Pepsin Hydrolysis of Orange By-Products for the Production of Bioactive Peptides with Gastrointestinal Resistant Properties
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Preparation of Orange Seed Protein Concentrates
2.3. Enzymatic Hydrolysis Conditions
2.4. DPPH Radical Scavenging Activity Assay
2.5. Ferric Reducing Antioxidant Power Assay
2.6. ACEI Inhibition Activity Assay
2.7. α-Amylase Inhibition Activity Assay
2.8. α-Glucosidase Inhibition Activity Assay
2.9. Size-Exclusion Chromatography (SEC) Separation of Hydrolyzed Protein
2.10. RP-HPLC Separation of SEC Fractions
2.11. In Vitro Gastrointestinal Digestion
2.12. Identification of Peptides Using Mass Spectrometry in Tandem
2.13. Statistical Analysis
3. Results
3.1. Evaluation of Biological Activity of the Hydrolyzed Proteins
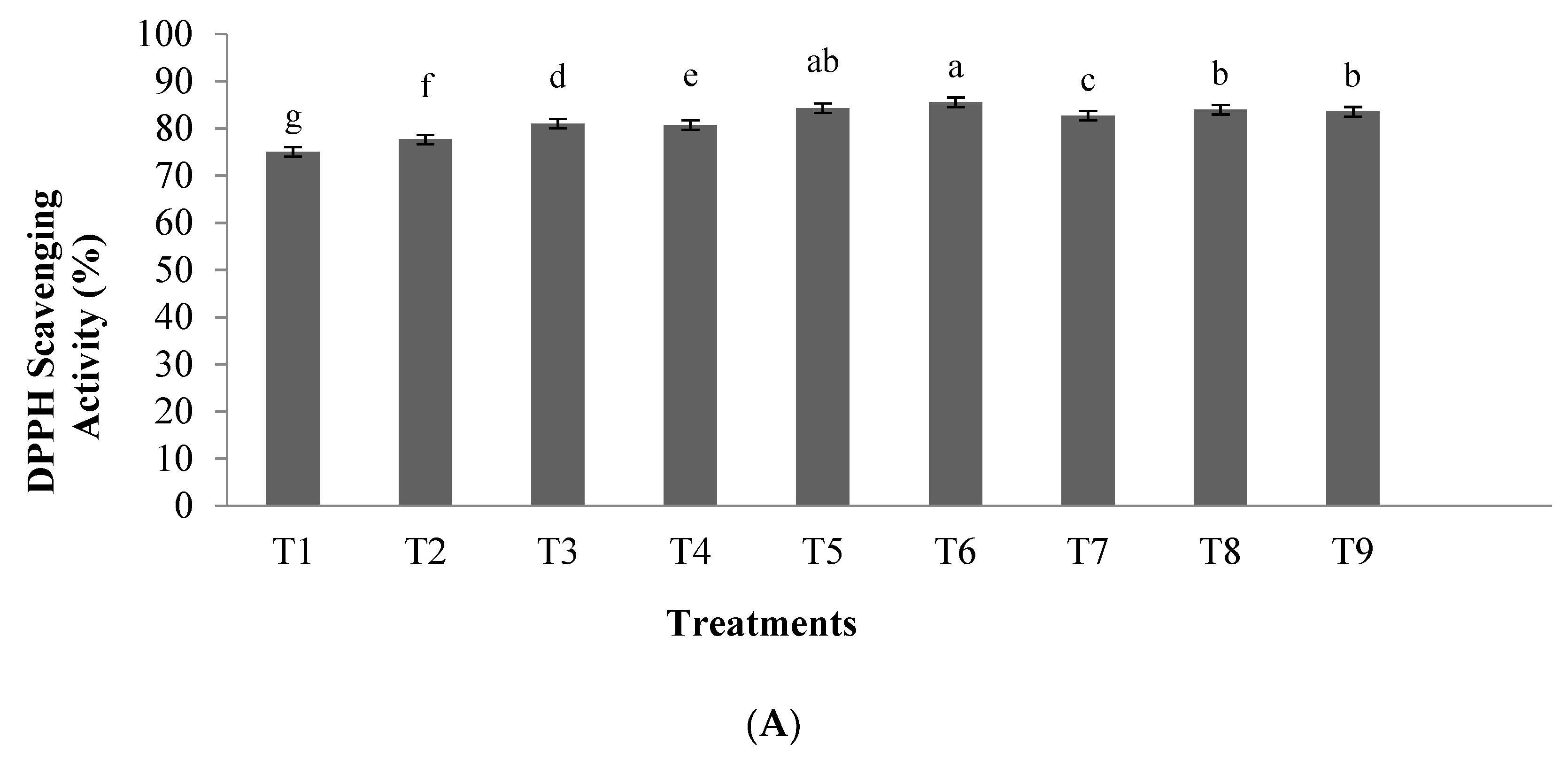
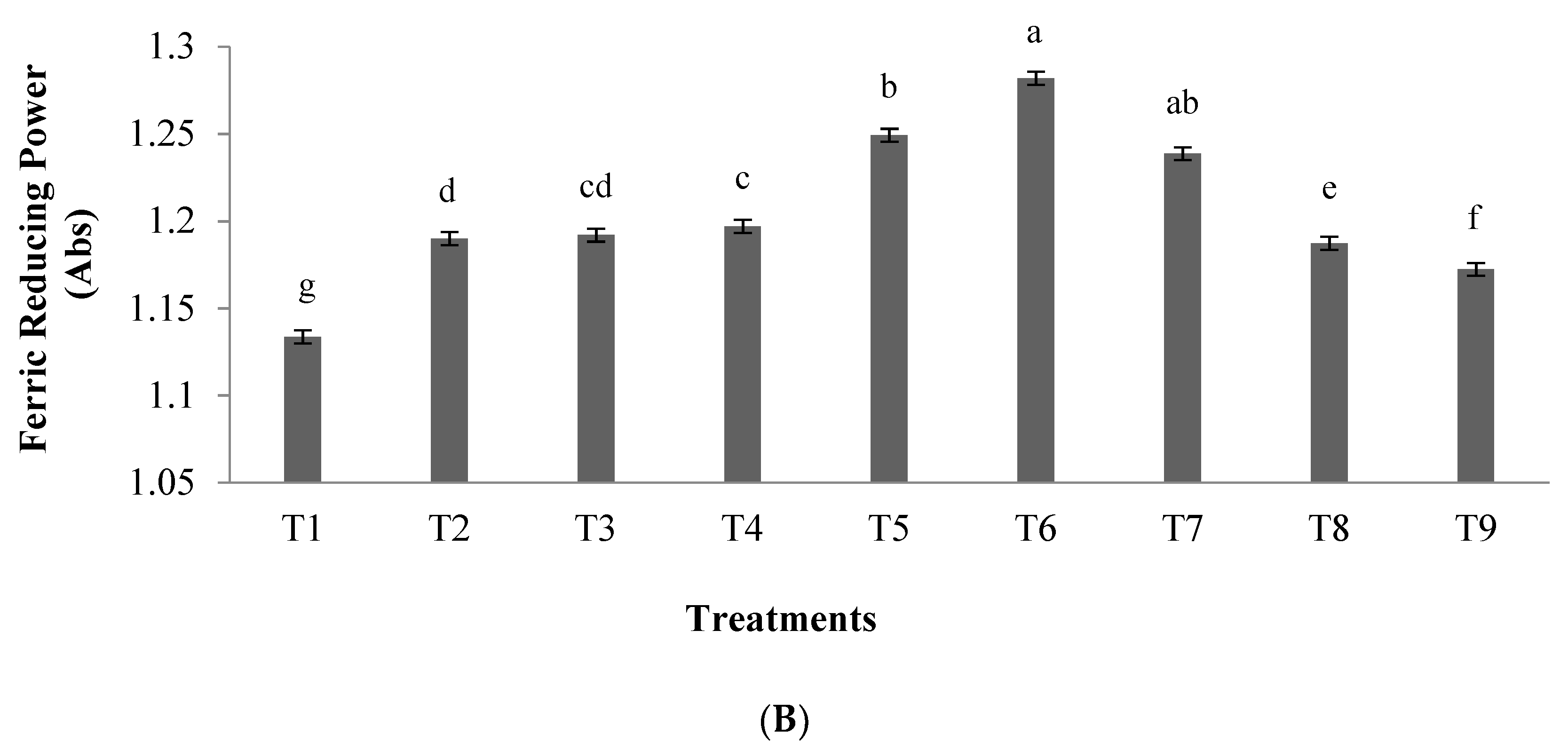
3.1.1. Antioxidant Activity
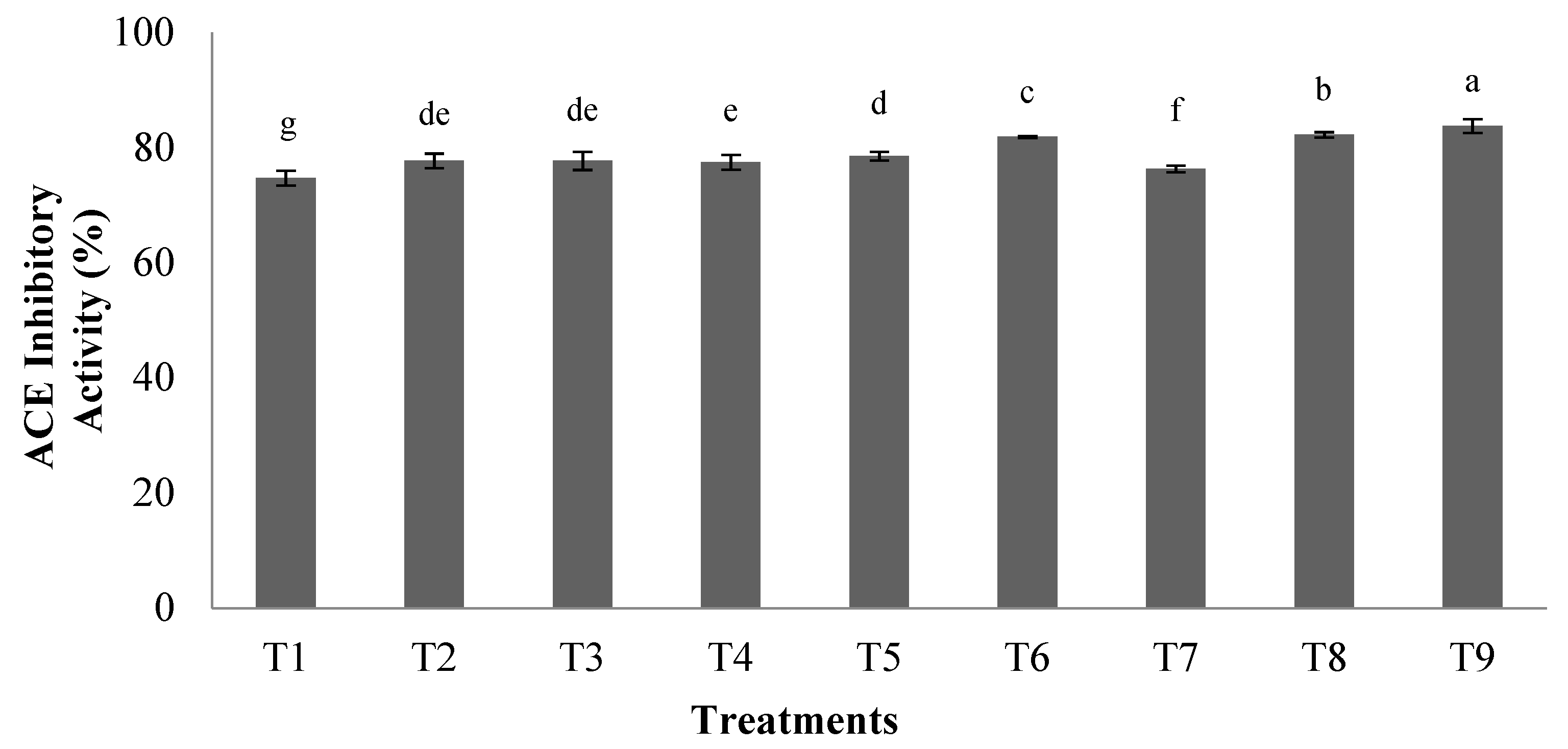
3.1.2. ACEI-Inhibitory Activity
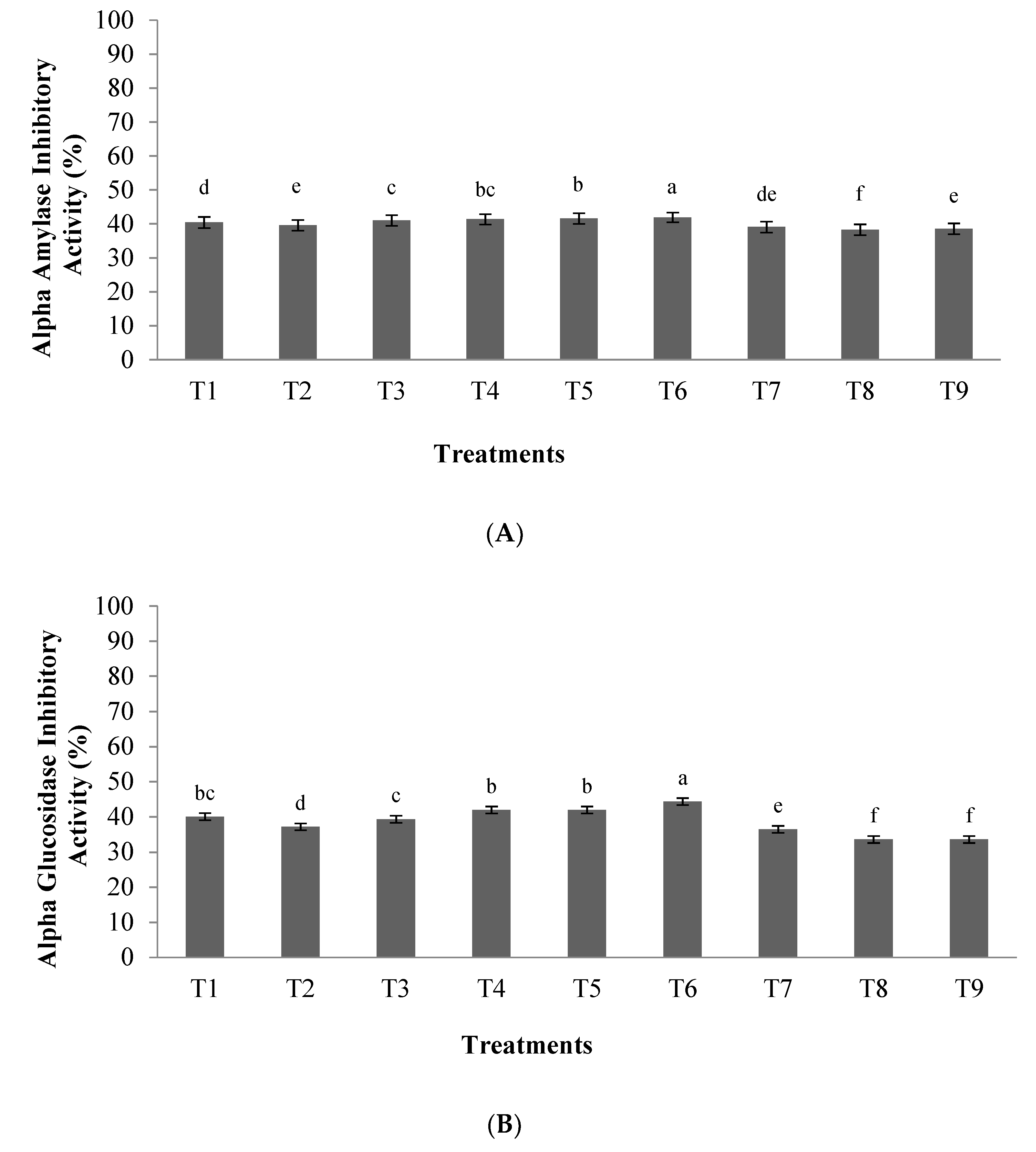
3.1.3. α-Amylase and α-Glucosidase Inhibitory Activity
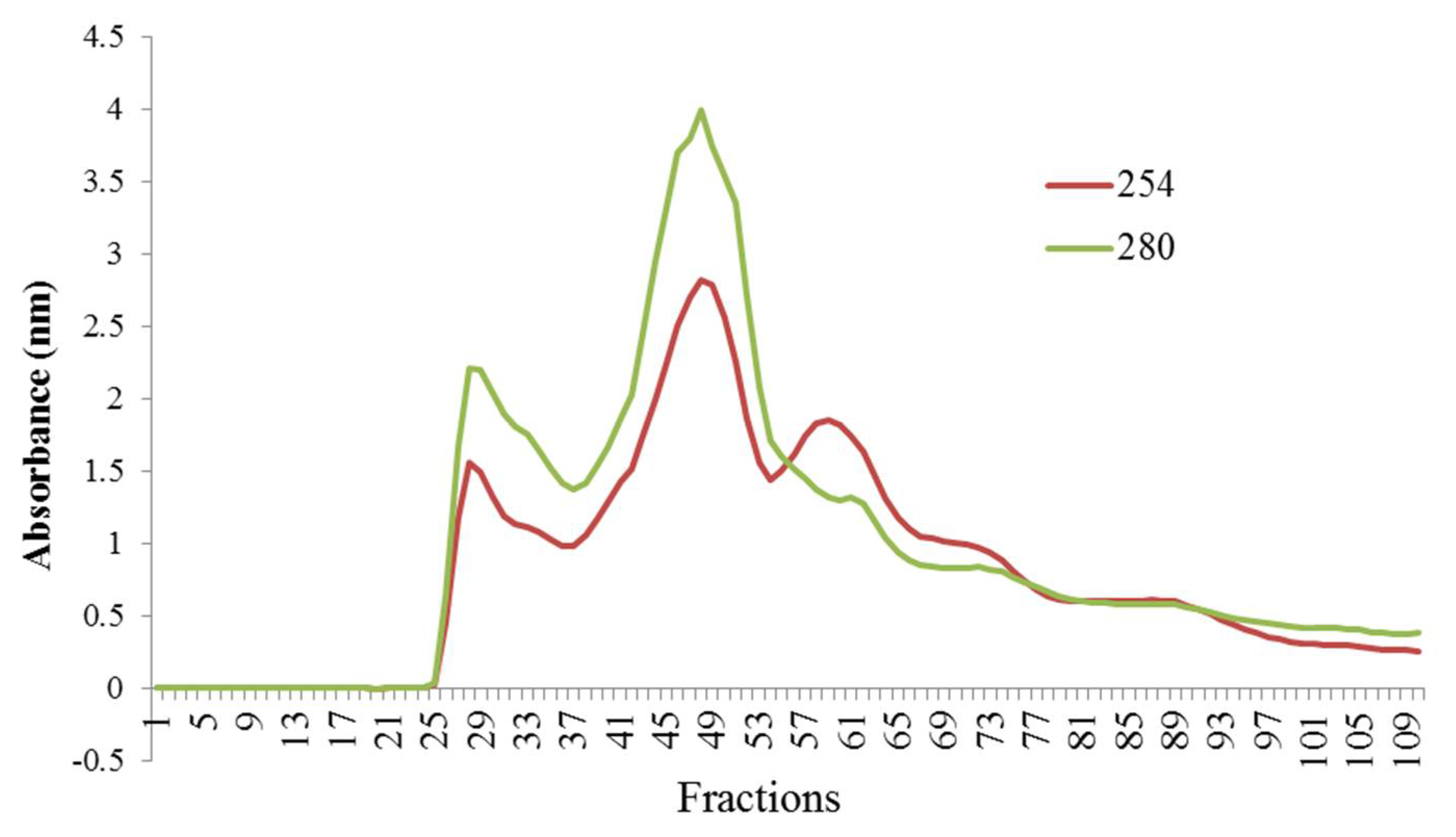
3.2. Fractionation of Hydrolyzed Orange Seed Proteins Using SEC
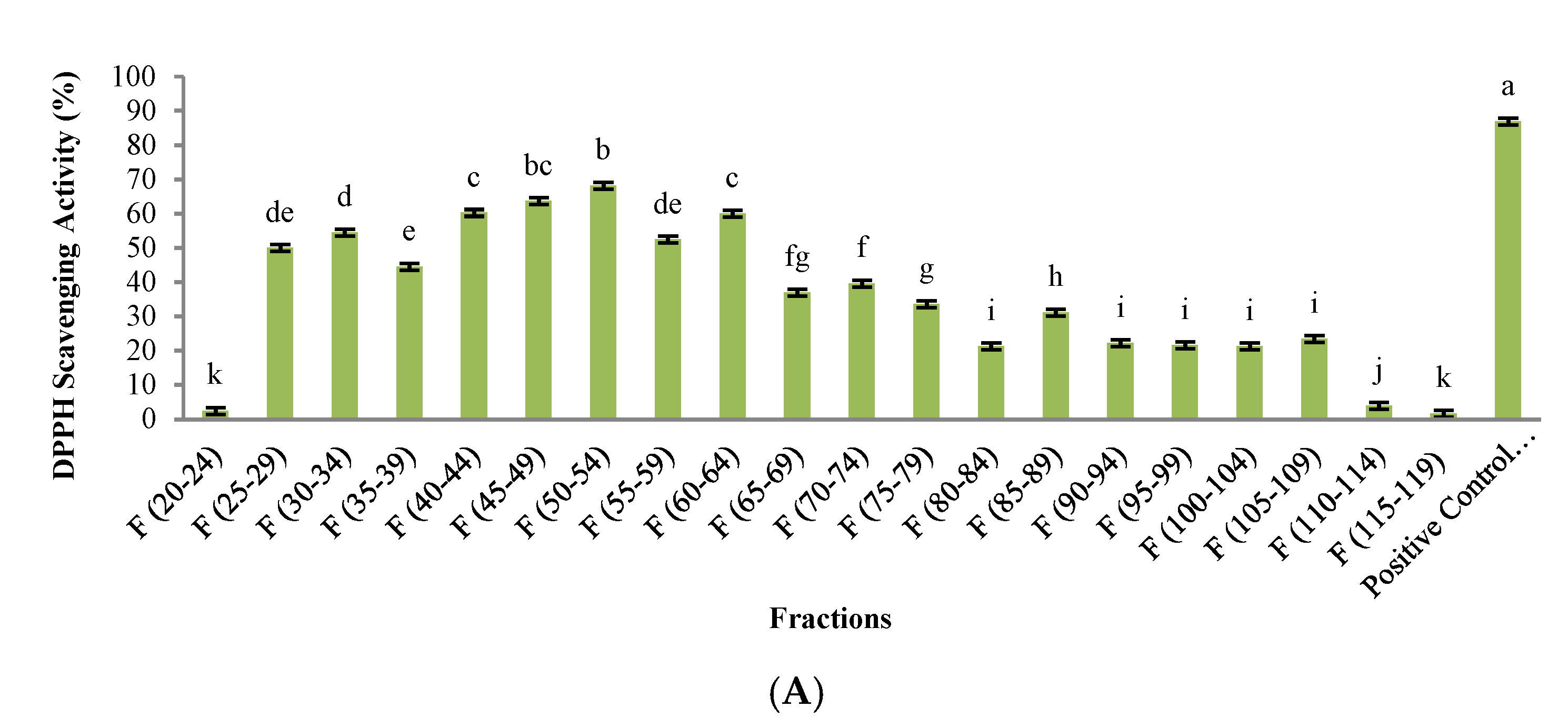
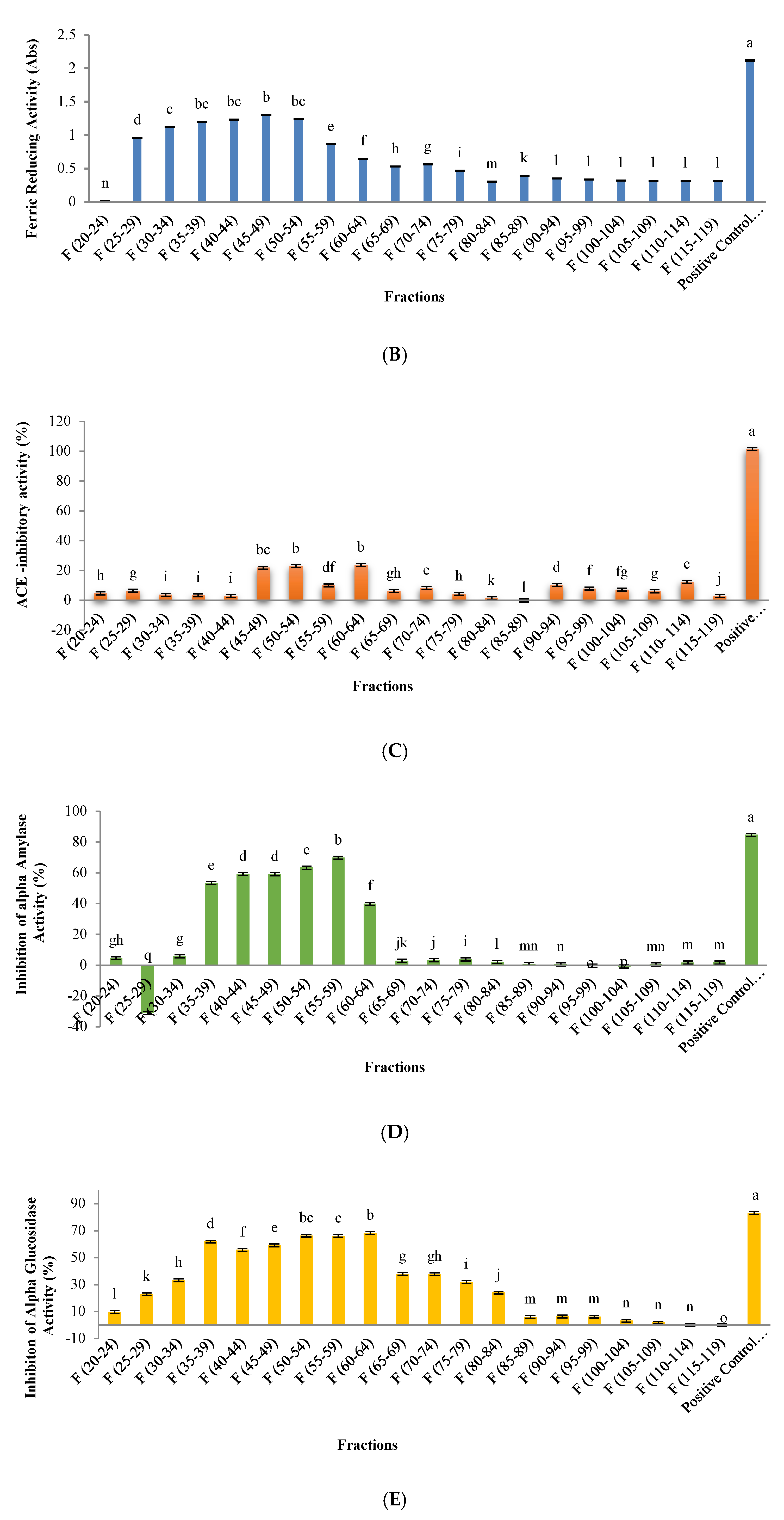
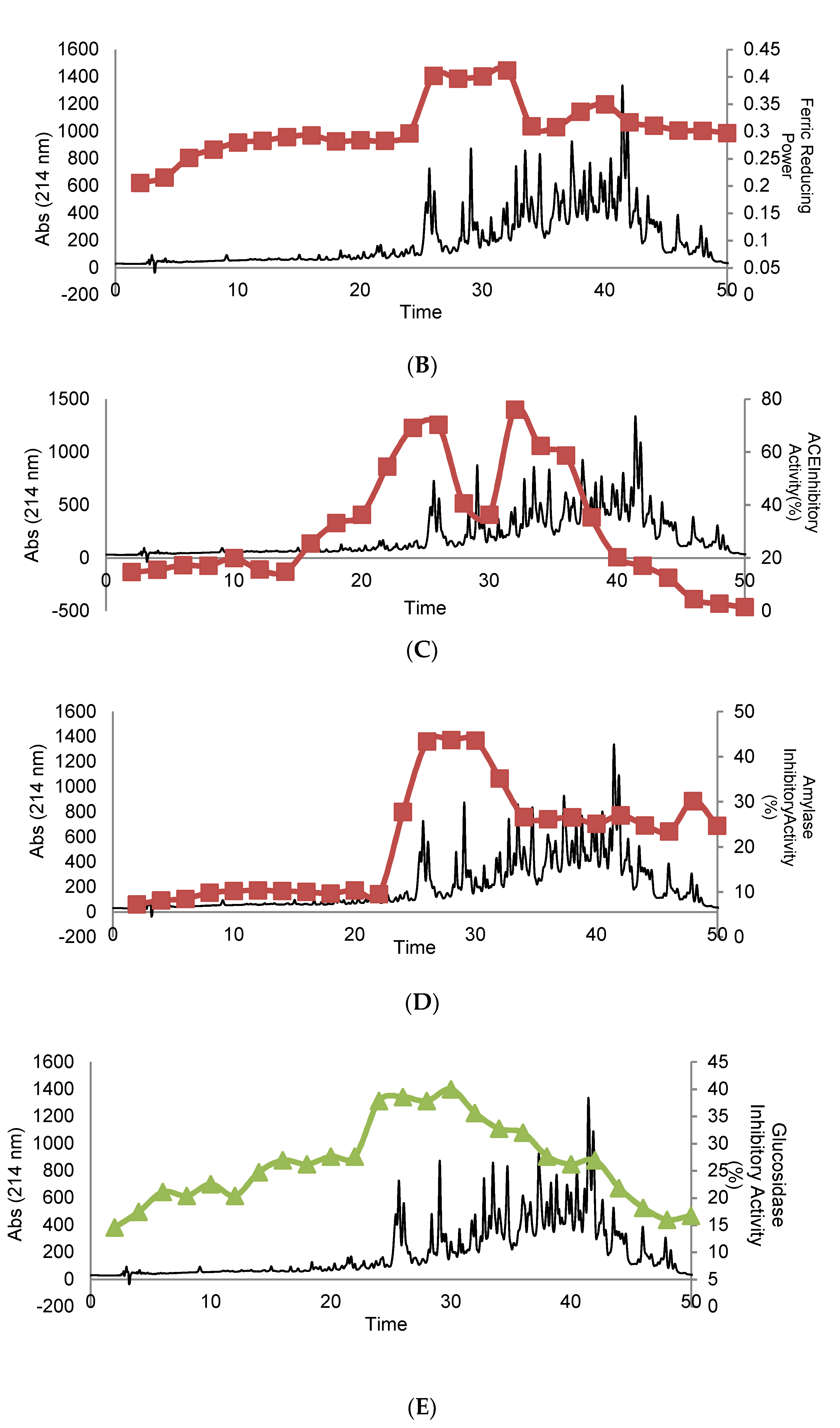
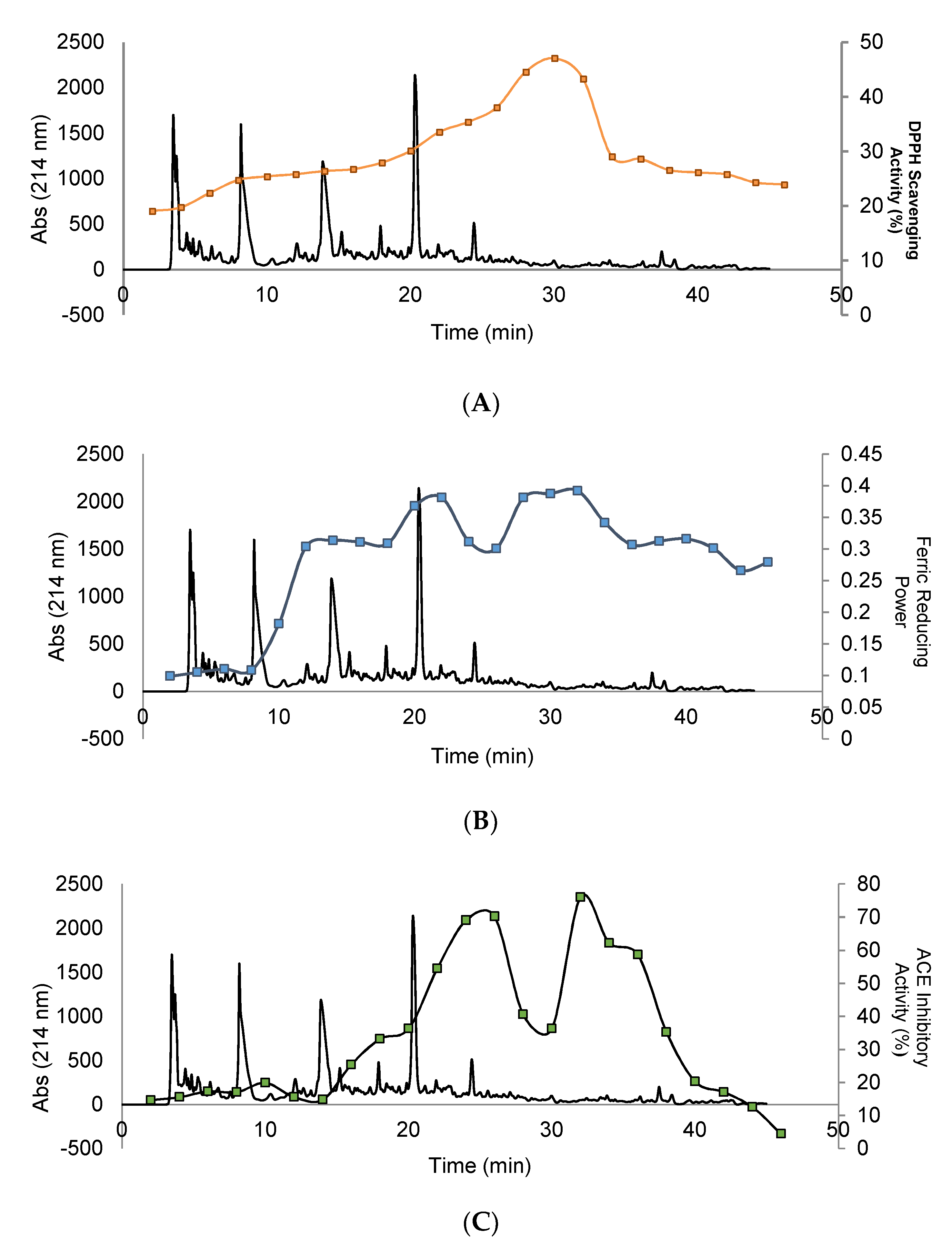
3.3. Evaluation of Biological Activity in Peptides Isolated from SEC
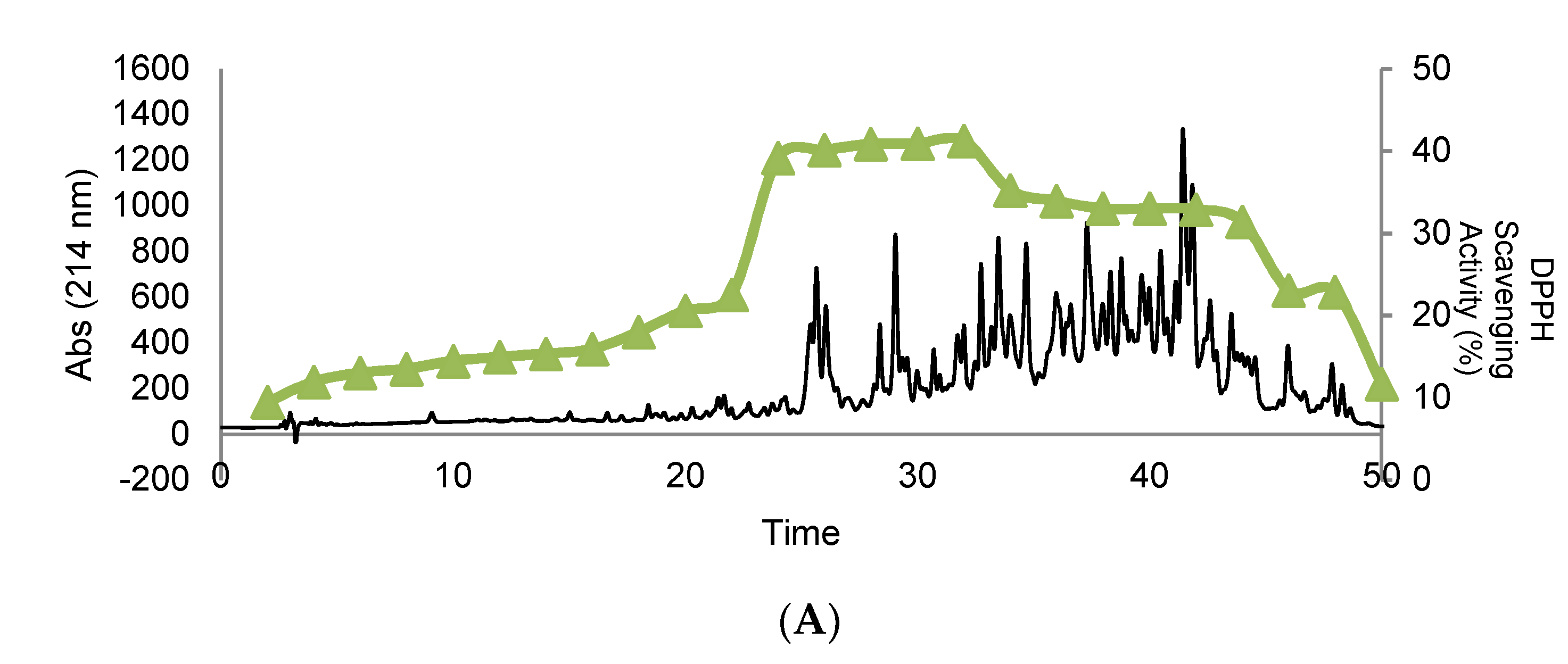
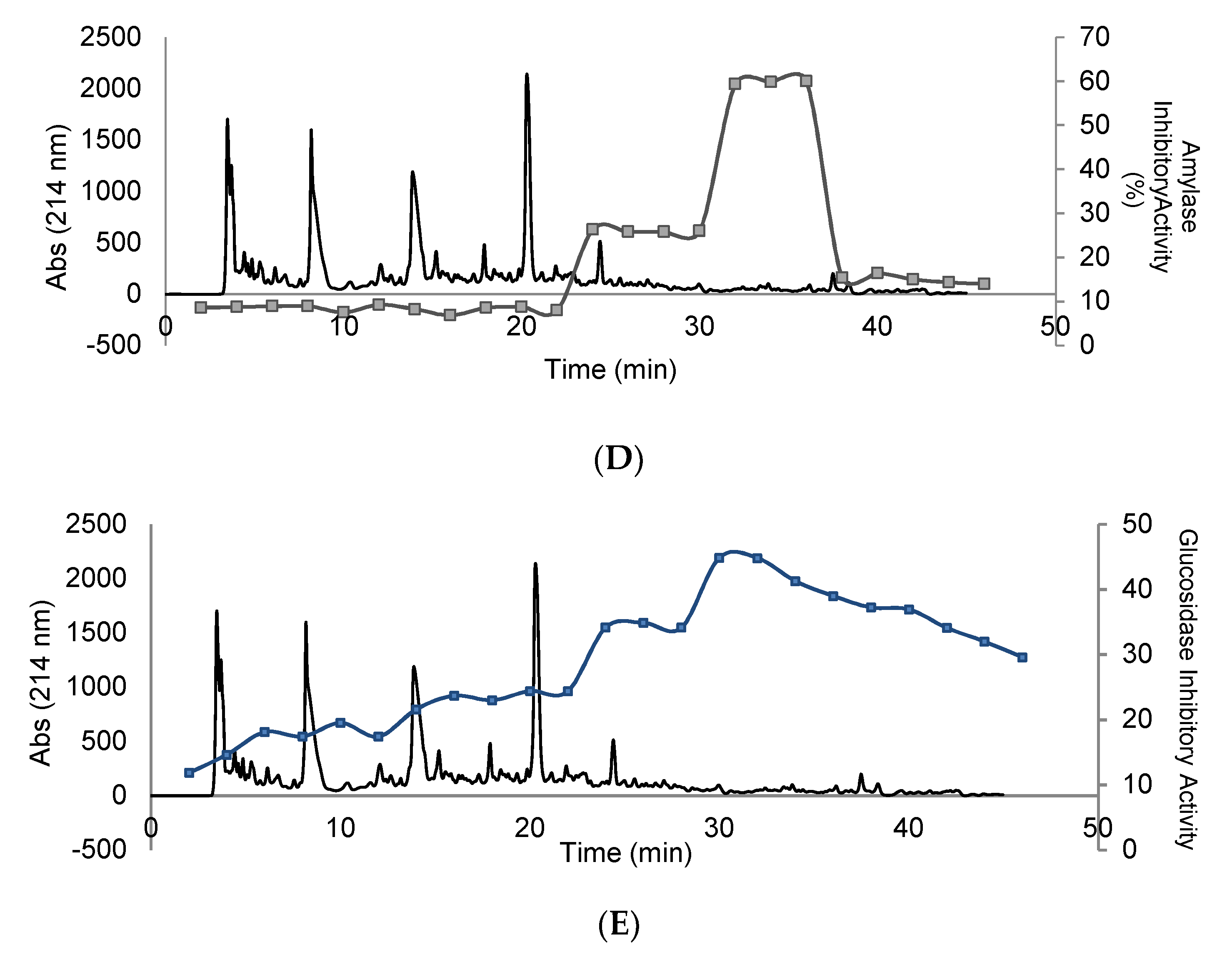
3.4. Purification and Isolation of Peptide Fractions Using RP-HPLC
3.5. Effect of In Vitro GI Digestion on the Peptides’ Bioactivity after RP-HPLC Separation
3.6. Identification of Peptides by Nano-LC-MS/MS
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- De Deckere, E.; Verschuren, P. Functional fats and spreads. In Functional Foods; Elsevier: Amsterdam, The Netherlands, 2000; pp. 233–257. [Google Scholar]
- Taha, F.S.; Mohamed, S.S.; Wagdy, S.M.; Mohamed, G.F. Antioxidant and antimicrobial activities of enzymatic hydrolysis products from sunflower protein isolate. World Appl. Sci. J. 2013, 21, 651–658. [Google Scholar]
- Selamassakul, O.; Laohakunjit, N.; Kerdchoechuen, O.; Yang, L.; Maier, C.S. Isolation and characterisation of antioxidative peptides from bromelain-hydrolysed brown rice protein by proteomic technique. Process Biochem. 2018, 70, 179–187. [Google Scholar] [CrossRef] [PubMed]
- Jo, C.; Khan, F.F.; Khan, M.I. Marine bioactive peptides: Types, structures, and physiological functions. Food Rev. Int. 2016, 33, 44–61. [Google Scholar] [CrossRef]
- De Domenico, S.; De Rinaldis, G.; Paulmery, M.; Piraino, S.; Leone, A. Barrel Jellyfish (Rhizostoma pulmo) as Source of Antioxidant Peptides. Mar. Drugs 2019, 17, 134. [Google Scholar] [CrossRef]
- Ngo, D.H.; Kim, S.K. Marine Bioactive Peptides as Potential Antioxidants. Curr. Protein Pept. Sci. 2013, 14, 189–198. [Google Scholar] [CrossRef] [PubMed]
- Szwajkowska, M.; Wolanciuk, A.; Barłowska, J.; Krol, J.; Litwińczuk, Z. Bovine milk proteins as the source of bioactive peptides influencing the consumers’ immune system—A review. Anim. Sci. Pap. Rep. 2011, 29, 269–280. [Google Scholar]
- Soleymanzadeh, N.; Mirdamadi, S.; Mirzaei, M.; Kianirad, M. Novel β-casein derived antioxidant and ACE-inhibitory active peptide from camel milk fermented by Leuconostoc lactis PTCC1899: Identification and molecular docking. Int. Dairy J. 2019, 97, 201–208. [Google Scholar] [CrossRef]
- Chakrabarti, S.; Jahandideh, F.; Wu, J. Food-derived bioactive peptides on inflammation and oxidative stress. BioMed Res. Int. 2014, 2014, 608979. [Google Scholar] [CrossRef]
- Yang, H.J.; Kwon, D.Y.; Kim, M.J.; Kang, S.; Park, S. Meju, unsalted soybeans fermented with Bacillus subtilis and Aspergilus oryzae, potentiates insulinotropic actions and improves hepatic insulin sensitivity in diabetic rats. Nutr. Metab. 2012, 9, 37. [Google Scholar] [CrossRef]
- Maqsoudlou, A.; Mahoonak, A.S.; Mora, L.; Mohebodini, H.; Toldrá, F.; Ghorbani, M. Peptide identification in alcalase hydrolysated pollen and comparison of its bioactivity with royal jelly. Int. Food Res. J. 2019, 116, 905–915. [Google Scholar] [CrossRef]
- Mazloomi, S.N.; Mahoonak, A.S.; Ghorbani, M.; Houshmand, G. Physicochemical properties of chitosan-coated nanoliposome loaded with orange seed protein hydrolysate. J. Food Eng. 2020, 280, 109976. [Google Scholar] [CrossRef]
- Mazloomi-Kiyapey, S.N.; Sadeghi-Mahoonak, A.; Ranjbar-Nedamani, E.; Nourmohammadi, E. Production of antioxidant peptides through hydrolysis of medicinal pumpkin seed protein using pepsin enzyme and the evaluation of their functional and nutritional properties. ARYA Atheroscler. 2019, 15, 218. [Google Scholar]
- Xie, J.; Du, M.; Shen, M.; Wu, T.; Lin, L. Physico-chemical properties, antioxidant activities and angiotensin-I converting enzyme inhibitory of protein hydrolysates from Mung bean (Vigna radiate). Food Chem. 2019, 270, 243–250. [Google Scholar] [CrossRef]
- Ramezanzade, L.; Hosseini, S.F.; Nikkhah, M. Biopolymer-coated nanoliposomes as carriers of rainbow trout skin-derived antioxidant peptides. Food Chem. 2017, 234, 220–229. [Google Scholar] [CrossRef]
- Mohan, A.; Rajendran, S.R.; Thibodeau, J.; Bazinet, L.; Udenigwe, C.C. Liposome encapsulation of anionic and cationic whey peptides: Influence of peptide net charge on properties of the nanovesicles. LWT 2018, 87, 40–46. [Google Scholar] [CrossRef]
- Meshginfar, N.; Mahoonak, A.S.; Hosseinian, F.; Tsopmo, A. Physicochemical, antioxidant, calcium binding, and angiotensin converting enzyme inhibitory properties of hydrolyzed tomato seed proteins. J. Food Biochem. 2019, 43, e12721. [Google Scholar] [CrossRef]
- Mohamed, B.; El-Shenawi, M. Functional properties and In-vitro digestibility of bitter orange (Citrus aurantium) seed flour. Merit Res. J. Agric. Sci. Soil Sci. 2013, 1, 47. [Google Scholar]
- Horax, R.; Hettiarachchy, N.; Over, K.; Chen, P.; Gbur, E. Extraction, fractionation and characterization of bitter melon seed proteins. J. Agric. Food Chem. 2010, 58, 1892–1897. [Google Scholar] [CrossRef]
- Matsuoka, T.; Kawashima, T.; Nakamura, T.; Kanamaru, Y.; Yabe, T. Isolation and characterization of proteases that hydrolyze royal jelly proteins from queen bee larvae of the honeybee, Apis mellifera. Apidologie 2012, 43, 685–697. [Google Scholar] [CrossRef]
- Bersuder, P.; Hole, M.; Smith, G. Antioxidants from a heated histidine-glucose model system. I: Investigation of the antioxidant role of histidine and isolation of antioxidants by high-performance liquid chromatography. J. Am. Oil Chem. Soc. 1998, 75, 181–187. [Google Scholar] [CrossRef]
- Bougatef, A.; Hajji, M.; Balti, R.; Lassoued, I.; Triki-Ellouz, Y.; Nasri, M. Antioxidant and free radical-scavenging activities of smooth hound (Mustelus mustelus) muscle protein hydrolysates obtained by gastrointestinal proteases. Food Chem. 2009, 114, 1198–1205. [Google Scholar] [CrossRef]
- Sentandreu, M.A.; Toldrá, F. A fluorescence-based protocol for quantifying angiotensin-converting enzyme activity. Nat. Protoc. 2006, 1, 2423. [Google Scholar] [CrossRef]
- Lassoued, I.; Mora, L.; Barkia, A.; Aristoy, M.C.; Nasri, M.; Toldrá, F. Angiotensin I-converting enzyme inhibitory peptides FQPSF and LKYPI identified in Bacillus subtilis A26 hydrolysate of thornback ray muscle. Int. J. Food Sci. Technol. 2016, 51, 1604–1609. [Google Scholar] [CrossRef]
- Lassoued, I.; Mora, L.; Barkia, A.; Aristoy, M.-C.; Nasri, M.; Toldrá, F. Bioactive peptides identified in thornback ray skin’s gelatin hydrolysates by proteases from Bacillus subtilis and Bacillus amyloliquefaciens. J. Proteom. 2015, 128, 8–17. [Google Scholar] [CrossRef]
- Minekus, M.; Alminger, M.; Alvito, P.; Ballance, S.; Bohn, T.; Bourlieu, C.; Carriere, F.; Boutrou, R.; Corredig, M.; Dupont, D. A standardised static in vitro digestion method suitable for food–an international consensus. Food Funct. 2014, 5, 1113–1124. [Google Scholar] [CrossRef]
- Jamdar, S.; Rajalakshmi, V.; Pednekar, M.; Juan, F.; Yardi, V.; Sharma, A. Influence of degree of hydrolysis on functional properties, antioxidant activity and ACE inhibitory activity of peanut protein hydrolysate. Food Chem. 2010, 121, 178–184. [Google Scholar] [CrossRef]
- Je, J.-Y.; Lee, K.-H.; Lee, M.H.; Ahn, C.-B. Antioxidant and antihypertensive protein hydrolysates produced from tuna liver by enzymatic hydrolysis. Int. Food Res. J. 2009, 42, 1266–1272. [Google Scholar] [CrossRef]
- Coscueta, E.R.; Amorim, M.M.; Voss, G.B.; Nerli, B.B.; Picó, G.A.; Pintado, M.E. Bioactive properties of peptides obtained from Argentinian defatted soy flour protein by Corolase PP hydrolysis. Food Chem. 2016, 198, 36–44. [Google Scholar] [CrossRef]
- Mora, L.; Aristoy, M.-C.; Toldrá, F. Bioactive peptides in foods. In Encyclopedia of Food and Health; Caballero, B., Finglas, P.M., Toldrá, F., Eds.; Academic Press: Cambridge, MA, USA, 2016; pp. 395–400. [Google Scholar]
- Ktari, N.; Nasri, R.; Mnafgui, K.; Hamden, K.; Belguith, O.; Boudaouara, T.; El Feki, A.; Nasri, M. Antioxidative and ACE inhibitory activities of protein hydrolysates from zebra blenny (Salaria basilisca) in alloxan-induced diabetic rats. Process. Biochem. 2014, 49, 890–897. [Google Scholar] [CrossRef]
- Nasri, R.; Younes, I.; Jridi, M.; Trigui, M.; Bougatef, A.; Nedjar-Arroume, N.; Dhulster, P.; Nasri, M.; Karra-Châabouni, M. ACE inhibitory and antioxidative activities of Goby (Zosterissessor ophiocephalus) fish protein hydrolysates: Effect on meat lipid oxidation. Int. Food Res. J. 2013, 54, 552–561. [Google Scholar] [CrossRef]
- Lacroix, I.M.; Li-Chan, E.C. Inhibition of dipeptidyl peptidase (DPP)-IV and α-glucosidase activities by pepsin-treated whey proteins. J. Agric. Food Chem. 2013, 61, 7500–7506. [Google Scholar] [CrossRef] [PubMed]
- Siow, H.-L.; Lim, T.S.; Gan, C.-Y. Development of a workflow for screening and identification of α-amylase inhibitory peptides from food source using an integrated Bioinformatics-phage display approach: Case study–Cumin seed. Food Chem. 2017, 214, 67–76. [Google Scholar] [CrossRef]
- Khantaphant, S.; Benjakul, S.; Ghomi, M.R. The effects of pretreatments on antioxidative activities of protein hydrolysate from the muscle of brownstripe red snapper (Lutjanus vitta). LWT Food Sci Technol. 2011, 44, 1139–1148. [Google Scholar] [CrossRef]
- Jemil, I.; Abdelhedi, O.; Mora, L.; Nasri, R.; Aristoy, M.-C.; Jridi, M.; Hajji, M.; Toldrá, F.; Nasri, M. Peptidomic analysis of bioactive peptides in zebra blenny (Salaria basilisca) muscle protein hydrolysate exhibiting antimicrobial activity obtained by fermentation with Bacillus mojavensis A21. Process. Biochem. 2016, 51, 2186–2197. [Google Scholar] [CrossRef]
- Lassoued, I.; Mora, L.; Nasri, R.; Aydi, M.; Toldrá, F.; Aristoy, M.-C.; Barkia, A.; Nasri, M. Characterization, antioxidative and ACE inhibitory properties of hydrolysates obtained from thornback ray (Raja clavata) muscle. J. Proteom. 2015, 128, 458–468. [Google Scholar] [CrossRef] [PubMed]
- Chen, N.; Yang, H.; Sun, Y.; Niu, J.; Liu, S. Purification and identification of antioxidant peptides from walnut (Juglans regia L.) protein hydrolysates. Peptides 2012, 38, 344–349. [Google Scholar] [CrossRef] [PubMed]
- Salem, R.B.S.-B.; Ktari, N.; Bkhairia, I.; Nasri, R.; Mora, L.; Kallel, R.; Hamdi, S.; Jamoussi, K.; Boudaouara, T.; El-Feki, A. In vitro and in vivo anti-diabetic and anti-hyperlipidemic effects of protein hydrolysates from Octopus vulgaris in alloxanic rats. Int. Food Res. J. 2018, 106, 952–963. [Google Scholar] [CrossRef]
- Andersen, M.L.; Lauridsen, R.K.; Skibsted, L.H. Optimising the Use of Phenolic Compounds in Foods; Woodhead Publishing Ltd.: Cambridge, UK, 2003. [Google Scholar]
- Moayedi, A.; Mora, L.; Aristoy, M.-C.; Hashemi, M.; Safari, M.; Toldrá, F. ACE-inhibitory and antioxidant activities of peptide fragments obtained from tomato processing by-products fermented using Bacillus subtilis: Effect of amino acid composition and peptides molecular mass distribution. Appl. Biochem. Biotechnol. 2017, 181, 48–64. [Google Scholar] [CrossRef]
- Power, O.; Jakeman, P.; FitzGerald, R. Antioxidative peptides: Enzymatic production, in vitro and in vivo antioxidant activity and potential applications of milk-derived antioxidative peptides. Amino Acids 2013, 44, 797–820. [Google Scholar] [CrossRef]
- Lee, S.-H.; Qian, Z.-J.; Kim, S.-K. A novel angiotensin I converting enzyme inhibitory peptide from tuna frame protein hydrolysate and its antihypertensive effect in spontaneously hypertensive rats. Food Chem. 2010, 118, 96–102. [Google Scholar] [CrossRef]
- Wijesekara, I.; Kim, S.-K. Angiotensin-I-converting enzyme (ACE) inhibitors from marine resources: Prospects in the pharmaceutical industry. Mar. Drugs 2010, 8, 1080–1093. [Google Scholar] [CrossRef]
- Megías, C.; Yust, M.d.M.; Pedroche, J.; Lquari, H.; Girón-Calle, J.; Alaiz, M.; Millán, F.; Vioque, J. Purification of an ACE inhibitory peptide after hydrolysis of sunflower (Helianthus annuus L.) protein isolates. J. Agric. Food Chem. 2004, 52, 1928–1932. [Google Scholar] [CrossRef] [PubMed]
- Marambe, H.K.; Shand, P.J.; Wanasundara, J.P. Release of angiotensin I-converting enzyme inhibitory peptides from flaxseed (Linum usitatissimum L.) protein under simulated gastrointestinal digestion. J. Agric. Food Chem. 2011, 59, 9596–9604. [Google Scholar] [CrossRef]
- Liu, R.; Zheng, W.; Li, J.; Wang, L.; Wu, H.; Wang, X.; Shi, L. Rapid identification of bioactive peptides with antioxidant activity from the enzymatic hydrolysate of Mactra veneriformis by UHPLC–Q-TOF mass spectrometry. Food Chem. 2015, 167, 484–489. [Google Scholar] [CrossRef] [PubMed]
- Torkova, A.; Kononikhin, A.; Bugrova, A.; Khotchenkov, V.; Tsentalovich, M.; Medvedeva, U. Effect of in vitro Gastrointestinal Digestion on Bioactivity of Poultry Protein Hydrolysate. Curr. Res. Nutr. Food Sci. 2016, 4, 77–86. [Google Scholar]
- Mazloomi, S.N.; Mora, L.; Aristoy, M.; Sadeghi Mahoonak, A.; Ghorbani, M.; Houshmand, G.H.; Toldrá, F. Impact of Simulated Gastrointestinal Digestion on the Biological Activity of an Alcalase Hydrolysate of Orange Seed (Siavaraze, Citrus sinensis) by-Products. Foods 2020, 9, 1217. [Google Scholar] [CrossRef] [PubMed]
| Sequence | Amphipathicity | Hydrophilicity | MW | Probable Allergenicity |
|---|---|---|---|---|
| IQLSAEKGNLYQNA | 0.53 | −0.11 | 1548.93 | NO |
| LVDVGNSDNQ | 0.12 | 0.21 | 1060.23 | YES |
| RVESEAGVTEF | 0.57 | 0.54 | 1223.45 | YES |
| YQGSQGGEGGDRS | 0.48 | 0.59 | 1297.45 | NO |
| YQGSQGGEGGD | 0.34 | 0.4 | 1054.16 | YES |
| QKVESEAGVTEF | 0.73 | 0.51 | 1323.59 | NO |
| SAEKGNLYPNAL | 0.41 | −0.02 | 1276.58 | NO |
| VDVGNSQNQLDQY | 0.29 | 0.02 | 1479.72 | NO |
| GIETVGGVMT | 0.13 | −0.39 | 963.27 | YES |
| GIETVGGVMTK | 0.45 | −0.08 | 1091.46 | NO |
| STASDNQNTVTIQV | 0.18 | −0.15 | 1477.75 | YES |
| GVNENEYKPEL | 0.68 | 0.62 | 1291.54 | YES |
| VNENEYKPEL | 0.75 | 0.68 | 1234.47 | YES |
| FSDEGFGPAPK | 0.45 | 0.35 | 1151.38 | NO |
| SFSDEGFGPAPK | 0.41 | 0.34 | 1238.47 | YES |
| FSQPDGPIMGN | 0.11 | −0.17 | 1162.44 | NO |
| VVDSIDQDEL | 0.25 | 0.59 | 1132.33 | NO |
| EGPSLLPDDPEK | 0.52 | 0.97 | 1296.56 | NO |
| HDLEIGPGAPT | 0.25 | 0.09 | 1106.36 | NO |
| EGMGGGGGAHDPF | 0.21 | 0.09 | 1188.43 | YES |
| VDSFHPGEPI | 0.27 | 0 | 1097.33 | NO |
| DFEDSHPDLPS | 0.25 | 0.71 | 1258.4 | NO |
| ISVGDKLPDAT | 0.33 | 0.3 | 1115.4 | YES |
| VFPAGPGGAA | 0 | −0.55 | 843.08 | YES |
| VIDANGNLVPH | 0.13 | −0.38 | 1148.45 | YES |
| DADGNGTIDFPE | 0.11 | 0.58 | 1250.4 | NO |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mazloomi, S.N.; Mahoonak, A.S.; Mora, L.; Ghorbani, M.; Houshmand, G.; Toldrá, F. Pepsin Hydrolysis of Orange By-Products for the Production of Bioactive Peptides with Gastrointestinal Resistant Properties. Foods 2021, 10, 679. https://doi.org/10.3390/foods10030679
Mazloomi SN, Mahoonak AS, Mora L, Ghorbani M, Houshmand G, Toldrá F. Pepsin Hydrolysis of Orange By-Products for the Production of Bioactive Peptides with Gastrointestinal Resistant Properties. Foods. 2021; 10(3):679. https://doi.org/10.3390/foods10030679
Chicago/Turabian StyleMazloomi, Seyadeh Narges, Alireza Sadeghi Mahoonak, Leticia Mora, Mohammad Ghorbani, Gholamreza Houshmand, and Fidel Toldrá. 2021. "Pepsin Hydrolysis of Orange By-Products for the Production of Bioactive Peptides with Gastrointestinal Resistant Properties" Foods 10, no. 3: 679. https://doi.org/10.3390/foods10030679
APA StyleMazloomi, S. N., Mahoonak, A. S., Mora, L., Ghorbani, M., Houshmand, G., & Toldrá, F. (2021). Pepsin Hydrolysis of Orange By-Products for the Production of Bioactive Peptides with Gastrointestinal Resistant Properties. Foods, 10(3), 679. https://doi.org/10.3390/foods10030679








