Genetic Diversity and Population Structure of Moroccan Beni Ahsen: Is This Endangered Ovine Breed One of the Ancestors of Merino?
Abstract
:1. Introduction
2. Materials and Methods
3. Results
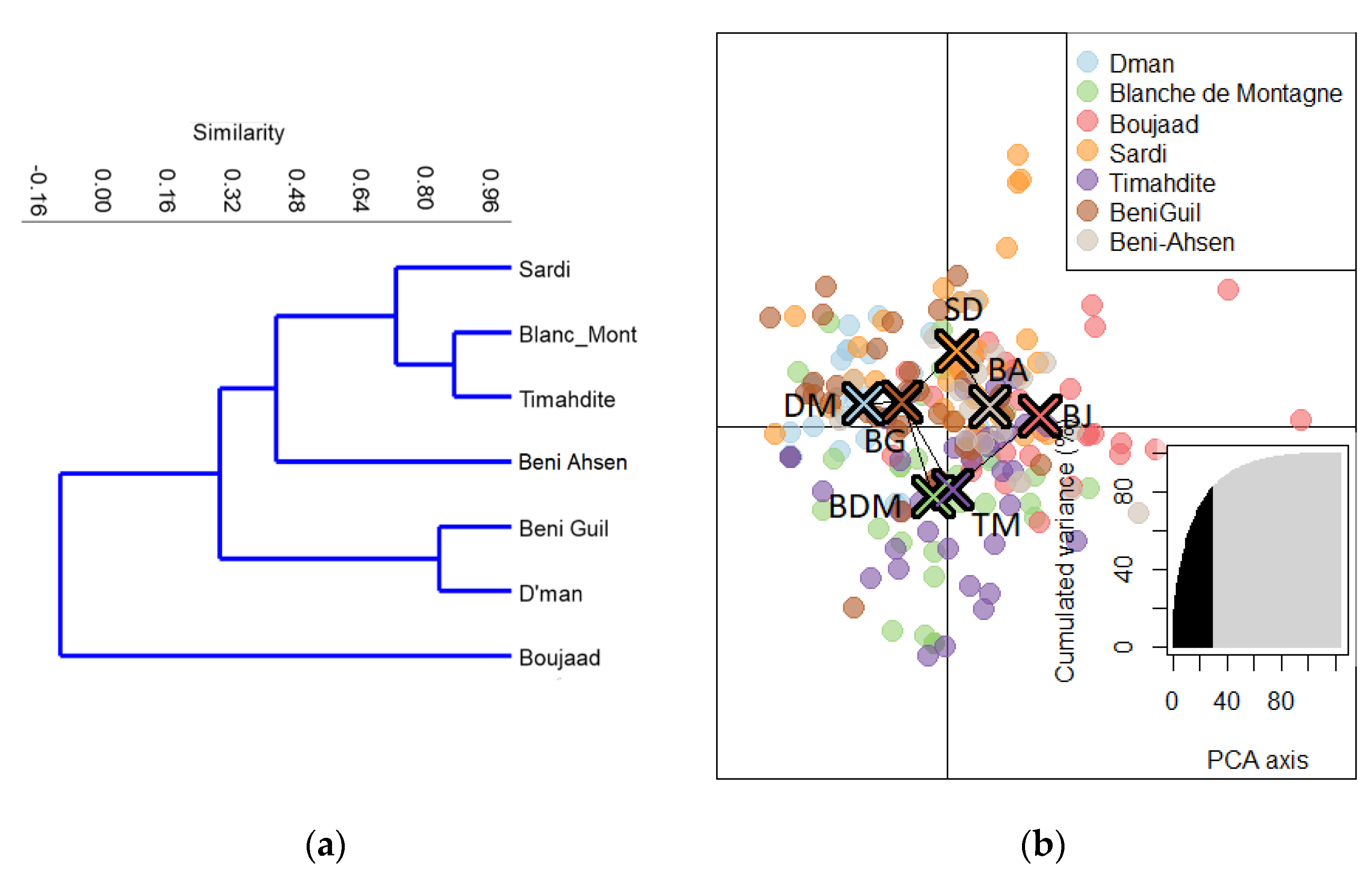
3.1. Relationships between Moroccan Breeds
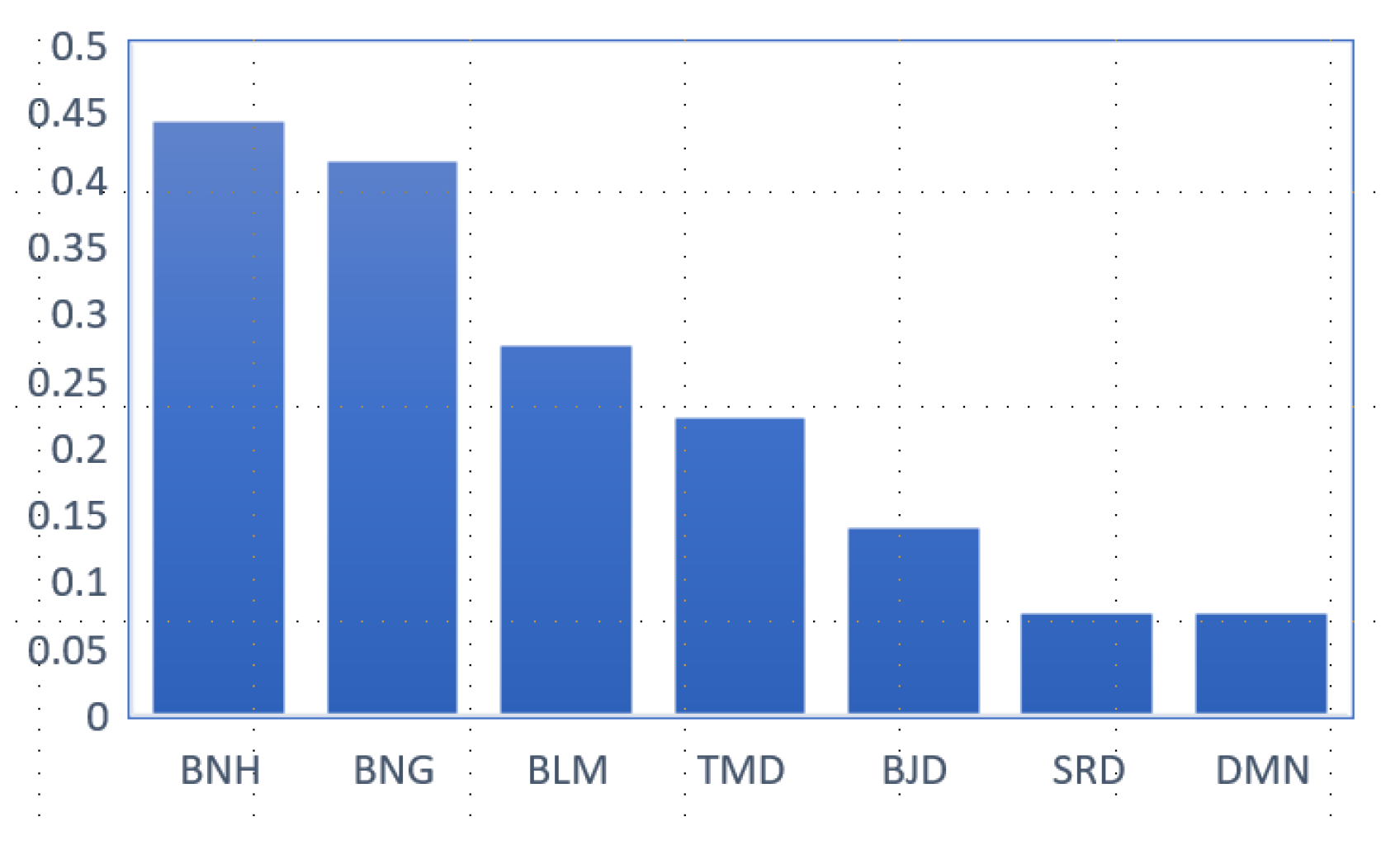
3.1.1. Polymorphism of Beni Ahsen Compared to Other Moroccan Breeds
3.1.2. Affinity Degree of Beni Ahsen to Other Moroccan Breeds
3.1.3. Breed Assignation
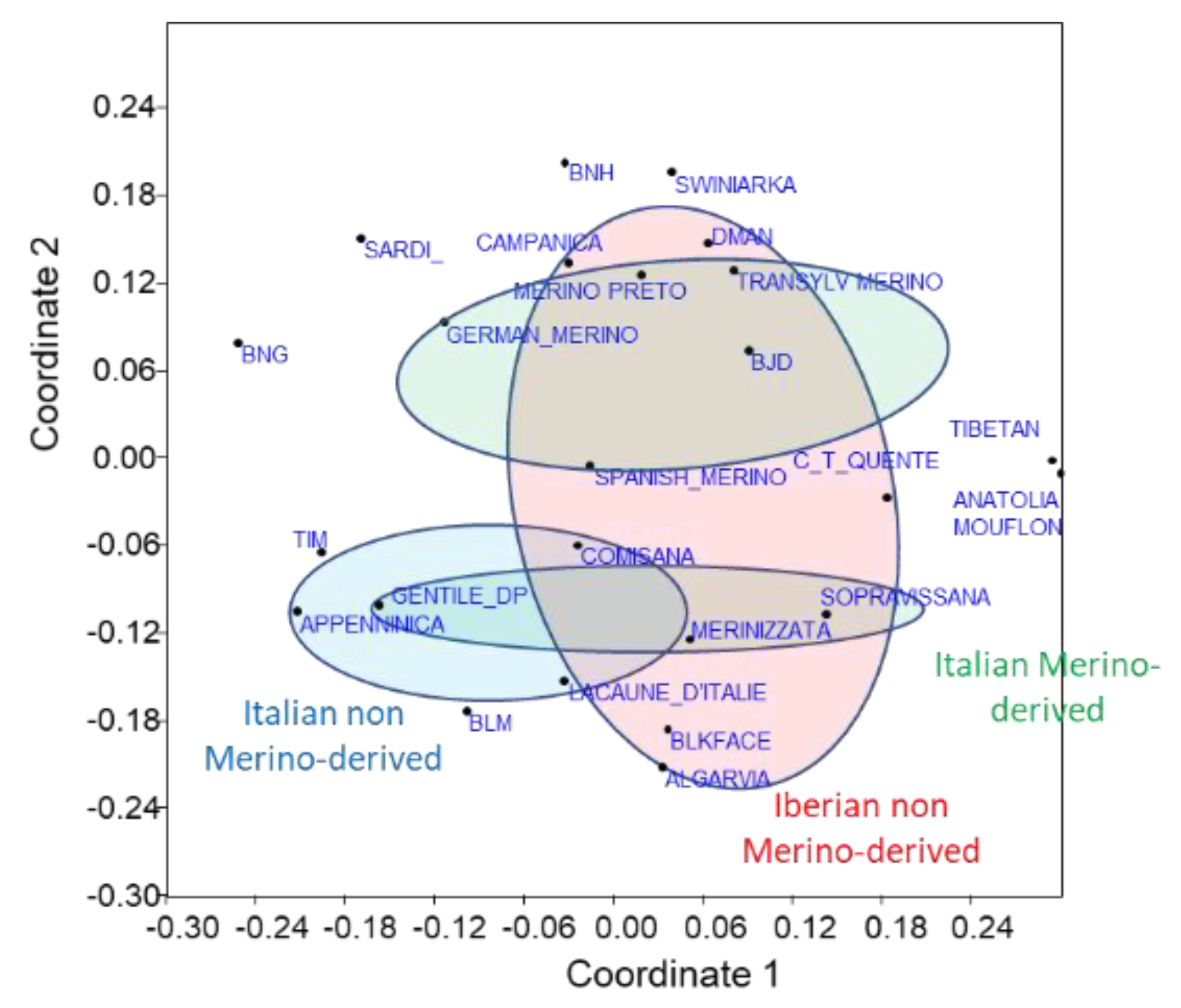
3.2. Relationships with Foreign Breeds
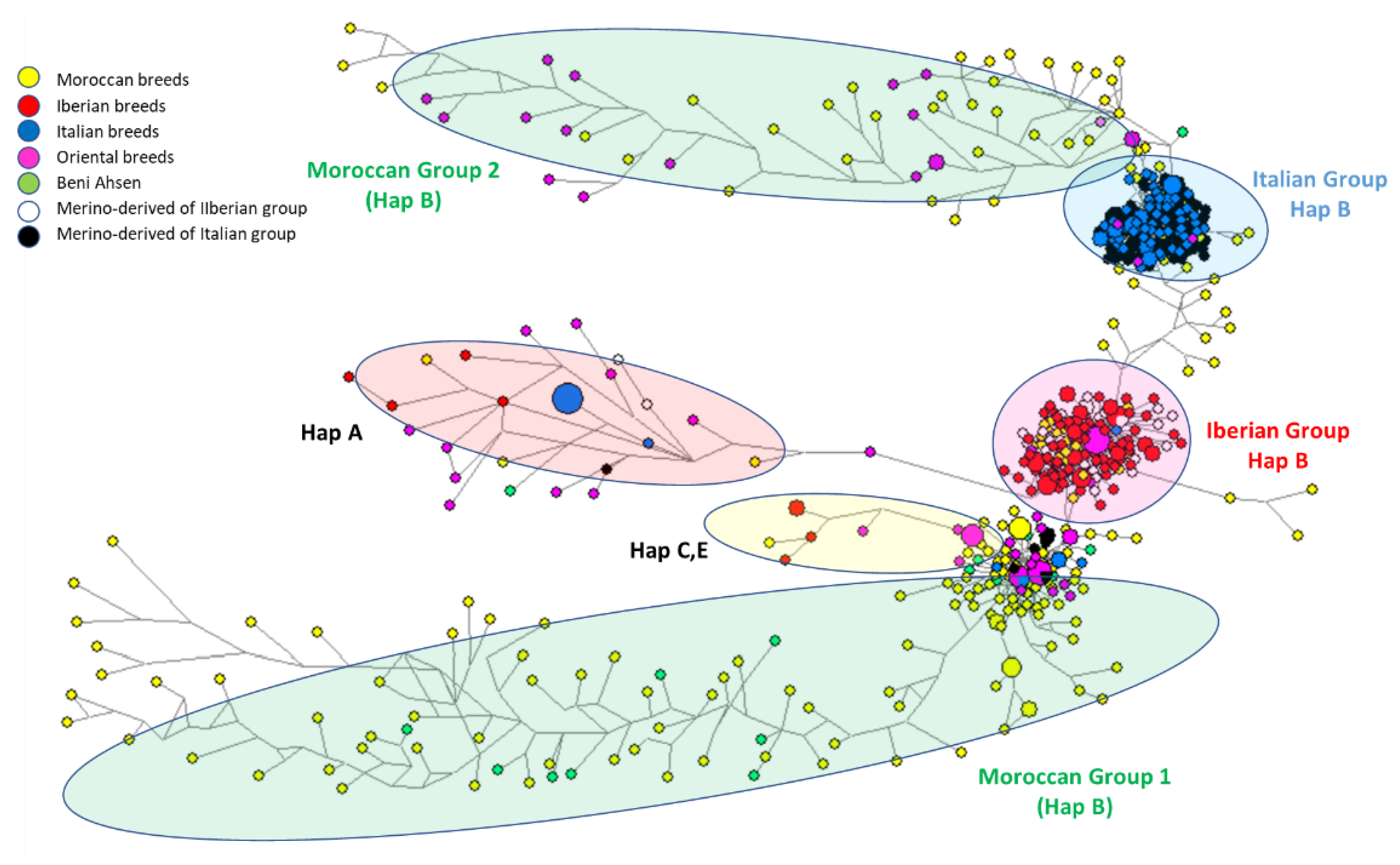
3.2.1. Network Approach
3.2.2. Phylogenetic Approach
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Diez-Tascón, C.; Littlejohn, R.P.; Almeida, P.A.R.; Crawford, A.M. Genetic variation within the Merino sheep breed: Analysis of closely related populations using microsatellites. Anim. Genet. 2000, 31, 243–251. [Google Scholar] [CrossRef] [PubMed]
- Ryder, M.L. Merino history in old wool: The use of wool remains in ancient skin and cloth to study the origin and history of the fine-woolled sheep that became the Spanish Merino. Text. Hist. 1987, 18, 117–132. [Google Scholar] [CrossRef]
- Pedrosa, S.; Arranz, J.J.; Brito, N.; Molina, A.; San Primitivo, F.; Bayón, Y. Mitochondrial diversity and the origin of Iberian sheep. Genet. Sel. Evol. 2007, 39, 91–103. [Google Scholar] [CrossRef] [PubMed]
- Columella, L.J.M. De re Rustica [De L’économie Rurale]; Transl. by Louis Du Bois; P. Panckoucke: Paris, France, 1845; Volume 7, p. 4. [Google Scholar]
- Parry, C.H. An Essay on the Nature, Produce, Origin, & Extension of the Merino Breed of Sheep; W. Bulmer & Company: London, UK, 1807. [Google Scholar]
- Geoffroy Saint-Hilaire, H. L’élevage dans l’Afrique du Nord, Maroc-Algérie-Tunisie; A. Challamel: Paris, France, 1919; pp. 253–257. [Google Scholar]
- Boujenane, I. Les Ressources Génétiques Ovines au Maroc; Actes Editions; IAV Hassan II: Rabat, Morocco, 1999; pp. 69–75. [Google Scholar]
- Sagne, J. L’histoire des populations ovines de l’Afrique du Nord. Zootechnia 1956, 5, 94–106. [Google Scholar]
- Eyraud. La Production Ovine au Maroc. In Les Journées Marocaines du Mouton: 13–15 October 1934; Direction Générale de l’Agriculture: Casablanca, Morocco, 1934; pp. 49–69. [Google Scholar]
- Miegeville, J. Les Ovins. In L’Elevage au Maroc; Vaysse, J., Ed.; La Terre Marocaine: Casablanca, Morocco, 1952; pp. 63–95. [Google Scholar]
- Bourfia, M. Standard de la Beni Ahsen, race ovine marocaine en péril. Anim. Genet. Resour. Inf. 1987, 6, 10–12. [Google Scholar] [CrossRef]
- Boujenane, I.; Petit, D. Between-and within-breed morphological variability in Moroccan sheep breeds. Anim. Genet. Resour. Inf. 2006, 58, 91–100. [Google Scholar] [CrossRef]
- Kandoussi, A.; Boujenane, I.; Auger, C.; Serranito, B.; Germot, A.; Piro, M.; Maftah, A.; Badaoui, B.; Petit, D. The origin of sheep settlement in Western Mediterranean. Sci. Rep. 2020, 10, 1–11. [Google Scholar] [CrossRef]
- Kandoussi, A.; Badaoui, B.; Boujenane, I.; Piro, M.; Petit, D. How have sheep breeds differentiated from each other in Morocco? Genetic structure and geographical distribution patterns. Genet. Sel. Evol. 2021, 53, 1–13. [Google Scholar] [CrossRef]
- Hiendleder, S.; Lewalski, H.; Wassmuth, R.; Janke, A. The complete mitochondrial DNA sequence of the domestic sheep (Ovis aries) and comparison with the other major ovine haplotype. J. Mol. Evol. 1998, 47, 441–448. [Google Scholar] [CrossRef]
- Librado, P.; Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 2009, 25, 1451–1452. [Google Scholar] [CrossRef] [Green Version]
- Hammer, Ø.; Harper, D.A.; Ryan, P.D. PAST: PAleontological STatistics software package for education and data analysis. Palaeontol. Electron. 2001, 4, 9. [Google Scholar]
- Lancioni, H.; Di Lorenzo, P.; Ceccobelli, S.; Perego, U.A.; Miglio, A.; Landi, V.; Antognoni, M.T.; Sarti, F.M.; Lasagna, E.; Achilli, A. Phylogenetic Relationships of Three Italian Merino-Derived Sheep Breeds Evaluated through a Complete Mitogenome Analysis. PLoS ONE 2013, 8, e73712. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Goudet, J. Hierfstat, a package for R to compute and test hierarchical F-statistics. Mol. Ecol. Notes 2005, 5, 184–186. [Google Scholar] [CrossRef] [Green Version]
- Jombart, T.; Devillard, S.; Balloux, F. Discriminant analysis of principal components: A new method for the analysis of genetically structured populations. BMC Genet. 2010, 11, 94. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Bandelt, H.J.; Forster, P.; Röhl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 1999, 16, 37–48. [Google Scholar] [CrossRef]
- Arranz, J.J.; Bayon, Y.; San Primitivo, F. Genetic variation at microsatellite loci in Spanish sheep. Small Rum. Res. 2001, 39, 3–10. [Google Scholar] [CrossRef]
- Ciani, E.; Mastrangelo, S.; Da Silva, A.; Marroni, F.; Ferenčaković, M.; Ajmone-Marsan, P.; Baird, H.; Barbato, M.; Colli, L.; Delvento, C.; et al. On the origin of European sheep as revealed by the diversity of the Balkan breeds and by optimizing population-genetic analysis tools. Gen. Sel. Evol. 2020, 52, 25. [Google Scholar] [CrossRef]
- Ciani, E.; Lasagna, E.; D’Andrea, M.; Alloggio, I.; Marroni, F.; Ceccobelli, S.; Bermejo, J.V.D.; Sarti, F.M.; Kijas, J.; Lenstra, J.A.; et al. Merino and Merino-derived sheep breeds: A genome-wide intercontinental study. Genet. Sel. Evol. 2015, 47, 64. [Google Scholar] [CrossRef] [Green Version]
- Demars, J.; Cano, M.; Drouilhet, L.; Plisson-Petit, F.; Bardou, P.; Fabre, S.; Servin, B.; Sarry, J.; Woloszyn, F.; Mulsant, P.; et al. Genome-wide identification of the mutation underlying fleece variation and discriminating ancestral hairy species from modern woolly sheep. Mol. Biol. Evol. 2017, 34, 1722–1729. [Google Scholar] [CrossRef] [Green Version]
- Koseniuk, A.; Słota, E. Mitochondrial control region diversity in Polish sheep breeds. Arch. Anim. Breed. 2016, 59, 227–233. [Google Scholar] [CrossRef]
- Valera, M.; Arrebola, F.; Juárez, M.; Molina, A. Genetic improvement of wool production in Spanish Merino sheep: Genetic parameters and simulation of selection strategies. Anim. Prod. Sci. 2009, 49, 43–47. [Google Scholar] [CrossRef]

| Morocco | Iberia | Italia | Oriental and Primitive | Other |
|---|---|---|---|---|
| D’man (27) | Churra de Terra Quente (25) | Appenninica (29) | Polish Swiniarka (25) | German Merino (29) |
| Bl. de Montagne (32) | Churra Algarvia (30) | Lacaune of Italy (30) | Tibetan sheep (35) | Transylvanian Merino (6) |
| Boujaad (31) | Latxa Black Face (31) | Comisana (30) | Awassi (3) | |
| Sardi (33) | Campaniça (20) | Gentile de Puglia (30) | Anatolia Mouflon (3) | |
| Timahdite (37) | Merino Preto (10) | Sopravissana (30) | Corsica Mouflon (2) | |
| Beni Guil (31) | Spanish Merino (26) | Merinizzata (30) | ||
| Beni Ahsen (20) |
| N | S | P | Sg | H | Hd | Π | Fu-Li’s F * | Fu-Li’s D * | Fu’s Fs | Tajima D | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Beni Ahsen | 20 | 65 | 22 | 43 | 19 | 0.995 | 0.02011 | −2.367 NS | −2.148 NS | −9.096 *** | −1.75292 NS |
| Bl. de Mont. | 32 | 44 | 16 | 28 | 24 | 0.97 | 0.0133 | −3.016 * | −2.943 * | −13.24 *** | −1.766 NS |
| Boujaad | 31 | 91 | 30 | 61 | 29 | 0.994 | 0.0226 | −3.280 ** | −3.105 * | −18.224 *** | −2.111 * |
| Beni Guil | 31 | 66 | 24 | 42 | 29 | 0.996 | 0.0164 | −2.931 * | −2.741 * | −22.713 *** | −1.953 * |
| D’man | 27 | 41 | 14 | 27 | 21 | 0.963 | 0.0151 | −0.880 NS | −0.634 NS | −7.424 * | −1.744 NS |
| Sardi | 33 | 76 | 42 | 34 | 30 | 0.994 | 0.0203 | −1.821 NS | −1.457 NS | −18.582 *** | −1.692 NS |
| Timahdite | 37 | 56 | 27 | 29 | 32 | 0.988 | 0.0215 | −2.502 NS | −2.242 NS | −25.845 *** | −1.855 * |
| Genetic Dist\FST | D’man | Blanche de Montagne | Boujaad | Sardi | Timahdite | Beni Guil | Beni Ahsen |
|---|---|---|---|---|---|---|---|
| D’man | 0 | 0.0622 | 0.1119 | 0.052 | 0.0517 | 0.0114 | 0.0878 |
| Bl. de Montagne | 0.0082 | 0 | 0.0852 | 0.0233 | 0.0223 | 0.0193 | 0.0394 |
| Boujaad | 0.0172 | 0.0141 | 0 | 0.0761 | 0.0727 | 0.0778 | 0.0927 |
| Sardi | 0.0083 | 0.0054 | 0.0149 | 0 | 0.0234 | 0.0098 | 0.0149 |
| Timahdite | 0.0076 | 0.0049 | 0.0133 | 0.0057 | 0 | 0.0207 | 0.0285 |
| Beni Guil | 0.0038 | 0.0045 | 0.0133 | 0.0042 | 0.0048 | 0 | 0.0374 |
| Beni Ahsen | 0.0126 | 0.0079 | 0.0179 | 0.0062 | 0.0070 | 0.0078 | 0 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kandoussi, A.; Boujenane, I.; Piro, M.; Petit, D.P. Genetic Diversity and Population Structure of Moroccan Beni Ahsen: Is This Endangered Ovine Breed One of the Ancestors of Merino? Ruminants 2022, 2, 201-211. https://doi.org/10.3390/ruminants2020013
Kandoussi A, Boujenane I, Piro M, Petit DP. Genetic Diversity and Population Structure of Moroccan Beni Ahsen: Is This Endangered Ovine Breed One of the Ancestors of Merino? Ruminants. 2022; 2(2):201-211. https://doi.org/10.3390/ruminants2020013
Chicago/Turabian StyleKandoussi, Asmae, Ismaïl Boujenane, Mohammed Piro, and Daniel Pierre Petit. 2022. "Genetic Diversity and Population Structure of Moroccan Beni Ahsen: Is This Endangered Ovine Breed One of the Ancestors of Merino?" Ruminants 2, no. 2: 201-211. https://doi.org/10.3390/ruminants2020013
APA StyleKandoussi, A., Boujenane, I., Piro, M., & Petit, D. P. (2022). Genetic Diversity and Population Structure of Moroccan Beni Ahsen: Is This Endangered Ovine Breed One of the Ancestors of Merino? Ruminants, 2(2), 201-211. https://doi.org/10.3390/ruminants2020013






