Visualising the Cardiovascular System of Embryos of Biomedical Model Organisms with High Resolution Episcopic Microscopy (HREM)
Abstract
:1. Introduction
2. What Is High-Resolution Episcopic Microscopy (HREM)
2.1. Sample Preparation
2.1.1. Fixation
2.1.2. Dehydration
2.1.3. Infiltration
2.1.4. Embedding
2.2. HREM Data Generation
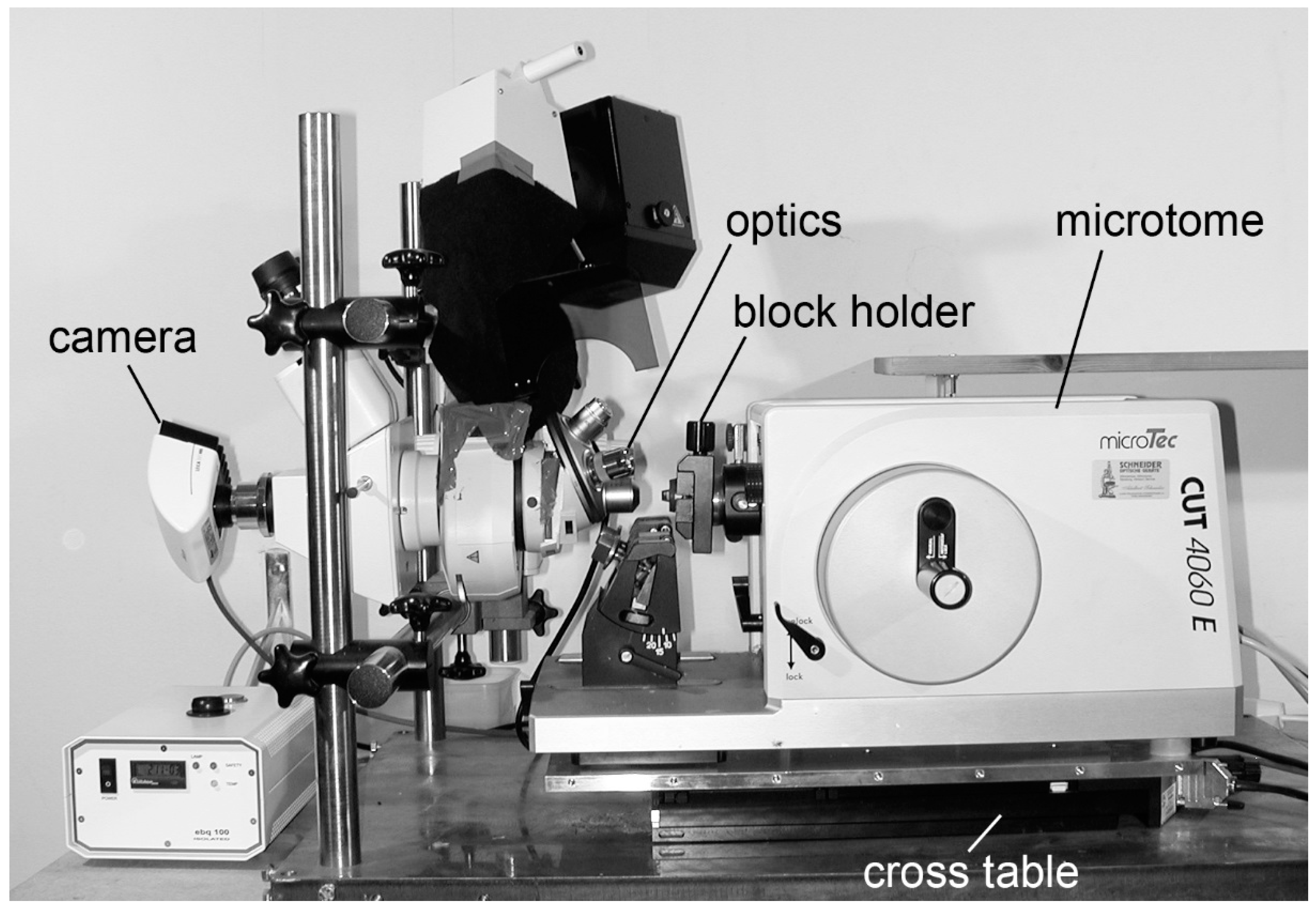
2.2.1. HREM Apparatus
2.2.2. Data Processing
2.2.3. Data Visualisation
3. Examples
3.1. Skin Vasculature
3.2. Biomedical Model Organisms
3.2.1. Zebrafish and Frog Embryos
3.2.2. Chick Embryos
3.2.3. Mouse Embryos
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Olson, E.N. A decade of discoveries in cardiac biology. Nat. Med. 2004, 10, 467–474. [Google Scholar] [CrossRef]
- Srivastava, D. Making or breaking the heart: from lineage determination to morphogenesis. Cell 2006, 126, 1037–1048. [Google Scholar] [CrossRef] [PubMed]
- Pierpont, M.E.; Basson, C.T.; Benson, D.W., Jr.; Gelb, B.D.; Giglia, T.M.; Goldmuntz, E.; McGee, G.; Sable, C.A.; Srivastava, D.; Webb, C.L. Genetic basis for congenital heart defects: Current knowledge: A scientific statement from the American Heart Association Congenital Cardiac Defects Committee, Council on Cardiovascular Disease in the Young: Endorsed by the American Academy of Pediatrics. Circulation 2007, 115, 3015–3038. [Google Scholar] [CrossRef] [PubMed]
- Andelfinger, G. Genetic factors in congenital heart malformation. Clin. Genet. 2008, 73, 516–527. [Google Scholar] [CrossRef] [PubMed]
- Dees, E.; Baldwin, H.S. Making a heart: Advances in understanding the mechanisms of cardiac development. Curr. Opin. Pediatr. 2016, 28, 584–589. [Google Scholar] [CrossRef] [PubMed]
- Ivanovitch, K.; Esteban, I.; Torres, M. Growth and Morphogenesis during Early Heart Development in Amniotes. J. Cardiovasc. Dev. Dis. 2017, 4. [Google Scholar] [CrossRef] [PubMed]
- Zaidi, S.; Brueckner, M. Genetics and Genomics of Congenital Heart Disease. Circ. Res. 2017, 120, 923–940. [Google Scholar] [CrossRef] [Green Version]
- Weninger, W.J.; Streicher, J.; Müller, G.B. 3-dimensional reconstruction of histological serial sections using a computer. Wien. Klin. Wochenschr. 1996, 108, 515–520. [Google Scholar]
- Streicher, J.; Weninger, W.J.; Müller, G.B. External marker-based automatic congruencing: A new method of 3D reconstruction from serial sections. Anat. Rec. 1997, 248, 583–602. [Google Scholar] [CrossRef] [Green Version]
- Streicher, J.; Donat, M.A.; Strauss, B.; Spörle, R.; Schughart, K.; Müller, G.B. Computer-based three-dimensional visualization of developmental gene expression. Nat. Genet. 2000, 25, 147–152. [Google Scholar] [CrossRef]
- Metscher, B. MicroCT for comparative morphology: Simple staining methods allow high-contrast 3D imaging of diverse non-mineralized animal tissues. BMC Physiol. 2009, 9, 11–24. [Google Scholar] [CrossRef] [PubMed]
- Metscher, B.D. MicroCT for developmental biology: A versatile tool for high-contrast 3D imaging at histological resolutions. Dev. Dyn. 2009, 238, 632–640. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Metscher, B.D.; Müller, G.B. MicroCT for molecular imaging: Quantitative visualization of complete three-dimensional distributions of gene products in embryonic limbs. Dev. Dyn. 2011, 240, 2301–2308. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gignac, P.M.; Kley, N.J.; Clarke, J.A.; Colbert, M.W.; Morhardt, A.C.; Cerio, D.; Cost, I.N.; Cox, P.G.; Daza, J.D.; Early, C.M.; et al. Diffusible iodine-based contrast-enhanced computed tomography (diceCT): An emerging tool for rapid, high-resolution, 3-D imaging of metazoan soft tissues. J. Anat. 2016, 228, 889–909. [Google Scholar] [CrossRef] [PubMed]
- Degenhardt, K.; Wright, A.C.; Horng, D.; Padmanabhan, A.; Epstein, J.A. Rapid 3D phenotyping of cardiovascular development in mouse embryos by micro-CT with iodine staining. Circ. Cardiovasc. Imaging 2010, 3, 314–322. [Google Scholar] [CrossRef] [PubMed]
- Wong, M.D.; Dorr, A.E.; Walls, J.R.; Lerch, J.P.; Henkelman, R.M. A novel 3D mouse embryo atlas based on micro-CT. Development 2012, 139, 3248–3256. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Johnson, J.T.; Hansen, M.S.; Wu, I.; Healy, L.J.; Johnson, C.R.; Jones, G.M.; Capecchi, M.R.; Keller, C. Virtual histology of transgenic mouse embryos for high-throughput phenotyping. PLoS Genet. 2006, 2, e61. [Google Scholar] [CrossRef]
- Hsu, C.-W.; Wong, L.; Rasmussen, T.L.; Kalaga, S.; McElwee, M.L.; Keith, L.C.; Bohat, R.; Seavitt, J.R.; Beaudet, A.L.; Dickinson, M.E. Three-dimensional microCT imaging of mouse development from early post-implantation to early postnatal stages. Dev. Biol. 2016, 419, 229–236. [Google Scholar] [CrossRef]
- Núñez, J.A.; Goring, A.; Hesse, E.; Thurner, P.J.; Schneider, P.; Clarkin, C.E. Simultaneous visualisation of calcified bone microstructure and intracortical vasculature using synchrotron X-ray phase contrast-enhanced tomography. Sci. Rep. 2017, 7, 13289. [Google Scholar] [CrossRef] [Green Version]
- Cleary, J.O.; Modat, M.; Norris, F.C.; Price, A.N.; Jayakody, S.A.; Martinez-Barbera, J.P.; Greene, N.D.E.; Hawkes, D.J.; Ordidge, R.J.; Scambler, P.J.; et al. Magnetic resonance virtual histology for embryos: 3D atlases for automated high-throughput phenotyping. Neuroimage 2011, 54, 769–778. [Google Scholar] [CrossRef]
- Zamyadi, M.; Baghdadi, L.; Lerch, J.P.; Bhattacharya, S.; Schneider, J.E.; Henkelman, R.M.; Sled, J.G. Mouse embryonic phenotyping by morphometric analysis of MR images. Physiol. Genet. 2010, 42A, 89–95. [Google Scholar] [CrossRef] [PubMed]
- Cleary, J.O.; Price, A.N.; Thomas, D.L.; Scambler, P.J.; Kyriakopoulou, V.; McCue, K.; Schneider, J.E.; Ordidge, R.J.; Lythgoe, M.F. Cardiac phenotyping in ex vivo murine embryos using microMRI. NMR Biomed. 2009, 22, 857–866. [Google Scholar] [CrossRef] [PubMed]
- Berrios-Otero, C.A.; Wadghiri, Y.Z.; Nieman, B.J.; Joyner, A.L.; Turnbull, D.H. Three-dimensional micro-MRI analysis of cerebral artery development in mouse embryos. Magn. Reson. Med. 2009, 62, 1431–1439. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Petiet, A.E.; Kaufman, M.H.; Goddeeris, M.M.; Brandenburg, J.; Elmore, S.A.; Johnson, G.A. High-resolution magnetic resonance histology of the embryonic and neonatal mouse: A 4D atlas and morphologic database. Proc. Natl. Acad. Sci. USA 2008, 105, 12331–12336. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schneider, J.E.; Bose, J.; Bamforth, S.D.; Gruber, A.D.; Broadbent, C.; Clarke, K.; Neubauer, S.; Lengeling, A.; Bhattacharya, S. Identification of cardiac malformations in mice lacking Ptdsr using a novel high-throughput magnetic resonance imaging technique. BMC Dev. Biol. 2004, 4, 16–27. [Google Scholar] [CrossRef]
- Schneider, J.E.; Bhattacharya, S. Making the mouse embryo transparent: Identifying developmental malformations using magnetic resonance imaging. Birth Defects Res. 2004, 72, 241–249. [Google Scholar] [CrossRef] [PubMed]
- Schneider, J.E.; Bamforth, S.D.; Farthing, C.R.; Clarke, K.; Neubauer, S.; Bhattacharya, S. Rapid identification and 3D reconstruction of complex cardiac malformations in transgenic mouse embryos using fast gradient echo sequence magnetic resonance imaging. J. Mol. Cell. Cardiol. 2003, 35, 217–222. [Google Scholar] [CrossRef]
- Smith, B.R. Magnetic resonance microscopy in cardiac development. Microsc. Res. Tech. 2001, 52, 323–330. [Google Scholar] [CrossRef] [Green Version]
- Dhenain, M.; Ruffins, S.W.; Jacobs, R.E. Three-dimensional digital mouse atlas using high-resolution MRI. Dev. Biol. 2001, 232, 458–470. [Google Scholar] [CrossRef]
- Jacobs, R.E.; Ahrens, E.T.; Dickinson, M.E.; Laidlaw, D. Towards a microMRI atlas of mouse development. Comput. Med. Imaging Graph. 1999, 23, 15–24. [Google Scholar] [CrossRef] [Green Version]
- Sharpe, J.; Ahlgren, U.; Perry, P.; Hill, B.; Ross, A.; Hecksher-Sorensen, J.; Baldock, R.; Davidson, D. Optical projection tomography as a tool for 3D microscopy and gene expression studies. Science 2002, 296, 541–545. [Google Scholar] [CrossRef] [PubMed]
- Sharpe, J. Optical projection tomography as a new tool for studying embryo anatomy. J. Anat. 2003, 202, 175–181. [Google Scholar] [CrossRef] [Green Version]
- Lickert, H.; Takeuchi, J.K.; Von Both, I.; Walls, J.R.; McAuliffe, F.; Adamson, S.L.; Henkelman, R.M.; Wrana, J.L.; Rossant, J.; Bruneau, B.G. Baf60c is essential for function of BAF chromatin remodelling complexes in heart development. Nature 2004, 432, 107–112. [Google Scholar] [CrossRef]
- Colas, J.F.; Sharpe, J. Live optical projection tomography. Organogenesis 2009, 5, 211–216. [Google Scholar] [CrossRef] [Green Version]
- Breckenridge, R.; Kotecha, S.; Towers, N.; Bennett, M.; Mohun, T. Pan-myocardial expression of Cre recombinase throughout mouse development. Genesis 2007, 45, 135–144. [Google Scholar] [CrossRef] [PubMed]
- Jeansson, M.; Gawlik, A.; Anderson, G.; Li, C.; Kerjaschki, D.; Henkelman, M.; Quaggin, S.E. Angiopoietin-1 is essential in mouse vasculature during development and in response to injury. J. Clin. Investig. 2011, 121, 2278–2289. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Anderson, G.A.; Wong, M.D.; Yang, J.; Henkelman, R.M. 3D imaging, registration, and analysis of the early mouse embryonic vasculature. Dev. Dyn. 2013, 242, 527–538. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, S.; Garcia, M.D.; Lopez, A.L.; Overbeek, P.A.; Larin, K.V.; Larina, I.V. Dynamic imaging and quantitative analysis of cranial neural tube closure in the mouse embryo using optical coherence tomography. Biomed. Opt. Express 2017, 8, 407–419. [Google Scholar] [CrossRef]
- Singh, M.; Raghunathan, R.; Piazza, V.; Davis-Loiacono, A.M.; Cable, A.; Vedakkan, T.J.; Janecek, T.; Frazier, M.V.; Nair, A.; Wu, C.; et al. Applicability, usability, and limitations of murine embryonic imaging with optical coherence tomography and optical projection tomography. Biomed. Opt. Express 2016, 7, 2295–2310. [Google Scholar] [CrossRef]
- Liu, M.; Maurer, B.; Hermann, B.; Zabihian, B.; Sandrian, M.G.; Unterhuber, A.; Baumann, B.; Zhang, E.Z.; Beard, P.C.; Weninger, W.J.; et al. Dual modality optical coherence and whole-body photoacoustic tomography imaging of chick embryos in multiple development stages. Biomed. Opt. Express 2014, 5, 3150–3159. [Google Scholar] [CrossRef]
- Raghunathan, R.; Singh, M.; Dickinson, M.E.; Larin, K.V. Optical coherence tomography for embryonic imaging: a review. J. Biomed. Opt. 2016, 21, 50902. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, Q.; Jin, T.; Qi, W.; Mo, X.; Xi, L. Label-free photoacoustic imaging of the cardio-cerebrovascular development in the embryonic zebrafish. Biomed. Opt. Express 2017, 8, 2359–2367. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ewald, A.J.; McBride, H.; Reddington, M.; Fraser, S.E.; Kerschmann, R. Surface imaging microscopy, an automated method for visualizing whole embryo samples in three dimensions at high resolution. Dev. Dyn. 2002, 225, 369–375. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sivaguru, M.; Fried, G.A.; Miller, C.A.; Fouke, B.W. Multimodal optical microscopy methods reveal polyp tissue morphology and structure in Caribbean reef building corals. J. Vis. Exp. 2014, e51824. [Google Scholar] [CrossRef] [PubMed]
- Srinivasan, S.; Baldwin, H.S.; Aristizabal, O.; Kwee, L.; Labow, M.; Artman, M.; Turnbull, D.H. Noninvasive, in utero imaging of mouse embryonic heart development with 40-MHz echocardiography. Circulation 1998, 98, 912–918. [Google Scholar] [CrossRef]
- Phoon, C.K.; Ji, R.P.; Aristizabal, O.; Worrad, D.M.; Zhou, B.; Baldwin, H.S.; Turnbull, D.H. Embryonic heart failure in NFATc1-/- mice: Novel mechanistic insights from in utero ultrasound biomicroscopy. Circ. Res. 2004, 95, 92–99. [Google Scholar] [CrossRef] [PubMed]
- Kulandavelu, S.; Qu, D.; Sunn, N.; Mu, J.; Rennie, M.Y.; Whiteley, K.J.; Walls, J.R.; Bock, N.A.; Sun, J.C.; Covelli, A.; et al. Embryonic and neonatal phenotyping of genetically engineered mice. Ilar. J. 2006, 47, 103–117. [Google Scholar] [CrossRef]
- Phoon, C.K. Imaging tools for the developmental biologist: Ultrasound biomicroscopy of mouse embryonic development. Pediatr. Res. 2006, 60, 14–21. [Google Scholar] [CrossRef]
- Golden, H.B.; Sunder, S.; Liu, Y.; Peng, X.; Dostal, D.E. In utero assessment of cardiovascular function in the embryonic mouse heart using high-resolution ultrasound biomicroscopy. In Cardiovascular Development; Humana Press: Totowa, NJ, USA, 2012; pp. 245–263. [Google Scholar]
- Weninger, W.J.; Meng, S.; Streicher, J.; Müller, G.B. A new episcopic method for rapid 3-D reconstruction: applications in anatomy and embryology. Anat. Embryol. 1998, 197, 341–348. [Google Scholar] [CrossRef]
- Weninger, W.J.; Mohun, T. Phenotyping transgenic embryos: A rapid 3-D screening method based on episcopic fluorescence image capturing. Nat. Genet. 2002, 30, 59–65. [Google Scholar] [CrossRef]
- Weninger, W.J.; Floro, K.L.; Bennett, M.B.; Withington, S.L.; Preis, J.I.; Barbera, J.P.; Mohun, T.J.; Dunwoodie, S.L. Cited2 is required both for heart morphogenesis and establishment of the left-right axis in mouse development. Development 2005, 132, 1337–1348. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Weninger, W.J.; Geyer, S.H.; Mohun, T.J.; Rasskin-Gutman, D.; Matsui, T.; Ribeiro, I.; Costa Lda, F.; Izpisua-Belmonte, J.C.; Müller, G.B. High-resolution episcopic microscopy: A rapid technique for high detailed 3D analysis of gene activity in the context of tissue architecture and morphology. Anat. Embryol. 2006, 211, 213–221. [Google Scholar] [CrossRef] [PubMed]
- Geyer, S.H.; Mohun, T.J.; Weninger, W.J. Visualizing vertebrate embryos with episcopic 3D imaging techniques. Sci. World J. 2009, 9, 1423–1437. [Google Scholar] [CrossRef] [PubMed]
- Mohun, T.J.; Weninger, W.J. Imaging heart development using high-resolution episcopic microscopy. Curr. Opin. Genet. Dev. 2011, 21, 573–578. [Google Scholar] [CrossRef] [PubMed]
- Geyer, S.H.; Maurer-Gesek, B.; Reissig, L.F.; Weninger, W.J. High-resolution Episcopic Microscopy (HREM)—Simple and Robust Protocols for Processing and Visualizing Organic Materials. J. Vis. Exp. 2017. [Google Scholar] [CrossRef] [PubMed]
- Denk, W.; Horstmann, H. Serial block-face scanning electron microscopy to reconstruct three-dimensional tissue nanostructure. PLoS Biol. 2004, 2, e329. [Google Scholar] [CrossRef] [PubMed]
- Kalson, N.S.; Holmes, D.F.; Herchenhan, A.; Lu, Y.; Starborg, T.; Kadler, K.E. Slow stretching that mimics embryonic growth rate stimulates structural and mechanical development of tendon-like tissue in vitro. Dev. Dyn. 2011, 240, 2520–2528. [Google Scholar] [CrossRef]
- Smith, D.; Starborg, T. Serial block face scanning electron microscopy in cell biology: Applications and technology. Tissue cell 2018. [Google Scholar] [CrossRef]
- Weninger, W.J.; Maurer, B.; Zendron, B.; Dorfmeister, K.; Geyer, S.H. Measurements of the diameters of the great arteries and semi-lunar valves of chick and mouse embryos. J. Microsc. 2009, 234, 173–190. [Google Scholar] [CrossRef]
- Geyer, S.H.; Maurer, B.; Pötz, L.; Singh, J.; Weninger, W.J. High-Resolution Episcopic Microscopy Data-Based Measurements of the Arteries of Mouse Embryos: Evaluation of Significance and Reproducibility under Routine Conditions. Cells Tissues Organs 2012, 195, 524–534. [Google Scholar] [CrossRef]
- Geyer, S.H.; Weninger, W.J. Some Mice Feature 5th Pharyngeal Arch Arteries and Double-Lumen Aortic Arch Malformations. Cells Tissues Organs 2012, 196, 90–98. [Google Scholar] [CrossRef] [PubMed]
- Geyer, S.H.; Weninger, W.J. Metric characterization of the aortic arch of early mouse fetuses and of a fetus featuring a double lumen aortic arch malformation. Ann. Anat. 2013, 195, 175–182. [Google Scholar] [CrossRef] [PubMed]
- Bruneel, B.; Matha, M.; Paesen, R.; Ameloot, M.; Weninger, W.J.; Huysseune, A. Imaging the zebrafish dentition: from traditional approaches to emerging technologies. Zebrafish 2015, 12, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Pokhrel, N.; Cohen, E.B.; Genin, O.; Ruzal, M.; Sela-Donenfeld, D.; Cinnamon, Y. Effects of storage conditions on hatchability, embryonic survival and cytoarchitectural properties in broiler from young and old flocks. Poult. Sci. 2018, 97, 1429–1440. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Perez-Garcia, V.; Fineberg, E.; Wilson, R.; Murray, A.; Mazzeo, C.I.; Tudor, C.; Sienerth, A.; White, J.K.; Tuck, E.; Ryder, E.J.; et al. Placentation defects are highly prevalent in embryonic lethal mouse mutants. Nature 2018, 555, 463–468. [Google Scholar] [CrossRef] [PubMed]
- Aiello, V.D.; Spicer, D.E.; Anderson, R.H.; Brown, N.A.; Mohun, T.J. The independence of the infundibular building blocks in the setting of double-outlet right ventricle. Cardiol. Young 2017, 27, 825–836. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lana-Elola, E.; Watson-Scales, S.; Slender, A.; Gibbins, D.; Martineau, A.; Douglas, C.; Mohun, T.; Fisher, E.M.; Tybulewicz, V. Genetic dissection of Down syndrome-associated congenital heart defects using a new mouse mapping panel. Elife 2016, 5. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Spicer, D.E.; Henderson, D.J.; Chaudhry, B.; Mohun, T.J.; Anderson, R.H. The anatomy and development of normal and abnormal coronary arteries. Cardiol. Young 2015, 25, 1493–1503. [Google Scholar] [CrossRef] [PubMed]
- Anderson, R.H.; Mohun, T.J.; Brown, N.A. Clarifying the morphology of the ostium primum defect. J. Anat. 2015, 226, 244–257. [Google Scholar] [CrossRef] [Green Version]
- Mohun, T.; Adams, D.J.; Baldock, R.; Bhattacharya, S.; Copp, A.J.; Hemberger, M.; Houart, C.; Hurles, M.E.; Robertson, E.; Smith, J.C.; et al. Deciphering the Mechanisms of Developmental Disorders (DMDD): A new programme for phenotyping embryonic lethal mice. Dis Model. Mech. 2013, 6, 562–566. [Google Scholar] [CrossRef]
- Weninger, W.J.; Geyer, S.H.; Martineau, A.; Galli, A.; Adams, D.J.; Wilson, R.; Mohun, T.J. Phenotyping structural abnormalities in mouse embryos using high-resolution episcopic microscopy. Dis Model. Mech. 2014, 7, 1143–1152. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wilson, R.; McGuire, C.; Mohun, T.; Adams, D.; Baldock, R.; Bhattacharya, S.; Collins, J.; Fineberg, E.; Firminger, L.; Galli, A.; et al. Deciphering the mechanisms of developmental disorders: Phenotype analysis of embryos from mutant mouse lines. Nucleic Acids Research 2016, 44, D855–D861. [Google Scholar] [CrossRef] [PubMed]
- Wilson, R.; Geyer, S.H.; Reissig, L.; Rose, J.; Szumska, D.; Hardman, E.; Prin, F.; McGuire, C.; Ramirez-Solis, R.; White, J.; et al. Highly variable penetrance of abnormal phenotypes in embryonic lethal knockout mice. Wellcome. Open Res. 2016, 1, 1. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mohun, T.J.; Weninger, W.J. Embedding Embryos for High-Resolution Episcopic Microscopy (HREM). Cold Spring Harb. Protoc. 2012, 2012, 678–680. [Google Scholar] [CrossRef] [PubMed]
- Geyer, S.H.; Nöhammer, M.M.; Mathä, M.; Reissig, L.; Tinhofer, I.E.; Weninger, W.J. High-Resolution Episcopic Microscopy (HREM): A Tool for Visualizing Skin Biopsies. Microsc. Microanal. 2014, 20, 1356–1364. [Google Scholar] [CrossRef] [PubMed]
- Geyer, S.H.; Mohun, T.J.; Kamolz, L.P.; Weninger, W.J. High resolution episcopic microscopy - current applications. Curr. Biotech. 2012, 1, 281–286. [Google Scholar] [CrossRef]
- Mohun, T.J.; Weninger, W.J. Generation of volume data by episcopic three-dimensional imaging of embryos. Cold Spring Harb. Protoc. 2012, 2012, 681–682. [Google Scholar] [CrossRef] [PubMed]
- Geyer, S.H.; Nohammer, M.M.; Tinhofer, I.E.; Weninger, W.J. The dermal arteries of the human thumb pad. J. Anat. 2013, 223, 603–609. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tinhofer, I.E.; Zaussinger, M.; Geyer, S.H.; Meng, S.; Kamolz, L.P.; Tzou, C.H.; Weninger, W.J. The dermal arteries in the cutaneous angiosome of the descending genicular artery. J. Anat. 2018. [Google Scholar] [CrossRef]
- Geyer, S.H.; Tinhofer, I.E.; Lumenta, D.B.; Kamolz, L.P.; Branski, L.; Finnerty, C.C.; Herndon, D.N.; Weninger, W.J. High-resolution episcopic microscopy (HREM): A useful technique for research in wound care. Ann. Anat. 2015, 197, 3–10. [Google Scholar] [CrossRef]
- Ertl, J.; Pichlsberger, M.; Tuca, A.-C.; Wurzer, P.; Fuchs, J.; Geyer, S.H.; Maurer-Gesek, B.; Weninger, W.J.; Pfeiffer, D.; Bubalo, V.; et al. Comparative study of regenerative effects of mesenchymal stem cells derived from placental amnion, chorion and umbilical cord on dermal wounds. Placenta 2018, 65, 37–46. [Google Scholar] [CrossRef]
- Wiedner, M.; Tinhofer, I.E.; Kamolz, L.P.; Seyedian Moghaddam, A.; Justich, I.; Liegl-Atzwanger, B.; Bubalo, V.; Weninger, W.J.; Lumenta, D.B. Simultaneous dermal matrix and autologous split-thickness skin graft transplantation in a porcine wound model: A three-dimensional histological analysis of revascularization. Wound Repair Regen. 2014, 22, 749–754. [Google Scholar] [CrossRef] [PubMed]
- Hamburger, V.; Hamilton, H.L. A series of normal stages in the development of the chick embryo. J. Morph. 1951, 88, 49–92. [Google Scholar] [CrossRef] [PubMed]
- Maurer, B.; Geyer, S.H.; Weninger, W.J. A Chick Embryo with a yet Unclassified Type of Cephalothoracopagus Malformation and a Hypothesis for Explaining its Genesis. Anat. Histol. Embryol. 2013, 42, 191–200. [Google Scholar] [CrossRef] [PubMed]
- Maurer-Gesek, B. Malformations of the Great Intrathoracic Arteries caused by Hemodynamic Alterations in Chick Embryos. Ph.D. Thesis, Medical University of Vienna, Vienna, Austria, 2016. [Google Scholar]
- Bamforth, S.D.; Chaudhry, B.; Bennett, M.; Wilson, R.; Mohun, T.J.; Van Mierop, L.H.; Henderson, D.J.; Anderson, R.H. Clarification of the identity of the mammalian fifth pharyngeal arch artery. Clin. Anat. 2013, 26, 173–182. [Google Scholar] [CrossRef] [PubMed]
- Anderson, R.H.; Brown, N.A.; Mohun, T.J. Insights regarding the normal and abnormal formation of the atrial and ventricular septal structures. Clin. Anat. 2016, 29, 290–304. [Google Scholar] [CrossRef] [PubMed]
- Paun, B.; Bijnens, B.; Cook, A.C.; Mohun, T.J.; Butakoff, C. Quantification of the detailed cardiac left ventricular trabecular morphogenesis in the mouse embryo. Med. Image Anal. 2018, 49, 89–104. [Google Scholar] [CrossRef]
- Le Garrec, J.F.; Dominguez, J.N.; Desgrange, A.; Ivanovitch, K.D.; Raphael, E.; Bangham, J.A.; Torres, M.; Coen, E.; Mohun, T.J.; Meilhac, S.M. A predictive model of asymmetric morphogenesis from 3D reconstructions of mouse heart looping dynamics. Elife 2017, 6. [Google Scholar] [CrossRef] [Green Version]
- Dunlevy, L.; Bennett, M.; Slender, A.; Lana-Elola, E.; Tybulewicz, V.L.; Fisher, E.M.; Mohun, T. Down’s syndrome-like cardiac developmental defects in embryos of the transchromosomic Tc1 mouse. Cardiovasc. Res. 2010, 88, 287–295. [Google Scholar] [CrossRef]
- Captur, G.; Wilson, R.; Bennett, M.F.; Luxan, G.; Nasis, A.; de la Pompa, J.L.; Moon, J.C.; Mohun, T.J. Morphogenesis of myocardial trabeculae in the mouse embryo. J. Anat. 2016, 229, 314–325. [Google Scholar] [CrossRef] [Green Version]
- Geyer, S.H.; Reissig, L.F.; Husemann, M.; Hofle, C.; Wilson, R.; Prin, F.; Szumska, D.; Galli, A.; Adams, D.J.; White, J.; et al. Morphology, topology and dimensions of the heart and arteries of genetically normal and mutant mouse embryos at stages S21–S23. J. Anat. 2017, 231, 600–614. [Google Scholar] [CrossRef] [PubMed]


© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Weninger, W.J.; Maurer-Gesek, B.; Reissig, L.F.; Prin, F.; Wilson, R.; Galli, A.; Adams, D.J.; White, J.K.; Mohun, T.J.; Geyer, S.H. Visualising the Cardiovascular System of Embryos of Biomedical Model Organisms with High Resolution Episcopic Microscopy (HREM). J. Cardiovasc. Dev. Dis. 2018, 5, 58. https://doi.org/10.3390/jcdd5040058
Weninger WJ, Maurer-Gesek B, Reissig LF, Prin F, Wilson R, Galli A, Adams DJ, White JK, Mohun TJ, Geyer SH. Visualising the Cardiovascular System of Embryos of Biomedical Model Organisms with High Resolution Episcopic Microscopy (HREM). Journal of Cardiovascular Development and Disease. 2018; 5(4):58. https://doi.org/10.3390/jcdd5040058
Chicago/Turabian StyleWeninger, Wolfgang J., Barbara Maurer-Gesek, Lukas F. Reissig, Fabrice Prin, Robert Wilson, Antonella Galli, David J. Adams, Jacqueline K. White, Timothy J. Mohun, and Stefan H. Geyer. 2018. "Visualising the Cardiovascular System of Embryos of Biomedical Model Organisms with High Resolution Episcopic Microscopy (HREM)" Journal of Cardiovascular Development and Disease 5, no. 4: 58. https://doi.org/10.3390/jcdd5040058
APA StyleWeninger, W. J., Maurer-Gesek, B., Reissig, L. F., Prin, F., Wilson, R., Galli, A., Adams, D. J., White, J. K., Mohun, T. J., & Geyer, S. H. (2018). Visualising the Cardiovascular System of Embryos of Biomedical Model Organisms with High Resolution Episcopic Microscopy (HREM). Journal of Cardiovascular Development and Disease, 5(4), 58. https://doi.org/10.3390/jcdd5040058



