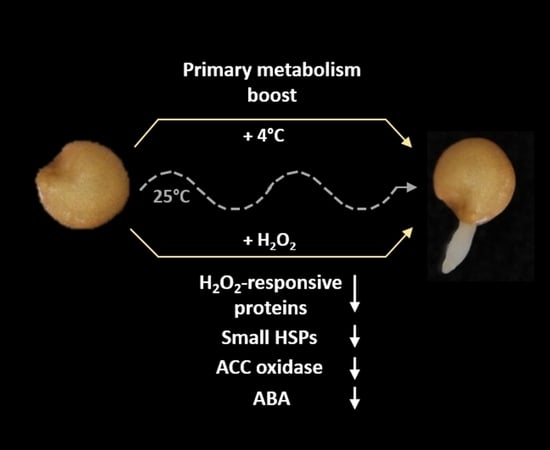
Eggplant Germination is Promoted by Hydrogen Peroxide and Temperature in an Independent but Overlapping Manner
Abstract
1. Introduction
2. Results
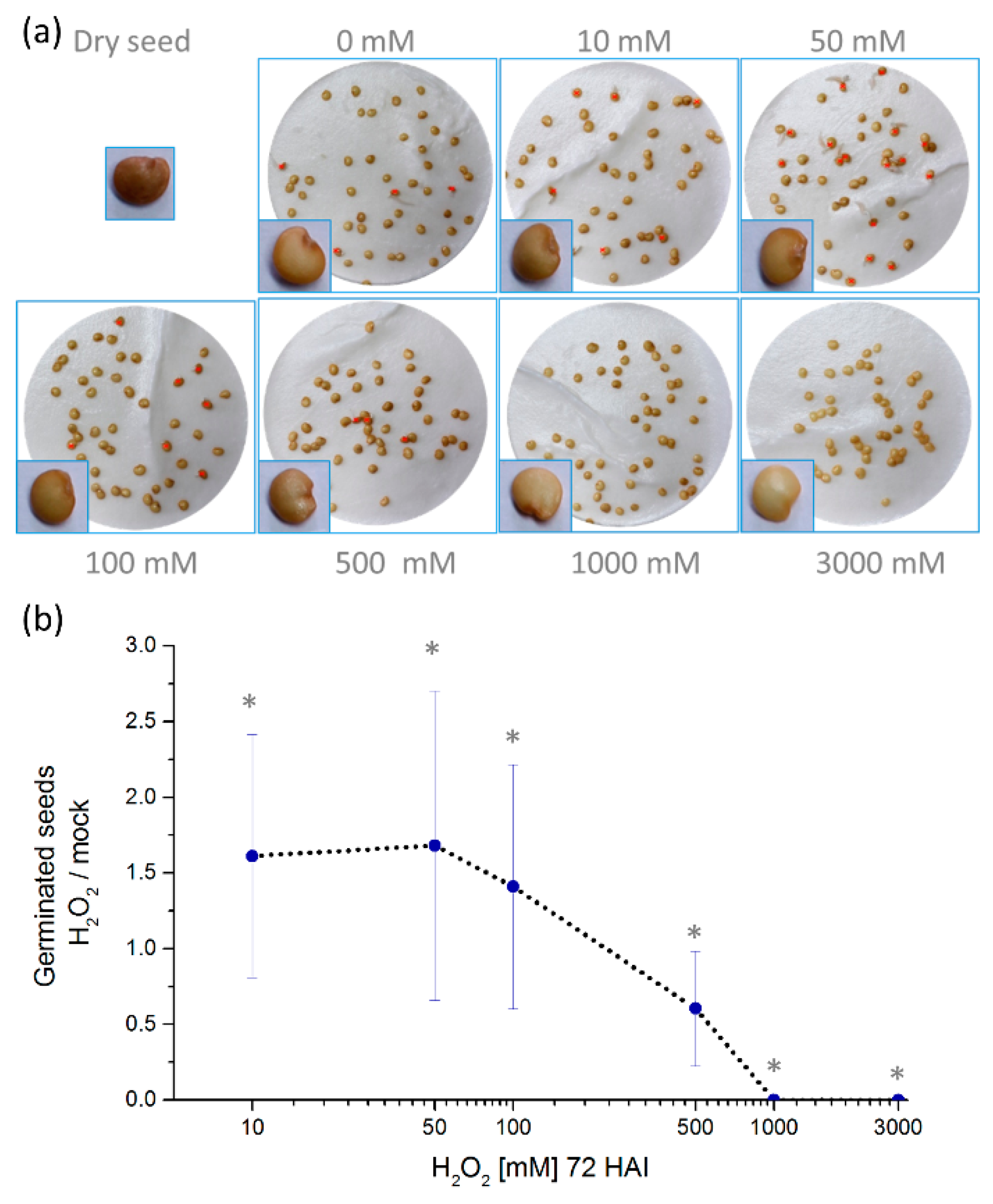
2.1. Hydrogen Peroxide Stimulates Eggplant Seed Germination
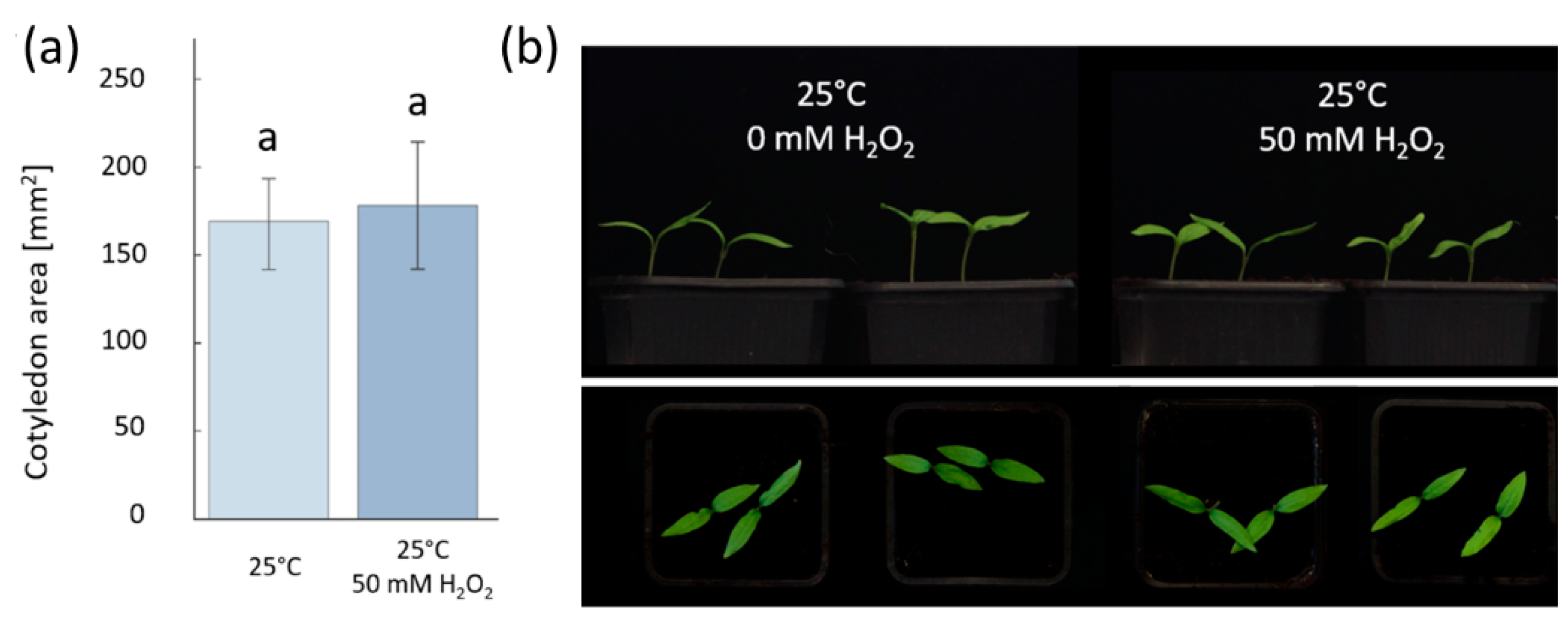
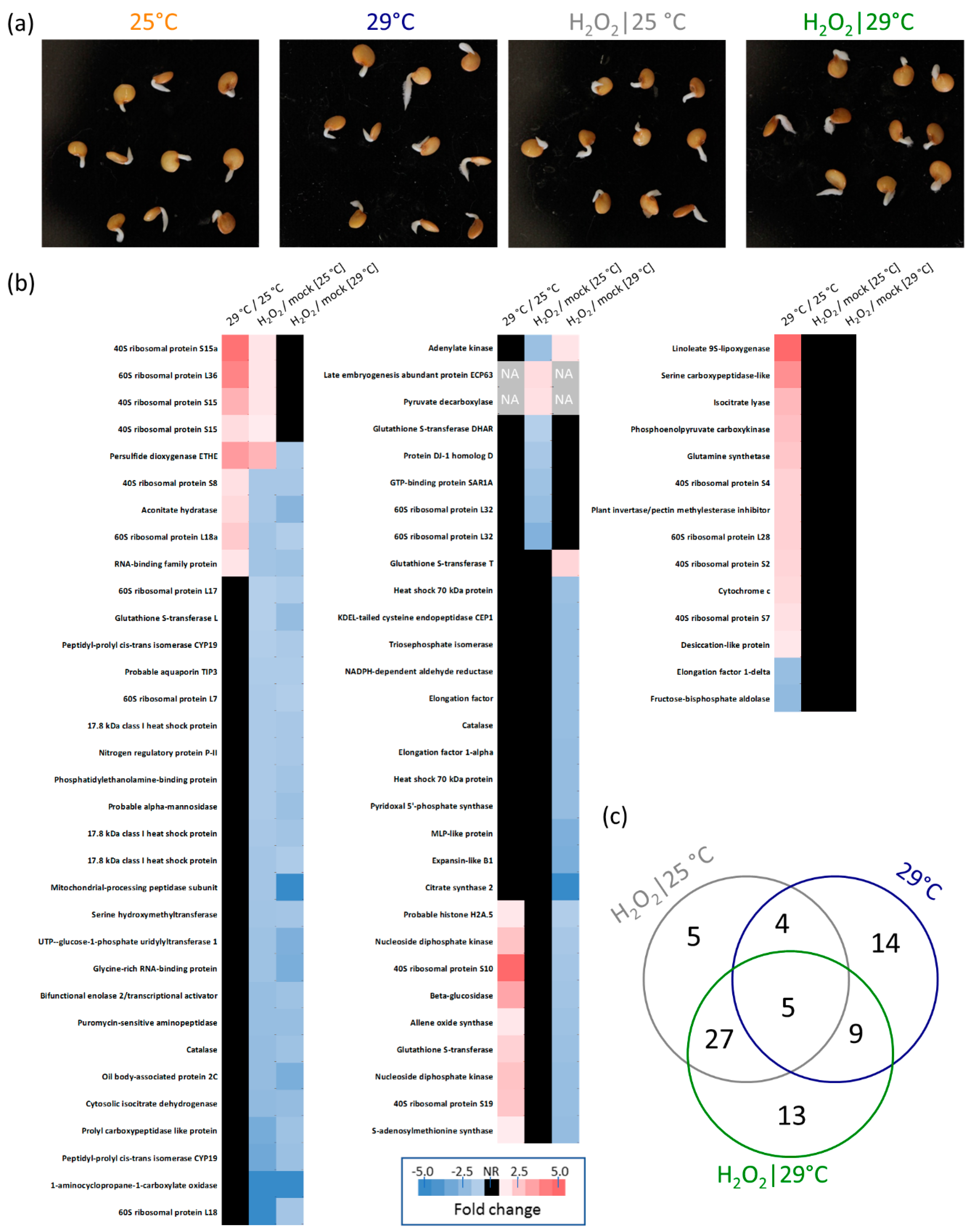
2.2. Hydrogen Peroxide-Induced Germination Rate does not Manifest Any Visible Seedlings Traits
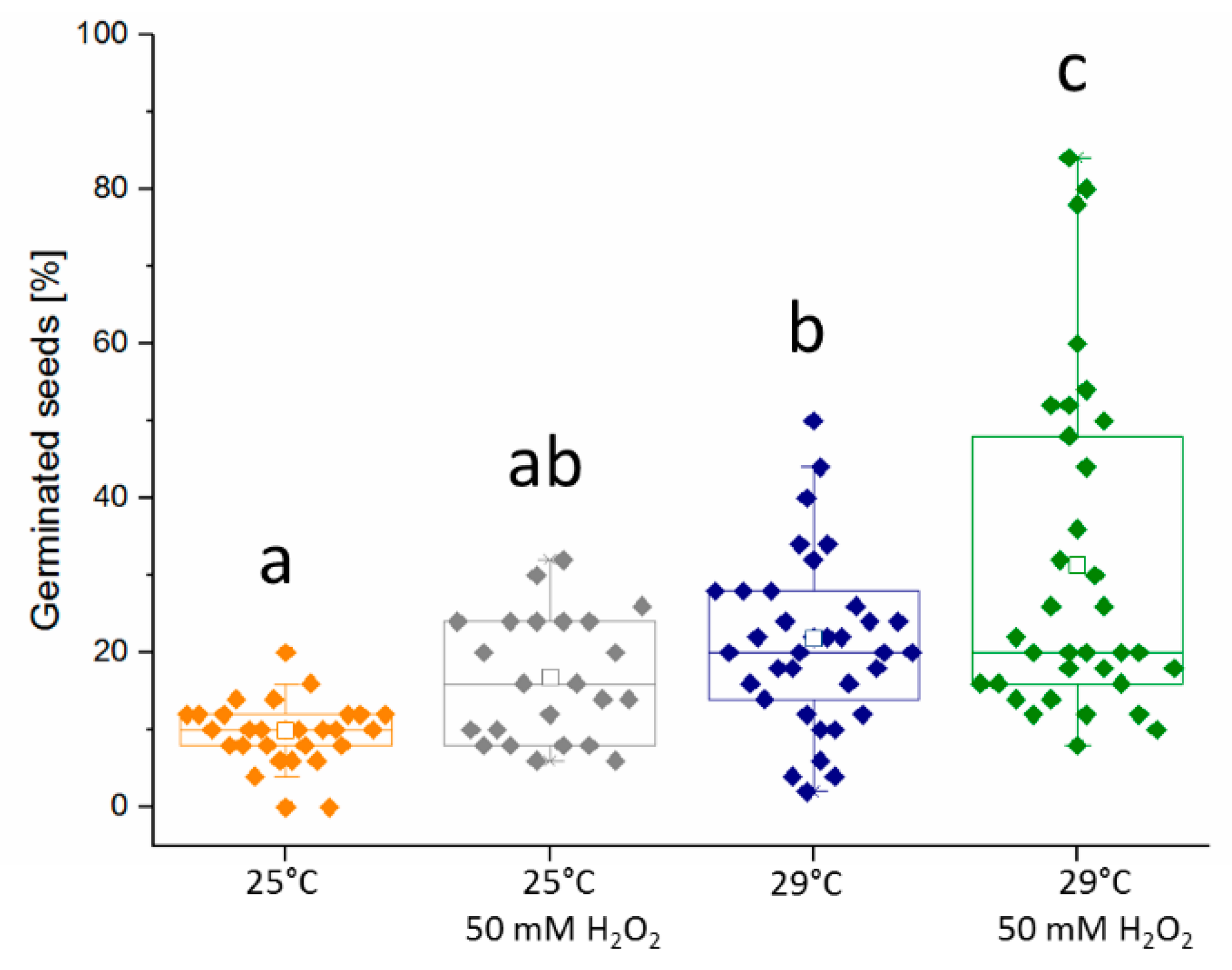
2.3. Hydrogen Peroxide Seemingly Mimics the Temperature Stimulus
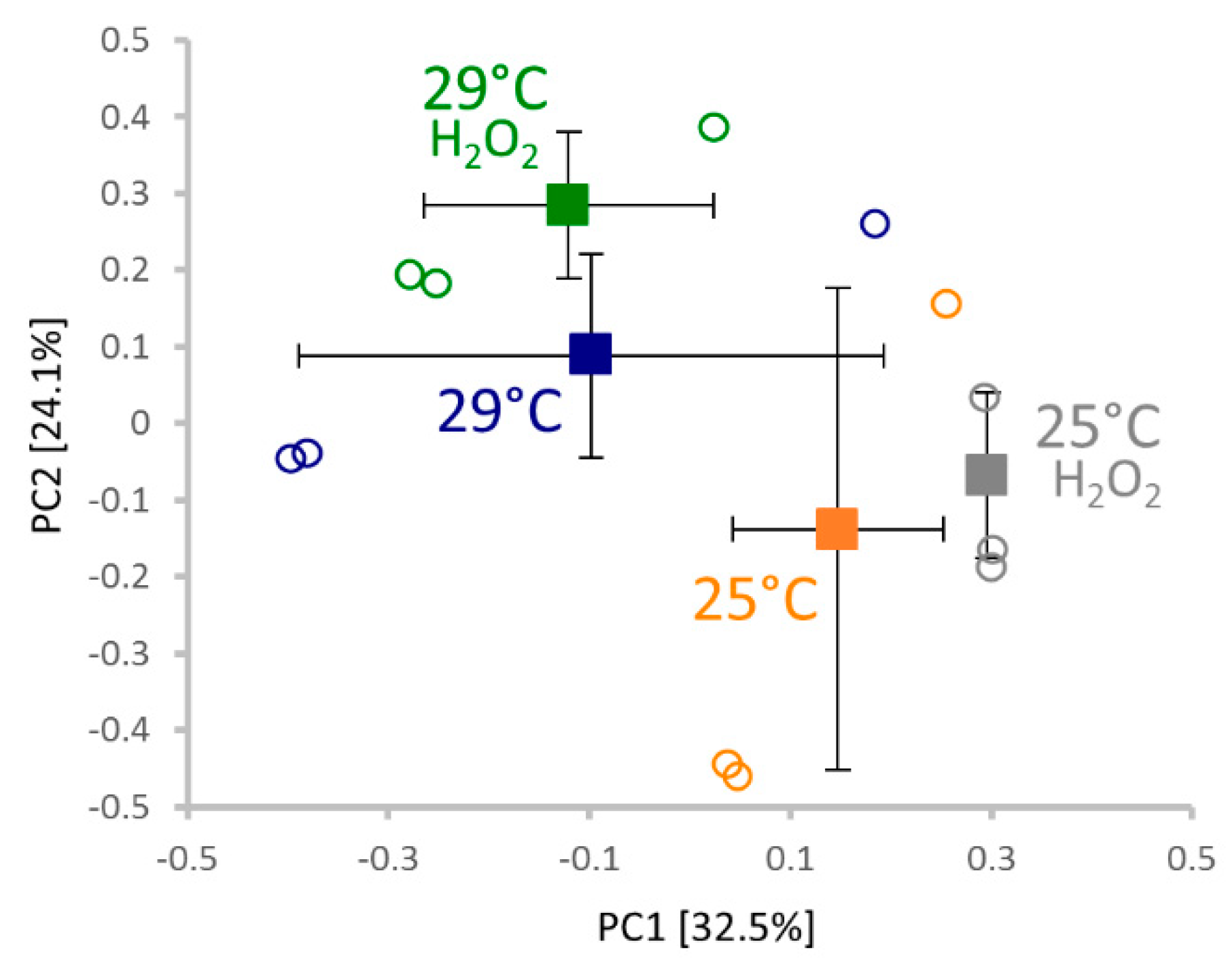
2.4. Proteomics Analysis
2.5. Response to Hydrogen Peroxide and Temperature is Predominantly Different
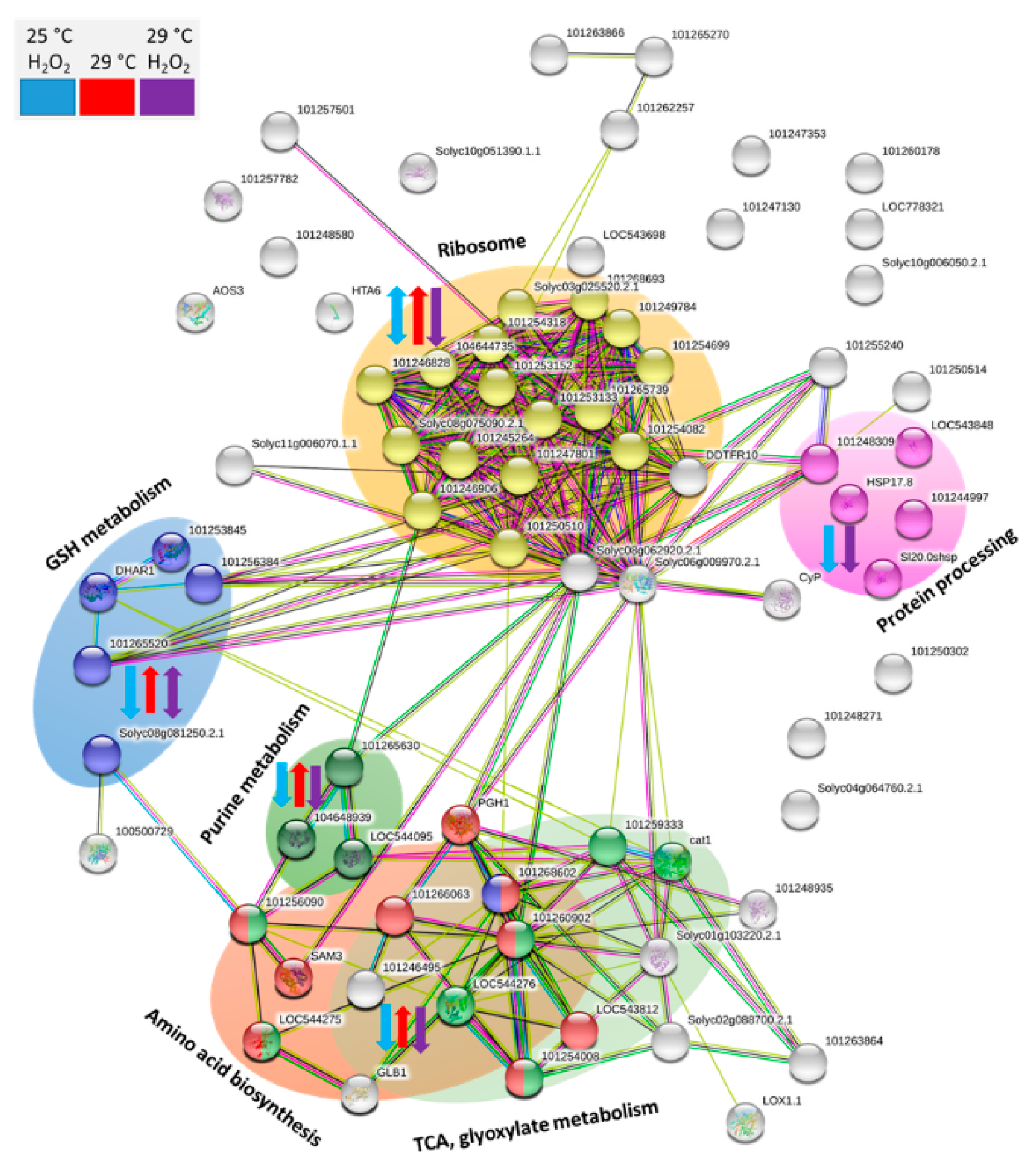
2.6. Expected Localization of Identified Proteins
2.7. Enzymes of Hydrogen Peroxide Metabolism
2.8. Putative Function of Identified Differentially Abundant Proteins
3. Discussion
3.1. Hydrogen Peroxide—A Potent Enhancer of Eggplant Germination
3.2. Omics Analyses Indicate a Low Effect of Temperature on Hydrogen Peroxide Metabolism and a Possible Effect of Hydrogen Peroxide on Temperature Perception
3.3. Majority of Identified Temperature-Responsive Proteins is not a Part of Stress Response Mechanisms
3.4. Hydrogen Peroxide Effect on Eggplant Seed Proteome Indicates Depletion of Hydrogen Peroxide Pool
3.5. Depletion of Heat Shock Proteins Comparison with Phytohormone-Responsive Proteomics Indicate a Putative Link to Abscisic Acid Sensitivity
4. Conclusions
5. Materials and Methods
5.1. Plant Material
5.2. Protein Extraction and LC–MS Analysis
5.3. Bioinformatics Analysis
5.4. Data Visualization and Statistics
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Wojtyla, Ł.; Lechowska, K.; Kubala, S.; Garnczarska, M. Different Modes of Hydrogen Peroxide Action During Seed Germination. Front. Plant Sci. 2016, 7, 66. [Google Scholar] [CrossRef] [PubMed]
- Waszczak, C.; Carmody, M.; Kangasjärvi, J. Reactive Oxygen Species in Plant Signaling. Annu. Rev. Plant Biol. 2018, 69, 209–236. [Google Scholar] [CrossRef] [PubMed]
- Černý, M.; Habánová, H.; Berka, M.; Luklová, M.; Brzobohatý, B. Hydrogen peroxide: Its role in plant biology and crosstalk with signaling networks. Int. J. Mol. Sci. 2018, 19, 2812. [Google Scholar] [CrossRef] [PubMed]
- Oracz, K.; Karpiński, S. Phytohormones Signaling Pathways and ROS Involvement in Seed Germination. Front. Plant Sci. 2016, 7, 864. [Google Scholar] [CrossRef]
- Kai, K.; Kasa, S.; Sakamoto, M.; Aoki, N.; Watabe, G.; Yuasa, T.; Iwaya-Inoue, M.; Ishibashi, Y. Role of reactive oxygen species produced by NADPH oxidase in gibberellin biosynthesis during barley seed germination. Plant Signal. Behav. 2016, 11, e1180492. [Google Scholar] [CrossRef]
- Ishibashi, Y.; Tawaratsumida, T.; Kondo, K.; Kasa, S.; Sakamoto, M.; Aoki, N.; Zheng, S.-H.; Yuasa, T.; Iwaya-Inoue, M. Reactive Oxygen Species Are Involved in Gibberellin/Abscisic Acid Signaling in Barley Aleurone Cells. Plant Physiol. 2012, 158, 1705–1714. [Google Scholar] [CrossRef]
- Ishibashi, Y.; Aoki, N.; Kasa, S.; Sakamoto, M.; Kai, K.; Tomokiyo, R.; Watabe, G.; Yuasa, T.; Iwaya-Inoue, M. The Interrelationship between Abscisic Acid and Reactive Oxygen Species Plays a Key Role in Barley Seed Dormancy and Germination. Front. Plant Sci. 2017, 8, 275. [Google Scholar] [CrossRef]
- Zhang, Y.; Chen, B.; Xu, Z.; Shi, Z.; Chen, S.; Huang, X.; Chen, J.; Wang, X. Involvement of reactive oxygen species in endosperm cap weakening and embryo elongation growth during lettuce seed germination. J. Exp. Bot. 2014, 65, 3189–3200. [Google Scholar] [CrossRef]
- Ullio, L. Eggplant growing. Agfacts 2003, H8.1.29. 1–4. Available online: https://www.dpi.nsw.gov.au/__data/assets/pdf_file/0004/126292/Eggplant-Growing-Agfact-H8.1.29.pdf (accessed on 1 November 2019).
- Chen, N.C.; Li, H.M. Cultivation and breeding of eggplant. In Training Workshop on Vegetable Cultivation and Seed Production; AVRDC World Vegetable Center: Shanhua, Taiwan, 1996. [Google Scholar]
- Barchi, L.; Pietrella, M.; Venturini, L.; Minio, A.; Toppino, L.; Acquadro, A.; Andolfo, G.; Aprea, G.; Avanzato, C.; Bassolino, L.; et al. A chromosome-anchored eggplant genome sequence reveals key events in Solanaceae evolution. Sci. Rep. 2019, 9. [Google Scholar] [CrossRef]
- Baldrianová, J.; Černý, M.; Novák, J.; Jedelský, P.L.; Divíšková, E.; Brzobohatý, B. Arabidopsis proteome responses to the smoke-derived growth regulator karrikin. J. Proteomics 2015, 120, 7–20. [Google Scholar] [CrossRef] [PubMed]
- Hooper, C.M.; Castleden, I.R.; Tanz, S.K.; Aryamanesh, N.; Millar, A.H. SUBA4: The interactive data analysis centre for Arabidopsis subcellular protein locations. Nucleic Acids Res. 2017, 45, D1064–D1074. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Mhamdi, A.; Noctor, G. Analysis of catalase mutants underscores the essential role of CATALASE2 for plant growth and day length-dependent oxidative signaling. Plant Cell Environ. 2019, 42, 688–700. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. Von STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef]
- Zhang, Y.; Liu, H.; Shen, S.; Zhang, X. Improvement of eggplant seed germination and seedling emergence at low temperature by seed priming with incorporation SA into KNO3 solution. Front. Agric. China 2011, 5, 534–537. [Google Scholar] [CrossRef]
- Zdravkovic, J.; Ristic, N.; Girek, Z.; Pavlovic, S.; Pavlovic, N.; Pavlovic, R.; Zdravkovic, M. Understanding and overcoming seed dormancy in eggplant (Solanum melongena L.) breeding lines. SABRAO J. Breed. Genet. 2013, 45, 211–220. [Google Scholar]
- Neto, F.J.D.; Dalanhol, S.J.; Machry, M.; Junior, A.P.; Rodrigues, J.D.; Ono, E.O. Effects of plant growth regulators on eggplant seed germination and seedling growth. Aust. J. Crop Sci. 2017, 11, 1277–1282. [Google Scholar] [CrossRef]
- Ali, M.; Hayat, S.; Ahmad, H.; Ghani, M.I.; Amin, B.; Atif, M.J.; Cheng, Z. Priming of Solanum melongena L. seeds enhances germination, alters antioxidant enzymes, modulates ROS, and improves early seedling growth: Indicating aqueous garlic extract as seed-priming bio-stimulant for eggplant production. Appl. Sci. 2019, 9, 2203. [Google Scholar] [CrossRef]
- Tank, J.; Dhamangaonkar, B.; Ukale, D.U.; Cukkemane, N.; Cukkemane, A.A. Biochemical and Microbiological Analysis of Different Organic Manures: Their Effect on Germination of Coriandrum sativum (Cilantro) and Solanum melongena (Eggplant). J. Bioprocess. Biotech. 2017, 7. [Google Scholar] [CrossRef]
- Diao, Q.; Song, Y.; Shi, D.; Qi, H. Interaction of Polyamines, Abscisic Acid, Nitric Oxide, and Hydrogen Peroxide under Chilling Stress in Tomato (Lycopersicon esculentum Mill.) Seedlings. Front. Plant Sci. 2017, 8, 203. [Google Scholar] [CrossRef]
- Sun, M.; Jiang, F.; Cen, B.; Wen, J.; Zhou, Y.; Wu, Z. Respiratory burst oxidase homologue-dependent H2O2 and chloroplast H2O2 are essential for the maintenance of acquired thermotolerance during recovery after acclimation. Plant Cell Environ. 2018, 41, 2373–2389. [Google Scholar] [CrossRef] [PubMed]
- Carpenter, C.D.; Kreps, J.A.; Simon, A.E. Genes encoding glycine-rich Arabidopsis thaliana proteins with RNA-binding motifs are influenced by cold treatment and an endogenous circadian rhythm. Plant Physiol. 1994, 104, 1015–1025. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Xiong, L.; Ishitani, M.; Zhu, J.K. An Arabidopsis mutation in translation elongation factor 2 causes superinduction of CBF/DREB1 transcription factor genes but blocks the induction of their downstream targets under low temperatures. Proc. Natl. Acad. Sci. USA 2002, 99, 7786–7791. [Google Scholar] [CrossRef] [PubMed]
- Wilkins, O.; Bräutigam, K.; Campbell, M.M. Time of day shapes Arabidopsis drought transcriptomes. Plant J. 2010, 63, 715–727. [Google Scholar] [CrossRef]
- He, H.; Yan, J.; Yu, X.; Liang, Y.; Fang, L.; Scheller, H.V.; Zhang, A. The NADPH-oxidase AtRbohI plays a positive role in drought-stress response in Arabidopsis thaliana. Biochem. Biophys. Res. Commun. 2017, 491, 834–839. [Google Scholar] [CrossRef]
- Schmidt, F.; Marnef, A.; Cheung, M.K.; Wilson, I.; Hancock, J.; Staiger, D.; Ladomery, M. A proteomic analysis of oligo(dT)-bound mRNP containing oxidative stress-induced Arabidopsis thaliana RNA-binding proteins ATGRP7 and ATGRP8. Mol. Biol. Rep. 2010, 37, 839–845. [Google Scholar] [CrossRef]
- Fukamatsu, Y.; Yabe, N.; Hasunuma, K. Arabidopsis NDK1 is a Component of ROS Signaling by Interacting with Three Catalases. Plant Cell Physiol. 2003, 44, 982–989. [Google Scholar] [CrossRef]
- Yu, J.; Jin, X.; Sun, X.; Gao, T.; Chen, X.; She, Y.; Jiang, T.; Chen, S.; Dai, S. Hydrogen peroxide response in leaves of poplar (populus simonii × populus nigra) revealed from physiological and proteomic analyses. Int. J. Mol. Sci. 2017, 18, 2085. [Google Scholar] [CrossRef]
- Khan, A.; Ali, M.; Khattak, A.M.; Gai, W.X.; Zhang, H.X.; Wei, A.M.; Gong, Z.H. Gong Heat Shock Proteins: Dynamic Biomolecules to Counter Plant Biotic and Abiotic Stresses. Int. J. Mol. Sci. 2019, 20, 5321. [Google Scholar] [CrossRef]
- Klepikova, A.V.; Logacheva, M.D.; Dmitriev, S.E.; Penin, A.A. RNA-seq analysis of an apical meristem time series reveals a critical point in Arabidopsis thaliana flower initiation. BMC Genomics 2015, 16. [Google Scholar] [CrossRef]
- Inzé, A.; Vanderauwera, S.; Hoeberichts, F.A.; Vandorpe, M.; van Gaever, T.; van Breusegem, F. A subcellular localization compendium of hydrogen peroxide-induced proteins. Plant Cell Environ. 2012, 35, 308–320. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.H.; Xu, Z.Y.; Hwang, I. AtHSP17.8 overexpression in transgenic lettuce gives rise to dehydration and salt stress resistance phenotypes through modulation of ABA-mediated signaling. Plant Cell Rep. 2013, 32, 1953–1963. [Google Scholar] [CrossRef] [PubMed]
- Arc, E.; Sechet, J.; Corbineau, F.; Rajjou, L.; Marion-Poll, A. ABA crosstalk with ethylene and nitric oxide in seed dormancy and germination. Front. Plant Sci. 2013, 4, 63. [Google Scholar] [CrossRef] [PubMed]
- Černý, M.; Novák, J.; Habánová, H.; Cerna, H.; Brzobohatý, B. Role of the proteome in phytohormonal signaling. Biochim. Biophys. Acta Proteins Proteomics 2016, 1864, 1003–1015. [Google Scholar] [CrossRef]
- Li, B.; Takahashi, D.; Kawamura, Y.; Uemura, M. Comparison of plasma membrane proteomic changes of Arabidopsis suspension-cultured cells (T87 Line) after cold and ABA treatment in association with freezing tolerance development. Plant Cell Physiol. 2012, 53, 543–554. [Google Scholar] [CrossRef]
- Komatsu, S.; Han, C.; Nanjo, Y.; Altaf-Un-Nahar, M.; Wang, K.; He, D.; Yang, P. Label-free quantitative proteomic analysis of abscisic acid effect in early-stage soybean under flooding. J. Proteome Res. 2013, 12, 4769–4784. [Google Scholar] [CrossRef]
- Böhmer, M.; Schroeder, J.I. Quantitative transcriptomic analysis of abscisic acid-induced and reactive oxygen species-dependent expression changes and proteomic profiling in Arabidopsis suspension cells. Plant J. 2011, 67, 105–118. [Google Scholar] [CrossRef]
- Zhu, M.; Zhu, N.; Song, W.; Harmon, A.C.; Assmann, S.M.; Chen, S. Thiol-based redox proteins in abscisic acid and methyl jasmonate signaling in Brassica napus guard cells. Plant J. 2014, 78, 491–515. [Google Scholar] [CrossRef]
- Černý, M.; Kuklová, A.; Hoehenwarter, W.; Fragner, L.; Novák, O.; Rotková, G.; Jedelský, P.L.P.L.; Žáková, K.K.; Šmehilová, M.; Strnad, M.; et al. Proteome and metabolome profiling of cytokinin action in Arabidopsis identifying both distinct and similar responses to cytokinin down- and up-regulation. J. Exp. Bot. 2013, 64, 4193–4206. [Google Scholar] [CrossRef]
- Cerna, H.; Černý, M.; Habánová, H.; Šafářová, D.; Abushamsiya, K.; Navrátil, M.; Brzobohatý, B. Proteomics offers insight to the mechanism behind Pisum sativum L. response to pea seed-borne mosaic virus (PSbMV). J. Proteomics 2017, 153, 78–88. [Google Scholar] [CrossRef]
- Hruz, T.; Laule, O.; Szabo, G.; Wessendorp, F.; Bleuler, S.; Oertle, L.; Widmayer, P.; Gruissem, W.; Zimmermann, P. Genevestigator V3: A Reference Expression Database for the Meta-Analysis of Transcriptomes. Adv. Bioinformatics 2008, 2008, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Nolte, H.; MacVicar, T.D.; Tellkamp, F.; Krüger, M. Instant Clue: A Software Suite for Interactive Data Visualization and Analysis. Sci. Rep. 2018, 8. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of the compounds are not available from the authors. |

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dufková, H.; Berka, M.; Luklová, M.; Rashotte, A.M.; Brzobohatý, B.; Černý, M. Eggplant Germination is Promoted by Hydrogen Peroxide and Temperature in an Independent but Overlapping Manner. Molecules 2019, 24, 4270. https://doi.org/10.3390/molecules24234270
Dufková H, Berka M, Luklová M, Rashotte AM, Brzobohatý B, Černý M. Eggplant Germination is Promoted by Hydrogen Peroxide and Temperature in an Independent but Overlapping Manner. Molecules. 2019; 24(23):4270. https://doi.org/10.3390/molecules24234270
Chicago/Turabian StyleDufková, Hana, Miroslav Berka, Markéta Luklová, Aaron M. Rashotte, Břetislav Brzobohatý, and Martin Černý. 2019. "Eggplant Germination is Promoted by Hydrogen Peroxide and Temperature in an Independent but Overlapping Manner" Molecules 24, no. 23: 4270. https://doi.org/10.3390/molecules24234270
APA StyleDufková, H., Berka, M., Luklová, M., Rashotte, A. M., Brzobohatý, B., & Černý, M. (2019). Eggplant Germination is Promoted by Hydrogen Peroxide and Temperature in an Independent but Overlapping Manner. Molecules, 24(23), 4270. https://doi.org/10.3390/molecules24234270