Structural Conservation and Adaptation of the Bacterial Flagella Motor
Abstract
1. Introduction
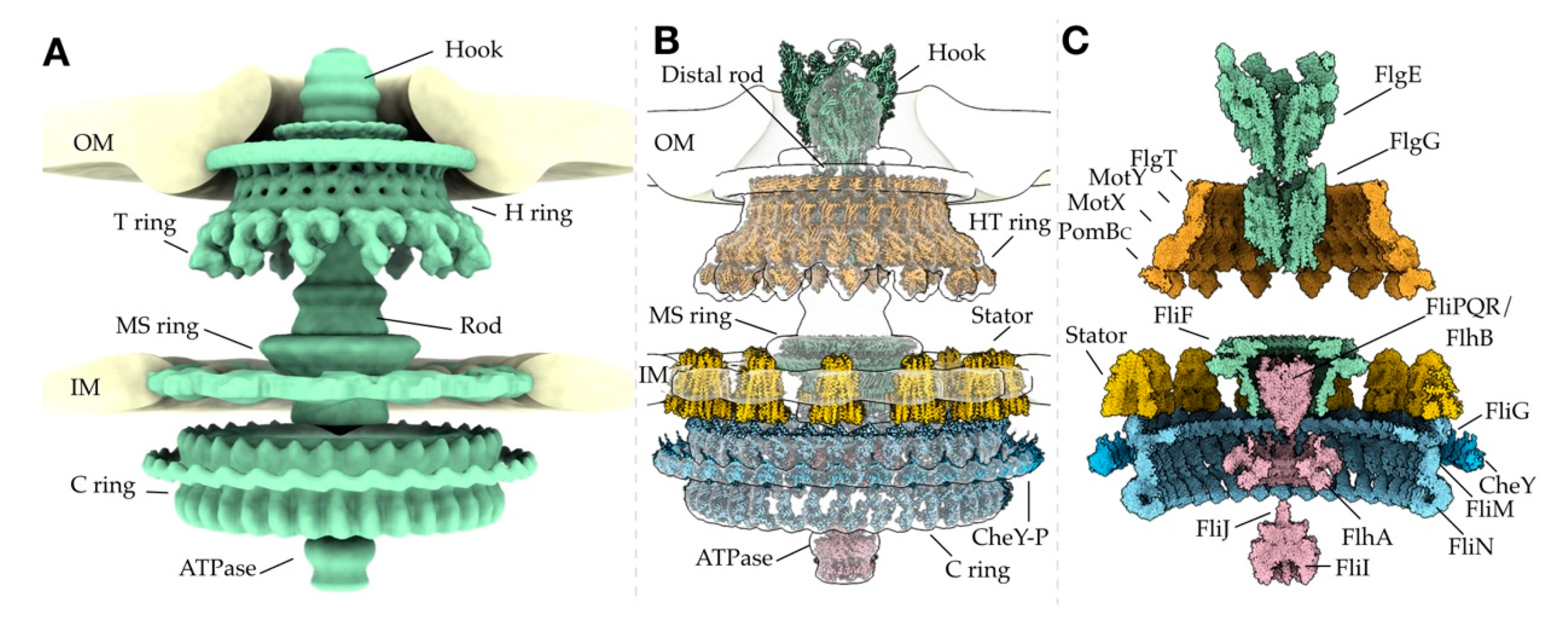
2. The Bacterial Flagellar Structure
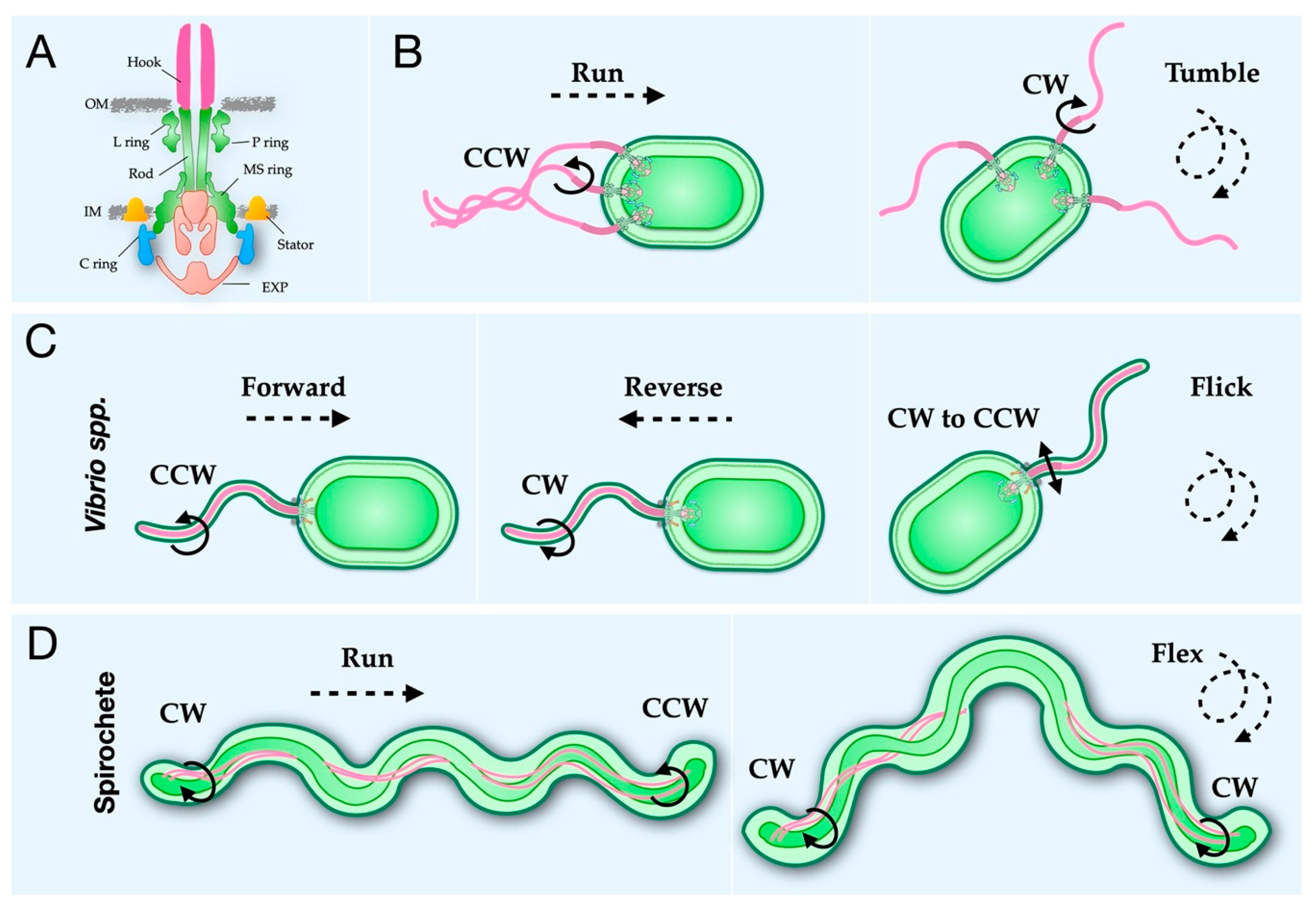
2.1. The Rod, Hook, and Filament Extend from the Cell Body
2.2. The Periplasmic P and L Rings Stabilize the Rod
2.3. The MS Ring is the Base of the Motor
2.4. The C Ring Acts as a Rotor Within the Cytosol
2.5. Torque is Generated by the Stator Through Ion Gradients
2.6. A Conserved Mechanism for Flagellar Rotational Switching
2.7. The Export Apparatus Secretes Flagellar Proteins for Assembly
3. Specific Examples of Adaptation within the Bacterial Flagellum
3.1. Vibrio Flagella Have Additional Rings in the Periplasm for Greater Torque Generation
3.2. The ε-Proteobacteria Flagellum Cage Traps Additional Stators
3.3. The Periplasmic Flagella of Spirochetes Uses a Collar to Stabilize Stators
4. Conclusions and Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Nakamura, S.; Minamino, T. Flagella-Driven Motility of Bacteria. Biomolecules 2019, 9, 279. [Google Scholar] [CrossRef]
- Minamino, T.; Imada, K.; Namba, K. Molecular motors of the bacterial flagella. Curr. Opin. Struct. Biol. 2008, 18, 693–701. [Google Scholar] [CrossRef]
- Pallen, M.J.; Matzke, N.J. From The Origin of Species to the origin of bacterial flagella. Nat. Rev. Genet. 2006, 4, 784–790. [Google Scholar] [CrossRef]
- Terashima, H.; Kawamoto, A.; Morimoto, Y.V.; Imada, K.; Minamino, T. Structural differences in the bacterial flagellar motor among bacterial species. Biophys. Physicobiology 2017, 14, 191–198. [Google Scholar] [CrossRef]
- Terashima, H.; Kojima, S.; Homma, M. Flagellar motility in bacteria structure and function of flagellar motor. Int. Rev. Cell. Mol. Biol. 2008, 270, 39–85. [Google Scholar] [CrossRef]
- Zhao, X.; Norris, S.J.; Liu, J. Molecular Architecture of the Bacterial Flagellar Motor in Cells. Biochemistry 2014, 53, 4323–4333. [Google Scholar] [CrossRef]
- Diepold, A.; Armitage, J.P. Type III secretion systems: The bacterial flagellum and the injectisome. Philos. Trans. R. Soc. B Biol. Sci. 2015, 370, 20150020. [Google Scholar] [CrossRef]
- McCarter, L.L. Dual Flagellar Systems Enable Motility under Different Circumstances. J. Mol. Microbiol. Biotechnol. 2004, 7, 18–29. [Google Scholar] [CrossRef]
- Chang, Y.; Liu, J. Architecture and Assembly of Periplasmic Flagellum. Microbiol. Spectr. 2019, 7, 10. [Google Scholar] [CrossRef]
- Blair, D.F. Flagellar movement driven by proton translocation. FEBS Lett. 2003, 545, 86–95. [Google Scholar] [CrossRef]
- Francis, N.R.; Irikura, V.M.; Yamaguchi, S.; DeRosier, D.J.; Macnab, R.M. Localization of the Salmonella typhimurium flagellar switch protein FliG to the cytoplasmic M-ring face of the basal body. Proc. Natl. Acad. Sci. USA 1992, 89, 6304–6308. [Google Scholar] [CrossRef]
- Francis, N.R.; Sosinsky, G.E.; Thomas, D.; DeRosier, D.J. Isolation, Characterization and Structure of Bacterial Flagellar Motors Containing the Switch Complex. J. Mol. Biol. 1994, 235, 1261–1270. [Google Scholar] [CrossRef]
- Homma, M.; Ohnishi, K.; Iino, T.; Macnab, R.M. Identification of flagellar hook and basal body gene products (FlaFV, FlaFVI, FlaFVII and FlaFVIII) in Salmonella typhimurium. J. Bacteriol. 1987, 169, 3617–3624. [Google Scholar] [CrossRef]
- Sato, K.; Homma, M. Multimeric Structure of PomA, a Component of the Na+-driven Polar Flagellar Motor ofVibrio alginolyticus. J. Biol. Chem. 2000, 275, 20223–20228. [Google Scholar] [CrossRef]
- Sato, K.; Homma, M. Functional Reconstitution of the Na+-driven Polar Flagellar Motor Component ofVibrio alginolyticus. J. Biol. Chem. 2000, 275, 5718–5722. [Google Scholar] [CrossRef]
- Ueno, T.; Oosawa, K.; Aizawa, S.-I. M ring, S ring and proximal rod of the flagellar basal body of Salmonella typhimurium are composed of subunits of a single protein, FliF. J. Mol. Biol. 1992, 227, 672–677. [Google Scholar] [CrossRef]
- Asai, Y.; Kojima, S.; Kato, H.; Nishioka, N.; Kawagishi, I.; Homma, M. Putative channel components for the fast-rotating sodium-driven flagellar motor of a marine bacterium. J. Bacteriol. 1997, 179, 5104–5110. [Google Scholar] [CrossRef]
- Braun, T.F.; Al-Mawsawi, L.Q.; Kojima, A.S.; Blair, D.F. Arrangement of Core Membrane Segments in the MotA/MotB Proton-Channel Complex ofEscherichia coli. Biochemistry 2004, 43, 35–45. [Google Scholar] [CrossRef]
- Kojima, S.; Blair, D.F. The Bacterial Flagellar Motor: Structure and Function of a Complex Molecular Machine. Int. Rev. Cytol. 2004, 233, 93–134. [Google Scholar] [CrossRef]
- Homma, M.; Kutsukake, K.; Hasebe, M.; Iino, T.; Macnab, R.M. FlgB, FlgC, FlgF and FlgG. A family of structurally related proteins in the flagellar basal body of Salmonella typhimurium. J. Mol. Biol. 1990, 211, 465–477. [Google Scholar] [CrossRef]
- Kubori, T.; Shimamoto, N.; Yamaguchi, S.; Namba, K.; Aizawa, S.-I. Morphological pathway of flagellar assembly in Salmonella typhimurium. J. Mol. Biol. 1992, 226, 433–446. [Google Scholar] [CrossRef]
- Minamino, T.; Yamaguchi, S.; Macnab, R.M. Interaction between FliE and FlgB, a Proximal Rod Component of the Flagellar Basal Body ofSalmonella. J. Bacteriol. 2000, 182, 3029–3036. [Google Scholar] [CrossRef]
- Karlinsey, J.; Pease, A.J.; Winkler, M.E.; Bailey, J.L.; Hughes, K.T. The flk gene of Salmonella typhimurium couples flagellar P- and L-ring assembly to flagellar morphogenesis. J. Bacteriol. 1997, 179, 2389–2400. [Google Scholar] [CrossRef][Green Version]
- Fukumura, T.; Makino, F.; Dietsche, T.; Kinoshita, M.; Kato, T.; Wagner, S.; Namba, K.; Imada, K.; Minamino, T. Assembly and stoichiometry of the core structure of the bacterial flagellar type III export gate complex. PLoS Biol. 2017, 15, e2002281. [Google Scholar] [CrossRef]
- Minamino, T.; Macnab, R.M. Components of the Salmonella Flagellar Export Apparatus and Classification of Export Substrates. J. Bacteriol. 1999, 181, 1388–1394. [Google Scholar] [CrossRef]
- Minamino, T.; Macnab, R.M. Interactions among components of the Salmonella flagellar export apparatus and its substrates. Mol. Microbiol. 2000, 35, 1052–1064. [Google Scholar] [CrossRef]
- Minamino, T. Protein export through the bacterial flagellar type III export pathway. Biochim. et Biophys. Acta (BBA)-Bioenerg. 2014, 1843, 1642–1648. [Google Scholar] [CrossRef]
- Lucic, V.; Rigort, A.; Baumeister, W. Cryo-electron tomography: The challenge of doing structural biology in situ. J. Cell. Biol. 2013, 202, 407–419. [Google Scholar] [CrossRef]
- Kim, M.I.; Lee, C.; Park, J.; Jeon, B.-Y.; Hong, M. Crystal structure of Bacillus cereus flagellin and structure-guided fusion-protein designs. Sci. Rep. 2018, 8, 5814. [Google Scholar] [CrossRef]
- Samatey, F.A.; Imada, K.; Nagashima, S.; Vonderviszt, F.; Kumasaka, T.; Yamamoto, M.; Namba, K. Structure of the bacterial flagellar protofilament and implications for a switch for supercoiling. Nat. Cell Biol. 2001, 410, 331–337. [Google Scholar] [CrossRef]
- Maruyama, Y.; Momma, M.; Mikami, B.; Hashimoto, W.; Murata, K. Crystal Structure of a Novel Bacterial Cell-Surface Flagellin Binding to a Polysaccharide. Biochemistry 2008, 47, 1393–1402. [Google Scholar] [CrossRef]
- Nithichanon, A.; Rinchai, D.; Gori, A.; Lassaux, P.; Peri, C.; Conchillio-Solé, O.; Ferrer-Navarro, M.; Gourlay, L.J.; Nardini, M.; Vila, J.; et al. Sequence- and Structure-Based Immunoreactive Epitope Discovery for Burkholderia pseudomallei Flagellin. PLoS Neglected Trop. Dis. 2015, 9, e0003917. [Google Scholar] [CrossRef]
- Song, W.S.; Yoon, S.-I. Crystal structure of FliC flagellin from Pseudomonas aeruginosa and its implication in TLR5 binding and formation of the flagellar filament. Biochem. Biophys. Res. Commun. 2014, 444, 109–115. [Google Scholar] [CrossRef]
- Evdokimov, A.G.; Phan, J.; Tropea, J.E.; Routzahn, K.M.; Peters, H.K.; Pokross, M.; Waugh, D.S. Similar modes of polypeptide recognition by export chaperones in flagellar biosynthesis and type III secretion. Nat. Struct. Mol. Biol. 2003, 10, 789–793. [Google Scholar] [CrossRef]
- Lee, C.; Kim, M.I.; Park, J.; Jeon, B.-Y.; Yoon, S.-I.; Hong, M. Crystal structure of the flagellar chaperone FliS from Bacillus cereus and an invariant proline critical for FliS dimerization and flagellin recognition. Biochem. Biophys. Res. Commun. 2017, 487, 381–387. [Google Scholar] [CrossRef]
- Lam, W.W.L.; Woo, E.J.; Kotaka, M.; Tam, W.K.; Leung, Y.C.; Ling, T.K.W.; Au, S.W.N. Molecular interaction of flagellar export chaperone FliS and cochaperone HP1076 in Helicobacter pylori. FASEB J. 2010, 24, 4020–4032. [Google Scholar] [CrossRef]
- Horstmann, J.A.; Lunelli, M.; Cazzola, H.; Heidemann, J.; Kühne, C.; Steffen, P.; Szefs, S.; Rossi, C.; Lokareddy, R.K.; Wang, C.; et al. Methylation of Salmonella Typhimurium flagella promotes bacterial adhesion and host cell invasion. Nat. Commun. 2020, 11, 1–11. [Google Scholar] [CrossRef]
- Gibson, K.H.; Trajtenberg, F.; Wunder, E.A.; Brady, M.R.; San Martin, F.; Mechaly, A.; Shang, Z.; Liu, J.; Picardeau, M.; Ko, A.; et al. An asymmetric sheath controls flagellar supercoiling and motility in the leptospira spirochete. eLife 2020, 9. [Google Scholar] [CrossRef]
- Altegoer, F.; Mukherjee, S.; Steinchen, W.; Bedrunka, P.; Linne, U.; Kearns, D.B.; Bange, G. FliS/flagellin/FliW heterotrimer couples type III secretion and flagellin homeostasis. Sci. Rep. 2018, 8, 11552. [Google Scholar] [CrossRef]
- Skorupka, K.; Han, S.K.; Nam, H.-J.; Kim, S.; Faham, S. Protein design by fusion: Implications for protein structure prediction and evolution. Acta Crystallogr. Sect. D Biol. Crystallogr. 2013, 69, 2451–2460. [Google Scholar] [CrossRef]
- Kekez, I.; Cendron, L.; Stojanović, M.; Zanotti, G.; Matković-Čalogović, D. Structure and Stability of FlgD from the Pathogenic 26695 Strain of Helicobacter pylori. Croat. Chem. Acta 2016, 89, 1–7. [Google Scholar] [CrossRef]
- Pulić, I.; Cendron, L.; Salamina, M.; De Laureto, P.P.; Matković-Čalogović, D.; Zanotti, G. Crystal structure of truncated FlgD from the human pathogen Helicobacter pylori. J. Struct. Biol. 2016, 194, 147–155. [Google Scholar] [CrossRef]
- Yoon, Y.-H.; Barker, C.S.; Bulieris, P.V.; Matsunami, H.; Samatey, F.A. Structural insights into bacterial flagellar hooks similarities and specificities. Sci. Rep. 2016, 6, 35552. [Google Scholar] [CrossRef]
- LoConte, V.; Kekez, I.; Matković-Čalogović, D.; Zanotti, G. Structural characterization of FlgE2 protein from Helicobacter pylori hook. FEBS J. 2017, 284, 4328–4342. [Google Scholar] [CrossRef]
- Samatey, F.A.; Matsunami, H.; Imada, K.; Nagashima, S.; Shaikh, T.R.; Thomas, D.R.; Chen, J.Z.; DeRosier, D.J.; Kitao, A.; Namba, K. Structure of the bacterial flagellar hook and implication for the molecular universal joint mechanism. Nat. Cell Biol. 2004, 431, 1062–1068. [Google Scholar] [CrossRef]
- Lynch, M.J.; Miller, M.; James, M.; Zhang, S.; Zhang, K.; Li, C.; Charon, N.W.; Pollack, L. Structure and chemistry of lysinoalanine crosslinking in the spirochaete flagella hook. Nat. Chem. Biol. 2019, 15, 959–965. [Google Scholar] [CrossRef]
- Bulieris, P.V.; Shaikh, N.H.; Freddolino, P.L.; Samatey, F.A. Structure of FlgK reveals the divergence of the bacterial Hook-Filament Junction of Campylobacter. Sci. Rep. 2017, 7, 15743. [Google Scholar] [CrossRef]
- Hong, H.J.; Kim, T.H.; Song, W.S.; Ko, H.-J.; Lee, G.-S.; Kang, S.G.; Kim, P.-H.; Yoon, S.-I. Crystal structure of FlgL and its implications for flagellar assembly. Sci. Rep. 2018, 8, 1–11. [Google Scholar] [CrossRef]
- Postel, S.; Deredge, D.; Bonsor, D.A.; Yu, X.; Diederichs, K.; Helmsing, S.; Vromen, A.; Friedler, A.; Hust, M.; Egelman, E.H.; et al. Bacterial flagellar capping proteins adopt diverse oligomeric states. eLife 2016, 5, e18857. [Google Scholar] [CrossRef]
- Cho, S.Y.; Song, W.S.; Oh, H.-B.; Kim, H.-U.; Jung, H.S.; Yoon, S.-I. Structural analysis of the flagellar capping protein FliD from Helicobacter pylori. Biochem. Biophys. Res. Commun. 2019, 514, 98–104. [Google Scholar] [CrossRef]
- Saijo-Hamano, Y.; Matsunami, H.; Namba, K.; Imada, K. Architecture of the Bacterial Flagellar Distal Rod and Hook of Salmonella. Biochemistry 2019, 9, 260. [Google Scholar] [CrossRef]
- Zaloba, P.; Bailey-Elkin, B.A.; Derksen, M.; Mark, B.L. Structural and Biochemical Insights into the Peptidoglycan Hydrolase Domain of FlgJ from Salmonella typhimurium. PLoS ONE 2016, 11, e0149204. [Google Scholar] [CrossRef]
- Matsunami, H.; Yoon, Y.-H.; Meshcheryakov, V.A.; Namba, K.; Samatey, F.A. Structural flexibility of the periplasmic protein, FlgA, regulates flagellar P-ring assembly in Salmonella enterica. Sci. Rep. 2016, 6, 27399. [Google Scholar] [CrossRef]
- Xue, C.; Lam, K.H.; Zhang, H.; Sun, K.; Lee, S.H.; Chen, X.; Au, S.W.N. Crystal structure of the FliF-FliG complex from Helicobacter pylori yields insight into the assembly of the motor MS-C ring in the bacterial flagellum. J. Biol. Chem. 2018, 293, 2066–2078. [Google Scholar] [CrossRef]
- Lynch, M.J.; Levenson, R.; Kim, E.A.; Sircar, R.; Blair, D.F.; Dahlquist, F.W.; Crane, B.R. Co-Folding of a FliF-FliG Split Domain Forms the Basis of the MS:C Ring Interface within the Bacterial Flagellar Motor. Struct. 2017, 25, 317–328. [Google Scholar] [CrossRef]
- Lee, L.K.; Ginsburg, M.A.; Crovace, C.; Donohoe, M.; Stock, D. Structure of the torque ring of the flagellar motor and the molecular basis for rotational switching. Nat. Cell Biol. 2010, 466, 996–1000. [Google Scholar] [CrossRef]
- Qin, Z.; Lin, W.-T.; Zhu, S.; Franco, A.T.; Liu, J. Imaging the Motility and Chemotaxis Machineries in Helicobacter pylori by Cryo-Electron Tomography. J. Bacteriol. 2016, 199, e00695-16. [Google Scholar] [CrossRef]
- Hou, Y.; Sun, W.; Zhang, C.; Wang, T.; Guo, X.; Wu, L.; Qin, L.; Liu, T. Meta-analysis of the correlation between Helicobacter pylori infection and autoimmune thyroid diseases. Oncotarget 2017, 8, 115691–115700. [Google Scholar] [CrossRef]
- Li, T.H.; Qin, Y.; Sham, P.C.; Lau, K.; Chu, K.-M.; Leung, W.K. Alterations in Gastric Microbiota After H. Pylori Eradication and in Different Histological Stages of Gastric Carcinogenesis. Sci. Rep. 2017, 7, 44935. [Google Scholar] [CrossRef]
- Liu, A.Q.; Xie, Z.; Chen, X.N.; Feng, J.; Chen, J.W.; Qin, F.J.; Ge, L.Y. Fas-associated factor 1 inhibits tumor growth by suppressing Helicobacter pylori-induced activation of NF-kappaB signaling in human gastric carcinoma. Oncotarget 2017, 8, 7999–8009. [Google Scholar] [CrossRef]
- Lam, K.-H.; Lam, W.W.L.; Wong, J.Y.-K.; Chan, L.-C.; Kotaka, M.; Ling, T.K.-W.; Jin, D.-Y.; Ottemann, K.M.; Au, S.W.N. Structural basis of FliG-FliM interaction inHelicobacter pylori. Mol. Microbiol. 2013, 88, 798–812. [Google Scholar] [CrossRef]
- Park, S.-Y.; Lowder, B.; Bilwes, A.M.; Blair, D.F.; Pollack, L. Structure of FliM provides insight into assembly of the switch complex in the bacterial flagella motor. Proc. Natl. Acad. Sci. USA 2006, 103, 11886–11891. [Google Scholar] [CrossRef]
- Lam, K.-H.; Xue, C.; Sun, K.; Zhang, H.; Lam, W.W.L.; Zhu, Z.; Ng, J.T.Y.; Sause, W.E.; Lertsethtakarn, P.; Lau, K.; et al. Three SpoA-domain proteins interact in the creation of the flagellar type III secretion system in Helicobacter pylori. J. Biol. Chem. 2018, 293, 13961–13973. [Google Scholar] [CrossRef]
- Brown, P.N.; Mathews, M.A.A.; Joss, L.A.; Hill, C.P.; Blair, D.F. Crystal Structure of the Flagellar Rotor Protein FliN from Thermotoga maritima. J. Bacteriol. 2005, 187, 2890–2902. [Google Scholar] [CrossRef]
- Sircar, R.; Greenswag, A.R.; Bilwes, A.M.; Gonzalez-Bonet, G.; Crane, B.R. Structure and Activity of the Flagellar Rotor Protein FliY. J. Biol. Chem. 2013, 288, 13493–13502. [Google Scholar] [CrossRef]
- Sircar, R.; Borbat, P.P.; Lynch, M.J.; Bhatnagar, J.; Beyersdorf, M.S.; Halkides, C.J.; Freed, J.H.; Crane, B.R. Assembly States of FliM and FliG within the Flagellar Switch Complex. J. Mol. Biol. 2015, 427, 867–886. [Google Scholar] [CrossRef]
- Vartanian, A.S.; Paz, A.; Fortgang, E.A.; Abramson, J.; Dahlquist, F.W. Structure of Flagellar Motor Proteins in Complex Allows for Insights into Motor Structure and Switching. J. Biol. Chem. 2012, 287, 35779–35783. [Google Scholar] [CrossRef]
- Paul, K.; Gonzalez-Bonet, G.; Bilwes, A.M.; Crane, B.R.; Blair, D.F. Architecture of the flagellar rotor. EMBO J. 2011, 30, 2962–2971. [Google Scholar] [CrossRef]
- Notti, R.Q.; Bhattacharya, S.; Lilic, M.; Stebbins, C.E. A common assembly module in injectisome and flagellar type III secretion sorting platforms. Nat. Commun. 2015, 6, 7125. [Google Scholar] [CrossRef]
- Couturier, C.; Silve, S.; Morales, R.; Pessegue, B.; Llopart, S.; Nair, A.; Bauer, A.; Scheiper, B.; Pöverlein, C.; Ganzhorn, A.; et al. Nanomolar inhibitors of Mycobacterium tuberculosis glutamine synthetase 1: Synthesis, biological evaluation and X-ray crystallographic studies. Bioorganic Med. Chem. Lett. 2015, 25, 1455–1459. [Google Scholar] [CrossRef]
- Zhang, H.; Lam, K.-H.; Lam, W.W.L.; Wong, S.Y.Y.; Chan, V.S.F.; Au, S.W.N. A putative spermidine synthase interacts with flagellar switch protein FliM and regulates motility in Helicobacter pylori. Mol. Microbiol. 2017, 106, 690–703. [Google Scholar] [CrossRef]
- Lee, S.Y.; Cho, H.; Pelton, J.G.; Yan, D.; Henderson, R.K.; King, D.S.; Huang, L.-S.; Kustu, S.; Berry, E.A.; Wemmer, D.E. Crystal structure of an activated response regulator bound to its target. Nat. Genet. 2001, 8, 52–56. [Google Scholar] [CrossRef]
- Dyer, C.M.; Quillin, M.L.; Campos, A.; Lu, J.; McEvoy, M.M.; Hausrath, A.C.; Westbrook, E.M.; Matsumura, P.; Matthews, B.W.; Dahlquist, F.W. Structure of the Constitutively Active Double Mutant CheYD13K Y106W Alone and in Complex with a FliM Peptide. J. Mol. Biol. 2004, 342, 1325–1335. [Google Scholar] [CrossRef]
- Dyer, C.M.; Dahlquist, F.W. Switched or Not?: The Structure of Unphosphorylated CheY Bound to the N Terminus of FliM. J. Bacteriol. 2006, 188, 7354–7363. [Google Scholar] [CrossRef]
- Ahn, D.-R.; Song, H.; Kim, J.; Lee, S.; Park, S. The crystal structure of an activated Thermotoga maritima CheY with N-terminal region of FliM. Int. J. Biol. Macromol. 2013, 54, 76–83. [Google Scholar] [CrossRef]
- Biswas, M.; Dey, S.; Khamrui, S.; Sen, U.; Dasgupta, J. Conformational Barrier of CheY3 and Inability of CheY4 to Bind FliM Control the Flagellar Motor Action in Vibrio cholerae. PLoS ONE 2013, 8, e73923. [Google Scholar] [CrossRef]
- Schuhmacher, J.S.; Rossmann, F.M.; Dempwolff, F.; Knauer, C.; Altegoer, F.; Steinchen, W.; Dörrich, A.K.; Klingl, A.; Stephan, M.; Linne, U.; et al. MinD-like ATPase FlhG effects location and number of bacterial flagella during C-ring assembly. Proc. Natl. Acad. Sci. USA 2015, 112, 3092–3097. [Google Scholar] [CrossRef]
- Kojima, S.; Takao, M.; Almira, G.; Kawahara, I.; Sakuma, M.; Homma, M.; Kojima, C.; Imada, K. The Helix Rearrangement in the Periplasmic Domain of the Flagellar Stator B Subunit Activates Peptidoglycan Binding and Ion Influx. Struct. 2018, 26, 590–598.e5. [Google Scholar] [CrossRef]
- Kojima, S.; Imada, K.; Sakuma, M.; Sudo, Y.; Kojima, C.; Minamino, T.; Homma, M.; Namba, K. Stator assembly and activation mechanism of the flagellar motor by the periplasmic region of MotB. Mol. Microbiol. 2009, 73, 710–718. [Google Scholar] [CrossRef]
- Zhu, S.; Takao, M.; Li, N.; Sakuma, M.; Nishino, Y.; Homma, M.; Kojima, S.; Imada, K. Conformational change in the periplamic region of the flagellar stator coupled with the assembly around the rotor. Proc. Natl. Acad. Sci. USA 2014, 111, 13523–13528. [Google Scholar] [CrossRef]
- Kojima, S.; Shinohara, A.; Terashima, H.; Yakushi, T.; Sakuma, M.; Homma, M.; Namba, K.; Imada, K. Insights into the stator assembly of the Vibrio flagellar motor from the crystal structure of MotY. Proc. Natl. Acad. Sci. USA 2008, 105, 7696–7701. [Google Scholar] [CrossRef]
- Takekawa, N.; Isumi, M.; Terashima, H.; Zhu, S.; Nishino, Y.; Sakuma, M.; Kojima, S.; Homma, M.; Imada, K. Structure of Vibrio FliL, a New Stomatin-like Protein That Assists the Bacterial Flagellar Motor Function. mBio 2019, 10, e00292-19. [Google Scholar] [CrossRef]
- Terashima, H.; Li, N.; Sakuma, M.; Koike, M.; Kojima, S.; Homma, M.; Imada, K. Insight into the assembly mechanism in the supramolecular rings of the sodium-driven Vibrio flagellar motor from the structure of FlgT. Proc. Natl. Acad. Sci. USA 2013, 110, 6133–6138. [Google Scholar] [CrossRef]
- Bange, G.; Kümmerer, N.; Engel, C.; Bozkurt, G.; Wild, K.; Sinning, I. FlhA provides the adaptor for coordinated delivery of late flagella building blocks to the type III secretion system. Proc. Natl. Acad. Sci. USA 2010, 107, 11295–11300. [Google Scholar] [CrossRef]
- Xing, Q.; Shi, K.; Portaliou, A.; Rossi, P.; Economou, A.; Kalodimos, C.G. Structures of chaperone-substrate complexes docked onto the export gate in a type III secretion system. Nat. Commun. 2018, 9, 1–9. [Google Scholar] [CrossRef]
- Inoue, Y.; Ogawa, Y.; Kinoshita, M.; Terahara, N.; Shimada, M.; Kodera, N.; Ando, T.; Namba, K.; Kitao, A.; Imada, K.; et al. Structural Insights into the Substrate Specificity Switch Mechanism of the Type III Protein Export Apparatus. Struct. 2019, 27, 965–976.e6. [Google Scholar] [CrossRef]
- Meshcheryakov, V.A.; Kitao, A.; Matsunami, H.; Samatey, F.A. Inhibition of a type III secretion system by the deletion of a short loop in one of its membrane proteins. Acta Crystallogr. Sect. D Biol. Crystallogr. 2013, 69, 812–820. [Google Scholar] [CrossRef]
- Bange, G.; Petzold, G.; Wild, K.; Parlitz, R.O.; Sinning, I. The crystal structure of the third signal-recognition particle GTPase FlhF reveals a homodimer with bound GTP. Proc. Natl. Acad. Sci. USA 2007, 104, 13621–13625. [Google Scholar] [CrossRef]
- Imada, K.; Minamino, T.; Tahara, A.; Namba, K. Structural similarity between the flagellar type III ATPase FliI and F1-ATPase subunits. Proc. Natl. Acad. Sci. USA 2007, 104, 485–490. [Google Scholar] [CrossRef]
- Ibuki, T.; Imada, K.; Minamino, T.; Kato, T.; Miyata, T.; Namba, K. Common architecture of the flagellar type III protein export apparatus and F- and V-type ATPases. Nat. Struct. Mol. Biol. 2011, 18, 277–282. [Google Scholar] [CrossRef]
- Kinoshita, M.; Nakanishi, Y.; Furukawa, Y.; Namba, K.; Imada, K.; Minamino, T. Rearrangements of α-helical structures of FlgN chaperone control the binding affinity for its cognate substrates during flagellar type III export. Mol. Microbiol. 2016, 101, 656–670. [Google Scholar] [CrossRef]
- Imada, K.; Minamino, T.; Uchida, Y.; Kinoshita, M.; Namba, K. Insight into the flagella type III export revealed by the complex structure of the type III ATPase and its regulator. Proc. Natl. Acad. Sci. USA 2016, 113, 3633–3638. [Google Scholar] [CrossRef]
- Galkin, V.E.; Yu, X.; Bielnicki, J.; Heuser, J.; Ewing, C.P.; Guerry, P.; Egelman, E.H. Divergence of Quaternary Structures Among Bacterial Flagellar Filaments. Science 2008, 320, 382–385. [Google Scholar] [CrossRef]
- Yonekura, K.; Maki-Yonekura, S.; Namba, K. Complete atomic model of the bacterial flagellar filament by electron cryomicroscopy. Nat. Cell Biol. 2003, 424, 643–650. [Google Scholar] [CrossRef]
- Maki-Yonekura, S.; Yonekura, K.; Namba, K. Conformational change of flagellin for polymorphic supercoiling of the flagellar filament. Nat. Struct. Mol. Biol. 2010, 17, 417–422. [Google Scholar] [CrossRef]
- Wang, F.; Burrage, A.M.; Postel, S.; Clark, R.E.; Orlova, A.; Sundberg, E.J.; Kearns, D.B.; Egelman, E.H. A structural model of flagellar filament switching across multiple bacterial species. Nat. Commun. 2017, 8, 960. [Google Scholar] [CrossRef]
- Yamaguchi, T.; Toma, S.; Terahara, N.; Miyata, T.; Ashihara, M.; Minamino, T.; Namba, K.; Kato, T. Structural and Functional Comparison of Salmonella Flagellar Filaments Composed of FljB and FliC. Biochemistry 2020, 10, 246. [Google Scholar] [CrossRef]
- Blum, T.B.; Filippidou, S.; Fatton, M.; Junier, P.; Abrahams, J.P. The wild-type flagellar filament of the Firmicute Kurthia at 2.8 Å resolution in vivo. Sci. Rep. 2019, 9, 14948. [Google Scholar] [CrossRef]
- Matsunami, H.; Barker, C.S.; Yoon, Y.-H.; Wolf, M.; Samatey, F.A. Complete structure of the bacterial flagellar hook reveals extensive set of stabilizing interactions. Nat. Commun. 2016, 7, 13425. [Google Scholar] [CrossRef]
- Shaikh, T.R.; Thomas, D.R.; Chen, J.Z.; Samatey, F.A.; Matsunami, H.; Imada, K.; Namba, K.; DeRosier, D.J. A partial atomic structure for the flagellar hook of Salmonella typhimurium. Proc. Natl. Acad. Sci. USA 2005, 102, 1023–1028. [Google Scholar] [CrossRef]
- Horváth, P.; Kato, T.; Miyata, T.; Namba, K. Structure of Salmonella Flagellar Hook Reveals Intermolecular Domain Interactions for the Universal Joint Function. Biochemistry 2019, 9, 462. [Google Scholar] [CrossRef]
- Fujii, T.; Kato, T.; Namba, K. Specific Arrangement of α-Helical Coiled Coils in the Core Domain of the Bacterial Flagellar Hook for the Universal Joint Function. Structure 2009, 17, 1485–1493. [Google Scholar] [CrossRef]
- Shibata, S.; Matsunami, H.; Aizawa, S.-I.; Wolf, M. Torque transmission mechanism of the curved bacterial flagellar hook revealed by cryo-EM. Nat. Struct. Mol. Biol. 2019, 26, 941–945. [Google Scholar] [CrossRef]
- Kato, T.; Makino, F.; Miyata, T.; Horváth, P.; Namba, K. Structure of the native supercoiled flagellar hook as a universal joint. Nat. Commun. 2019, 10, 5295–5298. [Google Scholar] [CrossRef]
- Maki-Yonekura, S.; Yonekura, K.; Namba, K. Domain movements of HAP2 in the cap-filament complex formation and growth process of the bacterial flagellum. Proc. Natl. Acad. Sci. USA 2003, 100, 15528–15533. [Google Scholar] [CrossRef] [PubMed]
- Thomas, D.R.; Francis, N.R.; Xu, C.; DeRosier, D.J. The Three-Dimensional Structure of the Flagellar Rotor from a Clockwise-Locked Mutant of Salmonella enterica Serovar Typhimurium. J. Bacteriol. 2006, 188, 7039–7048. [Google Scholar] [CrossRef]
- Johnson, S.; Kuhlen, L.; Deme, J.C.; Abrusci, P.; Lea, S.M. The Structure of an Injectisome Export Gate Demonstrates Conservation of Architecture in the Core Export Gate between Flagellar and Virulence Type III Secretion Systems. mBio 2019, 10, e00818-19. [Google Scholar] [CrossRef]
- Suzuki, H.; Yonekura, K.; Namba, K. Structure of the Rotor of the Bacterial Flagellar Motor Revealed by Electron Cryomicroscopy and Single-particle Image Analysis. J. Mol. Biol. 2004, 337, 105–113. [Google Scholar] [CrossRef] [PubMed]
- Takekawa, N.; Terahara, N.; Kato, T.; Gohara, M.; Mayanagi, K.; Hijikata, A.; Onoue, Y.; Kojima, S.; Shirai, T.; Namba, K.; et al. The tetrameric MotA complex as the core of the flagellar motor stator from hyperthermophilic bacterium. Sci. Rep. 2016, 6, 31526. [Google Scholar] [CrossRef]
- Santiveri, M.; Roa-Eguiara, A.; Kühne, C.; Wadhwa, N.; Berg, H.C.; Erhardt, M.; Taylor, N.M.I. Structure and Function of Stator Units of the Bacterial Flagellar Motor. SSRN Electron. J. 2020, 183, 244–257. [Google Scholar] [CrossRef]
- Deme, J.C.; Johnson, S.; Vickery, O.; Muellbauer, A.; Monkhouse, H.; Griffiths, T.; James, R.H.; Berks, B.C.; Coulton, J.W.; Stansfeld, P.J.; et al. Structures of the stator complex that drives rotation of the bacterial flagellum. Nat. Microbiol. 2020, 1–12. [Google Scholar] [CrossRef]
- Kuhlen, L.; Johnson, S.; Zeitler, A.; Bäurle, S.; Deme, J.C.; Caesar, J.J.E.; Debo, R.; Fisher, J.; Wagner, S.; Lea, S.M. The substrate specificity switch FlhB assembles onto the export gate to regulate type three secretion. Nat. Commun. 2020, 11, 1296. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Beeby, M.; Murphy, G.E.; Leadbetter, J.R.; Hendrixson, D.R.; Briegel, A.; Li, Z.; Shi, J.; Tocheva, I.E.; Müller, A.; et al. Structural diversity of bacterial flagellar motors. EMBO J. 2011, 30, 2972–2981. [Google Scholar] [CrossRef] [PubMed]
- Chaban, B.; Coleman, I.; Beeby, M. Evolution of higher torque in Campylobacter-type bacterial flagellar motors. Sci. Rep. 2018, 8, 1–11. [Google Scholar] [CrossRef]
- Zhang, K.; Qin, Z.; Chang, Y.; Liu, J.; Malkowski, M.G.; Shipa, S.; Li, L.; Qiu, W.; Zhang, J.; Li, C. Analysis of a flagellar filament cap mutant reveals that HtrA serine protease degrades unfolded flagellin protein in the periplasm of Borrelia burgdorferi. Mol. Microbiol. 2019, 111, 1652–1670. [Google Scholar] [CrossRef]
- Chang, Y.; Moon, K.H.; Zhao, X.; Norris, S.J.; Motaleb, A.; Liu, J. Structural insights into flagellar stator–rotor interactions. eLife 2019, 8. [Google Scholar] [CrossRef] [PubMed]
- Kudryashev, M.; Cyrklaff, M.; Wallich, R.; Baumeister, W.; Frischknecht, F. Distinct in situ structures of the Borrelia flagellar motor. J. Struct. Biol. 2010, 169, 54–61. [Google Scholar] [CrossRef]
- Qin, Z.; Tu, J.; Lin, T.; Norris, S.J.; Li, C.; Motaleb, A.; Liu, J. Cryo-electron tomography of periplasmic flagella in Borrelia burgdorferi reveals a distinct cytoplasmic ATPase complex. PLoS Biol. 2018, 16, e3000050. [Google Scholar] [CrossRef]
- Zhao, X.; Zhang, K.; Boquoi, T.; Hu, B.; Motaleb, A.; Miller, K.A.; James, M.E.; Charon, N.W.; Manson, M.D.; Norris, S.J.; et al. Cryoelectron tomography reveals the sequential assembly of bacterial flagella in Borrelia burgdorferi. Proc. Natl. Acad. Sci. USA 2013, 110, 14390–14395. [Google Scholar] [CrossRef]
- Chang, Y.; Zhang, K.; Carroll, B.; Zhao, X.; Charon, N.W.; Norris, S.J.; Motaleb, A.; Li, C.; Liu, J. Molecular mechanism for rotational switching of the bacterial flagellar motor. Nat. Struct. Mol. Biol. 2020, 1–7. [Google Scholar] [CrossRef]
- Beeby, M.; Ribardo, D.A.; Brennan, C.A.; Ruby, E.G.; Jensen, G.J.; Hendrixson, D.R. Diverse high-torque bacterial flagellar motors assemble wider stator rings using a conserved protein scaffold. Proc. Natl. Acad. Sci. USA 2016, 113, E1917–E1926. [Google Scholar] [CrossRef] [PubMed]
- Henderson, L.D.; Matthews-Palmer, T.R.S.; Gulbronson, C.J.; Ribardo, D.A.; Beeby, M.; Hendrixson, D.R. Diversification of Campylobacter jejuni Flagellar C-Ring Composition Impacts Its Structure and Function in Motility, Flagellar Assembly, and Cellular Processes. mBio 2020, 11. [Google Scholar] [CrossRef]
- Rossmann, F.M.; Hug, I.; Sangermani, M.; Jenal, U.; Beeby, M. In situ structure of the Caulobacter crescentus flagellar motor and visualization of binding of a CheY-homolog. Mol. Microbiol. 2020, 114, 443–453. [Google Scholar] [CrossRef]
- Kaplan, M.; Ghosal, D.; Subramanian, P.; Oikonomou, C.M.; Kjaer, A.; Pirbadian, S.; Ortega, D.R.; Briegel, A.; El-Naggar, M.Y.; Jensen, G.J. The presence and absence of periplasmic rings in bacterial flagellar motors correlates with stator type. eLife 2019, 8. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, J.L.; Gao, F.Z.; Rossmann, F.M.; Nans, A.; Brenzinger, S.; Hosseini, R.; Wilson, A.; Briegel, A.; Thormann, K.M.; Rosenthal, P.B.; et al. gamma-proteobacteria eject their polar flagella under nutrient depletion, retaining flagellar motor relic structures. PLoS Biol. 2019, 17, e3000165. [Google Scholar] [CrossRef] [PubMed]
- Kawamoto, A.; Morimoto, Y.V.; Miyata, T.; Minamino, T.; Hughes, K.T.; Kato, T.; Namba, K. Common and distinct structural features of Salmonella injectisome and flagellar basal body. Sci. Rep. 2013, 3, 3369. [Google Scholar] [CrossRef]
- Murphy, G.E.; Leadbetter, J.R.; Jensen, G.J. In situ structure of the complete Treponema primitia flagellar motor. Nat. Cell Biol. 2006, 442, 1062–1064. [Google Scholar] [CrossRef]
- Carroll, B.; Nishikino, T.; Guo, W.; Zhu, S.; Kojima, S.; Homma, M.; Liu, J. The flagellar motor of Vibrio alginolyticus undergoes major structural remodeling during rotational switching. eLife 2020, 9. [Google Scholar] [CrossRef] [PubMed]
- Imada, K. Bacterial flagellar axial structure and its construction. Biophys. Rev. 2017, 10, 559–570. [Google Scholar] [CrossRef]
- Turner, L.; Ryu, W.S.; Berg, H.C. Real-Time Imaging of Fluorescent Flagellar Filaments. J. Bacteriol. 2000, 182, 2793–2801. [Google Scholar] [CrossRef]
- Macnab, R.M.; Ornston, M.K. Normal-to-curly flagellar transitions and their role in bacterial tumbling. Stabilization of an alternative quaternary structure by mechanical force. J. Mol. Biol. 1977, 112, 1–30. [Google Scholar] [CrossRef]
- Macnab, R.M.; Koshland, D.E. The Gradient-Sensing Mechanism in Bacterial Chemotaxis. Proc. Natl. Acad. Sci. USA 1972, 69, 2509–2512. [Google Scholar] [CrossRef]
- Berg, H.C.; Anderson, R.A. Bacteria Swim by Rotating their Flagellar Filaments. Nat. Cell Biol. 1973, 245, 380–382. [Google Scholar] [CrossRef] [PubMed]
- Calladine, C.R. Construction of bacterial flagella. Nat. Cell Biol. 1975, 255, 121–124. [Google Scholar] [CrossRef] [PubMed]
- Chevance, F.F.; Takahashi, N.; Karlinsey, J.E.; Gnerer, J.; Hirano, T.; Samudrala, R.; Aizawa, S.-I.; Hughes, K.T. The mechanism of outer membrane penetration by the eubacterial flagellum and implications for spirochete evolution. Genes Dev. 2007, 21, 2326–2335. [Google Scholar] [CrossRef] [PubMed]
- Jones, C.J.; Macnab, R.M.; Okino, H.; Aizawa, S.-I. Stoichiometric analysis of the flagellar hook-(basal-body) complex of Salmonella typhimurium. J. Mol. Biol. 1990, 212, 377–387. [Google Scholar] [CrossRef]
- Müller, V.; Jones, C.J.; Kawagishi, I.; Aizawa, S.; Macnab, R.M. Characterization of the fliE genes of Escherichia coli and Salmonella typhimurium and identification of the FliE protein as a component of the flagellar hook-basal body complex. J. Bacteriol. 1992, 174, 2298–2304. [Google Scholar] [CrossRef]
- Osorio-Valeriano, M.; De La Mora, J.; Camarena, L.; Dreyfus, G. Biochemical Characterization of the Flagellar Rod Components of Rhodobacter sphaeroides: Properties and Interactions. J. Bacteriol. 2015, 198, 544–552. [Google Scholar] [CrossRef]
- Fujii, T.; Kato, T.; Hiraoka, K.D.; Miyata, T.; Minamino, T.; Chevance, F.F.V.; Hughes, K.T.; Namba, K. Identical folds used for distinct mechanical functions of the bacterial flagellar rod and hook. Nat. Commun. 2017, 8, 14276. [Google Scholar] [CrossRef]
- Cohen, E.J.; Hughes, K.T. Rod-to-Hook Transition for Extracellular Flagellum Assembly Is Catalyzed by the L-Ring-Dependent Rod Scaffold Removal. J. Bacteriol. 2014, 196, 2387–2395. [Google Scholar] [CrossRef]
- Kaplan, M.; Sweredoski, M.J.; Rodrigues, J.P.G.L.M.; Tocheva, E.I.; Chang, Y.-W.; Ortega, D.R.; Beeby, M.; Jensen, G.J. Bacterial flagellar motor PL-ring disassembly subcomplexes are widespread and ancient. Proc. Natl. Acad. Sci. USA 2020, 117, 8941–8947. [Google Scholar] [CrossRef]
- Kaplan, M.; Subramanian, P.; Ghosal, D.; Oikonomou, C.M.; Pirbadian, S.; Starwalt-Lee, R.; Mageswaran, S.K.; Ortega, D.R.; Gralnick, J.A.; El-Naggar, M.Y.; et al. In situ imaging of the bacterial flagellar motor disassembly and assembly processes. EMBO J. 2019, 38, e100957. [Google Scholar] [CrossRef] [PubMed]
- Zhu, S.; Schniederberend, M.; Zhitnitsky, D.; Jain, R.; Galán, J.E.; Kazmierczak, B.I.; Liu, J. In Situ Structures of Polar and Lateral Flagella Revealed by Cryo-Electron Tomography. J. Bacteriol. 2019, 201. [Google Scholar] [CrossRef]
- Liu, R.; Ochman, H. Stepwise formation of the bacterial flagellar system. Proc. Natl. Acad. Sci. USA 2007, 104, 7116–7121. [Google Scholar] [CrossRef] [PubMed]
- Ueno, T.; Oosawa, K.; Aizawa, S.-I. Domain Structures of the MS Ring Component Protein (FliF) of the Flagellar Basal Body of Salmonella typhimurium. J. Mol. Biol. 1994, 236, 546–555. [Google Scholar] [CrossRef] [PubMed]
- Johnson, S.; Fong, Y.H.; Deme, J.C.; Furlong, E.J.; Kuhlen, L.; Lea, S.M. Symmetry mismatch in the MS-ring of the bacterial flagellar rotor explains the structural coordination of secretion and rotation. Nat. Microbiol. 2020, 5, 966–975. [Google Scholar] [CrossRef]
- Wagner, S.; Königsmaier, L.; Lara-Tejero, M.; Lefebre, M.; Marlovits, T.C.; Galán, J.E. Organization and coordinated assembly of the type III secretion export apparatus. Proc. Natl. Acad. Sci. USA 2010, 107, 17745–17750. [Google Scholar] [CrossRef]
- Thomas, D.R.; Morgan, D.G.; DeRosier, D.J. Rotational symmetry of the C ring and a mechanism for the flagellar rotary motor. Proc. Natl. Acad. Sci. USA 1999, 96, 10134–10139. [Google Scholar] [CrossRef]
- Young, H.S.; Dang, H.; Lai, Y.; DeRosier, D.J.; Khan, S. Variable Symmetry in Salmonella typhimurium Flagellar Motors. Biophys. J. 2003, 84, 571–577. [Google Scholar] [CrossRef]
- Zhao, R.; Amsler, C.D.; Matsumura, P.; Khan, S. FliG and FliM distribution in the Salmonella typhimurium cell and flagellar basal bodies. J. Bacteriol. 1996, 178, 258–265. [Google Scholar] [CrossRef][Green Version]
- McDowell, M.A.; Marcoux, J.; McVicker, G.; Johnson, S.; Fong, Y.H.; Stevens, R.; Bowman, L.A.H.; Degiacomi, M.T.; Yan, J.; Wise, A.; et al. Characterisation ofShigella Spa33 andThermotoga FliM/N reveals a new model for C-ring assembly in T3SS. Mol. Microbiol. 2015, 99, 749–766. [Google Scholar] [CrossRef]
- Sarkar, M.K.; Paul, K.; Blair, D.F. Subunit Organization and Reversal-associated Movements in the Flagellar Switch of Escherichia coli. J. Biol. Chem. 2009, 285, 675–684. [Google Scholar] [CrossRef] [PubMed]
- Bai, F.; Branch, R.W.; Nicolau, D.V.; Pilizota, T.; Steel, B.C.; Maini, P.K.; Berry, R.M. Conformational Spread as a Mechanism for Cooperativity in the Bacterial Flagellar Switch. Science 2010, 327, 685–689. [Google Scholar] [CrossRef] [PubMed]
- Dos Santos, R.N.; Khan, S.; Morcos, F. Characterization of C-ring component assembly in flagellar motors from amino acid coevolution. R. Soc. Open Sci. 2018, 5, 171854. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, R.; Abe-Yoshizumi, R.; Kishi, T.; Homma, M.; Kojima, S. Interaction of the C-Terminal Tail of FliF with FliG from the Na+-Driven Flagellar Motor of Vibrio alginolyticus. J. Bacteriol. 2014, 197, 63–72. [Google Scholar] [CrossRef] [PubMed]
- Lloyd, S.A.; Blair, D.F. Charged residues of the rotor protein FliG essential for torque generation in the flagellar motor of Escherichia coli. J. Mol. Biol. 1997, 266, 733–744. [Google Scholar] [CrossRef] [PubMed]
- Yakushi, T.; Yang, J.; Fukuoka, H.; Homma, M.; Blair, D.F. Roles of Charged Residues of Rotor and Stator in Flagellar Rotation: Comparative Study using H+-Driven and Na+-Driven Motors in Escherichia coli. J. Bacteriol. 2006, 188, 1466–1472. [Google Scholar] [CrossRef] [PubMed]
- Takekawa, N.; Kojima, S.; Homma, M. Contribution of Many Charged Residues at the Stator-Rotor Interface of the Na+-Driven Flagellar Motor to Torque Generation in Vibrio alginolyticus. J. Bacteriol. 2014, 196, 1377–1385. [Google Scholar] [CrossRef]
- Paul, K.; Brunstetter, D.; Titen, S.; Blair, D.F. A molecular mechanism of direction switching in the flagellar motor of Escherichia coli. Proc. Natl. Acad. Sci. USA 2011, 108, 17171–17176. [Google Scholar] [CrossRef]
- Minamino, T.; Imada, K.; Kinoshita, M.; Nakamura, S.; Morimoto, Y.V.; Namba, K. Structural Insight into the Rotational Switching Mechanism of the Bacterial Flagellar Motor. PLoS Biol. 2011, 9, e1000616. [Google Scholar] [CrossRef]
- Brown, P.N.; Hill, C.P.; Blair, D.F. Crystal structure of the middle and C-terminal domains of the flagellar rotor protein FliG. EMBO J. 2002, 21, 3225–3234. [Google Scholar] [CrossRef] [PubMed]
- Brown, P.N.; Terrazas, M.; Paul, K.; Blair, D.F. Mutational Analysis of the Flagellar Protein FliG: Sites of Interaction with FliM and Implications for Organization of the Switch Complex. J. Bacteriol. 2006, 189, 305–312. [Google Scholar] [CrossRef] [PubMed]
- Mathews, M.A.A.; Tang, H.L.; Blair, D.F. Domain Analysis of the FliM Protein ofEscherichia coli. J. Bacteriol. 1998, 180, 5580–5590. [Google Scholar] [CrossRef]
- Paul, K.; Blair, D.F. Organization of FliN Subunits in the Flagellar Motor of Escherichia coli. J. Bacteriol. 2006, 188, 2502–2511. [Google Scholar] [CrossRef]
- Lowenthal, A.C.; Hill, M.; Sycuro, L.K.; Mehmood, K.; Salama, N.R.; Ottemann, K.M. Functional Analysis of the Helicobacter pylori Flagellar Switch Proteins. J. Bacteriol. 2009, 191, 7147–7156. [Google Scholar] [CrossRef] [PubMed]
- Häuser, R.; Ceol, A.; Rajagopala, S.V.; Mosca, R.; Siszler, G.; Wermke, N.; Sikorski, P.; Schwarz, F.; Schick, M.; Wuchty, S.; et al. A Second-generation Protein–Protein Interaction Network ofHelicobacter pylori. Mol. Cell. Proteom. 2014, 13, 1318–1329. [Google Scholar] [CrossRef] [PubMed]
- Parrish, J.R.; Yu, J.; Liu, G.; Hines, J.A.; Chan, J.E.; Mangiola, B.A.; Zhang, H.; Pacifico, S.; Fotouhi, F.; DiRita, V.J.; et al. A proteome-wide protein interaction map for Campylobacter jejuni. Genome Biol. 2007, 8, 1–19. [Google Scholar] [CrossRef]
- Li, N.; Kojima, S.; Homma, M. Sodium-driven motor of the polar flagellum in marine bacteria Vibrio. Genes Cells 2011, 16, 985–999. [Google Scholar] [CrossRef]
- Berg, H.C. The Rotary Motor of Bacterial Flagella. Annu. Rev. Biochem. 2003, 72, 19–54. [Google Scholar] [CrossRef]
- Kojima, S.; Blair, D.F. Solubilization and Purification of the MotA/MotB Complex ofEscherichia coli. Biochemistry 2004, 43, 26–34. [Google Scholar] [CrossRef]
- Leake, M.C.; Chandler, J.H.; Wadhams, G.H.; Bai, F.; Berry, R.M.; Armitage, J.P. Stoichiometry and turnover in single, functioning membrane protein complexes. Nat. Cell Biol. 2006, 443, 355–358. [Google Scholar] [CrossRef]
- Blair, D.F.; Berg, H.C. Restoration of torque in defective flagellar motors. Science 1988, 242, 1678–1681. [Google Scholar] [CrossRef] [PubMed]
- Block, S.M.; Berg, H.C. Successive incorporation of force-generating units in the bacterial rotary motor. Nat. Cell Biol. 1984, 309, 470–472. [Google Scholar] [CrossRef] [PubMed]
- Reid, S.W.; Leake, M.C.; Chandler, J.H.; Lo, C.-J.; Armitage, J.P.; Berry, R.M. The maximum number of torque-generating units in the flagellar motor of Escherichia coli is at least 11. Proc. Natl. Acad. Sci. USA 2006, 103, 8066–8071. [Google Scholar] [CrossRef]
- Baker, A.E.; O’Toole, G.A. Bacteria, Rev Your Engines: Stator Dynamics Regulate Flagellar Motility. J. Bacteriol. 2017, 199, e00088-17. [Google Scholar] [CrossRef] [PubMed]
- Yonekura, K.; Maki-Yonekura, S.; Homma, M. Structure of the Flagellar Motor Protein Complex PomAB: Implications for the Torque-Generating Conformation. J. Bacteriol. 2011, 193, 3863–3870. [Google Scholar] [CrossRef][Green Version]
- Coulton, J.W.; Murray, R.G. Cell envelope associations of Aquaspirillum serpens flagella. J. Bacteriol. 1978, 136, 1037–1049. [Google Scholar] [CrossRef]
- Khan, S.; Dapice, M.; Reese, T.S. Effects of mot gene expression on the structure of the flagellar motor. J. Mol. Biol. 1988, 202, 575–584. [Google Scholar] [CrossRef]
- Liu, J.; Howell, J.K.; Bradley, S.D.; Zheng, Y.; Zhou, Z.H.; Norris, S.J. Cellular Architecture of Treponema pallidum: Novel Flagellum, Periplasmic Cone, and Cell Envelope as Revealed by Cryo Electron Tomography. J. Mol. Biol. 2010, 403, 546–561. [Google Scholar] [CrossRef]
- Zhu, S.; Nishikino, T.; Hu, B.; Kojima, S.; Homma, M.; Liu, J. Molecular architecture of the sheathed polar flagellum in Vibrio alginolyticus. Proc. Natl. Acad. Sci. USA 2017, 114, 10966–10971. [Google Scholar] [CrossRef]
- Zhu, S.; Nishikino, T.; Takekawa, N.; Terashima, H.; Kojima, S.; Imada, K.; Homma, M.; Liu, J. In Situ Structure of the Vibrio Polar Flagellum Reveals a Distinct Outer Membrane Complex and Its Specific Interaction with the Stator. J. Bacteriol. 2019, 202. [Google Scholar] [CrossRef] [PubMed]
- Chevance, F.F.V.; Hughes, K.T. Coordinating assembly of a bacterial macromolecular machine. Nat. Rev. Genet. 2008, 6, 455–465. [Google Scholar] [CrossRef]
- Xie, L.; Altindal, T.; Chattopadhyay, S.; Wu, X.L. From the Cover: Bacterial flagellum as a propeller and as a rudder for efficient chemotaxis. Proc. Natl. Acad. Sci. USA 2011, 108, 2246–2251. [Google Scholar] [CrossRef]
- Charon, N.W.; Cockburn, A.; Li, C.; Liu, J.; Miller, K.A.; Miller, M.R.; Motaleb, A.; Wolgemuth, C.W. The Unique Paradigm of Spirochete Motility and Chemotaxis. Annu. Rev. Microbiol. 2012, 66, 349–370. [Google Scholar] [CrossRef]
- Charon, N.W.; Goldstein, S.F. Genetics of Motility and Chemotaxis of a Fascinating Group of Bacteria: The Spirochetes. Annu. Rev. Genet. 2002, 36, 47–73. [Google Scholar] [CrossRef] [PubMed]
- Goldstein, S.F.; Buttle, K.F.; Charon, N.W. Structural analysis of the Leptospiraceae and Borrelia burgdorferi by high-voltage electron microscopy. J. Bacteriol. 1996, 178, 6539–6545. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Minamino, T.; Kinoshita, M.; Namba, K. Directional Switching Mechanism of the Bacterial Flagellar Motor. Comput. Struct. Biotechnol. J. 2019, 17, 1075–1081. [Google Scholar] [CrossRef]
- Branch, R.W.; Sayegh, M.N.; Shen, C.; Nathan, V.S.; Berg, H.C. Adaptive Remodelling by FliN in the Bacterial Rotary Motor. J. Mol. Biol. 2014, 426, 3314–3324. [Google Scholar] [CrossRef]
- Lele, P.P.; Branch, R.W.; Nathan, V.S.J.; Berg, H.C. Mechanism for adaptive remodeling of the bacterial flagellar switch. Proc. Natl. Acad. Sci. USA 2012, 109, 20018–20022. [Google Scholar] [CrossRef]
- Delalez, N.J.; Wadhams, G.H.; Rosser, G.; Xue, Q.; Brown, M.T.; Dobbie, I.M.; Berry, R.M.; Leake, M.C.; Armitage, J.P. Signal-dependent turnover of the bacterial flagellar switch protein FliM. Proc. Natl. Acad. Sci. USA 2010, 107, 11347–11351. [Google Scholar] [CrossRef]
- Delalez, N.J.; Berry, R.M.; Armitage, J.P. Stoichiometry and Turnover of the Bacterial Flagellar Switch Protein FliN. mBio 2014, 5, e01216-14. [Google Scholar] [CrossRef] [PubMed]
- Sakai, T.; Miyata, T.; Terahara, N.; Mori, K.; Inoue, Y.; Morimoto, Y.V.; Kato, T.; Namba, K.; Minamino, T. Novel Insights into Conformational Rearrangements of the Bacterial Flagellar Switch Complex. mBio 2019, 10, e00079-19. [Google Scholar] [CrossRef] [PubMed]
- Erhardt, M.; Wheatley, P.; Kim, E.A.; Hirano, T.; Zhang, Y.; Sarkar, M.K.; Hughes, K.T.; Blair, D.F. Mechanism of type-III protein secretion: Regulation of FlhA conformation by a functionally critical charged-residue cluster. Mol. Microbiol. 2017, 104, 234–249. [Google Scholar] [CrossRef] [PubMed]
- Minamino, T.; Morimoto, Y.V.; Hara, N.; Aldridge, P.D.; Namba, K. The Bacterial Flagellar Type III Export Gate Complex Is a Dual Fuel Engine That Can Use Both H+ and Na+ for Flagellar Protein Export. PLoS Pathog. 2016, 12, e1005495. [Google Scholar] [CrossRef]
- Minamino, T.; Morimoto, Y.V.; Hara, N.; Namba, K. An energy transduction mechanism used in bacterial flagellar type III protein export. Nat. Commun. 2011, 2, 475. [Google Scholar] [CrossRef]
- Morimoto, Y.V.; Kami-Ike, N.; Miyata, T.; Kawamoto, A.; Kato, T.; Namba, K.; Minamino, T. High-Resolution pH Imaging of Living Bacterial Cells To Detect Local pH Differences. mBio 2016, 7, e01911-16. [Google Scholar] [CrossRef]
- Barker, C.S.; Samatey, F.A. Cross-Complementation Study of the Flagellar Type III Export Apparatus Membrane Protein FlhB. PLoS ONE 2012, 7, e44030. [Google Scholar] [CrossRef][Green Version]
- Hara, N.; Namba, K.; Minamino, T. Genetic Characterization of Conserved Charged Residues in the Bacterial Flagellar Type III Export Protein FlhA. PLoS ONE 2011, 6, e22417. [Google Scholar] [CrossRef][Green Version]
- Kihara, M.; Minamino, T.; Yamaguchi, S.; Macnab, R.M. Intergenic Suppression between the Flagellar MS Ring Protein FliF of Salmonella and FlhA, a Membrane Component of Its Export Apparatus. J. Bacteriol. 2001, 183, 1655–1662. [Google Scholar] [CrossRef]
- Ferris, H.U.; Furukawa, Y.; Minamino, T.; Kroetz, M.B.; Kihara, M.; Namba, K.; Macnab, R.M. FlhB Regulates Ordered Export of Flagellar Components via Autocleavage Mechanism. J. Biol. Chem. 2005, 280, 41236–41242. [Google Scholar] [CrossRef]
- Fraser, G.M.; Hirano, T.; Ferris, H.U.; Devgan, L.L.; Kihara, M.; Macnab, R.M. Substrate specificity of type III flagellar protein export in Salmonella is controlled by subdomain interactions in FlhB. Mol. Microbiol. 2003, 48, 1043–1057. [Google Scholar] [CrossRef] [PubMed]
- Kuhlen, L.; Abrusci, P.; Johnson, S.; Gault, J.; Deme, J.; Caesar, J.; Dietsche, T.; Mebrhatu, M.T.; Ganief, T.; Macek, B.; et al. Structure of the core of the type III secretion system export apparatus. Nat. Struct. Mol. Biol. 2018, 25, 583–590. [Google Scholar] [CrossRef] [PubMed]
- Katayama, E.; Shiraishi, T.; Oosawa, K.; Baba, N.; Aizawa, S. Geometry of the flagellar motor in the cytoplasmic membrane of Salmonella typhimurium as determined by stereo-photogrammetry of quick-freeze deep-etch replica images. J. Mol. Biol. 1996, 255, 458–475. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Lin, T.; Botkin, D.J.; McCrum, E.; Winkler, H.; Norris, S.J. Intact Flagellar Motor of Borrelia burgdorferi Revealed by Cryo-Electron Tomography: Evidence for Stator Ring Curvature and Rotor/C-Ring Assembly Flexion. J. Bacteriol. 2009, 191, 5026–5036. [Google Scholar] [CrossRef] [PubMed]
- Galperin, M.; Dibrov, P.A.; Glagolev, A.N. delta mu H+ is required for flagellar growth in Escherichia coli. FEBS Lett. 1982, 143, 319–322. [Google Scholar] [CrossRef]
- Ewilharm, G.; Lehmann, V.; Krauss, K.; Lehnert, B.; Richter, S.; Ruckdeschel, K.; Heesemann, J.; Trülzsch, K. Yersinia enterocolitica Type III Secretion Depends on the Proton Motive Force but Not on the Flagellar Motor Components MotA and MotB. Infect. Immun. 2004, 72, 4004–4009. [Google Scholar] [CrossRef]
- Paul, K.; Erhardt, M.; Hirano, T.; Blair, D.F.; Hughes, K.T. Energy source of flagellar type III secretion. Nat. Cell Biol. 2008, 451, 489–492. [Google Scholar] [CrossRef]
- Minamino, T.; Namba, K. Distinct roles of the FliI ATPase and proton motive force in bacterial flagellar protein export. Nat. Cell Biol. 2008, 451, 485–488. [Google Scholar] [CrossRef]
- Barker, C.S.; Inoue, T.; Meshcheryakova, I.V.; Kitanobo, S.; Samatey, F.A. Function of the conserved FHIPEP domain of the flagellar type III export apparatus, protein FlhA. Mol. Microbiol. 2016, 100, 278–288. [Google Scholar] [CrossRef]
- Butan, C.; Lara-Tejero, M.; Li, W.; Liu, J.; Galán, J.E. High-resolution view of the type III secretion export apparatus in situ reveals membrane remodeling and a secretion pathway. Proc. Natl. Acad. Sci. USA 2019, 116, 24786–24795. [Google Scholar] [CrossRef]
- Echazarreta, M.A.; Klose, K.E. Vibrio Flagellar Synthesis. Front. Cell. Infect. Microbiol. 2019, 9, 131. [Google Scholar] [CrossRef] [PubMed]
- Chu, J.; Liu, J.; Hoover, T.R. Phylogenetic Distribution, Ultrastructure, and Function of Bacterial Flagellar Sheaths. Biomol. 2020, 10, 363. [Google Scholar] [CrossRef] [PubMed]
- Kusumoto, A.; Shinohara, A.; Terashima, H.; Kojima, S.; Yakushi, T.; Homma, M. Collaboration of FlhF and FlhG to regulate polar-flagella number and localization in Vibrio alginolyticus. Microbiol. 2008, 154, 1390–1399. [Google Scholar] [CrossRef] [PubMed]
- Terashima, H.; Fukuoka, H.; Yakushi, T.; Kojima, S.; Homma, M. The Vibrio motor proteins, MotX and MotY, are associated with the basal body of Na+-driven flagella and required for stator formation. Mol. Microbiol. 2006, 62, 1170–1180. [Google Scholar] [CrossRef]
- Terashima, H.; Koike, M.; Kojima, S.; Homma, M. The Flagellar Basal Body-Associated Protein FlgT Is Essential for a Novel Ring Structure in the Sodium-Driven Vibrio Motor. J. Bacteriol. 2010, 192, 5609–5615. [Google Scholar] [CrossRef]
- Zhu, S.; Nishikino, T.; Kojima, S.; Homma, M.; Liu, J. The Vibrio H-Ring Facilitates the Outer Membrane Penetration of the Polar Sheathed Flagellum. J. Bacteriol. 2018, 200. [Google Scholar] [CrossRef]
- Motaleb, A.; Corum, L.; Bono, J.L.; Elias, A.F.; Rosa, P.; Samuels, D.; Charon, N.W. Borrelia burgdorferi periplasmic flagella have both skeletal and motility functions. Proc. Natl. Acad. Sci. USA 2000, 97, 10899–10904. [Google Scholar] [CrossRef]
- Sosinsky, G.E.; Francis, N.R.; Stallmeyer, M.; DeRosier, D.J. Substructure of the flagellar basal body of Salmonella typhimurium. J. Mol. Biol. 1992, 223, 171–184. [Google Scholar] [CrossRef]
- Moon, K.H.; Zhao, X.; Manne, A.; Wang, J.; Yu, Z.; Liu, J.; Motaleb, A. Spirochetes flagellar collar protein FlbB has astounding effects in orientation of periplasmic flagella, bacterial shape, motility, and assembly of motors inBorrelia burgdorferi. Mol. Microbiol. 2016, 102, 336–348. [Google Scholar] [CrossRef]
- Rajagopala, S.V.; Titz, B.; Goll, J.; Parrish, J.R.; Wohlbold, K.; McKevitt, M.T.; Palzkill, T.; Mori, H.; Jr, R.L.F.; Uetz, P. The protein network of bacterial motility. Mol. Syst. Biol. 2007, 3, 128. [Google Scholar] [CrossRef]
- Moon, K.H.; Zhao, X.; Xu, H.; Liu, J.; Motaleb, A. A tetratricopeptide repeat domain protein has profound effects on assembly of periplasmic flagella, morphology and motility of the lyme disease spirochete Borrelia burgdorferi. Mol. Microbiol. 2018, 110, 634–647. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; He, J.; Liu, J.; Motaleb, A. BB0326 is responsible for the formation of periplasmic flagellar collar and assembly of the stator complex in Borrelia burgdorferi. Mol. Microbiol. 2019, 113, 418–429. [Google Scholar] [CrossRef]

| Protein(s) | Species | PDB ID | Refs |
|---|---|---|---|
| Axial | |||
| Flagellin FliC | Bacillus cereus | 5Z7Q | [29] |
| Salmonella typhimurium | 1IO1 | [30] | |
| Sphingaomonas sp | 2ZBI, 3K8V, 3K8W | [31] | |
| Burkholderia psuedomallei | 4CFI | [32] | |
| Pseudomonas aeruginosa | 4NX9 | [33] | |
| FliS | Aquifex aeolicus | 1ORY, 1ORJ | [34] |
| Bacillus cereus | 5XEF | [35] | |
| Helicobacter pylori | 3IQC | [36] | |
| FliT | Salmonella typhimurium | 5GNA | |
| Yersinia enterocolitica | 3NKZ | ||
| FljB | Salmonella typhimurium | 6RGV | [37] |
| FcpA | Leptospira biflexa | 6NQY | [38] |
| FcpB | Leptospira interrogans | 6NQZ | [38] |
| Flagellin–FliS | Bacillus subtilis | 5MAW, 6GOW | [39] |
| FliC–FliS fusion | Aquifex aeolicus | 4IWB | [40] |
| FlgD | Helicobacter pylori | 4ZZF, 4ZZK, 5K5Y | [41,42] |
| Salmonella typhimurium | 6IEE, 6IEF | ||
| FlgE | Campylobacter jejuni | 5AZ4 | [43] |
| Caulobacter crescentus | 5AY6 | [43] | |
| Helicobacter pylori | 5NPY | [44] | |
| Salmonella typhimurium | 1WLG | [45] | |
| Treponema denticola | 6NDT, 6NDW, 6NDV, 6NDX | [46] | |
| FlgK | Campylobacter jejuni | 5XBJ | [47] |
| FlgL | Bacillus cereus | 5ZIY | [48] |
| Xanthomonas campestris | 5ZIZ, 5ZJ0 | [48] | |
| Legionella pneumophila | 5YTI | ||
| FliD (HAP2) | Pseudomonas aeruginosa | 5FHY | [49] |
| Helicobacter pylori | 6IWY | [50] | |
| FlgG | Salmonella typhimurium | 6JF2 | [51] |
| FlgJ | Salmonella typhimurium | 5DN4, 5DN5 | [52] |
| Basal Body | |||
| FlgA | Salmonella typhimurium | 3VKI, 3VJP, 3TEE | [53] |
| FliF–FliG | Helicobater pylori | 5WUJ | [54] |
| FliFc–FliGN | Thermotaoga maritima | 5TDY | [55] |
| FliG | Aquifex aeolicus | 3HJL | [56] |
| Helicobacter Pylori | 3USY, 3USW | [57] | |
| Thermotoga maritima | 1LKV, 1QC7, 3AJC | [58,59,60] | |
| FliM | Helicobacter pylori | 4GC8 | [61] |
| Thermotoga maritima | 2HP7 | [62] | |
| Helicobacter pylori | 5XRW | [63] | |
| FliN | Thermotaoga maritima | 1YAB, 1O6A | [64] |
| FliY | Thermotoga maritima | 4HYN | [65] |
| FliG–FliM | Helicobacter pylori | 4FQ0 | [61] |
| Thermotaoga maritima | 3SOH, 4FHR, 4QRM | [66,67,68] | |
| FliM–FliN | Salmonella typhimurium | 4XYB | [69] |
| FliM–FliN–FliH | Salmonella typhimurium | 4XYC | [70] |
| FliM–SpeE | Helicobacter pylori | 5X0Z | [71] |
| CheY | Escherichia coli | 1U8T, 1ZDM, 2B1J, 2ID7, 2ID9, 2IDM, 6TG7 | [72,73] [74] |
| Thermotoga maritima | 4IGA | [75] | |
| CheY3 | Vibrio cholerae | 3TO5, 4H60, 4HNQ, 4JP1, 4LX8 | [76] |
| CheY4 | Vibrio cholerae | 4HNR, 4HNS | [76] |
| CheY–FliM | Escherichia coli | 1F4V | [72] |
| FlhG | Geobacillus thermodenitrificans | 4RZ2, 4RZ3 | [77] |
| MotB | Salmonella typhimurium | 5Y3Z, 5Y40, 2ZVY, 2ZVZ, 2ZOV | [78,79] |
| PomBc | Vibrio alginolyticus | 3WPW, 3WPX | [80] |
| MotY | Vibrio alginolyticus | 2ZF8 | [81] |
| FliL | Vibrio alginolyticus | 6AHQ, 6AHP | [82] |
| FlgT | Vibrio alginolyticus | 3W1E | [83] |
| Export Apparatus | |||
| FlhA | Bacillus subtilis | 3MIX | [84] |
| Salmonella typhimurium | 6CH1, 6AI0, 6AI1, 6AI2, 6AI3 | [85,86] | |
| FlhA FliT–FliD complex | Salmonella typhimurium | 6CH2 | [85] |
| FlhA FliS–FliC complex | Salmonella typhimurium | 6CH3 | [85] |
| FlhB | Aquifex aeolicus | 3B1S | [87] |
| Salmonella typhimurium | 3B0Z | [87] | |
| FlhF | Bacillus subtilis | 2PX0, 2PX3 | [88] |
| FliI | Salmonella typhimurium | 2DPY | [89] |
| FliJ | Salmonella typhimurium | 3AJW | [90] |
| FlgN | Pseudomonas aeruginosa | 2FUP | |
| Salmonella typhimurium | 5B3D | [91] | |
| FliH–FliI | Salmonella typhimurium | 5B0O | [92] |
| Protein(s) | Species | PDB ID | EMDB ID | Refs |
|---|---|---|---|---|
| Axial | ||||
| Flagellin | Campylobacter jejuni | 5007 | [93] | |
| Salmonella typhimurium | 1UCU, 3A5X | 1641 | [94,95] | |
| Bacillus subtilis | 5WJT, 5WJU, 5WJV, 5WJW, 5WJX, 5WJY, 5WJZ | 8447, 8848, 8849, 8850, 8851, 8852, 8853 | [96] | |
| Pseudomonas aeruginosa | 5WK5, 5WK6 | 8855, 8856 | [96] | |
| Leptospira biflexa | 6PWB | 20504 | [38] | |
| Salmonella typhimurium | 6JY0 | 9896 | [97] | |
| Kurthia spp. | 6T17 | 10362 | [98] | |
| FlgE | Helicobacter pylori | 5JXL | 8179 | [99] |
| Caulobacter crescentus | 2BGY | 1132 | [100] | |
| Salmonella typhimurium | 2BGZ, 3A69, 6JZT, 6KFK, 6K3I | 1132, 1647, 9974, 9909 | [100,101] [51,102,103] | |
| Salmonella enterica | 6K9Q | 9952 | [104] | |
| FliD (HAP2) | Escherichia coli | 1873 | [105] | |
| FlgG | Salmonella typhimurium | 6JZR | 6683 | [51] |
| Basal Body | ||||
| Salmonella typhimurium | 1887 | [106] | ||
| FliF | Salmonella typhimurium | 6SCN, 6SD1, 6SD2, 6SD3, 6SD4, 6SD5, 6TRE | 10143, 10145, 10146, 10147, 10148, 10149, 10560, 6715 | [107,108] |
| FliF–FliG | Salmonella typhimurium | 6716 | [108] | |
| MotA | Aquifex aeolicus | 3417 | [109] | |
| MotA/B | Campylobacter jejuni | 6YKM, 6YKP, 6YKR | 10828, 10829, 10830 | [110] |
| Clostridium sporogenes | 6YSF | 10895, 10897 | [111] | |
| Bacillus subtilis | 6YSL | 10899 | [111] | |
| PomA/PomB | Vibrio mimicus | 10901 | [111] | |
| Export Apparatus | ||||
| FliPQR | Salmonella typhimurium | 6R69, 6F2D | 4733, 4173 | [107] |
| Vibrio mimicus | 6S3S | 10096 | [112] | |
| Pseudomonas savastanoi | 6S3R | 10095 | [112] | |
| FliPQR–FlhB | Vibrio mimicus | 6S3L | 10093 | [112] |
| SctRST | Salmonella typhimurium | 6R6B | 4734 | [107] |
| Species | EMDB ID | Refs |
|---|---|---|
| Acetonema longum | 5297 | [113] |
| Arcobacter butzleri | 3910 | [114] |
| Borrelia burgdorferi | 0525, 0534, 0536, 0537, 0538, 1644, 5298, 5627, 5628, 5629, 5630, 5631, 5632, 5633, 6088, 6089, 6090, 6091, 6092, 6093, 6094, 6095, 6096, 6097, 6098, 9123, 21885, 21884, 21886 | [113,115,116,117,118,119,120] |
| Bdellovibrio bacteriovorus | 3911 | [114] |
| Campylbacter jejuni | 3150, 3157, 3158, 3159, 3160, 3161, 5300, 10341, 10342, 10343, 10345, 10454, 10455, 10456, 10457 | [113,121,122] |
| Caulobacter crescentus | 5312, 10943, 10945, 10949, 10950, 10955, 10956, 10957 | [113,123] |
| Escherichia coli | 5311 | [113] |
| Helicobacter pylori | 8459 | [57] |
| Heliobacter Hepaticus | 5299 | [113] |
| Hylemonella gracilis | 5309 | [113] |
| Hyphomonas neptunium | 5313 | [113] |
| Legionella pneumophila | 0464 | [124] |
| Leptospira biflexa | 20503, 20504 | [38] |
| Leptospira interrogans | 5912, 5913, 5914 | [6] |
| Plesiomonas shigelloides | 4569, 10057 | [125] |
| Pseudomonas aeruginosa | 0465 | [124] |
| Salmonella enterica | 2520, 2521, 3154, 3813, 5310 | [113,121,126] |
| Shewanella oneidensis | 0467 | [124] |
| Treponema primitia | 1235 | [127] |
| Vibrio cholerae | 5308 | [113] |
| Vibrio fischeri | 3155, 3156, 3162 | [121] |
| Vibrio alginolyticus | 21819, 21837 | [128] |
| Wolinella succinogenes | 3912 | [114] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Carroll, B.L.; Liu, J. Structural Conservation and Adaptation of the Bacterial Flagella Motor. Biomolecules 2020, 10, 1492. https://doi.org/10.3390/biom10111492
Carroll BL, Liu J. Structural Conservation and Adaptation of the Bacterial Flagella Motor. Biomolecules. 2020; 10(11):1492. https://doi.org/10.3390/biom10111492
Chicago/Turabian StyleCarroll, Brittany L., and Jun Liu. 2020. "Structural Conservation and Adaptation of the Bacterial Flagella Motor" Biomolecules 10, no. 11: 1492. https://doi.org/10.3390/biom10111492
APA StyleCarroll, B. L., & Liu, J. (2020). Structural Conservation and Adaptation of the Bacterial Flagella Motor. Biomolecules, 10(11), 1492. https://doi.org/10.3390/biom10111492






