Conservation and Selection of Genes Related to Environmental Adaptation in Native Small Ruminant Breeds: A Review
Abstract
:1. Introduction
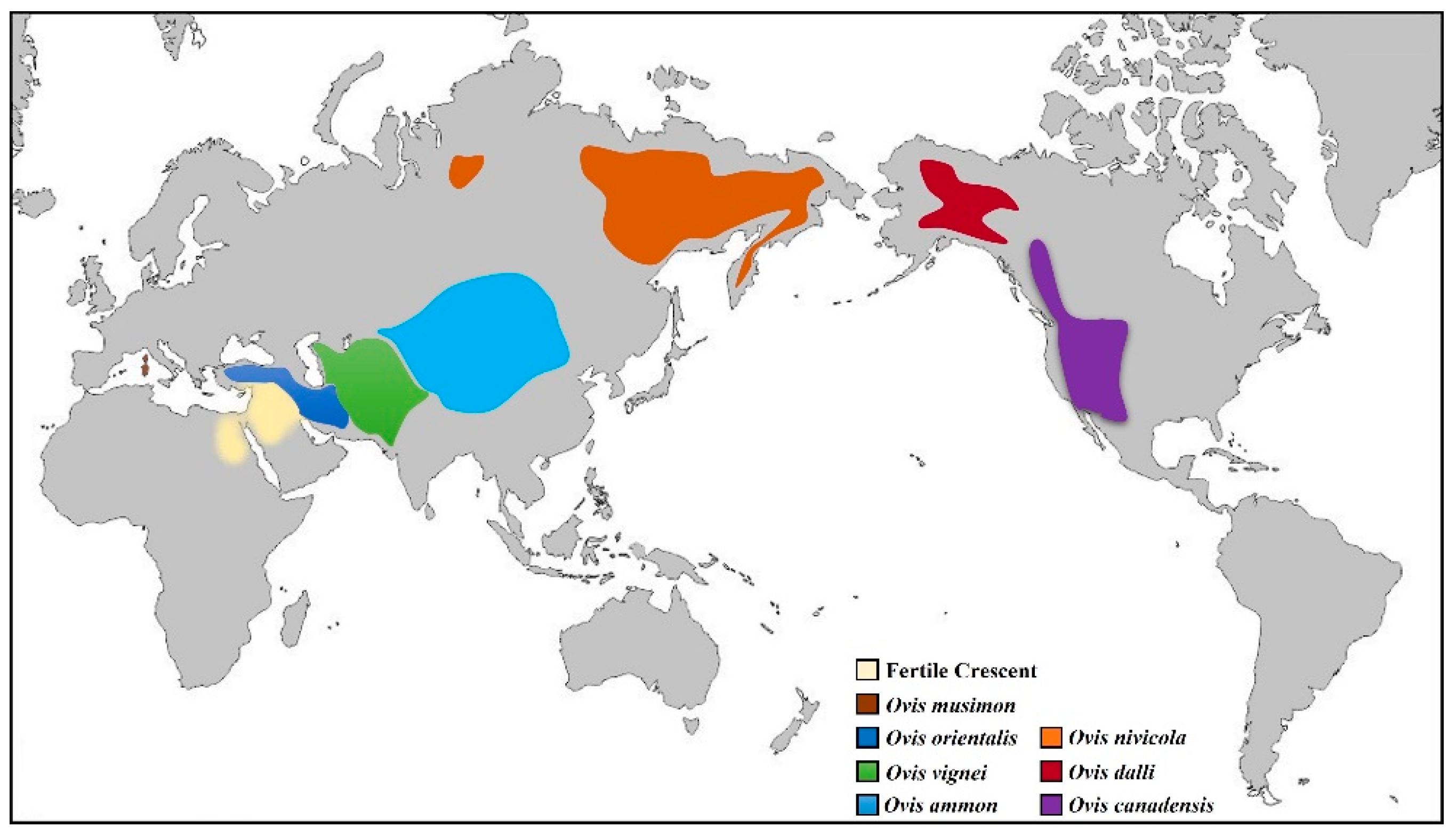
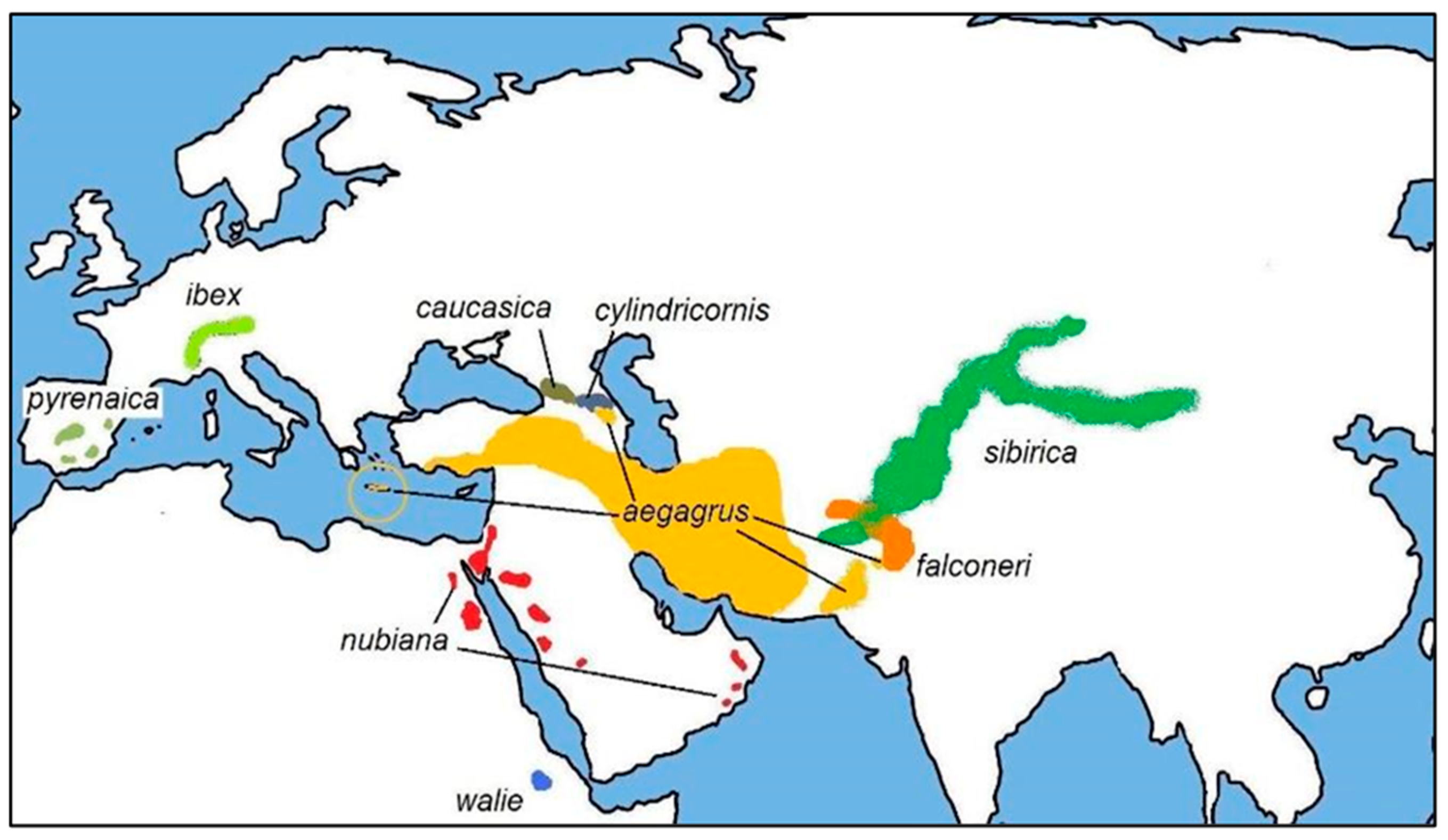
2. Domestication and Evolution of Native Small Ruminant Breeds
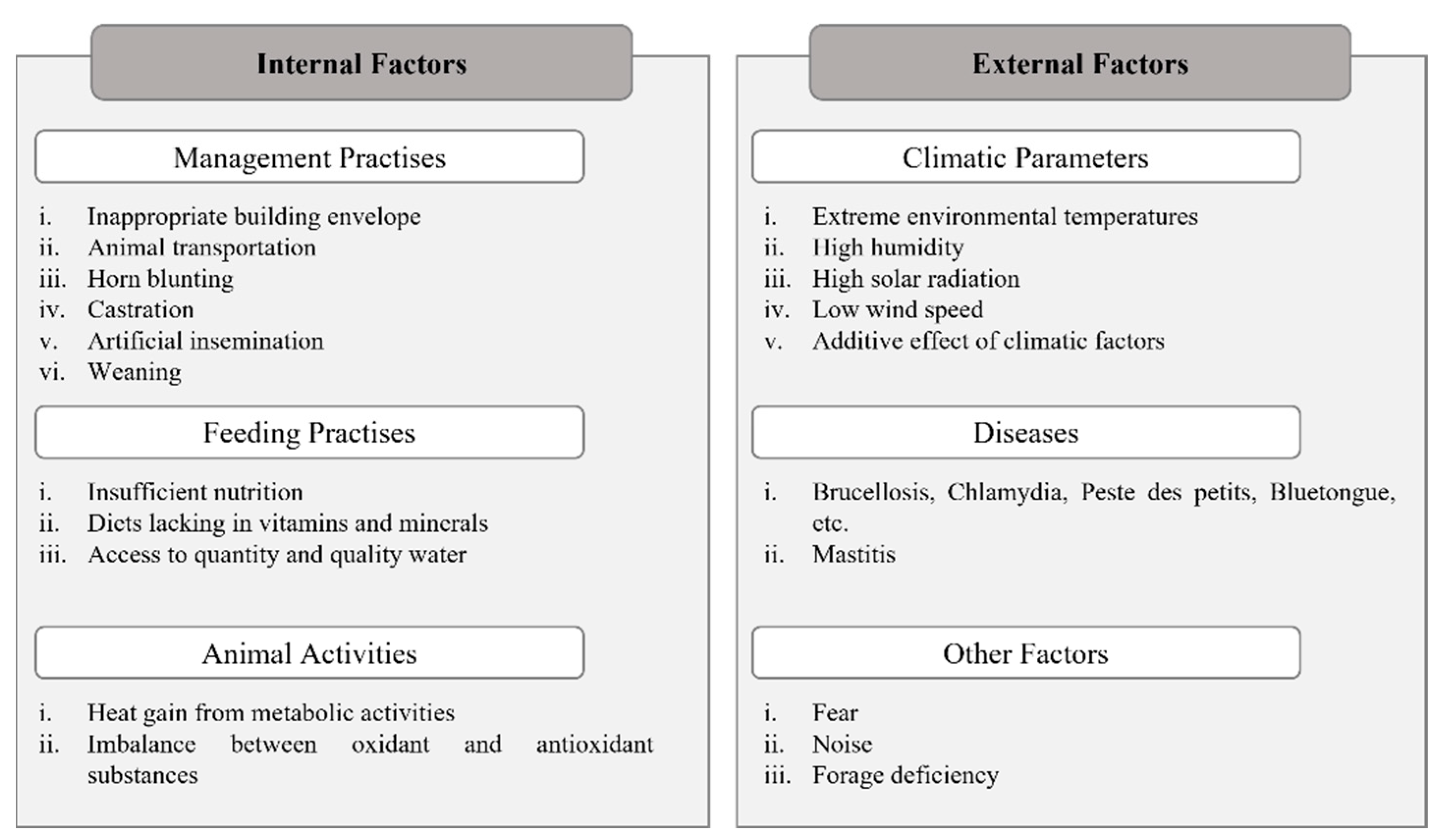
3. Adaptation to Environmental Stressors in Native Small Ruminant Breeds
4. Genes Involved in Environmental Adaptation and Acclimation
5. Conservation Strategies and Current Selection Practices for Genes Related to Environmental Adaptation
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Wilkins, A.S.; Wrangham, R.W.; Fitch, W.T. The “domestication syndrome” in mammals: A unified explanation based on neural crest cell behavior and genetics. Genetics 2014, 197, 795–808. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Amills, M.; Capote, J.; Tosser-Klopp, G. Goat domestication and breeding: A jigsaw of historical, biological and molecular data with missing pieces. Anim. Genet. 2017, 48, 631–644. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rezaei, H.R.; Naderi, S.; Chintauan-Marquier, I.C.; Jordan, S.; Taberlet, P.; Virk, A.T.; Naghash, H.R.; Rioux, D.; Kaboli, M.; Luikart, G.; et al. Evolution and taxonomy of the wild species of the genus Ovis (Mammalia, Artiodactyla, Bovidae). Mol. Phylogenet. Evol. 2010, 54, 315–326. [Google Scholar] [CrossRef] [PubMed]
- Schaller, G.B. Mountain Monarchs: Wild Sheep and Goats of the Himalaya; University of Chicago Press: Chicago, IL, USA; London, UK, 1977. [Google Scholar]
- Shackleton, D.M. Wild Sheep and Goats and Their Relatives: Status, Survey and Conservation Action Plan for Caprinae; IUCN: Gland, Switzerland, 1997.
- Hassanin, A.; Delsuc, F.; Ropiquet, A.; Hammer, C.; van Vuuren, B.J.; Matthee, C.; Ruiz-Garcia, M.; Catzeflis, F.; Areskoug, V.; Nguyen, T.T.; et al. Pattern and timing of diversification of Cetartiodactyla (Mammalia, Laurasiatheria), as revealed by a comprehensive analysis of mitochondrial genomes. C. R. Biol. 2012, 335, 32–50. [Google Scholar] [CrossRef]
- Alberto, F.J.; Boyer, F.; Orozco-terWengel, P.; Streeter, I.; Servin, B.; de Villemereuil, P.; Benjelloun, B.; Librado, P.; Biscarini, F.; Colli, L.; et al. Convergent genomic signatures of domestication in sheep and goats. Nat. Commun. 2018, 9, 813. [Google Scholar] [CrossRef]
- Nomura, K.; Yonezawa, T.; Mano, S.; Kawakami, S.; Shedlock, A.M.; Hasegawa, M.; Amano, T. Domestication process of the goat revealed by an analysis of the nearly complete mitochondrial protein-encoding genes. PLoS ONE 2013, 8, e67775. [Google Scholar] [CrossRef] [Green Version]
- Chessa, B.; Pereira, F.; Arnaud, F.; Amorim, A.; Goyache, F.; Mainland, I.; Kao, R.R.; Pemberton, J.M.; Beraldi, D.; Stear, M.J.; et al. Revealing the history of sheep domestication using retrovirus integrations. Science 2009, 324, 532–536. [Google Scholar] [CrossRef] [Green Version]
- Zeder, M.A. Domestication and early agriculture in the Mediterranean Basin: Origins, diffusion, and impact. Proc. Natl. Acad. Sci. USA 2008, 105, 11597–11604. [Google Scholar] [CrossRef] [Green Version]
- Di Lorenzo, P.; Lancioni, H.; Ceccobelli, S.; Curcio, L.; Panella, F.; Lasagna, E. Uniparental genetic systems: A male and a female perspective in the domestic cattle origin and evolution. Electron. J. Biotech. 2016, 23, 69–78. [Google Scholar] [CrossRef] [Green Version]
- Lancioni, H.; Di Lorenzo, P.; Ceccobelli, S.; Perego, U.A.; Miglio, A.; Landi, V.; Antognoni, M.T.; Sarti, F.M.; Lasagna, E.; Achilli, A. Phylogenetic relationships of three Italian merino-derived sheep breeds evaluated through a complete mitogenome analysis. PLoS ONE 2013, 8, e73712. [Google Scholar] [CrossRef] [Green Version]
- Wanjala, G.; Bagi, Z.; Kusza, S. Meta-Analysis of Mitochondrial DNA Control Region Diversity to Shed Light on Phylogenetic Relationship and Demographic History of African Sheep (Ovis aries) Breeds. Biology 2021, 10, 762. [Google Scholar] [CrossRef] [PubMed]
- Meadows, J.R.; Cemal, I.; Karaca, O.; Gootwine, E.; Kijas, J.W. Five ovine mitochondrial lineages identified from sheep breeds of the near East. Genetics 2007, 175, 1371–1379. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tapio, M.; Marzanov, N.; Ozerov, M.; Ćinkulov, M.; Gonzarenko, G.; Kiselyova, T.; Murawski, M.; Viinalass, H.; Kantanen, J. Sheep mitochondrial DNA variation in European, Caucasian, and Central Asian areas. Mol. Biol. Evol. 2006, 23, 1776–1783. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.H.; Xu, Y.X.; Xie, X.L.; Wang, D.F.; Aguilar-Gómez, D.; Liu, G.J.; Li, X.; Esmailizadeh, A.; Rezaei, V.; Kantanen, J.; et al. Whole-genome sequence analysis unveils different origins of European and Asiatic mouflon and domestication-related genes in sheep. Commun. Biol. 2021, 4, 1307. [Google Scholar] [CrossRef] [PubMed]
- Lv, F.H.; Cao, Y.H.; Liu, G.J.; Luo, L.Y.; Lu, R.; Liu, M.J.; Li, W.R.; Zhou, P.; Wang, X.H.; Shen, M.; et al. Whole-genome resequencing of worldwide wild and domestic sheep elucidates genetic diversity, introgression and agronomically important loci. Mol. Biol. Evol. 2022, 39, msab353. [Google Scholar] [CrossRef]
- Daly, K.G.; Maisano Delser, P.; Mullin, V.E.; Scheu, A.; Mattiangeli, V.; Teasdale, M.D.; Hare, A.; Burger, J.; Pereira Verdugo, M.; Collins, M.J.; et al. Ancient goat genomes reveal mosaic domestication in the Fertile Crescent. Science 2018, 361, 85–88. [Google Scholar] [CrossRef] [Green Version]
- Colli, L.; Lancioni, H.; Cardinali, I.; Olivieri, A.; Capodiferro, M.R.; Pellecchia, M.; Rzepus, M.; Zamani, W.; Naderi, S.; Gandini, F.; et al. Whole mitochondrial genomes unveil the impact of domestication on goat matrilineal variability. BMC Genom. 2015, 16, 1115. [Google Scholar] [CrossRef] [Green Version]
- Naderi, S.; Rezaei, H.R.; Pompanon, F.; Blum, M.G.; Negrini, R.; Naghash, H.R.; Balkiz, Ö.; Mashkour, M.; Gaggiotti, O.E.; Ajmone-Marsan, P.; et al. The goat domestication process inferred from large-scale mitochondrial DNA analysis of wild and domestic individuals. Proc. Natl. Acad. Sci. USA 2008, 105, 17659–17664. [Google Scholar] [CrossRef] [Green Version]
- Zheng, Z.; Wang, X.; Li, M.; Li, Y.; Yang, Z.; Wang, X.; Pan, X.; Gong, M.; Zhang, Y.; Guo, Y.; et al. The origin of domestication genes in goats. Sci. Adv. 2020, 6, eaaz5216. [Google Scholar] [CrossRef]
- Collier, R.J.; Baumgard, L.H.; Zimbelman, R.B.; Xiao, Y. Heat stress: Physiology of acclimation and adaptation. Anim. Front. 2019, 9, 12–19. [Google Scholar] [CrossRef] [Green Version]
- Sunil Kumar, B.V.; Ajeet, K.; Meena, K. Effect of heat stress in tropical livestock and different strategies for its amelioration. J. Stress Physiol. Biochem. 2011, 7, 45–54. [Google Scholar]
- Niyas, P.A.A.; Chaidanya, K.; Shaji, S.; Sejian, V.; Bhatta, R. Adaptation of livestock to environmental challenges. J. Vet. Sci. Med. Diagn. 2015, 4, 3. [Google Scholar]
- Baxevanou, C.; Fidaros, D.; Papanastasiou, D.; Bartzanas, T.; Kittas, C. 2017. Numerical simulation of internal microclimate inside a livestock building. Environ. Eng. Manag. J. 2017, 16, 2513–2523. [Google Scholar] [CrossRef]
- Demir, E.; Bilginer, U.; Balcioglu, M.S.; Karsli, T. Direct and indirect contributions of molecular genetics to farm animal welfare: A review. Anim. Health Res. Rev. 2021, 22, 177–186. [Google Scholar] [CrossRef]
- Petherick, J.C.; Phillips, C.J.C. Space allowances for confined livestock and their determination from allometric principles. Appl. Anim. Behav. Sci. 2009, 117, 1–12. [Google Scholar] [CrossRef]
- Rovelli, G.; Ceccobelli, S.; Perini, F.; Demir, E.; Mastrangelo, S.; Conte, G.; Abeni, F.; Marletta, D.; Ciampolini, R.; Cassandro, M.; et al. The genetics of phenotypic plasticity in livestock in the era of climate change: A review. Ital. J. Anim. Sci. 2020, 19, 997–1014. [Google Scholar] [CrossRef]
- Adams, R.M.; Hurd, B.H.; Lenhart, S.; Leary, N. Effects of global climate change on agriculture: An interpretative review. Clim. Res. 1998, 11, 19–30. [Google Scholar] [CrossRef] [Green Version]
- Perini, F.; Cendron, F.; Rovelli, G.; Castellini, C.; Cassandro, M.; Lasagna, E. Emerging genetic tools to investigate molecular pathways related to heat stress in chickens: A review. Animals 2021, 11, 46. [Google Scholar] [CrossRef]
- Angrecka, S.; Herbut, P.; Mishra, G.; Jena, B.; Dar, M.; Bhat, A. Conditions for cold stress development in dairy cattle kept in free stall barn during severe frosts. Czech. J. Anim. Sci. 2015, 60, 81–87. [Google Scholar] [CrossRef]
- Belhadj Slimen, I.; Najar, T.; Ghram, A.; Abdrrabba, M. Heat stress effects on livestock: Molecular, cellular and metabolic aspects, a review. J. Anim. Physiol. Anim. Nutr. 2016, 100, 401–412. [Google Scholar] [CrossRef] [Green Version]
- McManus, C.M.; Faria, D.A.; Lucci, C.M.; Louvandini, H.; Pereira, S.A.; Paiva, S.R. Heat stress effects on sheep: Are hair sheep more heat resistant? Theriogenology 2020, 155, 157–167. [Google Scholar] [CrossRef] [PubMed]
- Berihulay, H.; Abied, A.; He, X.; Jiang, L.; Ma, Y. Adaptation mechanisms of small ruminants to environmental heat stress. Animals 2019, 9, 75. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Passamonti, M.M.; Somenzi, E.; Barbato, M.; Chillemi, G.; Colli, L.; Joost, S.; Milanesi, M.; Negrini, R.; Santini, M.; Vajana, E.; et al. The Quest for Genes Involved in Adaptation to Climate Change in Ruminant Livestock. Animals 2021, 11, 2833. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, I. Livestock genetic diversity and climate change adaptation. In Proceedings of the International Conference on Livestock and Global Climate Change 2008, Hammamet, Tunisia, 17–20 May 2008; Rowlinson, P., Steele, M., Nefzaoui, A., Eds.; Cambridge University Press: Cambridge, UK, 2008; pp. 76–80. [Google Scholar]
- St-Pierre, N.R.; Cobanov, B.; Schnitkey, G. Economic losses from heat stress by US livestock industries. J. Dai. Sci. 2003, 86, E52–E77. [Google Scholar] [CrossRef] [Green Version]
- Nardone, A.; Ronchi, B.; Lacetera, N.; Ranieri, M.S.; Bernabucci, U. Effects of climate changes on animal production and sustainability of livestock systems. Livest. Sci. 2010, 130, 57–69. [Google Scholar] [CrossRef]
- Sejian, V.; Bhatta, R.; Soren, N.M.; Malik, P.K.; Ravindra, J.P.; Prasad, C.S.; Lal, R. Introduction to concepts of climate change impact on livestock and its adaptation and mitigation. In Climate Change Impact on Livestock: Adaptation and Mitigation; Sejian, V., Gaughan, J., Baumgard, L., Prasad, C., Eds.; Springer: New Delhi, India, 2015; pp. 1–23. [Google Scholar]
- Rojas-Downing, M.M.; Nejadhashemi, A.P.; Harrigan, T.; Woznicki, S.A. Climate change and livestock: Impacts, adaptation, and mitigation. Clim. Risk Manag. 2017, 16, 145–163. [Google Scholar] [CrossRef]
- Escarcha, J.F.; Lassa, J.A.; Zander, K.K. Livestock under climate change: A systematic review of impacts and adaptation. Climate 2018, 6, 54. [Google Scholar] [CrossRef] [Green Version]
- Wreford, A.; Topp, C.F. Impacts of climate change on livestock and possible adaptations: A case study of the United Kingdom. Agric. Syst. 2020, 178, 102737. [Google Scholar] [CrossRef]
- Sejian, V.; Silpa, M.V.; Lees, A.M.; Krishnan, G.; Devaraj, C.; Bagath, M.; Anisha, J.P.; Reshma Nair, M.R.; Manimaran, A.; Bhatta, R.; et al. Opportunities, Challenges, and Ecological Footprint of Sustaining Small Ruminant Production in the Changing Climate Scenario. In Agroecological Footprints Management for Sustainable Food System; Banerjee, A., Meena, R.S., Jhariya, M.K., Yadav, D.K., Eds.; Springer: Singapore, 2021; pp. 365–396. [Google Scholar]
- Moran, D.S.; Eli-Berchoer, L.; Heled, Y.; Mendel, L.; Schocina, M.; Horowitz, M. Heat intolerance: Does gene transcription contribute? J. Appl. Physiol. 2006, 100, 1370–1376. [Google Scholar] [CrossRef] [Green Version]
- Cavalcanti, L.C.G.; Moraes, J.C.F.; Faria, D.A.D.; McManus, C.M.; Nepomuceno, A.R.; Souza, C.J.H.D.; Caetano, A.R.; Paiva, S.R. Genetic characterization of coat color genes in Brazilian Crioula sheep from a conservation nucleus. Pesq. Agropec. Brasil. 2017, 52, 615–622. [Google Scholar] [CrossRef]
- Wei, C.; Wang, H.; Liu, G.; Zhao, F.; Kijas, J.W.; Ma, Y.; Lu, J.; Zhang, L.; Cao, J.; Wu, M.; et al. Genome-wide analysis reveals adaptation to high altitudes in Tibetan sheep. Sci. Rep. 2016, 6, 26770. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yudin, N.; Larkin, D.M. Shared Signatures of Selection Related to Adaptation and Acclimation in Local Cattle and Sheep Breeds from Russia. Russ. J. Genet. 2019, 55, 1008–1014. [Google Scholar] [CrossRef]
- Gorkhali, N.A.; Dong, K.; Yang, M.; Song, S.; Kader, A.; Shrestha, B.S.; He, X.; Zhao, Q.; Pu, Y.; Li, X.; et al. Genomic analysis identified a potential novel molecular mechanism for high-altitude adaptation in sheep at the Himalayas. Sci. Rep. 2016, 6, 29963. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yurchenko, A.A.; Deniskova, T.E.; Yudin, N.S.; Dotsev, A.V.; Khamiruev, T.N.; Selionova, M.I.; Egorov, S.V.; Reyer, H.; Wimmers, K.; Brem, G.; et al. High-density genotyping reveals signatures of selection related to acclimation and economically important traits in 15 local sheep breeds from Russia. BMC Genom. 2019, 20, 294. [Google Scholar] [CrossRef] [PubMed]
- Rochus, C.M.; Tortereau, F.; Plisson-Petit, F.; Restoux, G.; Moreno-Romieux, C.; Tosser-Klopp, G.; Servin, B. Revealing the selection history of adaptive loci using genome-wide scans for selection: An example from domestic sheep. BMC Genom. 2018, 19, 71. [Google Scholar] [CrossRef] [Green Version]
- Lv, F.H.; Agha, S.; Kantanen, J.; Colli, L.; Stucki, S.; Kijas, J.W.; Joost, S.; Li, M.H.; Ajmone Marsan, P. Adaptations to climate-mediated selective pressures in sheep. Mol. Biol. Evol. 2014, 31, 3324–3343. [Google Scholar] [CrossRef] [Green Version]
- Fariello, M.I.; Servin, B.; Tosser-Klopp, G.; Rupp, R.; Moreno, C.; International Sheep Genomics Consortium; San Cristobal, M.; Boitard, S. Selection signatures in worldwide sheep populations. PLoS ONE 2014, 9, e103813. [Google Scholar] [CrossRef] [Green Version]
- Singh, K.M.; Singh, S.; Ganguly, I.; Nachiappan, R.K.; Ganguly, A.; Venkataramanan, R.; Chopra, A.; Narula, H.K. Association of heat stress protein 90 and 70 gene polymorphism with adaptability traits in Indian sheep (Ovis aries). Cell Stress Chaperones 2017, 22, 675–684. [Google Scholar] [CrossRef] [Green Version]
- Wen, Y.; Hu, J.; Wang, J.; Liu, X.; Li, S.; Luo, Y. Effect of glycolysis and heat shock proteins on hypoxia adaptation of Tibetan sheep at different altitude. Gene 2021, 803, 145893. [Google Scholar] [CrossRef]
- Wiener, P.; Robert, C.; Ahbara, A.; Salavati, M.; Abebe, A.; Kebede, A.; Wragg, D.; Friedrich, J.; Vasoya, D.; Hume, D.A.; et al. Whole-Genome Sequence Data Suggest Environmental Adaptation of Ethiopian Sheep Populations. Genome Biol. Evol. 2021, 13, evab014. [Google Scholar] [CrossRef]
- Kaushik, R.; Dige, M.S.; Rout, P.K. Molecular characterization and expression profiling of ENOX2 gene in response to heat stress in goats. Cell Dev. Biol. 2016, 5, 1–5. [Google Scholar]
- Kim, E.S.; Elbeltagy, A.R.; Aboul-Naga, A.M.; Rischkowsky, B.; Sayre, B.; Mwacharo, J.M.; Rothschild, M.F. Multiple genomic signatures of selection in goats and sheep indigenous to a hot arid environment. Heredity 2016, 116, 255–264. [Google Scholar] [CrossRef] [PubMed]
- Onzima, R.B.; Upadhyay, M.R.; Doekes, H.P.; Brito, L.; Bosse, M.; Kanis, E.; Groenen, M.A.M.; Crooijmans, R.P.M.A. Genome-wide characterization of selection signatures and runs of homozygosity in Ugandan goat breeds. Front. Genet. 2018, 9, 318. [Google Scholar] [CrossRef] [PubMed]
- Yadav, V.P.; Dangi, S.S.; Chouhan, V.S.; Gupta, M.; Dangi, S.K.; Singh, G.; Maurya, V.P.; Kumar, P.; Sarkar, M. Expression analysis of NOS family and HSP genes during thermal stress in goat (Capra hircus). Int. J. Biometeorol. 2016, 60, 381–389. [Google Scholar] [CrossRef]
- Kumar, D.; Yadav, B.; Choudhury, S.; Kumari, P.; Madan, A.K.; Singh, S.P.; Rout, P.K.; Ramchandran, N.; Yadav, S. Evaluation of adaptability to different seasons in goat breeds of semi-arid region in India through differential expression pattern of heat shock protein genes. Biol. Rhythm Res. 2018, 49, 466–478. [Google Scholar] [CrossRef]
- Aleena, J.; Sejian, V.; Bagath, M.; Krishnan, G.; Beena, V.; Bhatta, R. Resilience of three indigenous goat breeds to heat stress based on phenotypic traits and pbmc hsp70 expression. Int. J. Biometeorol. 2018, 62, 1995–2005. [Google Scholar] [CrossRef]
- Bertolini, F.; Servin, B.; Talenti, A.; Rochat, E.; Kim, E.S.; Oget, C.; Palhière, I.; Crisà, A.; Catillo, G.; Steri, R.; et al. Signatures of selection and environmental adaptation across the goat genome post-domestication. Genet. Sel. Evol. 2018, 50, 57. [Google Scholar] [CrossRef]
- Roti Roti, J.L. Cellular responses to hyperthermia (40–46 °C): Cell killing and molecular events. Int. J. Hyperth. 2008, 24, 3–15. [Google Scholar] [CrossRef]
- Wang, Q.; Wang, J.; Wang, G.; Wu, C.; Li, J. Molecular cloning, sequencing, and expression profiles of heat shock protein 90 (HSP90) in Hyriopsis cumingii exposed to different stressors: Temperature, cadmium and Aeromonas hydrophila. Aquac. Fish. 2017, 2, 59–66. [Google Scholar] [CrossRef]
- Hoffmann, A.A.; Sørensen, J.G.; Loeschcke, V. Adaptation of Drosophila to temperature extremes: Bringing together quantitative and molecular approaches. J. Therm. Biol. 2003, 28, 175–216. [Google Scholar] [CrossRef]
- Banerjee, D.; Upadhyay, R.C.; Chaudhary, U.B.; Kumar, R.; Singh, S.; Polley, S.; Mukherjee, A.; Das, T.K.; De, S. Seasonal variation in expression pattern of genes under HSP70. Cell Stress Chaperones 2014, 19, 401–408. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hooper, H.B.; dos Santos Silva, P.; de Oliveira, S.A.; Merighe, G.K.F.; Negrão, J.A. Acute heat stress induces changes in physiological and cellular responses in Saanen goats. Int. J. Biometeorol. 2018, 62, 2257–2265. [Google Scholar] [CrossRef] [PubMed]
- Mohanarao, G.J.; Mukherjee, A.; Banerjee, D.; Gohain, M.; Dass, G.; Brahma, B.; Datta, T.K.; Upadhyay, R.C.; De, S. Hsp70 family genes and hsp27 expression in response to heat and cold stress in vitro in peripheral blood mononuclear cells of goat (capra hircus). Small Rumin. Res. 2014, 116, 94–99. [Google Scholar] [CrossRef]
- Volloch, V.; Rits, S. A natural extracellular factor that induces Hsp72, inhibits apoptosis, and restores stress resistance in aged human cells. Exp. Cell Res. 1999, 253, 483–492. [Google Scholar] [CrossRef] [PubMed]
- Rhoads, R.P.; Baumgard, L.H.; Suagee, J.K.; Sanders, S.R. Nutritional interventions to alleviate the negative consequences of heat stress. Adv. Nutr. 2013, 4, 267–276. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Silanikove, N. Effects of heat stress on the welfare of extensively managed domestic ruminants. Livest. Prod. Sci. 2000, 67, 1–18. [Google Scholar] [CrossRef]
- Naandam, J.; Assan, I.K. Effect of coat color, ecotype, location and sex on hair density of West African Dwarf (WAD) goats in Northern Ghana. Sky J. Agric. Res. 2014, 3, 25–30. [Google Scholar]
- Hubbard, J.K.; Uy, J.A.K.; Hauber, M.E.; Hoekstra, H.E.; Safran, R.J. Vertebrate pigmentation: From underlying genes to adaptive function. Trends Genet. 2010, 26, 231–239. [Google Scholar] [CrossRef]
- Cieslak, M.; Reissmann, M.; Hofreiter, M.; Ludwig, A. Colours of domestication. Biol. Rev. 2011, 86, 885–899. [Google Scholar] [CrossRef]
- McManus, C.; Paludo, G.R.; Louvandini, H.; Gugel, R.; Sasaki, L.C.B.; Paiva, S.R. Heat tolerance in Brazilian sheep: Physiological and blood parameters. Trop. Anim. Health Prod. 2009, 41, 95–101. [Google Scholar] [CrossRef]
- Ross, T.T.; Goode, L.; Linnerud, A.C. Effects of high ambient temperature on respiration rate, rectal temperature, fetal development and thyrold gland activity in tropical and temperate breeds of sheep. Theriogenology 1985, 24, 259–269. [Google Scholar] [CrossRef]
- Groeneveld, L.F.; Lenstra, J.A.; Eding, H.; Toro, M.A.; Scherf, B.; Pilling, D.; Negrini, R.; Finlay, E.K.; Jianlin, H.; Groeneveld, E.; et al. Genetic diversity in farm animals—A review. Anim. Genet. 2010, 41, 6–31. [Google Scholar] [CrossRef] [Green Version]
- Mahmoudi, B.; Bayat, M.; Sadeghi, R.; Babayev, M.S.; Abdollahi, H. Genetic diversity among three goat populations assessed by microsatellite DNA markers in Iran. Glob. Vet. 2010, 4, 118–124. [Google Scholar]
- Fabuel, E.; Barragan, C.; Silio, L.; Rodriguez, M.C.; Toro, M.A. Analysis of genetic diversity and conservation priorities in Iberian pigs based on microsatellite markers. Heredity 2004, 93, 104–113. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Karsli, T.; Demir, E.; Fidan, H.G.; Aslan, M.; Karsli, B.A.; Arik, I.Z.; Semerci, E.S.; Karabag, K.; Balcioglu, M.S. Determination of genetic variability, population structure and genetic differentiation of indigenous Turkish goat breeds based on SSR loci. Small Rumin. Res. 2020, 190, 106147. [Google Scholar] [CrossRef]
- Toro, M.A.; Caballero, A. Characterization and conservation of genetic diversity in subdivided populations. Philos. Trans. R. Soc. B 2005, 360, 1367–1378. [Google Scholar] [CrossRef] [Green Version]
- Karsli, A.B.; Demir, E.; Fidan, H.G.; Karsli, T. Assessment of genetic diversity and differentiation among four indigenous Turkish sheep breeds using microsatellites. Arch. Anim. Breed. 2020, 63, 165–172. [Google Scholar] [CrossRef]
- Malhi, G.S.; Kaur, M.; Kaushik, P. Impact of climate change on agriculture and its mitigation strategies: A review. Sustainability 2021, 13, 1318. [Google Scholar] [CrossRef]
- Cheng, M.; McCarl, B.; Chengcheng, F. Climate Change and Livestock Production: A Literature Review. Atmosphere 2022, 13, 140. [Google Scholar] [CrossRef]
- Lewis Baida, B.E.; Swinbourne, A.M.; Barwick, J.; Leu, S.T.; van Wettere, W.H. Technologies for the automated collection of heat stress data in sheep. Anim. Biotelemetry 2021, 9, 1–15. [Google Scholar] [CrossRef]
- Reynecke, D.P.; Waghorn, T.S.; Oliver, A.M.B.; Miller, C.M.; Vlassoff, A.; Leathwick, D.M. Dynamics of the free-living stages of sheep intestinal parasites on pasture in the North Island of New Zealand. 2. Weather variables associated with development. N. Z. Vet. J. 2011, 59, 287–292. [Google Scholar] [CrossRef] [PubMed]
- Guerrero, A.; Campo, M.D.M.; Olleta, J.L.; Sanudo, C. Carcass and meat quality in goat. In Goat Science; Kukovics, S., Ed.; IntechOpen: London, UK, 2018; pp. 154–196. [Google Scholar]
- Mohapatra, A.; Shinde, A.K. Fat-tailed sheep-an important sheep genetic resource for meat production in tropical countries: An overview. Indian J. Small Rumin. 2018, 24, 1–17. [Google Scholar] [CrossRef]
- Mapiye, C.; Chikwanha, O.C.; Chimonyo, M.; Dzama, K. Strategies for sustainable use of indigenous cattle genetic resources in Southern Africa. Diversity 2019, 11, 214. [Google Scholar] [CrossRef] [Green Version]
- Monau, P.; Raphaka, K.; Zvinorova-Chimboza, P.; Gondwe, T. Sustainable utilization of indigenous goats in southern Africa. Diversity 2020, 12, 20. [Google Scholar] [CrossRef] [Green Version]
- Aslan, M.; Demir, E.; Karsli, T. Microsatellite diversity and restriction enzyme-based polymorphisms of MHC loci in some native Turkish goats. J. Agric. Sci. 2022, in press. [Google Scholar]
- Ajmone-Marsan, P.; Negrini, R.; Crepaldi, P.; Milanesi, E.; Gorni, C.; Valentini, A.; Cicogna, M. Assessing genetic diversity in Italian goat populations using AFLP® markers. Anim. Genet. 2001, 32, 281–288. [Google Scholar] [CrossRef]
- Bozzi, R.; Alvarez, I.; Crovetti, A.; Fernande, I.; De Petris, D.; Goyache, F. Assessing priorities for conservation in Tuscan cattle breeds using microsatellites. Animal 2012, 6, 203–211. [Google Scholar] [CrossRef] [Green Version]
- Karsli, T. Assessment of genetic diversity and conservation priorities in some Turkish indigenous Hair goat populations by microsatellite loci. Indian J. Anim. Sci. 2020, 90, 728–733. [Google Scholar]
- Mateescu, R.G.; Thonney, M.L. Genetic mapping of quantitative trait loci for milk production in sheep. Anim. Genet. 2010, 41, 460–466. [Google Scholar] [CrossRef]
- Chen, X.; Sun, H.; Tian, S.; Xiang, H.; Zhou, L.; Dun, W.; Zhao, X. Increasing litter size in a sheep breed by marker-assisted selection of BMPR1B A746G mutation. J. Genet. 2015, 94, 139–142. [Google Scholar] [CrossRef]
- Allendorf, F.; Hohenlohe, P.; Luikart, G. Genomics and the future of conservation genetics. Nat. Rev. Genet. 2010, 11, 697–709. [Google Scholar] [CrossRef] [PubMed]
- Yan, Z.; Wang, Z.; Zhang, Q.; Yue, S.; Yin, B.; Jiang, Y.; Shi, K. Identification of whole-genome significant single nucleotide polymorphisms in candidate genes associated with body conformation traits in Chinese Holstein cattle. Anim. Genet. 2020, 51, 141–146. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Weller, J.I.; Seroussi, E.; Ron, M. Estimation of the number of genetic markers required for individual animal identification accounting for genotyping errors. Anim. Genet. 2006, 37, 387–389. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, M.E.; Goszczynski, D.E.; Liron, J.; Villegas-Castagnasso, E.E.; Carino, M.H. Comparison of the effectiveness of microsatellites and SNP panels for genetic identification, traceability and assessment of parentage in an inbred Angus herd. Genet. Mol. Biol. 2013, 36, 185–191. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Edea, Z.; Dadi, H.; Dessie, T.; Kim, K.S. Genomic signatures of high-altitude adaptation in Ethiopian sheep populations. Genes Genom. 2019, 41, 973–981. [Google Scholar] [CrossRef]
- Cao, Y.H.; Xu, S.S.; Shen, M.; Chen, Z.H.; Gao, L.; Lv, F.H.; Xie, X.L.; Wang, X.H.; Yang, H.; Liu, C.B.; et al. Historical Introgression from Wild Relatives Enhanced Climatic Adaptation and Resistance to Pneumonia in Sheep. Mol. Biol. Evol. 2021, 38, 838–855. [Google Scholar] [CrossRef]
- Garner, J.; Douglas, M.; Williams, S.R.O.; Wales, W.J.; Marett, L.C.; Nguyen, T.T.T.; Reich, C.M.; Hayes, B.J. Genomic Selection Improves Heat Tolerance in Dairy Cattle. Sci. Rep. 2016, 6, 34114. [Google Scholar] [CrossRef]
- Jin, M.; Lu, J.; Fei, X.; Lu, Z.; Quan, K.; Liu, Y.; Chu, M.; Di, R.; Wei, C.; Wang, H. Selection Signatures Analysis Reveals Genes Associated with High-Altitude Adaptation in Tibetan Goats from Nagqu, Tibet. Animals 2020, 10, 1599. [Google Scholar] [CrossRef]
- Ahmad, S.F.; Mehrotra, A.; Charles, S.; Ganai, N.A. Analysis of selection signatures reveals important insights into the adaptability of high-altitude Indian sheep breed Changthangi. Gene 2021, 799, 145809. [Google Scholar] [CrossRef]
- Yang, J.; Li, W.R.; Lv, F.H.; He, S.G.; Tian, S.L.; Peng, W.F.; Sun, Y.W.; Zhao, Y.X.; Tu, X.L.; Zhang, M.; et al. Whole-Genome Sequencing of Native Sheep Provides Insights into Rapid Adaptations to Extreme Environments. Mol. Biol. Evol. 2016, 13, 2576–2592. [Google Scholar] [CrossRef] [Green Version]
- Sweet-Jones, J.; Lenis, V.P.; Yurchenko, A.A.; Yudin, N.S.; Swain, M.; Larkin, D.M. Genotyping and Whole-Genome Resequencing of Welsh Sheep Breeds Reveal Candidate Genes and Variants for Adaptation to Local Environment and Socioeconomic Traits. Front. Genet. 2021, 12, 612492. [Google Scholar] [CrossRef] [PubMed]
- Perez-Enciso, M.; Rincon, J.C.; Legarra, A. Sequence- vs. chip-assisted genomic selection: Accurate biological information is advised. Genet. Sel. Evol. 2015, 47, 43. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Eusebi, P.G.; Martinez, A.; Cortes, O. Genomic Tools for Effective Conservation of Livestock Breed Diversity. Diversity 2020, 12, 8. [Google Scholar] [CrossRef] [Green Version]
- Stranden, I.; Kantanen, J.; Russo, I.R.M.; Orozco-terWengel, P.; Bruford, M.W.; The Climgen Consortium. Genomic selection strategies for breeding adaptation and production in dairy cattle under climate change. Heredity 2019, 123, 307–317. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Osei-Amponsah, R.; Chauhan, S.S.; Leury, B.J.; Cheng, L.; Cullen, B.; Clarke, I.J.; Dunshea, F.R. Genetic selection for thermotolerance in ruminants. Animals 2019, 9, 948. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wakchaure, R.; Ganguly, S.; Praveen, P.K.; Kumar, A.; Sharma, S.; Mahajan, T. Marker assisted selection (MAS) in animal breeding: A review. J. Drug. Metab. Toxicol. 2015, 6, e127. [Google Scholar] [CrossRef]
- Chang, L.Y.; Toghiani, S.; Ling, A.; Aggrey, S.E.; Rekaya, R. High density marker panels, SNPs prioritizing and accuracy of genomic selection. BMC Genet. 2018, 19, 4. [Google Scholar]
- Zhang, F.; Zhu, F.; Yang, F.X.; Hao, J.P.; Hou, Z.C. Genomic selection for meat quality traits in Pekin duck. Anim. Genet. 2022, 53, 94–100. [Google Scholar] [CrossRef]
- Zhang, M.; Luo, H.; Xu, L.; Shi, Y.; Zhou, J.; Wang, D.; Zhang, X.; Huang, X.; Wang, Y. Genomic Selection for Milk Production Traits in Xinjiang Brown Cattle. Animals 2022, 12, 136. [Google Scholar] [CrossRef]
- Granleese, T.; Clark, S.A.; van der Werf, J.H. Genotyping strategies of selection candidates in livestock breeding programmes. J. Anim. Breed. Genet. 2019, 136, 91–101. [Google Scholar] [CrossRef]
- Zvinorova, P.I.; Halimani, T.E.; Muchadeyi, F.C.; Matika, O.; Riggio, V.; Dzama, K. Breeding for resistance to gastrointestinal nematodes–the potential in low-input/output small ruminant production systems. Vet. Parasitol. 2016, 225, 19–28. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schiavo, G.; Bovo, S.; Ribani, A.; Moscatelli, G.; Bonacini, M.; Prandi, M.; Mancin, E.; Mantovani, R.; Dall’Olio, S.; Fontanesi, L. Comparative analysis of inbreeding parameters and runs of homozygosity islands in 2 Italian autochthonous cattle breeds mainly raised in the Parmigiano-Reggiano cheese production region. J. Dairy Sci. 2022, 105, 2408–2425. [Google Scholar] [CrossRef] [PubMed]
| Features | Species | Breed | Related Gene | Detection Method | References |
|---|---|---|---|---|---|
| Coat Colour | Sheep | Crioula sheep | MCIR, ASIP, TYRP1 | Sequencing | [45] |
| High-altitude hypoxia adaptation | Sheep | Tibetan sheep | EPAS1, CRYAA, LONP1, NF1, DPP4, SOD1, PPARG, SOCS2 | Sequencing | [46] |
| Adaptation to cold climate | Sheep | Russian sheep | NEB, APOB | SNP array | [47] |
| Protection against pulmonary injuries | Sheep | Nepal, Asian and Middle East breeds | FGF-7 | SNP array | [48] |
| Coat Colour | Sheep | Lezgin, Karachaev, Karakul, Edilbai, Romanov, Russian Longhaired, Groznensk, Salsk, Volgograd, Buubei, Tuva, Altai Mountain, Krasnoyarsk, Baikal, and Kulundin | KIT, KITLG | SNP array | [49] |
| Reproduction | CMTM6, HTRA1, GNAQ, UBQLN1, IFT88 | ||||
| Environmental adaptations | EGFR, HSPH1, NMUR1, EDNRB, PRL, TSHR, ADAMTS5 | ||||
| Coat Colour | Sheep | French sheep | ASIP, MC1R, TYRP1, MITF, EDN3, BNC2 | SNP array | [50] |
| Stature and morphology | NPR2, MSTN (GDF-8), LCORL, NCAPG, ALX4, EXT2, PALLD | ||||
| Horns | RXFP2 | ||||
| Wool | IRF2BP2 | ||||
| Energy and regulation activities | Sheep | Autochthonous sheep | TBC1D12 | SNP array | [51] |
| Cold tolerance | Sheep | Worldwide Sheep Populations | TRPM8 | Sequencing | [52] |
| Heat Stress | Sheep | Chokla, Magra, Marwari, and Madras Red | HSP90AA1, HSPA1A, HSPA8 | RT-PCR | [53] |
| Tibetan sheep | HSP27, HSP60 | RT-PCR | [54] | ||
| Altitude adaptation | Sheep | Ethiopian sheep populations | PLCB1, PLCE1, FHAD1, GNA12, KLF12, SUSD4, ZNF407 | Sequencing | [55] |
| Heat Stress | Goat | Barbari, Jakhrana, Sirohi and Jamunapari | ENOX2 | RT-PCR | [56] |
| Barki | EIF2B3 | SNP array | [57] | ||
| Karamojong | KPNA4, MTOR, SH2B1, MAPK3 | SNP array | [58] | ||
| Barbari | HSP90, HSP70 | RT-PCR | [59] | ||
| Barbari, Sirohi, and Jhakrana | HSP60, HSP70, HSP90 | RT-PCR | [60] | ||
| Osmanabadi, Malabari, and Salem Black | HSP70 | RT-PCR | [61] | ||
| Coat Colour | Goat | Unspecified | ADAMTS20, MC1R, ASIP, SOX18, TIMP3 | SNP array | [62] |
| Against oxidative stress | GPR37L1, INS | ||||
| Adaptation to hot arid environments | Sheep and Goat | Barki | BMP2, FGF | SNP array | [57] |
| Thermo-tolerance (melanogenesis), body size and development, energy, and digestive metabolism | Sheep and Goat | Barki | FGF2, GNAI3, PLCB1, BMP2, BMP4, GJA3, GJB2, MYH, TRHDE, ALDH1A3, GRIA1, IL2, IL7, IL21, IL1R1 | SNP array | [57] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Demir, E.; Ceccobelli, S.; Bilginer, U.; Pasquini, M.; Attard, G.; Karsli, T. Conservation and Selection of Genes Related to Environmental Adaptation in Native Small Ruminant Breeds: A Review. Ruminants 2022, 2, 255-270. https://doi.org/10.3390/ruminants2020017
Demir E, Ceccobelli S, Bilginer U, Pasquini M, Attard G, Karsli T. Conservation and Selection of Genes Related to Environmental Adaptation in Native Small Ruminant Breeds: A Review. Ruminants. 2022; 2(2):255-270. https://doi.org/10.3390/ruminants2020017
Chicago/Turabian StyleDemir, Eymen, Simone Ceccobelli, Umit Bilginer, Marina Pasquini, George Attard, and Taki Karsli. 2022. "Conservation and Selection of Genes Related to Environmental Adaptation in Native Small Ruminant Breeds: A Review" Ruminants 2, no. 2: 255-270. https://doi.org/10.3390/ruminants2020017
APA StyleDemir, E., Ceccobelli, S., Bilginer, U., Pasquini, M., Attard, G., & Karsli, T. (2022). Conservation and Selection of Genes Related to Environmental Adaptation in Native Small Ruminant Breeds: A Review. Ruminants, 2(2), 255-270. https://doi.org/10.3390/ruminants2020017