Monitoring Dormancy Transition in Almond [Prunus Dulcis (Miller) Webb] during Cold and Warm Mediterranean Seasons through the Analysis of a DAM (Dormancy-Associated MADS-Box) Gene
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. Chilling Requirements Evaluation
2.3. cDNA Isolation and Cloning
2.4. Phylogenetic Analysis
2.5. Gene Expression Analysis
3. Results
3.1. Chilling Requirements Evaluation
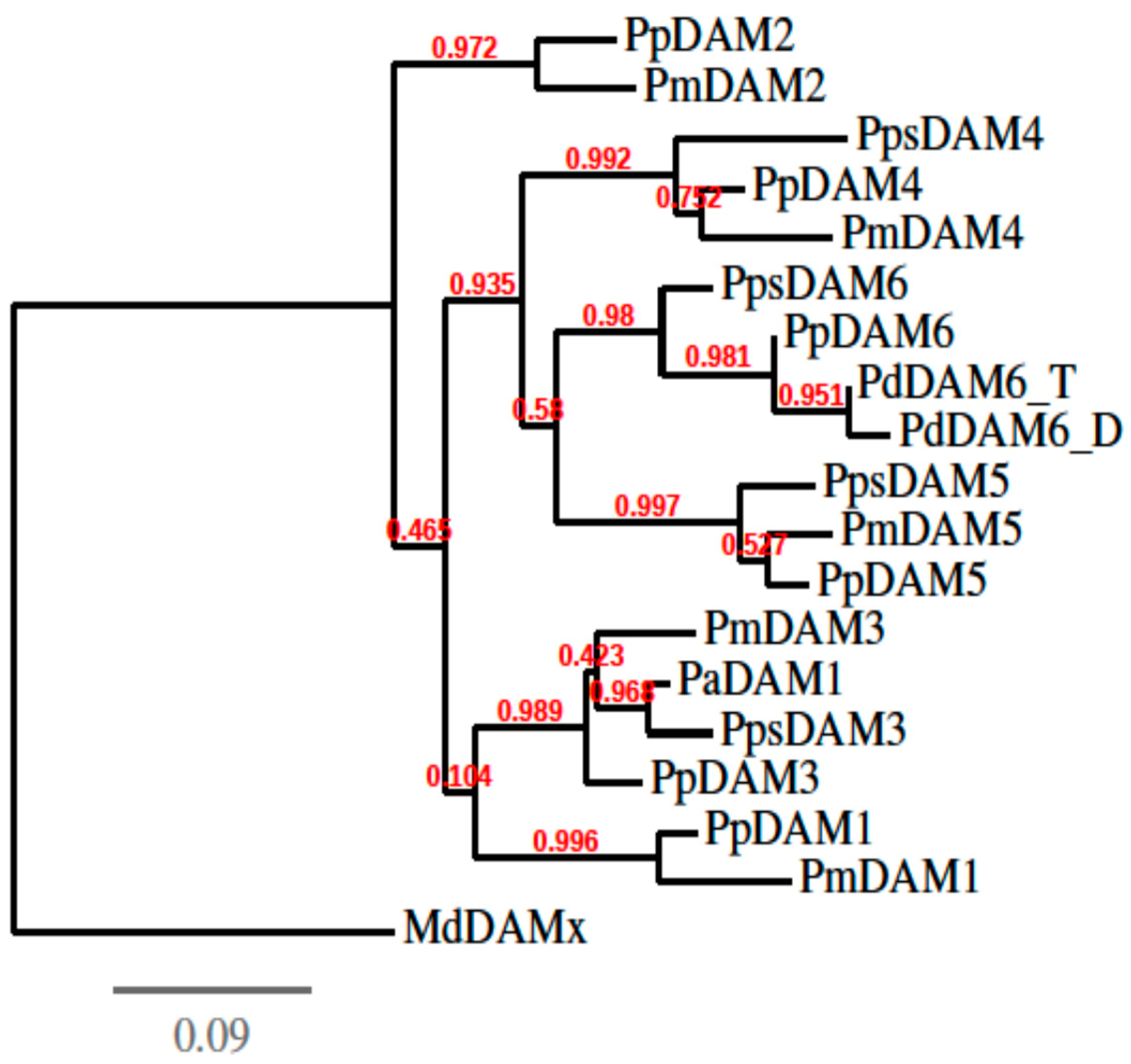
3.2. cDNA Isolation, Cloning and Phylogenetic Analysis
3.3. Expression Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Lloret, A.; Badenes, M.L.; Ríos, G. Modulation of Dormancy and Growth Responses in Reproductive Buds of Temperate Trees. Front. Plant Sci. 2018, 9, 1368. [Google Scholar] [CrossRef] [PubMed]
- Lang, B.A.; Early, J.D.; Martin, G.C.; Darnell, R.L. Endo-, para- and ecodormancy: Physiological terminology and classification for dormancy research. Hort. Sci 1987, 22, 371–377. [Google Scholar]
- Egea, J.; Ortega, E.; Martínez-Gómez, P.; Dicenta, F. Chilling and heat requirements of almond cultivars for flowering. Environ. Exp. Bot. 2003, 50, 79–85. [Google Scholar] [CrossRef]
- Campoy, J.A.; Ruiz, D.; Egea, J. Dormancy in temperate fruit trees in a global warming context: A review. Sci. Hort. 2011, 130, 357–372. [Google Scholar] [CrossRef]
- Martínez-Gómez, P.; Prudencio, A.S.; Gradziel, T.M.; Dicenta, F. The delay of flowering time in almond: A review of the combined effect of adaptation, 406 mutation and breeding. Euphytica 2017, 213, 197. [Google Scholar] [CrossRef]
- Prudencio, A.S.; Martínez-Gómez, P.; Dicenta, F. Evaluation of breaking dormancy, flowering and productivity of extra-late and ultra-late flowering almond cultivars during cold and warm seasons in South-East of Spain. Sci. Hort. 2018, 235, 39–46. [Google Scholar] [CrossRef]
- Dicenta, F.; Sánchez-Pérez, P.; Batlle, I.; Martínez-Gómez, P. Late-blooming almond cultivar development. In Almond: Botany, Production and Uses; Rafael Socias i Company, Gradizel, T.M., Ed.; CABI: Boston, MA, USA, 2017. [Google Scholar]
- Sánchez-Pérez, R.; Dicenta, F.; Martínez-Gómez, P. Inheritance of chilling and heat requirements for flowering in almond and QTL analysis. Tree Genet. Gen. 2012, 8, 379–389. [Google Scholar] [CrossRef]
- Sánchez-Pérez, R.; Del Cueto, J.; Dicenta, F.; Martínez-Gómez, P. Recent advancements to study flowering time in almond and other Prunus species. Front. Plant Sci. 2014, 5, 334. [Google Scholar] [PubMed]
- Bianchi, V.; Rubio, M.; Trainotti, L.; Verde, I.; Bonghi, C.; Martínez-Gómez, P. Prunus transcription factors: Breeding perspectives. Front. Plant Sci. 2015, 6, 443. [Google Scholar] [PubMed]
- Bielenberg, D.G.; Wang, Y.; Fan, S.; Reighard, R.; Abbott, A.G. A Deletion Affect ing Several Gene Candidates is Present in the Evergrowing Peach Mutant. J. Hered. 2004, 95, 436–444. [Google Scholar] [CrossRef] [PubMed]
- Bielenberg, D.G.; Wang, Y.; Li, Z.; Zhebentyayeva, T.; Fan, S.; Reighard, G.L. Sequencing and annotation of the evergrowing locus in peach [Prunus persica (L.) Batsch] reveals a cluster of six MADS-box transcription factors as candidate genes for regulation of terminal bud formation. Tree Genet. Gen. 2008, 4, 495–507. [Google Scholar] [CrossRef]
- Leida, C.; Terol, J.; Martí, G.; Agustí, M.; Llácer, G.; Badenes, M.L. Identification of genes associated with bud dormancy release in Prunus persica by suppression subtractive hybridization. Tree Physiol. 2010, 30, 655–666. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Leida, C.; Conesa, A.; Llácer, G.; Badenes, M.L.; Ríos, G. Histone modifications and expression of DAM6 gene in peach are modulated during bud dormancy release in a cultivar-dependent manner. New Phytol. 2012, 193, 67–80. [Google Scholar] [CrossRef] [PubMed]
- Jiménez, S.; Reighard, G.L.; Bielenberg, D.G. Gene expression of DAM5 and DAM6 is suppressed by chilling temperatures and inversely correlated with bud break rate. Plant Mol. Biol. 2010, 73, 157–167. [Google Scholar] [CrossRef] [PubMed]
- Yamane, H.; Ooka, T.; Jotatsu, H.; Sasaki, R.; Tao, R. Expression analysis of PpDAM5 and PpDAM6 during flower bud development in peach (Prunus persica). Sci. Hort. 2011, 129, 844–848. [Google Scholar] [CrossRef]
- Sasaki, R.; Yamane, H.; Ooka, T.; Jotatsu, H.; Kitamura, Y.; Akagi, T.; Tao, R. Functional and Expressional Analyses of PmDAM Genes Associated with Endodormancy in Japanese Apricot. Plant Physiol. 2011, 157, 485–497. [Google Scholar] [CrossRef] [PubMed]
- Kitamura, Y.; Takeuchi, T.; Yamane, H.; Tao, R. Simultaneous down-regulation of DORMANCY-ASSOCIATED MADS-box6 and SOC1 during dormancy release in Japanese apricot (Prunus mume) flower buds. J. Hort. Sci. Biotechnol. 2016, 5, 476–482. [Google Scholar] [CrossRef]
- Felipe, A.J. Phenological states of almond (In Italian). In Proceedings of the Third GREMPA Colloquium, Bari, Italy, 3–7 October 1977; pp. 101–103. [Google Scholar]
- Richardson, E.A. ; S.D. Seeley, D.R.; Walker, J.L.; Anderson, M.; Ashcroft, G.L. Pheno-climatology of spring peach bud development. Hort. Sci. 1975, 10, 236–237. [Google Scholar]
- Erez, A.; Couvillon, G.A.; Hendershott, C.H. The effect of cycle length on chilling negation by high temperatures in dormant peach leaf buds. J. Am. Soc. Hort. Sci. 1979, 104, 573–576. [Google Scholar]
- Fishman, S.; Erez, A.; Couvillon, G.A. The temperature dependence of dormancy breaking in plants: Computer simulation of processes studied under controlled temperatures. J. Theor. Biol. 1987, 126, 309–321. [Google Scholar] [CrossRef]
- Fishman, S.; Erez, A.; Couvillon, G.A. The temperature dependence of dormancy breaking in plants: Mathematical analysis of a two-step model involving a cooperative transition. J. Theorl Biol. 1987, 124, 473–483. [Google Scholar] [CrossRef]
- Le Provost, G.; Herrera, R.; Paiva, J.A.; Chaumeil, P.; Salin, F.; Plomion, C. A micromethod for high throughput RNA extraction in forest trees. Biol. Res. 2007, 40, 291–297. [Google Scholar] [CrossRef] [PubMed]
- Tong, Z.; Gao, Z.; Wang, F.; Zhou, J.; Zang, Z. Selection of reliable reference genes for gene expression studies in peach using real-time PCR. BMC Mol. Biol. 2009, 10, 71. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Ionescu, I.A.; Moller, B.L.; Sánchez-Pérez, R. Chemical control of Flowering time. J. Exp. Bot. 2017, 68, 369–382. [Google Scholar] [CrossRef] [PubMed]
- Ionescu, I.A.; López-Ortega, G.; Burow, M.; Bayo-Canha, A.; Junge, A.; Gericke, O.; Moller, B.L.; Sánchez-Pérez, R. Transcriptome and Metabolite Changes during Hydrogen Cyanimide-Induced Floral Bud break in Sweet Cherry. Front. Plant Sci. 2017, 8, 1233. [Google Scholar] [CrossRef] [PubMed]
- Erez, A. Means to compensate for insufficient chilling to improve bloom and leafing. Acta Hort. 1995, 395, 81–95. [Google Scholar] [CrossRef]
- Kauffman, H.; Blanke, M. Substitution of winter chilling by spring forcing for flowering using sweet cherry as model crop. Sci. Hort. 2018, 244, 75–81. [Google Scholar] [CrossRef]
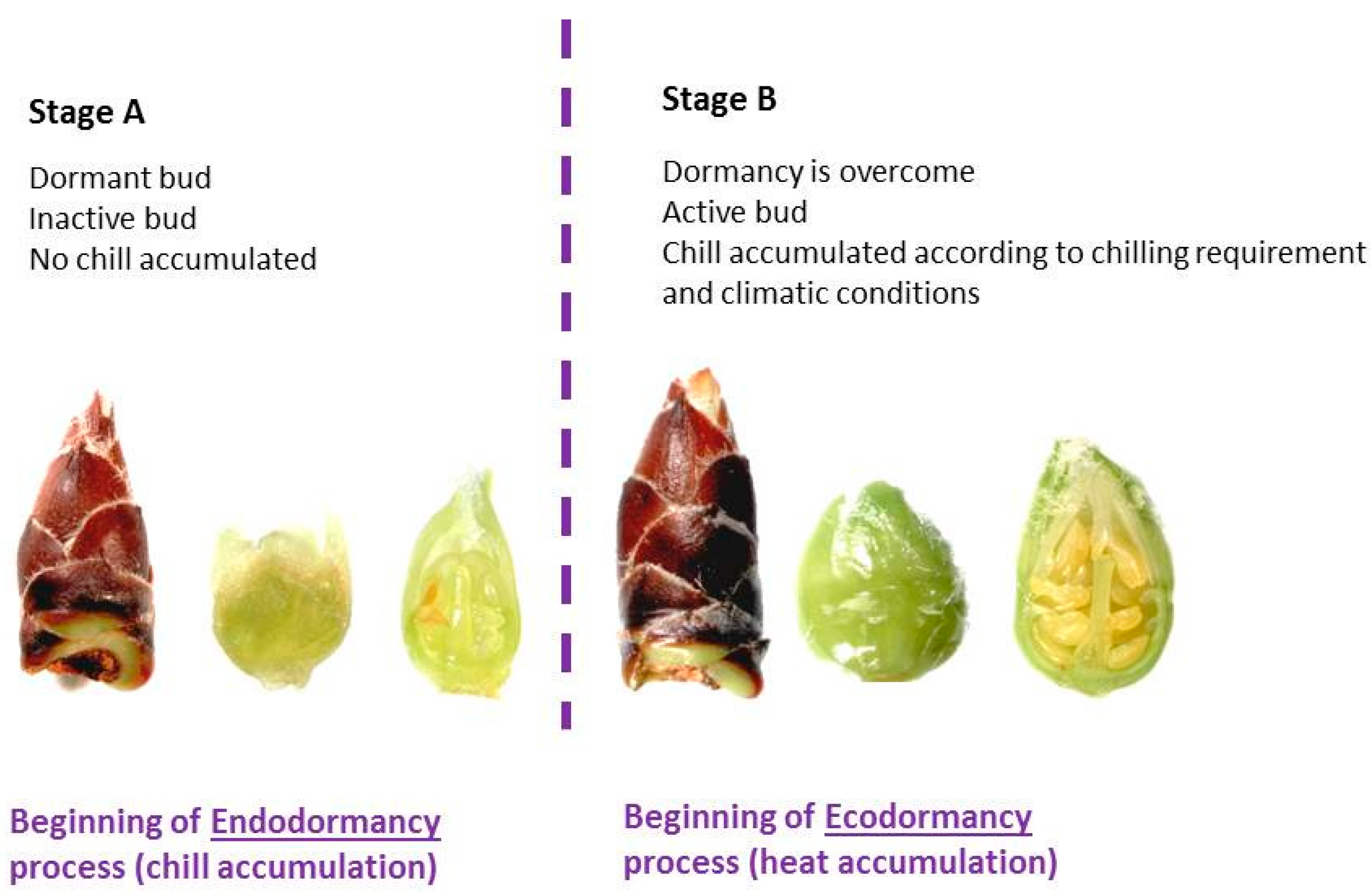


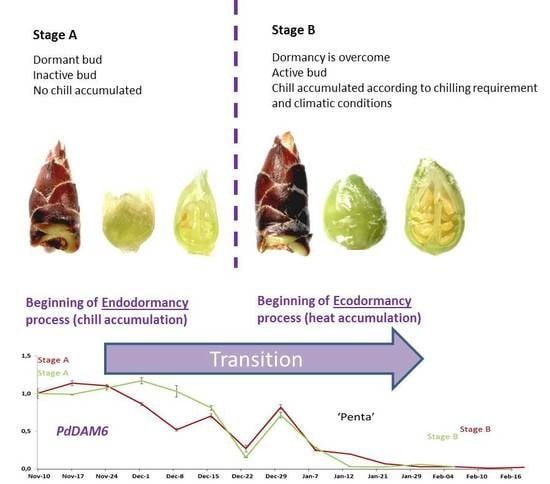
| Season | Stage A | Stage B | Flowering Date | |||
|---|---|---|---|---|---|---|
| Date | Chill Accumulation | Date | Chill Accumulation | |||
| ‘Desmayo Largueta’ | 2015/2016 | November 10 | 0 | December 21 | 16 | January 28 |
| 2016/2017 | November 10 | 0 | December 15 | 24 | January 27 | |
| ‘Penta’ | 2015/2016 | November 10 | 0 | February 10 | 41 | March 25 |
| 2016/2017 | November 10 | 0 | February 2 | 54 | March 12 | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Prudencio, Á.S.; Dicenta, F.; Martínez-Gómez, P. Monitoring Dormancy Transition in Almond [Prunus Dulcis (Miller) Webb] during Cold and Warm Mediterranean Seasons through the Analysis of a DAM (Dormancy-Associated MADS-Box) Gene. Horticulturae 2018, 4, 41. https://doi.org/10.3390/horticulturae4040041
Prudencio ÁS, Dicenta F, Martínez-Gómez P. Monitoring Dormancy Transition in Almond [Prunus Dulcis (Miller) Webb] during Cold and Warm Mediterranean Seasons through the Analysis of a DAM (Dormancy-Associated MADS-Box) Gene. Horticulturae. 2018; 4(4):41. https://doi.org/10.3390/horticulturae4040041
Chicago/Turabian StylePrudencio, Ángela S., Federico Dicenta, and Pedro Martínez-Gómez. 2018. "Monitoring Dormancy Transition in Almond [Prunus Dulcis (Miller) Webb] during Cold and Warm Mediterranean Seasons through the Analysis of a DAM (Dormancy-Associated MADS-Box) Gene" Horticulturae 4, no. 4: 41. https://doi.org/10.3390/horticulturae4040041
APA StylePrudencio, Á. S., Dicenta, F., & Martínez-Gómez, P. (2018). Monitoring Dormancy Transition in Almond [Prunus Dulcis (Miller) Webb] during Cold and Warm Mediterranean Seasons through the Analysis of a DAM (Dormancy-Associated MADS-Box) Gene. Horticulturae, 4(4), 41. https://doi.org/10.3390/horticulturae4040041