Gluten Assessment in Beers: Comparison by Different Commercial ELISA Kits and Evaluation of NIR Analysis as a Complementary Technique
Abstract
1. Introduction
2. Materials and Methods
2.1. Beer Sampling
2.2. Gluten Determination by Enzyme-Linked Immunosorbent Assay
2.3. NIR Spectroscopy Analysis
2.4. Statistical Analysis
3. Results
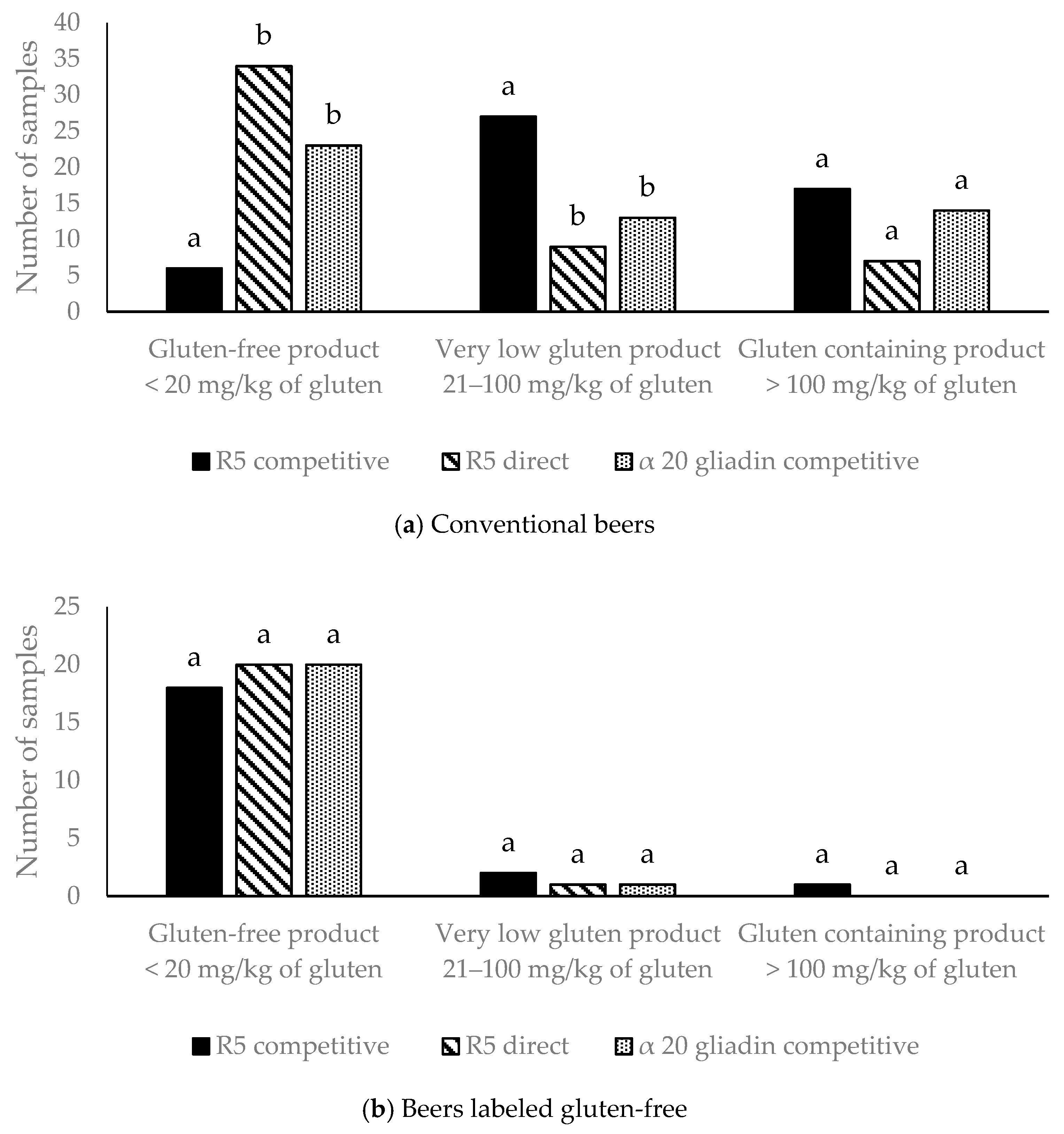
3.1. Gluten Analysis in Beer by Three Different ELISA Kits
3.2. ELISA Results and NIR Analysis Correlation
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Colgrave, M.L.; Byrne, K.; Howitt, C.A. Liquid Chromatography-Mass Spectrometry Analysis Reveals Hydrolyzed Gluten in Beers Crafted To Remove Gluten. J. Agric. Food Chem 2017, 65, 9715–9725. [Google Scholar] [CrossRef] [PubMed]
- FAO. FAOSTAT. Available online: http://www.fao.org/faostat/en/#home (accessed on 9 May 2021).
- Bustamante, M.; Simón, E. Gluten-Free Spirits and Drinks. In Advances in the Understanding of Gluten Related Pathology and the Evolution of Gluten-Free Foods; Arranz, E., Fernández-Bañares, F., Rosell, C., Rodrigo, L., Peña, A., Eds.; OmniaScience: Barcelona, Spain, 2015; pp. 645–673. [Google Scholar]
- Kok, Y.J.; Ye, L.; Muller, J.; Ow, D.S.; Bi, X. Brewing with malted barley or raw barley: What makes the difference in the processes? Appl. Microbiol. Biotechnol. 2019, 103, 1059–1067. [Google Scholar] [CrossRef]
- Humia, B.V.; Santos, K.S.; Barbosa, A.M.; Sawata, M.; Mendonça, M.D.C.; Padilha, F.F. Beer Molecules and Its Sensory and Biological Properties: A Review. Molecules 2019, 24, 1568. [Google Scholar] [CrossRef]
- Cela, N.; Condelli, N.; Caruso, M.C.; Perretti, G.; Di Cairano, M.; Tolve, R.; Galgano, F. Gluten-Free Brewing: Issues and Perspectives. Fermentation 2020, 6, 53. [Google Scholar] [CrossRef]
- Donadini, G.; Bertuzzi, T.; Kordialik-Bogacka, E.; Cywińska, D.; Rossi, F.; Spigno, G.; Porretta, S. Investigating patterns of millennials’ interest in gluten-free beer in Poland: A question of beer price and alcohol content. J. Food Sci. 2020, 85, 182–191. [Google Scholar] [CrossRef]
- Watson, H.G.; Vanderputten, D.; Van Landschoot, A.; Decloedt, A.I. Applicability of different brewhouse technologies and gluten-minimization treatments for the production of gluten-free (barley) malt beers: Pilot- to industrial-scale. J. Food Eng. 2019, 245, 33–42. [Google Scholar] [CrossRef]
- Hager, A.-S.; Taylor, J.P.; Waters, D.M.; Arendt, E.K. Gluten free beer—A review. Trends Food Sci. Technol. 2014, 36, 44–54. [Google Scholar] [CrossRef]
- Comino, I.; Real, A.; Moreno, M.e.L.; Montes, R.; Cebolla, A.; Sousa, C. Immunological determination of gliadin 33-mer equivalent peptides in beers as a specific and practical analytical method to assess safety for celiac patients. J. Sci. Food Agric. 2013, 93, 933–943. [Google Scholar] [CrossRef]
- European Commission; (EU). Commission Implementing Regulation (EU) No 828/2014 of 30 July 2014: Requirements for the provision of information to consumers on the absence or reduced presence of gluten in food. Off. J. Eur. Union L228 2014, 5–8. [Google Scholar]
- Guerdrum, L.J.; Bamforth, C.W. Prolamin Levels through Brewing and the Impact of Prolyl Endoproteinase. J. Am. Soc. Brew. Chem. 2012, 70, 35–38. [Google Scholar] [CrossRef]
- Fanari, M.; Porcu, M.; Zinellu, M.; Farina, D.; Scognamillo, S.; Forteschi, M.; Luca, P. A preliminary study about gluten levels in Sardinian craft beers. J. Microbiol. Biotechnol. Food Sci. 2017, 6, 1195–1198. [Google Scholar] [CrossRef]
- Watson, H.G.; Decloedt, A.I.; Hemeryck, L.Y.; Van Landschoot, A.; Prenni, J. Peptidomics of an industrial gluten-free barley malt beer and its non-gluten-free counterpart: Characterisation and immunogenicity. Food Chem. 2021, 355, 129597. [Google Scholar] [CrossRef] [PubMed]
- Tanner, G.J.; Blundell, M.J.; Colgrave, M.L.; Howitt, C.A. Quantification of Hordeins by ELISA: The correct standard makes a magnitude of difference. PLoS ONE 2013, 8, e56456. [Google Scholar] [CrossRef] [PubMed]
- Alimentarius, C. Revised version of Standard for Foods for Special Dietary Use for Persons Intolerant to Gluten. CODEX STAN 118-2008 2008, 1–3. [Google Scholar]
- Allred, L.K.; Lesko, K.; McKiernan, D.; Kupper, C.; Guandalini, S. The Celiac Patient Antibody Response to Conventional and Gluten-Removed Beer. J. AOAC Int. 2017, 100, 485–491. [Google Scholar] [CrossRef]
- Panda, R.; Garber, E.A.E. Detection and Quantitation of Gluten in Fermented-Hydrolyzed Foods by Antibody-Based Methods: Challenges, Progress, and a Potential Path Forward. Front. Nutr. 2019, 6, 97. [Google Scholar] [CrossRef]
- Sileoni, V.; Marconi, O.; Perretti, G. Near-infrared Spectroscopy in the Brewing Industry. Crit. Rev. Food Sci. Nutr. 2015, 55, 1771–1791. [Google Scholar] [CrossRef]
- Osborne, B.G. Near-Infrared Spectroscopy in Food Analysis. In Encyclopedia of Analytical Chemistry; Meyers, R.A., McGorrin, R.J., Eds.; John Wiley & Sons, Ltd: Hoboken, NJ, USA, 2006; pp. 1–14. [Google Scholar]
- Revilla, I.; Escuredo, O.; González-Martín, M.I.; Palacios, C. Fatty acids and fat-soluble vitamins in ewe’s milk predicted by near infrared reflectance spectroscopy. Determination of seasonality. Food Chem. 2017, 214, 468–477. [Google Scholar] [CrossRef]
- García-Molina, M.D.; García-Olmo, J.; Barro, F. Effective Identification of Low-Gliadin Wheat Lines by Near Infrared Spectroscopy (NIRS): Implications for the Development and Analysis of Foodstuffs Suitable for Celiac Patients. PLoS ONE 2016, 11, e0152292. [Google Scholar] [CrossRef]
- Albanell, E.; Miñarro, B.; Carrasco, N. Detection of low-level gluten content in flour and batter by near infrared reflectance spectroscopy (NIRS). J. Cereal Sci. 2012, 56, 490–495. [Google Scholar] [CrossRef]
- Elli, L.; Bascuñán, K.; di Lernia, L.; Bardella, M.T.; Doneda, L.; Soldati, L.; Orlando, S.; Ferretti, F.; Lombardo, V.; Barigelletti, G.; et al. Safety of occasional ingestion of gluten in patients with celiac disease: A real-life study. BMC Med. 2020, 18, 42. [Google Scholar] [CrossRef]
- Don, C.; Halbmayr-Jech, E.; Rogers, A.; Koehler, P. AACCI Approved Methods Technical Committee Report: Collaborative Study on the Immunochemical Quantitation of Intact Gluten in Rice Flour and Rice-Based Products Using G12 Sandwich ELISA. Cereal Foods World 2014, 59, 187–193. [Google Scholar] [CrossRef]
- Kanerva, P.; Sontag-Strohm, T.; Lehtonen, P. Determination of Prolamins in Beers by ELISA and SDS-PAGE. J. Inst. Brew. 2005, 111, 61–64. [Google Scholar] [CrossRef]
- Haas-Lauterbach, S.; Immer, U.; Richter, M.; Oehler, E. Gluten Fragment Detection with a Competitive ELISA. J. AOAC Int. 2012, 95, 377–381. [Google Scholar] [CrossRef]
- Mitea, C.; Kooy-Winkelaar, Y.; van Veelen, P.; de Ru, A.; Drijfhout, J.W.; Koning, F.; Dekking, L. Fine specificity of monoclonal antibodies against celiac disease-inducing peptides in the gluteome. Am. J. Clin. Nutr. 2008, 88, 1057–1066. [Google Scholar] [CrossRef]
- Watson, H.G.; Decloedt, A.I.; Vanderputten, D.; Van Landschoot, A. Variation in gluten protein and peptide concentrations in Belgian barley malt beers. J. Inst. Brew. 2018, 124, 148–157. [Google Scholar] [CrossRef]
- Scherf, K.A.; Poms, R.E. Recent developments in analytical methods for tracing gluten. J. Cereal Sci. 2016, 67, 112–122. [Google Scholar] [CrossRef]
- Mujico, J.R.; Dekking, L.; Kooy-Winkelaar, Y.; Verheijen, R.; van Wichen, P.; Streppel, L.; Sajic, N.; Drijfhout, J.W.; Koning, F. Validation of a new enzyme-linked immunosorbent assay to detect the triggering proteins and peptides for celiac disease: Interlaboratory study. J. AOAC Int. 2012, 95, 206–215. [Google Scholar] [CrossRef] [PubMed]
- Belitz, H.-D.; Grosch, W.; Schieberle, P. Food Chemistry, 4th revised and extended ed.; Springer: Berlin/Heidelberg, Germany, 2009. [Google Scholar]
- Tanner, G.J.; Colgrave, M.L.; Blundell, M.J.; Goswami, H.P.; Howitt, C.A. Measuring hordein (gluten) in beer--a comparison of ELISA and mass spectrometry. PLoS ONE 2013, 8, e56452. [Google Scholar] [CrossRef] [PubMed]
- Panda, R.; Boyer, M.; Garber, E.A.E. A multiplex competitive ELISA for the detection and characterization of gluten in fermented-hydrolyzed foods. Anal. Bioanal. Chem. 2017, 409, 6959–6973. [Google Scholar] [CrossRef]
- Van Landschoot, A. Gluten-free barley malt beers. Cerevisia 2011, 36, 93–97. [Google Scholar] [CrossRef]
- Guerdrum, L.J.; Bamforth, C.W. Levels of gliadin in commercial beers. Food Chem. 2011, 129, 1783–1784. [Google Scholar] [CrossRef]
- Scherf, K.A.; Catassi, C.; Chirdo, F.G.; Ciclitira, P.J.; Feighery, C.F.; Gianfrani, C.; Koning, F.; Lundin, K.E.A.; Masci, S.; Schuppan, D.; et al. Statement of the Prolamin Working Group on the Determination of Gluten in Fermented Foods Containing Partially Hydrolyzed Gluten. Front. Nutr. 2020, 7, 626712. [Google Scholar] [CrossRef] [PubMed]
- Department of Agriculture, U.S (USDA). Agricultural Marketing Service. Available online: https://www.ams.usda.gov/services/fgis/standardization/wheat-protein (accessed on 9 May 2021).
- Wesley, I.J.; Larroque, O.; Osborne, B.G.; Azudin, N.; Allen, H.; Skerritt, J.H. Measurement of Gliadin and Glutenin Content of Flour by NIR Spectroscopy. J. Cereal Sci. 2001, 34, 125–133. [Google Scholar] [CrossRef]
- Wimonsiri, L.; Ritthiruangdej, P.; Kasemsumran, S.; Therdthai, N.; Chanput, W.; Ozaki, Y. Rapid analysis of chemical composition in intact and milled rice cookies using near infrared spectroscopy. J. Near Infrared Spectrosc. 2017, 25, 330–337. [Google Scholar] [CrossRef]
- Giovenzana, V.; Beghi, R.; Guidetti, R. Rapid evaluation of craft beer quality during fermentation process by vis/NIR spectroscopy. J. Food Eng. 2014, 142, 80–86. [Google Scholar] [CrossRef]
- Cozzolino, D.; Delucchi, I.; Kholi, M.; Vázquez, D. Uso de la espectroscopía de reflectancia en el infrarrojo cercano para evaluar características de calidad en trigo. Agric. Técnica 2006, 66, 370–375. [Google Scholar]
| Beer Descriptor | Sample Number Labelled GF | Sample Number Conventional | |
|---|---|---|---|
| Manufacturer | |||
| Craft | 16 | 21 | |
| Industrial | 25 | 44 | |
| Yeast style | |||
| Ale | 23 | 37 | |
| Lager | 18 | 28 | |
| Ingredients * | |||
| Made from gluten-containing cereals only | 25 | 53 | |
| Made from gluten-containing and GF cereals | 13 | 11 | |
| Include wheat | 3 | 20 | |
| Without wheat | 36 | 43 | |
| Original extract (OE) ** | |||
| Traditional beer (<13 grams of OE per 100 grams of wort) | 20 | 41 | |
| Special beer (13–15 grams of OE per 100 grams of wort) | 10 | 14 | |
| Extra-special beer (>15 grams of OE per 100 grams of wort) | 4 | 8 | |
| Country of origin | |||
| Spain | 19 | 45 | |
| United Kingdom | 4 | 0 | |
| Italy | 1 | 0 | |
| Germany | 2 | 8 | |
| Belgium | 11 | 4 | |
| Norway | 1 | 0 | |
| Netherlands | 0 | 4 | |
| France | 1 | 0 | |
| Check Republic Estonia | 1 1 | 4 0 | |
| Overall beer | 41 | 65 |
| ELISA Kit | Calibrant | Extraction Volume | Approximate Time Requirement | ||
|---|---|---|---|---|---|
| (Reference) | (10 samples) | Sample Preparation (10 samples) | Implementation (10 samples) | Total Time 10 samples | |
| RIDASCREEN Gliadin Competitive (R7021) | Hydrolysate prolamin from mixture of wheat (PGW gliadin), rye, and barley. | 60%ethanol + 10% fish gelatin (90 mL) | 30 min | 45 min | 75 min |
| INGEZIM Gluten Hidrolizado (R.30.GLH.K2) | Gliadin European standard * | Polyvinylpyrrolidone (1 g) Extraction buffer (25 mL) 80% ethanol (75 mL) | 25 min | 215 min | 240 min |
| GLUTEN TEC (5171GT(10)03.20) | α-20 gliadin peptide | 60%ethanol (45 mL) | 25 min | 80 min | 105 min |
| Gluten Content mg/kg | Manufacturer | Yeast Style | Include Wheat | Original Extract * | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Industrial | Craft | p | Ale | Lager | p | Yes | No | p | Beer | Special/extra | p | |
| R5 competitive | ||||||||||||
| Labeled GF | ||||||||||||
| Mean ± SD | 9.723 ± 16.50 | 47.17 ± 119.6 | 0.597 | 8.900 ± 16.01 | 44.11 ± 112.4 | 0.678 | 3.400 ± 0.001 | 27.42 ± 80.01 | 0.379 | 32.58 ± 97.80 | 7.867 ± 10.94 | 1.000 |
| Conventional | ||||||||||||
| Mean ± SD | 502.5 ± 972.0 | 148.4 ± 338.4 | 0.832 | 650.1 ± 1064 | 73.11 ± 134.4 | 0.001 | 1145 ± 1206 | 44.11 ± 30.64 | <0.001 | 549.4 ± 1032 | 153.3 ± 270.1 | 0.771 |
| R5 direct Labeled GF | ||||||||||||
| Mean ± SD | 1.073 ± 2.615 | 6.300 ± 14.82 | 0.526 | 1.542 ± 3.102 | 5.095 ± 14.07 | 0.756 | 0.250 ± 0.001 | 3.533 ± 10.08 | 0.379 | 4.283 ± 12.19 | 0.458 ± 0.510 | 0.885 |
| Conventional | ||||||||||||
| Mean ± SD | 70.29 ± 129.6 | 22.76 ± 41.67 | 0.649 | 89.15 ± 140.6 | 13.89 ± 27.23 | 0.001 | 158.4 ± 157.2 | 7.872 ± 8.045 | <0.001 | 76.32 ± 138.3 | 22.22 ± 26.95 | 0.838 |
| α20gliadin competitive | ||||||||||||
| Labeled GF | ||||||||||||
| Mean ± SD | 2.415 ± 1.982 | 11.95 ± 27.65 | 0.698 | 2.133 ± 1.787 | 11.27 ± 25.94 | 0.324 | 1.400 ± 0.001 | 6.822 ± 18.43 | 0.313 | 8.433 ± 22.66 | 3.067 ± 2.594 | 0.750 |
| Conventional | ||||||||||||
| Mean ± SD | 329.5 ± 554.0 | 108.1 ± 308.7 | 0.403 | 425.4 ± 615.0 | 56.58 ± 137.1 | 0.001 | 784.0 ± 626.8 | 18.00 ± 15.12 | <0.001 | 357.0 ± 601.4 | 116.6 ± 209.9 | 0.804 |
| Gluten Range (mg/kg) | Sample Number | R2 | Outlier Sample Number |
|---|---|---|---|
| 0–19 | 214 | 0.139 | 23 |
| 20–100 | 87 | 0.184 | 8 |
| >100 | 34 | 0.592 | 3 |
| 0->100 | 335 | 0.167 | 21 |
| Codex categorizing * (GF, LG, GC) | 335 | 0.201 | 31 |
| Dichotomic categorizing (GF, non-GF) | 335 | 0.185 | 27 |
| R5 Competitive | R5 Direct | α20gliadin Competitive | ||||||
|---|---|---|---|---|---|---|---|---|
| Evaluated Sample Number | NIR R2 | ELISA Gluten Mean ± SD | NIR R2 | ELISA Gluten Mean ± SD | NIR R2 | ELISA Gluten Mean ± SD | ELISA ANOVA | |
| No categorizing | 71 | 0.435 | 286.1 a ± 729.0 | 0.450 | 40.37 b ± 97.30 | 0.482 | 187.1 a ± 435.3 | <0.001 |
| Codex categorizing (GF, LG, GC) | 71 | 0.500 | 0.505 | 0.520 | ||||
| Dichotomic categorizing (GF, non-GF) | 71 | 0.341 | 0.472 | 0.450 | ||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fernández-Gil, M.d.P.; Simon, E.; Gibert, A.; Miranda, J.; Roger Alcoba, E.; Martínez, O.; Vilchez Cerezo, E.; Bustamante, M.Á. Gluten Assessment in Beers: Comparison by Different Commercial ELISA Kits and Evaluation of NIR Analysis as a Complementary Technique. Foods 2021, 10, 1170. https://doi.org/10.3390/foods10061170
Fernández-Gil MdP, Simon E, Gibert A, Miranda J, Roger Alcoba E, Martínez O, Vilchez Cerezo E, Bustamante MÁ. Gluten Assessment in Beers: Comparison by Different Commercial ELISA Kits and Evaluation of NIR Analysis as a Complementary Technique. Foods. 2021; 10(6):1170. https://doi.org/10.3390/foods10061170
Chicago/Turabian StyleFernández-Gil, María del Pilar, Edurne Simon, Anna Gibert, Jonatan Miranda, Esther Roger Alcoba, Olaia Martínez, Elisenda Vilchez Cerezo, and María Ángeles Bustamante. 2021. "Gluten Assessment in Beers: Comparison by Different Commercial ELISA Kits and Evaluation of NIR Analysis as a Complementary Technique" Foods 10, no. 6: 1170. https://doi.org/10.3390/foods10061170
APA StyleFernández-Gil, M. d. P., Simon, E., Gibert, A., Miranda, J., Roger Alcoba, E., Martínez, O., Vilchez Cerezo, E., & Bustamante, M. Á. (2021). Gluten Assessment in Beers: Comparison by Different Commercial ELISA Kits and Evaluation of NIR Analysis as a Complementary Technique. Foods, 10(6), 1170. https://doi.org/10.3390/foods10061170