho, L.; Bento, A. Table olives from Portugal: Phenolic compounds, antioxidant potential, and antimicrobial activity. J. Agric. Food Chem. 2006, 54, 8425–8431. [Google Scholar] [CrossRef]
- Arslan, D.; Özcan, M.M. Phenolic profile and antioxidant activity of olive fruits of the Turkish variety “Sarıulak” from different locations. Grasas Aceites 2011, 62, 453–461. [Google Scholar] [CrossRef]
- Falk, J.; Munne-Bosch, S. Tocochromanol functions in plants: Antioxidation and beyond. J. Exp. Bot. 2010, 61, 1549–1566. [Google Scholar] [CrossRef] [PubMed]
- Grigoriadou, D.; Androulaki, A.; Psomiadou, E.; Tsimidou, M. Solid phase extraction in the analysis of squalene and tocopherols in olive oil. Food Chem. 2007, 105, 675–680. [Google Scholar] [CrossRef]
- Martinez-Beamonte, R.; Sanclemente, T.; Surra, J.C.; Osada, J. Could squalene be an added value to use olive by-products? J. Sci. Food Agric. 2020, 100, 915–925. [Google Scholar] [CrossRef] [PubMed]
- Ramírez-Torres, A.; Gabás-Rivera, C.; Osada, J. Squalene: A trove of metabolic actions. Products Olive Tree 2016, 1. [Google Scholar] [CrossRef]
- Gandul-Rojas, B.; Gallardo-Guerrero, L. Pigment changes during preservation of green table olive specialities treated with alkali and without fermentation: Effect of thermal treatments and storage conditions. Food Res. Int. 2018, 108, 57–67. [Google Scholar] [CrossRef]
- Gandul-Rojas, B.; Roca, M.; Gallardo-Guerrero, L. Chlorophylls and carotenoids in food products from olive tree. Products Olive Tree 2016, 1. [Google Scholar] [CrossRef]
- Buscemi, S.; Corleo, D.; Di Pace, F.; Petroni, M.L.; Satriano, A.; Marchesini, G. The effect of lutein on eye and extra-eye health. Nutrients 2018, 10, 1321. [Google Scholar] [CrossRef]
- Eroglu, A.; Harrison, E.H. Carotenoid metabolism in mammals, including man: Formation, occurrence, and function of apocarotenoids. J. Lipid Res. 2013, 54, 1719–1730. [Google Scholar] [CrossRef] [PubMed]
- Minguez-Mosquera, M.I.; Gallardo-Guerrero, L. Anomalous transformation of chloroplastic pigments in Gordal variety olives during processing for table olives. J. Food Prot. 1995, 58, 1241–1248. [Google Scholar] [CrossRef] [PubMed]
- López, A.; Montaño, A.; Garrido, A. Provitamin A carotenoids in table olives according to processing styles, cultivars, and commercial presentations. Eur. Food Res. Technol. 2005, 221, 406–411. [Google Scholar] [CrossRef]
- Mastralexi, A.; Mantzouridou, F.T.; Tsimidou, M.Z. Evolution of safety and other quality parameters of the Greek PDO table olives “Prasines Elies Chalkidikis” during industrial scale processing and storage. Eur. J. Lipid Sci. Technol. 2018, 121, 1800171. [Google Scholar] [CrossRef]
- Montaño, A.; Casado, F.J.; de Castro, A.; Sánchez, A.H.; Rejano, L. Influence of processing, storage time, and pasteurisation upon the tocopherol and amino acid contents of treated green table olives. Eur. Food Res. Technol. 2004, 220, 255–260. [Google Scholar] [CrossRef]
- Ramirez, E.; Gandul-Rojas, B.; Romero, C.; Brenes, M.; Gallardo-Guerrero, L. Composition of pigments and colour changes in green table olives related to processing type. Food Chem. 2015, 166, 115–124. [Google Scholar] [CrossRef] [PubMed]
- Sagratini, G.; Allegrini, M.; Caprioli, G.; Cristalli, G.; Giardina, D.; Maggi, F.; Ricciutelli, M.; Sirocchi, V.; Vittori, S. Simultaneous determination of squalene, α-tocopherol and β-carotene in table olives by solid phase extraction and high-performance liquid chromatography with diode array detection. Food Anal. Methods 2012, 6, 54–60. [Google Scholar] [CrossRef]
- Anniva, C.; Tsimidou, M.Z. On the quality control of ‘olive paste’, a specialty based on olives and olive oil. Eur. J. Lipid Sci. Technol. 2009, 111, 328–336. [Google Scholar] [CrossRef]
- Sakouhi, F.; Harrabi, S.; Absalon, C.; Sbei, K.; Boukhchina, S.; Kallel, H. alpha-Tocopherol and fatty acids contents of some Tunisian table olives (Olea europea L.): Changes in their composition during ripening and processing. Food Chem. 2008, 108, 833–839. [Google Scholar] [CrossRef] [PubMed]
- Kalogiouri, N.P.; Aalizadeh, R.; Thomaidis, N.S. Investigating the organic and conventional production type of olive oil with target and suspect screening by LC-QTOF-MS, a novel semi-quantification method using chemical similarity and advanced chemometrics. Anal. Bioanal. Chem. 2017, 409, 5413–5426. [Google Scholar] [CrossRef]
- Kalogiouri, N.P.; Aalizadeh, R.; Thomaidis, N.S. Application of an advanced and wide scope non-target screening workflow with LC-ESI-QTOF-MS and chemometrics for the classification of the Greek olive oil varieties. Food Chem. 2018, 256, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Kalogiouri, N.P.; Alygizakis, N.A.; Aalizadeh, R.; Thomaidis, N.S. Olive oil authenticity studies by target and nontarget LC-QTOF-MS combined with advanced chemometric techniques. Anal. Bioanal. Chem. 2016, 408, 7955–7970. [Google Scholar] [CrossRef]
- Martakos, I.; Kostakis, M.; Dasenaki, M.; Pentogennis, M.; Thomaidis, N. Simultaneous determination of pigments, tocopherols, and squalene in Greek olive oils: A study of the influence of cultivation and oil-production parameters. Foods 2019, 9, 31. [Google Scholar] [CrossRef]
- Kanakis, P.; Termentzi, A.; Michel, T.; Gikas, E.; Halabalaki, M.; Skaltsounis, A.L. From olive drupes to olive oil. An HPLC-orbitrap-based qualitative and quantitative exploration of olive key metabolites. Planta Med. 2013, 79, 1576–1587. [Google Scholar] [CrossRef]
- Ryan, D.; Antolovich, M.; Herlt, T.; Prenzler, P.D.; Lavee, S.; Robards, K. Identification of phenolic compounds in tissues of the novel olive cultivar hardy’s mammoth. J. Agric. Food Chem. 2002, 50, 6716–6724. [Google Scholar] [CrossRef] [PubMed]
- Ben Ghorbal, A.; Leventdurur, S.; Agirman, B.; Boyaci-Gunduz, C.P.; Kelebek, H.; Carsanba, E.; Darici, M.; Erten, H. Influence of geographic origin on agronomic traits and phenolic content of cv. Gemlik olive fruits. J. Food Compost. Anal. 2018, 74, 1–9. [Google Scholar] [CrossRef]
- Jerman, T.; Trebše, P.; Mozetič Vodopivec, B. Ultrasound-assisted solid liquid extraction (USLE) of olive fruit (Olea europaea) phenolic compounds. Food Chem. 2010, 123, 175–182. [Google Scholar] [CrossRef]
- Vinha, A.F.; Ferreres, F.; Silva, B.M.; Valentão, P.c.; Gonçalves, A.; Pereira, J.A.; Oliveira, M.B.; Seabra, R.M.; Andrade, P.B. Phenolic profiles of Portuguese olive fruits (Olea europaea L.): Influences of cultivar and geographical origin. Food Chem. 2005, 89, 561–568. [Google Scholar] [CrossRef]
- Tóth, G.; Alberti, Á.; Sólyomváry, A.; Barabás, C.; Boldizsár, I.; Noszál, B. Phenolic profiling of various olive bark-types and leaves: HPLC–ESI/MS study. Ind. Crops Prod. 2015, 67, 432–438. [Google Scholar] [CrossRef]
- Alipieva, K.; Korkina, L.; Orhan, I.E.; Georgiev, M.I. Verbascoside—A review of its occurrence, (bio)synthesis and pharmacological significance. Biotechnol. Adv. 2014, 32, 1065–1076. [Google Scholar] [CrossRef]
- Bouaziz, M.; Chamkha, M.; Sayadi, S. Comparative study on phenolic content and antioxidant activity during maturation of the olive cultivar Chemlali from Tunisia. J. Agric. Food Chem. 2004, 52, 5476–5481. [Google Scholar] [CrossRef]
- Gómez-Rico, A.; Fregapane, G.; Salvador, M.D. Effect of cultivar and ripening on minor components in Spanish olive fruits and their corresponding virgin olive oils. Food Res. Int. 2008, 41, 433–440. [Google Scholar] [CrossRef]
- Ganeshpurkar, A.; Saluja, A.K. The pharmacological potential of rutin. Saudi Pharm. J. 2017, 25, 149–164. [Google Scholar] [CrossRef] [PubMed]
- Horai, H.; Arita, M.; Kanaya, S.; Nihei, Y.; Ikeda, T.; Suwa, K.; Ojima, Y.; Tanaka, K.; Tanaka, S.; Aoshima, K.; et al. MassBank: A public repository for sharing mass spectral data for life sciences. J. Mass Spectrom. 2010, 45, 703–714. [Google Scholar] [CrossRef] [PubMed]
- Wolf, S.; Schmidt, S.; Müller-Hannemann, M.; Neumann, S. In silico fragmentation for computer assisted identification of metabolite mass spectra. BMC Bioinform. 2010, 11, 148. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, S.M.; Guyot, S.; Marnet, N.; Lopes-da-Silva, J.A.; Renard, C.M.G.C.; Coimbra, M.A. Characterisation of phenolic extracts from olive pulp and olive pomace by electrospray mass spectrometry. J. Sci. Food Agric. 2005, 85, 21–32. [Google Scholar] [CrossRef]
- Silva, S.; Gomes, L.; Leitão, F.; Coelho, A.V.; Boas, L.V. Phenolic compounds and antioxidant activity of Olea europaea L. fruits and leaves. Food Sci. Technol. Int. 2016, 12, 385–395. [Google Scholar] [CrossRef]
- Shawan, M.M.A.K.; Halder, S.K.; Hasan, M.A. Luteolin and abyssinone II as potential inhibitors of SARS-CoV-2: An in silico molecular modeling approach in battling the COVID-19 outbreak. Bull. Natl. Res. Cent. 2021, 45, 27. [Google Scholar] [CrossRef]
- Ambra, R.; Natella, F.; Bello, C.; Lucchetti, S.; Forte, V.; Pastore, G. Phenolics fate in table olives (Olea europaea L. cv. Nocellara del Belice) debittered using the Spanish and Castelvetrano methods. Food Res. Int. 2017, 100, 369–376. [Google Scholar] [CrossRef] [PubMed]
- Klen, T.J. Olive Fruit Phenols in Olive Oil Processing: The Fate and Antioxidant Potential. Ph.D. Thesis, Univeristy of Nova Gorica, Nova Gorica, Slovenia, 2014. [Google Scholar]
- Ryan, D.; Robards, K.; Lavee, S. Determination of phenolic compounds in olives by reversed-phase chromatography and mass spectrometry. J. Chromatogr. A 1999, 832, 87–96. [Google Scholar] [CrossRef]
- Deng, J.; Xu, Z.; Xiang, C.; Liu, J.; Zhou, L.; Li, T.; Yang, Z.; Ding, C. Comparative evaluation of maceration and ultrasonic-assisted extraction of phenolic compounds from fresh olives. Ultrason. Sonochem. 2017, 37, 328–334. [Google Scholar] [CrossRef] [PubMed]
- Bruno, L.; Chiappetta, A.; Muzzalupo, I.; Gagliardi, C.; Iaria, D.; Bruno, A.; Greco, M.; Giannino, D.; Perri, E.; Bitonti, M.B. Role of geranylgeranyl reductase gene in organ development and stress response in olive (Olea europaea) plants. Funct. Plant Biol. 2009, 36, 370–381. [Google Scholar] [CrossRef]
- Roca, M.; Leon, L.; de la Rosa, R. Pigment metabolism of ‘Sikitita’ olive (Olea europaea L.): A new cultivar obtained by cross-breeding. J. Agric. Food Chem. 2011, 59, 2049–2055. [Google Scholar] [CrossRef] [PubMed]
- Roca, M.; Mínguez-Mosquera, M.I. Changes in chloroplast pigments of olive varieties during fruit ripening. J. Agric. Food. Chem. 2001, 49, 832–839. [Google Scholar] [CrossRef] [PubMed]
- Bodoira, R.; Torres, M.; Pierantozzi, P.; Taticchi, A.; Servili, M.; Maestri, D. Oil biogenesis and antioxidant compounds from “Arauco” olive (Olea europaea L.) cultivar during fruit development and ripening. Eur. J. Lipid Sci. Technol. 2015, 117, 377–388. [Google Scholar] [CrossRef]
- Hassapidou, M.N.; Balatsouras, G.D.; Manoukas, A.G. Effect of processing upon the tocopherol and tocotrienol composition of table olives. Food Chem. 1994, 50, 111–114. [Google Scholar] [CrossRef]
- Georgiadou, E.C.; Goulas, V.; Ntourou, T.; Manganaris, G.A.; Kalaitzis, P.; Fotopoulos, V. Regulation of on-tree vitamin E biosynthesis in olive fruit during successive growing years: The impact of fruit development and environmental cues. Front. Plant Sci. 2016, 7, 1656. [Google Scholar] [CrossRef] [PubMed]
- Habibi, M.; Golmakani, M.T.; Mesbahi, G.; Majzoobi, M.; Farahnaky, A. Ultrasound-accelerated debittering of olive fruits. Innov. Food Sci. Emerg. Technol. 2015, 31, 105–115. [Google Scholar] [CrossRef]
- Bouaziz, M.; Grayer, R.J.; Simmonds, M.S.; Damak, M.; Sayadi, S. Identification and antioxidant potential of flavonoids and low molecular weight phenols in olive cultivar chemlali growing in Tunisia. J. Agric. Food Chem. 2005, 53, 236–241. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; Xia, W. Analysis of phenolic compounds in Chinese olive (Canarium album L.) fruit by RPHPLC–DAD–ESI–MS. Food Chem. 2007, 105, 1307–1311. [Google Scholar] [CrossRef]
- Bouaziz, M.; Jemai, H.; Khabou, W.; Sayadi, S. Oil content, phenolic profiling and antioxidant potential of Tunisian olive drupes. J. Sci. Food Agric. 2010, 90, 1750–1758. [Google Scholar] [CrossRef] [PubMed]
- Hashmi, M.A.; Khan, A.; Hanif, M.; Farooq, U.; Perveen, S. Traditional uses, phytochemistry, and pharmacology of Olea europaea (Olive). Evid.-Based Complementary Altern. Med. 2015, 2015, 541591. [Google Scholar] [CrossRef] [PubMed]
- Guodong, R.; Xiaoxia, L.; Weiwei, Z.; Wenjun, W.; Jianguo, Z. Metabolomics reveals variation and correlation among different tissues of olive (Olea europaea L.). Biol. Open 2017, 6, 1317–1323. [Google Scholar] [CrossRef] [PubMed]
- Aalizadeh, R.; Nika, M.C.; Thomaidis, N.S. Development and application of retention time prediction models in the suspect and non-target screening of emerging contaminants. J. Hazard. Mater. 2019, 363, 277–285. [Google Scholar] [CrossRef] [PubMed]
- Minguez-Mosquera, M.I.; Garrido-Fernandez, J. Chlorophyll and carotenoid presence in olive fruit (Olea europaea). J. Agric. Food. Chem. 1989, 37, 1–7. [Google Scholar] [CrossRef]
- Gliszczyńska-Świgło, A.; Sikorska, E. Simple reversed-phase liquid chromatography method for determination of tocopherols in edible plant oils. J. Chromatogr. A 2004, 1048, 195–198. [Google Scholar] [CrossRef] [PubMed]
- Magnusson, B.; Örnemark, U. (Eds.) Eurachem Guide: The Fitness for Purpose of Analytical Methods—A Laboratory Guide to Method Validation and Related Topics, 2nd ed.; Echo Library: Teddington, Middlesex, UK, 2014. [Google Scholar]

| Analytes | Kolovi Olive Fruit Samples | ||||||
|---|---|---|---|---|---|---|---|
| Sample 1 | Sample 2 | Sample 3 | Sample 4 | Sample 5 | Sample 6 | Sample 7 | |
| 1-acetoxypinoresinol | 12 | 17 | 31 | 10 | 12 | 3.9 | 5.7 |
| 10-hydroxy decarboxymethyl oleuropein aglycone | 4.9 | 23 | 2.0 | 15 | 17 | 374 | 214 |
| 10-hydroxyoleuropein aglycone | 1.8 | 2.4 | 0.99 | 1.6 | 0.50 | 0.53 | 2.1 |
| apigenin | 0.037 | 0.19 | 0.85 | ND | 0.93 | ND | ND |
| caffeic acid | 0.59 | 1.7 | 0.50 | 0.59 | 0.56 | 2.1 | 1.9 |
| elenolic acid | 17 | 45 | 22 | 15 | 23 | 30 | 24 |
| eriodictyol | 2.3 | n.d | 4.4 | 3.1 | 13 | 0.95 | 1.1 |
| hydroxytyrosol | 266 | 760 | 928 | 277 | 392 | 208 | 187 |
| lingstroside aglycone | 3.9 | 7.3 | 5.8 | ND | 5.3 | 8.2 | ND |
| luteolin | 48 | 291 | 450 | 149 | 284 | 150 | 75 |
| oleokoronal | 13 | 16 | 8.0 | 9.0 | 1.8 | 7.3 | 11 |
| oleuropein | 811 | 855 | 324 | 7.0 | 3.5 | 10 | 5.4 |
| oleuropein aglycone | 13 | 82 | 16 | 84 | 87 | 114 | 54 |
| quercetin | ND | 0.58 | ND | 0.46 | ND | 1.5 | ND |
| tyrosol | 48 | 78 | 123 | 19 | 29 | 17 | 19 |
| vanillic acid | 3.2 | ND | 4.3 | 1.0 | 1.5 | 0.38 | 0.46 |
| vanillin | ND | ND | ND | ND | ND | 3.8 | 5 |
| oleacein | 4.0 | 512 | 115 | 12 | 388 | 2447 | 434 |
| oleocanthal | 0.73 | 9.3 | 2.5 | ND | 6.2 | 24 | 7.7 |
| oleocanthalic acid | 0.36 | 4.2 | 0.59 | ND | 5.1 | 17 | 6.2 |
| oleomissional | 462 | 579 | 265 | 395 | 183 | 223 | 362 |
| p-coumaric acid | ND | 1.1 | 1.7 | 1.0 | 1.7 | 2.3 | 0.93 |
| rutin | 254 | 376 | 324 | 142 | 195 | 234 | 177 |
| salycilic acid | ND | 0.58 | ND | 0.15 | ND | ND | ND |
| verbascoside | 1255 | 9895 | 8821 | 6479 | 8970 | 6263 | |
| Compound | Molecular Formula | Ion | Mass Error (mDa) | tR Predicted (min) | tR Experimental (min) | Isotopic Fitting (mSigma) | m/z (Fragment ions) | # of Samples Detected |
|---|---|---|---|---|---|---|---|---|
| 3,4 dihydroxyphenylacetic acid | C8H8O4 | [M-H]− | 0.4 | 0.39 | 3.1 | 12 | 123.0439 | 7 |
| beta hydroxyacteoside | C29H36O16 | [M-H]− | 0.8 | 7.79 | 4.3 | 50 | 161.0243; 179.0349; 459.1505; 529.156; 621.1807 | 7 |
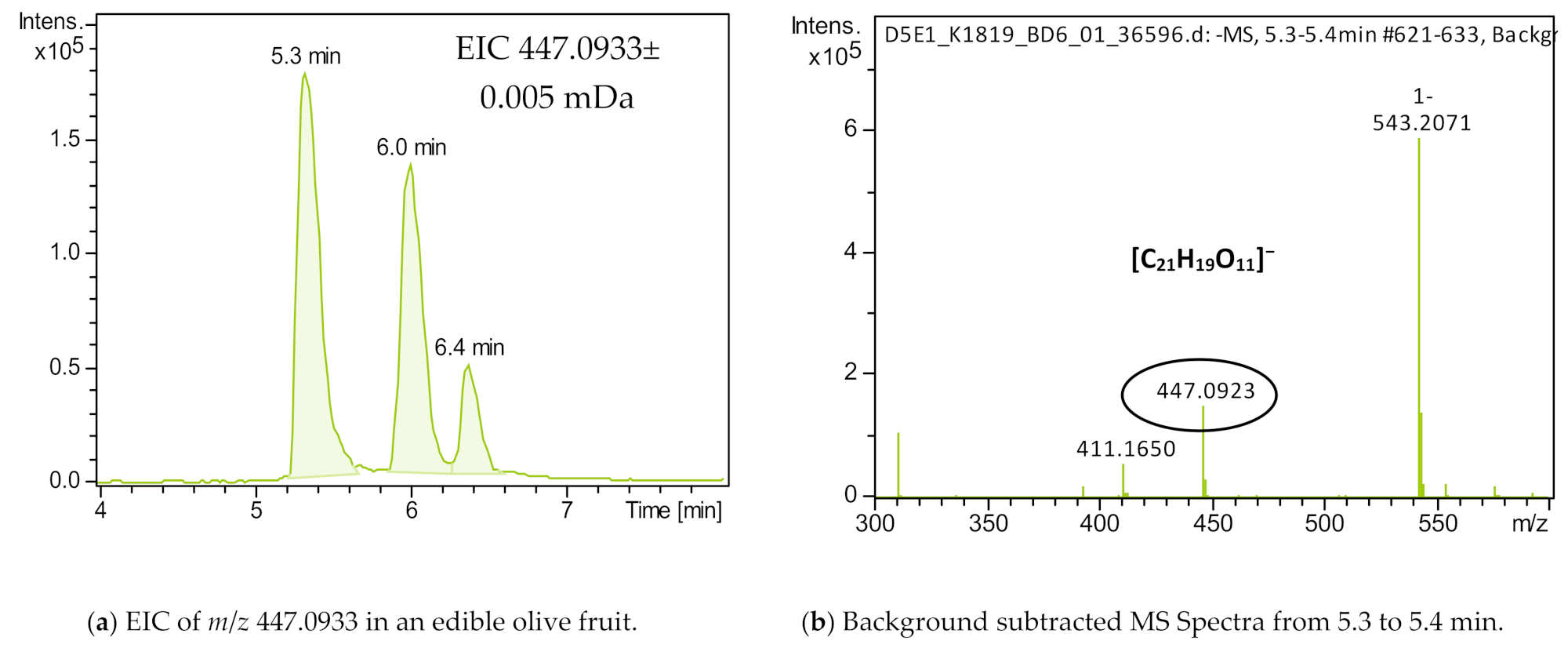
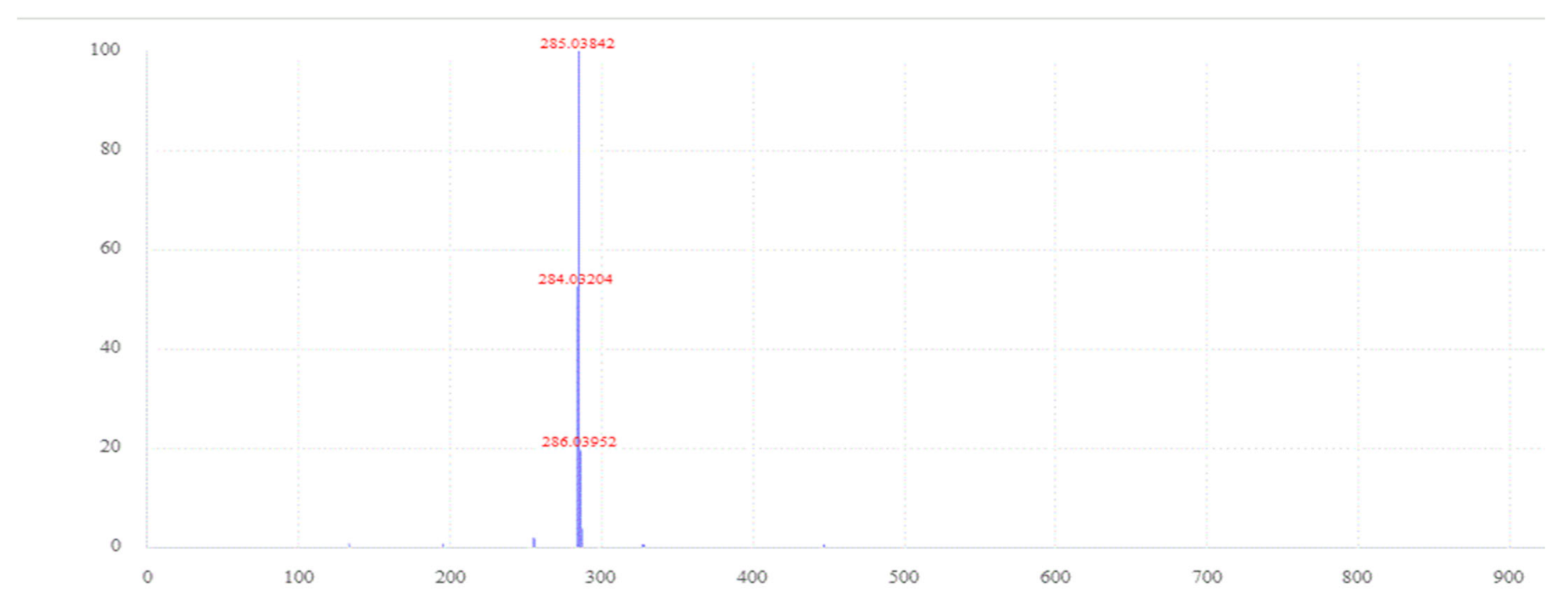
| luteolin-7-o-glucoside | C21H20O11 | [M-H]− | 1.0 | 6.61 | 5.3 | 16 | 285.0400 | 7 |
| luteolin-3-o-glucoside | C21H20O11 | [M-H]− | 1.0 | 6.58 | 6.4 | 16 | 285.0400 | 7 |
| luteolin-4-o-glucoside | C21H20O11 | [M-H]− | 1.0 | 6.54 | 6 | 16 | 285.0400 | 7 |
| caffeoyl-6 secologanoside | C25H28O14 | [M-H]− | 0.7 | - | 4.2 | 28 | 161.0242; 281.0659; 389.1077; 507.1492 | 7 |
| comselogoside | C25H28O13 | [M-H]− | 0.2 | - | 4.6 | 24 | 145.0291; 265.0715; 491.1550 | 7 |
| dihydrooleuropein | C29H36O13 | [M-H]− | 1.2 | - | F5.4 | 24 | 313.1287; 357.1186; 377.1448; 389.1447; 525.1971 | 7 |
| isoverbasoside | C29H36O15 | [M-H]− | 0.7 | 8.49 | 5.3 | 14 | 161.02423; 461.1659 | 7 |
| oleoside | C16H22O11 | [M-H]− | 0.2 | 2.91 | 1.8 | 8.8 | 165.0558; 183.0666; 209.0460; 345.1186 | 3 |
| apigenin 7-glucoside | C21H20O10 | [M-H]− | 0.6 | 6.39 | 5.9 | 31 | 269.0455; 268.0470 | 7 |
| isorhoifolin | C27 H30O14 | [M-H]− | 0.9 | 6.81 | 5.7 | 32 | 269.0453; 270.0483 | 7 |
| Analyte | Concentration Range (mg/L) | Calibration Curve | Correlation Factor Can | LOD (mg/kg) (n = 10) | LOQ (mg/kg) (n = 10) |
|---|---|---|---|---|---|
| lutein | 0.05–6.00 | y = 207.24–x − 8.0363 | 0.998 | 0.15 | 0.51 |
| β-carotene | 0.30–4.00 | y = 105.7–x − 6.7561 | 0.995 | 0.07 | 0.23 |
| α-tocopherol | 2.00–75.0 | y = 7.496–x − 13.526 | 0.991 | 1.1 | 3.7 |
| γ-tocopherol | 0.60–10.0 | y = 9.8181x + 3.0003 | 0.991 | 0.55 | 1.8 |
| δ-tocopherol | 1.00–24.0 | y = 7.623–x − 0.2302 | 0.994 | 1.0 | 3.4 |
| chlorophyll | 0.50–15.0 | y = 74.15–x − 6.949 | 0.996 | 0.21 | 0.71 |
| squalene | 150–700 | y = 38.318x + 3137.2 | 0.9996 | 4.2 | 13.9 |
| %RecoveryEvaluation of Accuracy | RSDrEvaluation of Repeatability | RSDREvaluation of Intermediate Precision | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Low Concentration (%R) (n = 18) | Medium Concentration (%R) (n = 18) | High Concentration (%R) (n = 6) | Low Concentration(%RSD) (n = 6) | Medium Concentration (%RSD) (n = 6) | High Concentration(%RSD) (n = 6) | Low Concentration (%RSD) (n = 18) | Medium Concentration(%RSD) (n = 18) | High Concentration(%RSD) (n = 18) | |
| lutein | 89.2 | 88.4 | 81.7 | 4.7 | 5.1 | 3.6 | 9.8 | 7.3 | 6.2 |
| β-carotene | 100.4 | 91.9 | 93.4 | 2.2 | 3.9 | 3.7 | 6.4 | 13.5 | 14.7 |
| α-tocopherol | 99.6 | 99.3 | 99.4 | 3.5 | 5.3 | 5.4 | 4.5 | 4.3 | 5.4 |
| γ-tocopherol | 97.6 | 93.0 | 89.3 | 3.2 | 2.5 | 9.5 | 12.0 | 6.7 | 6.8 |
| δ-tocopherol | 101.1 | 95.4 | 93.5 | 3.2 | 1.8 | 4.8 | 12.3 | 5.6 | 9.1 |
| chlorophyll | 88.3 | 88.0 | 83.7 | 5.9 | 5.1 | 4.8 | 9.0 | 5.8 | 5.5 |
| squalene | 93.2 | 84.3 | 81.0 | 6.7 | 7.0 | 4.2 | 11.2 | 9.5 | 10.8 |
| Chlorophyll | Lutein | β-Carotene | α-Tocopherol | (β + γ)-Tocopherols | Squalene | |
|---|---|---|---|---|---|---|
| mg/kg Raw Sample | ||||||
| Sample 1 | 12.6 | 2.39 | 2.08 | 54 | 2.34 | 634 |
| Sample 2 | 11.8 | 2.71 | 1.88 | 96 | 3.29 | 489 |
| Sample 3 | 12.2 | 2.95 | 1.92 | 84 | 3.41 | 659 |
| Sample 4 | 12.0 | 3.29 | 2.39 | 95 | 2.89 | 632 |
| Sample 5 | 8.2 | 2.33 | 1.39 | 57 | 3.60 | 734 |
| Sample 6 | 9.3 | <LOQ | <LOD | 91 | 4.30 | 529 |
| Sample 7 | <LOQ | 2.17 | 1.53 | 76 | 3.74 | 619 |
| Average concentration | 9.6 | 2.64 | 1.87 | 79 | 3.37 | 614 |
| Concentration range | <LOQ −12.6 | <LOQ −3.29 | <LOD−2.39 | 54–96 | 2.34–4.30 | 489–734 |
| SD* | 3.7 | 0.66 | 0.72 | 16 | 0.58 | 76 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Martakos, I.; Katsianou, P.; Koulis, G.; Efstratiou, E.; Nastou, E.; Nikas, S.; Dasenaki, M.; Pentogennis, M.; Thomaidis, N. Development of Analytical Strategies for the Determination of Olive Fruit Bioactive Compounds Using UPLC-HRMS and HPLC-DAD. Chemical Characterization of Kolovi Lesvos Variety as a Case Study. Molecules 2021, 26, 7182. https://doi.org/10.3390/molecules26237182
Martakos I, Katsianou P, Koulis G, Efstratiou E, Nastou E, Nikas S, Dasenaki M, Pentogennis M, Thomaidis N. Development of Analytical Strategies for the Determination of Olive Fruit Bioactive Compounds Using UPLC-HRMS and HPLC-DAD. Chemical Characterization of Kolovi Lesvos Variety as a Case Study. Molecules. 2021; 26(23):7182. https://doi.org/10.3390/molecules26237182
Chicago/Turabian StyleMartakos, Ioannis, Panagiota Katsianou, Georgios Koulis, Elvira Efstratiou, Eleni Nastou, Stylianos Nikas, Marilena Dasenaki, Michalis Pentogennis, and Nikolaos Thomaidis. 2021. "Development of Analytical Strategies for the Determination of Olive Fruit Bioactive Compounds Using UPLC-HRMS and HPLC-DAD. Chemical Characterization of Kolovi Lesvos Variety as a Case Study" Molecules 26, no. 23: 7182. https://doi.org/10.3390/molecules26237182
APA StyleMartakos, I., Katsianou, P., Koulis, G., Efstratiou, E., Nastou, E., Nikas, S., Dasenaki, M., Pentogennis, M., & Thomaidis, N. (2021). Development of Analytical Strategies for the Determination of Olive Fruit Bioactive Compounds Using UPLC-HRMS and HPLC-DAD. Chemical Characterization of Kolovi Lesvos Variety as a Case Study. Molecules, 26(23), 7182. https://doi.org/10.3390/molecules26237182








