Anti-Biofilm Molecules Targeting Functional Amyloids
Abstract
1. Introduction
1.1. Biofilm Related Infections
1.2. Amyloids as Structural Scaffolds of the Biofilm Matrix
2. Functional Amyloids of the Biofilm Matrix
2.1. Intrinsic Amyloids
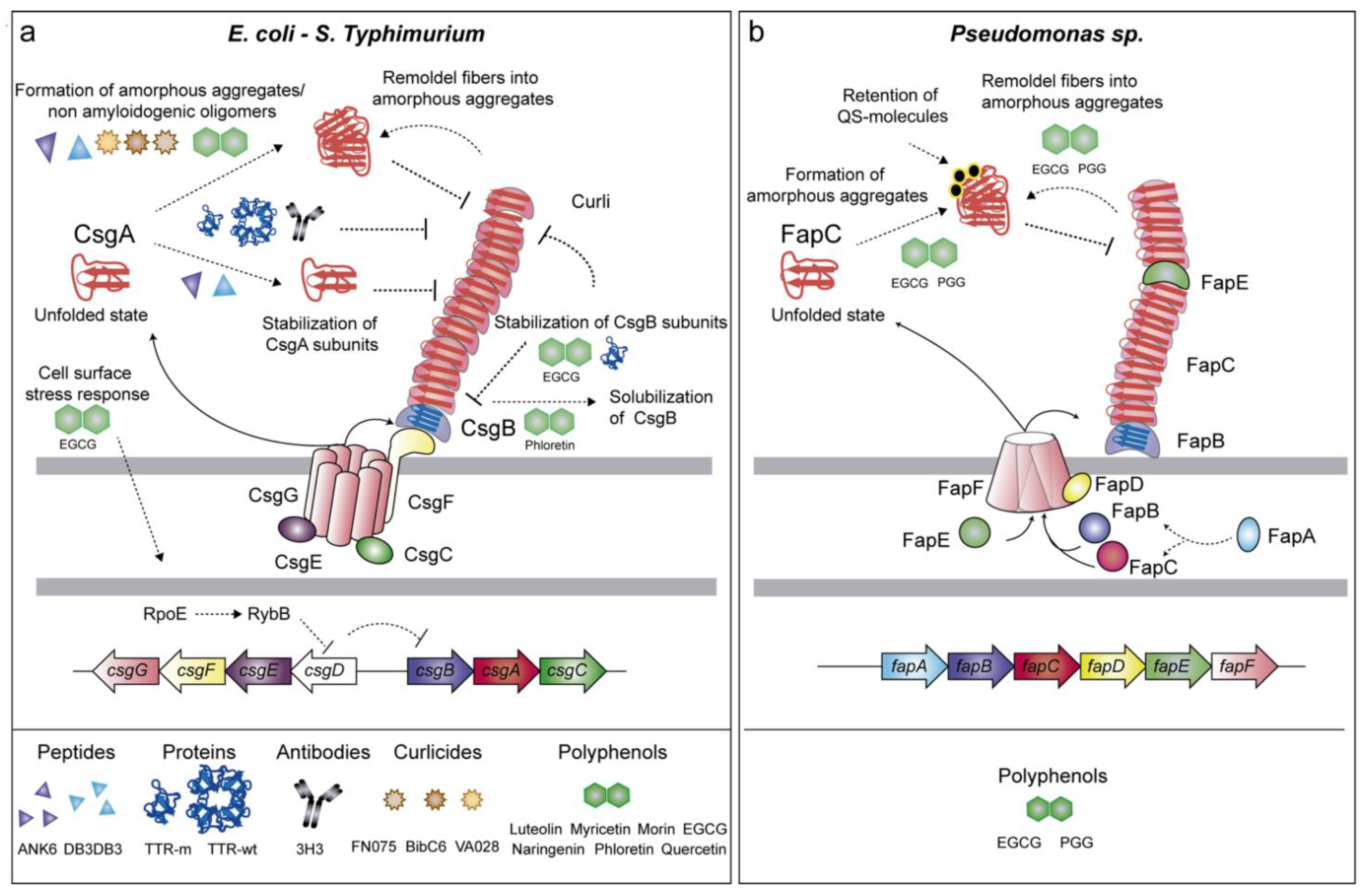
2.1.1. Curli (csgBAC-csgDEFG Genes)
2.1.2. Fap (fapA-F Genes)
2.1.3. Chaplins and Rodlins
2.2. Facultative Amyloids
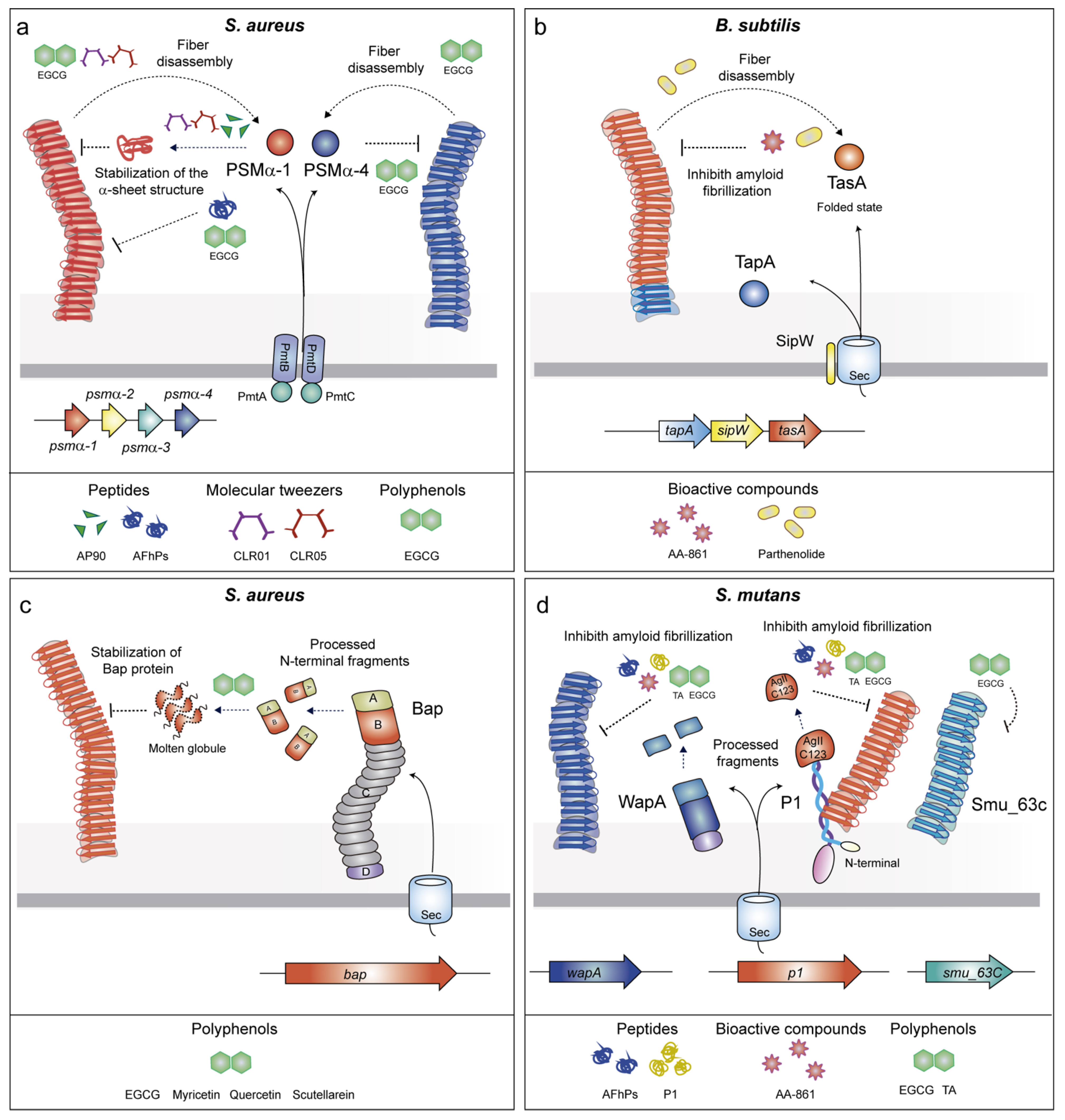
2.2.1. Phenol-Soluble Modulins (PSMs)
2.2.2. TasA
2.2.3. Biofilm Associated Proteins (BAPs)
2.2.4. P1 Adhesin/WapA/SMU_63c
3. Drugs Targeting Amyloid-Structured Biofilms
3.1. Anti-Amyloid Peptides
3.2. Anti-Amyloid Proteins
3.3. Antibodies as Native-State Stabilizing Agents
3.4. Molecular Tweezers
3.5. Anti-Amyloids Based on Pilicides and Curlicides
3.6. Bioactive Compounds with Anti-Amyloid Properties
3.7. Polyphenols
4. Final Remarks
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Sikora, A.; Zahra, F. Nosocomial Infections; StatPearls Publishing: Bethesda, MD, USA; Treasure Island, FL, USA, 2020. [Google Scholar]
- Ma, Y.X.; Wang, C.Y.; Li, Y.Y.; Li, J.; Wan, Q.Q.; Chen, J.H.; Tay, F.R.; Niu, L.N. Considerations and caveats in combating ESKAPE pathogens against nosocomial infections. Adv. Sci. 2020, 7, 1901872. [Google Scholar] [CrossRef]
- Laxminarayan, R.; Matsoso, P.; Pant, S.; Brower, C.; Rottingen, J.-A.; Klugman, K.; Davies, S. Access to effective antimicrobials: A worldwide challenge. Lancet 2016, 387, 168–175. [Google Scholar] [CrossRef]
- Holmes, A.H.; Moore, L.S.P.; Sundsfjord, A.; Steinbakk, M.; Regmi, S.; Karkey, A.; Guerin, P.J.; Piddock, L.J.V. Understanding the mechanisms and drivers of antimicrobial resistance. Lancet 2016, 387, 176–187. [Google Scholar] [CrossRef]
- Prestinaci, F.; Pezzotti, P.; Pantosti, A. Antimicrobial resistance: A global multifaceted phenomenon. Pathog. Glob. Health 2015, 109, 309–318. [Google Scholar] [CrossRef]
- Jefferson, K.K.; Goldmann, D.A.; Pier, G.B. Use of confocal microscopy to analyze the rate of vancomycin penetration through Staphylococcus aureus biofilms. Antimicrob. Agents Chemother. 2005, 49, 2467–2473. [Google Scholar] [CrossRef] [PubMed]
- Lewis, K. Persister cells, dormancy and infectious disease. Nat. Rev. Micro. 2007, 5, 48–56. [Google Scholar] [CrossRef] [PubMed]
- Flemming, H.-C.; Wingender, J. The biofilm matrix. Nat. Rev. Micro. 2010, 8, 623–633. [Google Scholar] [CrossRef]
- Hall, C.W.; Mah, T.-F. Molecular mechanisms of biofilm-based antibiotic resistance and tolerance in pathogenic bacteria. FEMS Microbiol. Rev. 2017, 41, 276–301. [Google Scholar] [CrossRef] [PubMed]
- Stewart, P.S.; Costerton, J.W. Antibiotic resistance of bacteria in biofilms. Lancet 2001, 358, 135–138. [Google Scholar] [CrossRef]
- Blanco, L.P.; Evans, M.L.; Smith, D.R.; Badtke, M.P.; Chapman, M.R. Diversity, biogenesis and function of microbial amyloids. Trends Microbiol. 2012, 20, 66–73. [Google Scholar] [CrossRef]
- Taglialegna, A.; Lasa, I.; Valle, J. Amyloid structures as biofilm matrix scaffolds. J. Bacteriol. 2016, 198, 2579–2588. [Google Scholar] [CrossRef]
- Álvarez-Mena, A.; Cámara-Almirón, J.; de Vicente, A.; Romero, D. Multifunctional Amyloids in the Biology of Gram-Positive Bacteria. Microorganisms 2020, 8, 2020. [Google Scholar] [CrossRef]
- Erskine, E.; Macphee, C.E.; Stanley-Wall, N.R. Functional Amyloid and Other Protein Fibers in the Biofilm Matrix. J. Mol. Biol. 2018, 430, 3642–3656. [Google Scholar] [CrossRef]
- Levkovich, S.A.; Gazit, E.; Bar-Yosef, D.L. Two Decades of Studying Functional Amyloids in Microorganisms. Trends Microbiol. 2020, 1–15. [Google Scholar] [CrossRef]
- Uversky, V.N.; Fink, A.L. Conformational constraints for amyloid fibrillation: The importance of being unfolded. Biochim. Biophys Acta 2004, 1698, 131–153. [Google Scholar] [CrossRef]
- Tayeb-Fligelman, E.; Tabachnikov, O.; Moshe, A.; Goldshmidt-Tran, O.; Sawaya, M.R.; Coquelle, N.; Colletier, J.-P.; Landau, M. The cytotoxic Staphylococcus aureus PSMα3 reveals a cross-α amyloid-like fibril. Science 2017, 355, 831–833. [Google Scholar] [CrossRef] [PubMed]
- Salinas, N.; Tayeb-Fligelman, E.; Sammito, M.D.; Bloch, D.; Jelinek, R.; Noy, D.; Usón, I.; Landau, M. The amphibian antimicrobial peptide uperin 3.5 is a cross-α/cross-β chameleon functional amyloid. Proc. Natl Acad Sci USA 2021, 118. [Google Scholar] [CrossRef] [PubMed]
- Groenning, M. Binding mode of Thioflavin T and other molecular probes in the context of amyloid fibrils-current status. J. Chem. Biol 2010, 3, 1–18. [Google Scholar] [CrossRef]
- Howie, A.J.; Brewer, D.B.; Howell, D.; Jones, A.P. Physical basis of colors seen in Congo red-stained amyloid in polarized light. Lab. Investig. 2008, 88, 232–242. [Google Scholar] [CrossRef] [PubMed]
- Navarro, S.; Ventura, S. Fluorescent dye ProteoStat to detect and discriminate intracellular amyloid-like aggregates in Escherichia coli. Biotechnol. J. 2014, 9, 1259–1266. [Google Scholar] [CrossRef] [PubMed]
- Shewmaker, F.; McGlinchey, R.P.; Wickner, R.B. Structural insights into functional and pathological amyloid. J. Biol. Chem. 2011, 286, 16533–16540. [Google Scholar] [CrossRef]
- Hufnagel, D.A.; DePas, W.H.; Chapman, M.R. The biology of the Escherichia coli extracellular matrix. MicroBiol. Spectr. 2015, 3, 23. [Google Scholar] [CrossRef]
- Van Gerven, N.; Van der Verren, S.E.; Reiter, D.M.; Remaut, H. The role of functional amyloids in bacterial virulence. J. Mol. Biol. 2018, 430, 3657–3684. [Google Scholar] [CrossRef]
- Hammar, M.; Arnqvist, A.; Bian, Z.; Olsen, A.; Normark, S. Expression of two csg operons is required for production of fibronectin- and congo red-binding curli polymers in Escherichia coli K-12. Mol. Microbiol. 1995, 18, 661–670. [Google Scholar] [CrossRef]
- Hammer, N.D.; McGuffie, B.A.; Zhou, Y.; Badtke, M.P.; Reinke, A.A.; Brännström, K.; Gestwicki, J.E.; Olofsson, A.; Almqvist, F.; Chapman, M.R. The C-terminal repeating units of CsgB direct bacterial functional amyloid nucleation. J. Mol. Biol. 2012, 422, 376–389. [Google Scholar] [CrossRef][Green Version]
- Hobley, L.; Harkins, C.; Macphee, C.E.; Stanley-Wall, N.R. Giving structure to the biofilm matrix: An overview of individual strategies and emerging common themes. FEMS MicroBiol. Rev. 2015, 39, 649–669. [Google Scholar] [CrossRef] [PubMed]
- Van Gerven, N.; Klein, R.D.; Hultgren, S.J.; Remaut, H. Bacterial amyloid formation: Structural insights into curli biogensis. Trends Microbiol. 2015, 23, 693–706. [Google Scholar] [CrossRef]
- Robinson, L.S.; Ashman, E.M.; Hultgren, S.J.; Chapman, M.R. Secretion of curli fibre subunits is mediated by the outer membrane-localized CsgG protein. Mol. Microbiol. 2006, 59, 870–881. [Google Scholar] [CrossRef]
- Nenninger, A.A.; Robinson, L.S.; Hammer, N.D.; Epstein, E.A.; Badtke, M.P.; Hultgren, S.J.; Chapman, M.R. CsgE is a curli secretion specificity factor that prevents amyloid fibre aggregation. Mol. Microbiol. 2011, 81, 486–499. [Google Scholar] [CrossRef] [PubMed]
- Brombacher, E.; Baratto, A.; Dorel, C.; Landini, P. Gene expression regulation by the Curli activator CsgD protein: Modulation of cellulose biosynthesis and control of negative determinants for microbial adhesion. J. Bacteriol. 2006, 188, 2027–2037. [Google Scholar] [CrossRef]
- Zakikhany, K.; Harrington, C.R.; Nimtz, M.; Hinton, J.C.D.; Römling, U. Unphosphorylated CsgD controls biofilm formation in Salmonella enterica serovar Typhimurium. Mol. Microbiol. 2010, 77, 771–786. [Google Scholar] [CrossRef] [PubMed]
- Jubelin, G.; Vianney, A.; Beloin, C.; Ghigo, J.-M.; Lazzaroni, J.-C.; Lejeune, P.; Dorel, C. CpxR/OmpR interplay regulates curli gene expression in response to osmolarity in Escherichia coli. J. Bacteriol. 2005, 187, 2038–2049. [Google Scholar] [CrossRef]
- Khambhati, K.; Patel, J.; Saxena, V.; Jain, N. Gene regulation of biofilm-associated functional amyloids. Pathogens 2021, 10, 490. [Google Scholar] [CrossRef]
- Gophna, U.; Oelschlaeger, T.A.; Hacker, J.; Ron, E.Z. Role of fibronectin in curli-mediated internalization. FEMS MicroBiol. Lett. 2002, 212, 55–58. [Google Scholar] [CrossRef] [PubMed]
- Pawar, D.M.; Rossman, M.L.; Chen, J. Role of curli fimbriae in mediating the cells of enterohaemorrhagic Escherichia coli to attach to abiotic surfaces. J. Appl. Microbiol. 2005, 99, 418–425. [Google Scholar] [CrossRef] [PubMed]
- Saldaña, Z.; Xicohténcatl-Cortes, J.; Avelino, F.; Phillips, A.D.; Kaper, J.B.; Puente, J.L.; Girón, J.A. Synergistic role of curli and cellulose in cell adherence and biofilm formation of attaching and effacing Escherichia coli and identification of Fis as a negative regulator of curli. Environ. Microbiol. 2009, 11, 992–1006. [Google Scholar] [CrossRef]
- Goulter-Thorsen, R.M.; Taran, E.; Gentle, I.R.; Gobius, K.S.; Dykes, G.A. CsgA production by Escherichia coli O157:H7 alters attachment to abiotic surfaces in some growth environments. Appl. Environ. Microbiol. 2011, 77, 7339–7344. [Google Scholar] [CrossRef] [PubMed]
- DeBenedictis, E.P.; Liu, J.; Keten, S. Adhesion mechanisms of curli subunit CsgA to abiotic surfaces. Sci. Adv. 2016, 2. [Google Scholar] [CrossRef]
- Chapman, M.R.; Robinson, L.S.; Pinkner, J.S.; Roth, R.; Heuser, J.; Hammar, M.; Normark, S.; Hultgren, S.J. Role of Escherichia coli curli operons in directing amyloid fiber formation. Science 2002, 295, 851–855. [Google Scholar] [CrossRef] [PubMed]
- Dueholm, M.S.; Petersen, S.V.; Sønderkaer, M.; Larsen, P.; Christiansen, G.; Hein, K.L.; Enghild, J.J.; Nielsen, J.L.; Nielsen, K.L.; Nielsen, P.H.; et al. Functional amyloid in Pseudomonas. Mol. Microbiol. 2010, 77, 1009–1020. [Google Scholar] [CrossRef] [PubMed]
- Alteri, C.J.; Xicohténcatl-Cortes, J.; Hess, S.; Caballero-Olín, G.; Girón, J.A.; Friedman, R.L. Mycobacterium tuberculosis produces pili during human infection. Proc. Natl. Acad. Sci. USA 2007, 104, 5145–5150. [Google Scholar] [CrossRef] [PubMed]
- Elliot, M.A.; Talbot, N.J. Building filaments in the air: Aerial morphogenesis in bacteria and fungi. Curr. Opin. MicroBiol. 2004, 7, 594–601. [Google Scholar] [CrossRef]
- Willey, J.M.; Willems, A.; Kodani, S.; Nodwell, J.R. Morphogenetic surfactants and their role in the formation of aerial hyphae in Streptomyces coelicolor. Mol. Microbiol. 2006, 59, 731–742. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, K.; Syed, A.K.; Stephenson, R.E.; Rickard, A.H.; Boles, B.R. Functional amyloids composed of phenol soluble modulins stabilize Staphylococcus aureus biofilms. PLoS Pathogens 2012, 8, e1002744. [Google Scholar] [CrossRef]
- Romero, D.; Aguilar, C.; Losick, R.; Kolter, R. Amyloid fibers provide structural integrity to Bacillus subtilis biofilms. Proc. Natl. Acad. Sci. USA 2010, 107, 2230–2234. [Google Scholar] [CrossRef]
- Romero, D.; Vlamakis, H.; Losick, R.; Kolter, R. An accessory protein required for anchoring and assembly of amyloid fibres in B. subtilis biofilms. Mol. Microbiol. 2011, 80, 1155–1168. [Google Scholar] [CrossRef]
- Taglialegna, A.; Navarro, S.; Ventura, S.; Garnett, J.A.; Matthews, S.; Penadés, J.R.; Lasa, I.; Valle, J. Staphylococcal Bap proteins build amyloid scaffold biofilm matrices in response to environmental signals. PLoS Pathog 2016, 12, e1005711. [Google Scholar] [CrossRef] [PubMed]
- Taglialegna, A.; Matilla-Cuenca, L.; Dorado-Morales, P.; Navarro, S.; Ventura, S.; Garnett, J.A.; Lasa, I.; Valle, J. The biofilm-associated surface protein Esp of Enterococcus faecalis forms amyloid-like fibers. NPJ Biofilms Microbiomes 2020, 6, 1–12. [Google Scholar] [CrossRef]
- Oli, M.W.; Otoo, H.N.; Crowley, P.J.; Heim, K.P.; Nascimento, M.M.; Ramsook, C.B.; Lipke, P.N.; Brady, L.J. Functional amyloid formation by Streptococcus mutans. Microbiology 2012, 158, 2903–2916. [Google Scholar] [CrossRef]
- Besingi, R.N.; Wenderska, I.B.; Senadheera, D.B.; Cvitkovitch, D.G.; Long, J.R.; Wen, Z.T.; Brady, L.J. Functional amyloids in Streptococcus mutans, their use as targets of biofilm inhibition and initial characterization of SMU_63c. Microbiology 2017, 163, 488–501. [Google Scholar] [CrossRef]
- Dueholm, M.S.; Søndergaard, M.T.; Nilsson, M.; Christiansen, G.; Stensballe, A.; Overgaard, M.T.; Givskov, M.; Tolker-Nielsen, T.; Otzen, D.E.; Nielsen, P.H. Expression of Fap amyloids in Pseudomonas aeruginosa, P. fluorescens, and P. putida results in aggregation and increased biofilm formation. Microbiologyopen 2013, 2, 365–382. [Google Scholar] [CrossRef]
- Rouse, S.L.; Hawthorne, W.J.; Berry, J.-L.; Chorev, D.S.; Ionescu, S.A.; Lambert, S.; Stylianou, F.; Ewert, W.; Mackie, U.; Morgan, R.M.L.; et al. A new class of hybrid secretion system is employed in Pseudomonas amyloid biogenesis. Nat. Commun. 2017, 8, 263. [Google Scholar] [CrossRef] [PubMed]
- Zeng, G.; Vad, B.S.; Dueholm, M.S.; Christiansen, G.; Nilsson, M.; Tolker-Nielsen, T.; Nielsen, P.H.; Meyer, R.L.; Otzen, D.E. Functional bacterial amyloid increases Pseudomonas biofilm hydrophobicity and stiffness. Front. Microbiol. 2015, 6, 1099. [Google Scholar] [CrossRef] [PubMed]
- Perrett, S.; Buell, A.K.; Knowles, T.P.J. Biological and Bio-Inspired Nanomaterials; Springer Nature: Singapore, 2020. [Google Scholar]
- Sawyer, E.B.; Claessen, D.; Haas, M.; Hurgobin, B.; Gras, S.L. The assembly of individual chaplin peptides from Streptomyces coelicolor into functional amyloid fibrils. PLoS ONE 2011, 6, e18839. [Google Scholar] [CrossRef] [PubMed]
- Gebbink, M.F.B.G.; Claessen, D.; Bouma, B.; Dijkhuizen, L.; Wösten, H.A.B. Amyloids—A functional coat for microorganisms. Nat. Rev. Micro 2005, 3, 333–341. [Google Scholar] [CrossRef] [PubMed]
- Dokouhaki, M.; Hung, A.; Day, L.; Gras, S.L. The pH-dependent assembly of Chaplin E from Streptomyces coelicolor. J. Struct. Biol. 2017, 198, 82–91. [Google Scholar] [CrossRef]
- Yang, W.; Willemse, J.; Sawyer, E.B.; Lou, F.; Gong, W.; Zhang, H.; Gras, S.L.; Claessen, D.; Perrett, S. The propensity of the bacterial rodlin protein RdlB to form amyloid fibrils determines its function in Streptomyces coelicolor. Sci. Rep. 2017, 7, 42867. [Google Scholar] [CrossRef]
- Periasamy, S.; Chatterjee, S.S.; Cheung, G.Y.C.; Otto, M. Phenol-soluble modulins in staphylococci: What are they originally for? Commun. Integr. Biol. 2012, 5, 275–277. [Google Scholar] [CrossRef]
- Peschel, A.; Otto, M. Phenol-soluble modulins and staphylococcal infection. Nat. Rev. Micro. 2013, 11, 667–673. [Google Scholar] [CrossRef]
- Cheung, G.Y.C.; Kretschmer, D.; Queck, S.Y.; Joo, H.-S.; Wang, R.; Duong, A.C.; Nguyen, T.H.; Bach, T.-H.L.; Porter, A.R.; DeLeo, F.R.; et al. Insight into structure-function relationship in phenol-soluble modulins using an alanine screen of the phenol-soluble modulin (PSM) α3 peptide. FASEB J. 2014, 28, 153–161. [Google Scholar] [CrossRef] [PubMed]
- Queck, S.Y.; Jameson-Lee, M.; Villaruz, A.E.; Bach, T.-H.L.; Khan, B.A.; Sturdevant, D.E.; Ricklefs, S.M.; Li, M.; Otto, M. RNAIII-independent target gene control by the agr quorum-sensing system: Insight into the evolution of virulence regulation in Staphylococcus aureus. Mol. Cell 2008, 32, 150–158. [Google Scholar] [CrossRef]
- Wang, R.; Braughton, K.R.; Kretschmer, D.; Bach, T.-H.L.; Queck, S.Y.; Li, M.; Kennedy, A.D.; Dorward, D.W.; Klebanoff, S.J.; Peschel, A.; et al. Identification of novel cytolytic peptides as key virulence determinants for community-associated MRSA. Nat. Med. 2007, 13, 1510–1514. [Google Scholar] [CrossRef]
- Chatterjee, S.S.; Joo, H.-S.; Duong, A.C.; Dieringer, T.D.; Tan, V.Y.; Song, Y.; Fischer, E.R.; Cheung, G.Y.C.; Li, M.; Otto, M. Essential Staphylococcus aureus toxin export system. Nat. Med. 2013, 19, 364–367. [Google Scholar] [CrossRef] [PubMed]
- Cogen, A.L.; Yamasaki, K.; Sanchez, K.M.; Dorschner, R.A.; Lai, Y.; MacLeod, D.T.; Torpey, J.W.; Otto, M.; Nizet, V.; Kim, J.E.; et al. Selective antimicrobial action is provided by phenol-soluble modulins derived from Staphylococcus epidermidis, a normal resident of the skin. J. Invest. Dermatol. 2010, 130, 192–200. [Google Scholar] [CrossRef]
- Joo, H.-S.; Cheung, G.Y.C.; Otto, M. Antimicrobial activity of community-associated methicillin-resistant Staphylococcus aureus is caused by phenol-soluble modulin derivatives. J. Biol. Chem. 2011, 286, 8933–8940. [Google Scholar] [CrossRef]
- Nakamura, Y.; Oscherwitz, J.; Cease, K.B.; Chan, S.M.; Muñoz-Planillo, R.; Hasegawa, M.; Villaruz, A.E.; Cheung, G.Y.C.; McGavin, M.J.; Travers, J.B.; et al. Staphylococcus δ-toxin induces allergic skin disease by activating mast cells. Nature 2013, 503, 397–401. [Google Scholar] [CrossRef] [PubMed]
- Richardson, J.R.; Armbruster, N.S.; Günter, M.; Biljecki, M.; Klenk, J.; Heumos, S.; Autenrieth, S.E. PSM Peptides from community-associated methicillin-resistant Staphylococcus aureus impair the adaptive immune response via modulation of dendritic cell subsets in vivo. Front. Immunol. 2019, 10, 995. [Google Scholar] [CrossRef]
- Terra, R.; Stanley-Wall, N.R.; Cao, G.; Lazazzera, B.A. Identification of Bacillus subtilis SipW as a bifunctional signal peptidase that controls surface-adhered biofilm formation. J. Bacteriol. 2012, 194, 2781–2790. [Google Scholar] [CrossRef] [PubMed]
- Chai, L.; Romero, D.; Kayatekin, C.; Akabayov, B.; Vlamakis, H.; Losick, R.; Kolter, R. Isolation, characterization, and aggregation of a structured bacterial matrix precursor. J. Biol. Chem. 2013, 288, 17559–17568. [Google Scholar] [CrossRef] [PubMed]
- Cámara-Almirón, J.; Navarro, Y.; Díaz-Martínez, L.; Magno-Pérez-Bryan, M.C.; Molina-Santiago, C.; Pearson, J.R.; de Vicente, A.; Pérez-García, A.; Romero, D. Dual functionality of the amyloid protein TasA in Bacillus physiology and fitness on the phylloplane. Nat. Commun. 2020, 11, 1–21. [Google Scholar] [CrossRef]
- Lasa, I.; Penadés, J.R. Bap: A family of surface proteins involved in biofilm formation. Res. Microbiol. 2006, 157, 99–107. [Google Scholar] [CrossRef] [PubMed]
- Valle, J.; Fang, X.; Lasa, I. Revisiting Bap multidomain protein: More than sticking bacteria together. Front. Microbiol. 2020, 11, 7490–7499. [Google Scholar] [CrossRef]
- Cucarella, C.; Solano, C.; Valle, J.; Amorena, B.; Lasa, I.; Penadés, J.R. Bap, a Staphylococcus aureus surface protein involved in biofilm formation. J. Bacteriol. 2001, 183, 2888–2896. [Google Scholar] [CrossRef]
- Arrizubieta, M.J.; Toledo-Arana, A.; Amorena, B.; Penadés, J.R.; Lasa, I. Calcium inhibits bap-dependent multicellular behavior in Staphylococcus aureus. J. Bacteriol. 2004, 186, 7490–7498. [Google Scholar] [CrossRef] [PubMed]
- Cucarella, C.; Tormo, M.A.; Ubeda, C.; Trotonda, M.P.; Monzón, M.; Peris, C.; Amorena, B.; Lasa, I.; Penadés, J.R. Role of biofilm-associated protein Bap in the pathogenesis of bovine Staphylococcus aureus. Infect. Immun. 2004, 72, 2177–2185. [Google Scholar] [CrossRef] [PubMed]
- Valle, J.; Latasa, C.; Gil, C.; Toledo-Arana, A.; Solano, C.; Penadés, J.R.; Lasa, I. Bap, a biofilm matrix protein of Staphylococcus aureus prevents cellular internalization through binding to GP96 host receptor. PLoS Pathog. 2012, 8, e1002843. [Google Scholar] [CrossRef]
- Toledo-Arana, A.; Valle, J.; Solano, C.; Arrizubieta, M.J.; Cucarella, C.; Lamata, M.; Amorena, B.; Leiva, J.; Penadés, J.R.; Lasa, I. The enterococcal surface protein, Esp, is involved in Enterococcus faecalis biofilm formation. Appl. Environ. Microbiol. 2001, 67, 4538–4545. [Google Scholar] [CrossRef]
- Shankar, V.; Baghdayan, A.S.; Huycke, M.M.; Lindahl, G.; Gilmore, M.S. Infection-derived Enterococcus faecalis strains are enriched in esp, a gene encoding a novel surface protein. Infect. Immun. 1999, 67, 193–200. [Google Scholar] [CrossRef]
- Tendolkar, P.M.; Baghdayan, A.S.; Shankar, N. The N-terminal domain of enterococcal surface protein, Esp, is sufficient for Esp-mediated biofilm enhancement in Enterococcus faecalis. J. Bacteriol. 2005, 187, 6213–6222. [Google Scholar] [CrossRef]
- Petersen, F.C.; Assev, S.; van der Mei, H.C.; Busscher, H.J.; Scheie, A.A. Functional variation of the antigen I/II surface protein in Streptococcus mutans and Streptococcus intermedius. Infect. Immun. 2002, 70, 249–256. [Google Scholar] [CrossRef]
- Kelemen, L.; Rizk, S.; Debreczeny, M.; Ogier, J.; Szalontai, B. Streptococcal antigen I/II binds to extracellular proteins through intermolecular beta-sheets. FEBS Lett. 2004, 566, 190–194. [Google Scholar] [CrossRef] [PubMed]
- Nobbs, A.H.; Lamont, R.J.; Jenkinson, H.F. Streptococcus adherence and colonization. MicroBiol. Mol. Biol. Rev. 2009, 73, 407–450. [Google Scholar] [CrossRef] [PubMed]
- Heim, K.P.; Sullan, R.M.A.; Crowley, P.J.; El-Kirat-Chatel, S.; Beaussart, A.; Tang, W.; Besingi, R.; Dufrêne, Y.F.; Brady, L.J. Identification of a supramolecular functional architecture of Streptococcus mutans adhesin P1 on the bacterial cell. J. Biol. Chem. 2015, 290, 9002–9019. [Google Scholar] [CrossRef] [PubMed]
- Tang, W.; Bhatt, A.; Smith, A.N.; Crowley, P.J.; Brady, L.J.; Long, J.R. Specific binding of a naturally occurring amyloidogenic fragment of Streptococcus mutans adhesin P1 to intact P1 on the cell surface characterized by solid state NMR spectroscopy. J. BioMol. NMR 2016, 64, 153–164. [Google Scholar] [CrossRef]
- Perov, S.; Lidor, O.; Salinas, N.; Golan, N.; Tayeb-Fligelman, E.; Deshmukh, M.; Willbold, D.; Landau, M. Structural Insights into Curli CsgA cross-β fibril architecture inspire repurposing of anti-amyloid compounds as anti-biofilm agents. PLoS Pathog. 2019, 15, e1007978-31. [Google Scholar] [CrossRef] [PubMed]
- Bleem, A.; Francisco, R.; Bryers, J.D.; Daggett, V. Designed α-sheet peptides suppress amyloid formation in Staphylococcus aureus biofilms. NPJ Biofilms Microbiomes 2017, 3, 16. [Google Scholar] [CrossRef]
- Chen, D.; Li, J.; Pan, T.; Wu, R.; Tao, Y.; Lin, H. The broad-spectrum antibiofilm activity of amyloid-forming hexapeptides. Microb. Biotechnol. 2021, 14, 656–667. [Google Scholar] [CrossRef]
- Tursi, S.A.; Puligedda, R.D.; Szabo, P.; Nicastro, L.K.; Miller, A.L.; Qiu, C.; Gallucci, S.; Relkin, N.R.; Buttaro, B.A.; Dessain, S.K.; et al. Salmonella Typhimurium biofilm disruption by a human antibody that binds a pan-amyloid epitope on curli. Nat. Commun. 2020, 11, 1–13. [Google Scholar] [CrossRef]
- Malishev, R.; Salinas, N.; Gibson, J.; Eden, A.B.; Mieres-Perez, J.; Ruiz-Blanco, Y.B.; Malka, O.; Kolusheva, S.; Klärner, F.-G.; Schrader, T.; et al. Inhibition of Staphylococcus aureus biofilm-forming functional amyloid by molecular tweezers. Cell Chemical. Biol. 2021, 1–17. [Google Scholar] [CrossRef]
- Cegelski, L.; Pinkner, J.S.; Hammer, N.D.; Cusumano, C.K.; Hung, C.S.; Chorell, E.; Aberg, V.; Walker, J.N.; Seed, P.C.; Almqvist, F.; et al. Small-molecule inhibitors target Escherichia coli amyloid biogenesis and biofilm formation. Nat. Chem. Biol. 2009, 5, 913–919. [Google Scholar] [CrossRef]
- Romero, D.; Sanabria-Valentín, E.; Vlamakis, H.; Kolter, R. Biofilm inhibitors that target amyloid proteins. Chem. Biol. 2013, 20, 102–110. [Google Scholar] [CrossRef]
- Serra, D.O.; Mika, F.; Richter, A.M.; Hengge, R. The green tea polyphenol EGCG inhibits E. coli biofilm formation by impairing amyloid curli fibre assembly and down-regulating the biofilm regulator CsgD via the σ E-dependent sRNA RybB. Mol. Microbiol. 2016. [Google Scholar] [CrossRef]
- Najarzadeh, Z.; Mohammad-Beigi, H.; Nedergaard Pedersen, J.; Christiansen, G.; Sønderby, T.V.; Shojaosadati, S.A.; Morshedi, D.; Strømgaard, K.; Meisl, G.; Sutherland, D.; et al. Plant polyphenols inhibit functional amyloid and biofilm formation in Pseudomonas strains by directing monomers to off-pathway oligomers. Biomolecules 2019, 9, 659. [Google Scholar] [CrossRef] [PubMed]
- Stenvang, M.; Dueholm, M.S.; Vad, B.S.; Seviour, T.W.; Zeng, G.; Geifman-Shochat, S.; Søndergaard, M.T.; Christiansen, G.; Meyer, R.L.; Kjelleberg, S.; et al. Epigallocatechin gallate remodels overexpressed functional amyloids in Pseudomonas aeruginosa and increases biofilm susceptibility to antibiotic treatment. J. Biol. Chem. 2016. [Google Scholar] [CrossRef] [PubMed]
- Marinelli, P.; Pallares, I.; Navarro, S.; Ventura, S. Dissecting the contribution of Staphylococcus aureus α-phenol-soluble modulins to biofilm amyloid structure. Sci. Rep. 2016, 6, 34552. [Google Scholar] [CrossRef]
- Pruteanu, M.; Hernández Lobato, J.I.; Stach, T.; Hengge, R. Common plant flavonoids prevent the assembly of amyloid curli fibres and can interfere with bacterial biofilm formation. Environ. Microbiol. 2020, 22, 5280–5299. [Google Scholar]
- Matilla-Cuenca, L.; Gil, C.; Cuesta, S.; Rapun-Araiz, B.; Žiemytė, M.; Mira, A.; Lasa, I.; Valle, J. Antibiofilm activity of flavonoids on staphylococcal biofilms through targeting BAP amyloids. Sci. Rep. 2020, 10, 1–12. [Google Scholar] [CrossRef]
- Ventura, S. Curing bacterial infections with protein aggregates. Mol. Microbiol. 2016, 99, 827–830. [Google Scholar] [CrossRef]
- Bednarska, N.G.; Van Eldere, J.; Gallardo, R.; Ganesan, A.; Ramakers, M.; Vogel, I.; Baatsen, P.; Staes, A.; Goethals, M.; Hammarström, P.; et al. Protein aggregation as an antibiotic design strategy. Mol. Microbiol. 2016, 99, 849–865. [Google Scholar] [CrossRef] [PubMed]
- Heisig, M.; Abraham, N.M.; Liu, L.; Neelakanta, G.; Mattessich, S.; Sultana, H.; Shang, Z.; Ansari, J.M.; Killiam, C.; Walker, W.; et al. Antivirulence properties of an antifreeze protein. Cell Rep. 2014, 9, 417–424. [Google Scholar] [CrossRef] [PubMed]
- Ansari, J.M.; Abraham, N.M.; Massaro, J.; Murphy, K.; Smith-Carpenter, J.; Fikrig, E. Anti-biofilm activity of a self-aggregating peptide against Streptococcus mutans. Front. Microbiol. 2017, 8, 488. [Google Scholar] [CrossRef] [PubMed]
- Hamilton, J.A.; Benson, M.D. Transthyretin: A review from a structural perspective. Cell Mol. Life Sci. 2001, 58, 1491–1521. [Google Scholar] [CrossRef]
- Taylor, J.D.; Hawthorne, W.J.; Lo, J.; Dear, A.; Jain, N.; Meisl, G.; Andreasen, M.; Fletcher, C.; Koch, M.; Darvill, N.; et al. Electrostatically-guided inhibition of curli amyloid nucleation by the CsgC-like family of chaperones. Sci. Rep. 2016, 6, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhang, X.; Ladiwala, A.R.A.; Du, D.; Yadav, J.K.; Tessier, P.M.; Wright, P.E.; Kelly, J.W.; Buxbaum, J.N. Mechanisms of transthyretin inhibition of β-amyloid aggregation in vitro. J. Neurosci. 2013, 33, 19423–19433. [Google Scholar] [CrossRef]
- Cascella, R.; Conti, S.; Mannini, B.; Li, X.; Buxbaum, J.N.; Tiribilli, B.; Chiti, F.; Cecchi, C. Transthyretin suppresses the toxicity of oligomers formed by misfolded proteins in vitro. BBA Mol. Basis Dis. 2013, 1832, 2302–2314. [Google Scholar] [CrossRef] [PubMed]
- Wasana Jayaweera, S.; Surano, S.; Pettersson, N.; Oskarsson, E.; Lettius, L.; Gharibyan, A.L.; Anan, I.; Olofsson, A. Mechanisms of transthyretin inhibition of IAPP amyloid formation. Biomolecules 2021, 11, 411. [Google Scholar] [CrossRef]
- Jain, N.; Åden, J.; Nagamatsu, K.; Evans, M.L.; Li, X.; McMichael, B.; Ivanova, M.I.; Almqvist, F.; Buxbaum, J.N.; Chapman, M.R. Inhibition of curli assembly and Escherichia coli biofilm formation by the human systemic amyloid precursor transthyretin. Proc. Natl. Acad. Sci. USA 2017, 114, 12184–12189. [Google Scholar] [CrossRef] [PubMed]
- Panza, F.; Lozupone, M.; Seripa, D.; Imbimbo, B.P. Amyloid-β immunotherapy for Alzheimer disease: Is it now a long shot? Ann. Neurol. 2019, 85, 303–315. [Google Scholar] [CrossRef]
- Fu, H.J.; Liu, B.; Frost, J.L.; Lemere, C.A. Amyloid-β immunotherapy for Alzheimer’s disease. CNS Neurol. Disord. Drug Targets 2010, 9, 197–206. [Google Scholar] [CrossRef] [PubMed]
- Härd, T.; Lendel, C. Inhibition of amyloid formation. J. Mol. Biol. 2012, 421, 441–465. [Google Scholar] [CrossRef]
- Knowles, T.P.J.; Vendruscolo, M.; Dobson, C.M. The amyloid state and its association with protein misfolding diseases. Nat. Rev. Mol. Cell Biol. 2014, 15, 384–396. [Google Scholar] [CrossRef]
- Mbarek, A.; Moussa, G.; Chain, J.L. Pharmaceutical applications of molecular tweezers, clefts and clips. Molecules 2019, 24, 1803. [Google Scholar] [CrossRef] [PubMed]
- Hadrovic, I.; Rebmann, P.; Klärner, F.-G.; Bitan, G.; Schrader, T. Molecular lysine tweezers counteract aberrant protein aggregation. Front. Chem. 2019, 7, 657. [Google Scholar] [CrossRef] [PubMed]
- Di, J.; Siddique, I.; Li, Z.; Malki, G.; Hornung, S.; Dutta, S.; Hurst, I.; Ishaaya, E.; Wang, A.; Tu, S.; et al. The molecular tweezer CLR01 improves behavioral deficits and reduces tau pathology in P301S-tau transgenic mice. Alzheimer’s Res. Ther. 2020, 13, 1–20. [Google Scholar]
- Pinkner, J.S.; Remaut, H.; Buelens, F.; Miller, E.; Aberg, V.; Pemberton, N.; Hedenström, M.; Larsson, A.; Seed, P.; Waksman, G.; et al. Rationally designed small compounds inhibit pilus biogenesis in uropathogenic bacteria. Proc. Natl. Acad. Sci. USA 2006, 103, 17897–17902. [Google Scholar] [CrossRef]
- Aberg, V.; Norman, F.; Chorell, E.; Westermark, A.; Olofsson, A.; Elisabeth Sauer-Eriksson, A.; Almqvist, F. Microwave-assisted decarboxylation of bicyclic 2-pyridone scaffolds and identification of Aβ-peptide aggregation inhibitors. Org. BioMol. Chem. 2005, 3, 2817–2823. [Google Scholar] [CrossRef] [PubMed]
- Horvath, I.; Weise, C.F.; Andersson, E.K.; Chorell, E.; Sellstedt, M.; Bengtsson, C.; Olofsson, A.; Hultgren, S.J.; Chapman, M.; Wolf-Watz, M.; et al. Mechanisms of protein oligomerization: Inhibitor of functional amyloids templates α-synuclein fibrillation. J. Am. Chem. Soc. 2012, 134, 3439–3444. [Google Scholar] [CrossRef] [PubMed]
- Velander, P.; Wu, L.; Henderson, F.; Zhang, S.; Bevan, D.R.; Xu, B. Natural product-based amyloid inhibitors. Biochem. Pharmacol. 2017, 139, 40–55. [Google Scholar] [CrossRef]
- Freyssin, A.; Page, G.; Fauconneau, B.; Rioux Bilan, A. Natural polyphenols effects on protein aggregates in Alzheimer’s and Parkinson’s prion-like diseases. Neural. Regen Res. 2018, 13, 955–961. [Google Scholar]
- Kobayashi, H.; Murata, M.; Kawanishi, S.; Oikawa, S. Polyphenols with anti-amyloid β aggregation show potential risk of toxicity via pro-oxidant properties. Int. J. Mol. Sci. 2020, 21, 3561. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Liu, Y.; Wang, S.; Dong, S.; Chang, P.; Jiang, Z. Structural characteristics of (−)-epigallocatechin-3-gallate inhibiting amyloid Aβ42 aggregation and remodeling amyloid fibers. RSC Adv. 2015, 5, 62402–62413. [Google Scholar] [CrossRef]
- Bieschke, J.; Russ, J.; Friedrich, R.P.; Ehrnhoefer, D.E.; Wobst, H.; Neugebauer, K.; Wanker, E.E. EGCG remodels mature alpha-synuclein and amyloid-beta fibrils and reduces cellular toxicity. Proc. Natl. Acad. Sci. USA 2010, 107, 7710–7715. [Google Scholar] [CrossRef] [PubMed]
- Ehrnhoefer, D.E.; Bieschke, J.; Boeddrich, A.; Herbst, M.; Masino, L.; Lurz, R.; Engemann, S.; Pastore, A.; Wanker, E.E. EGCG redirects amyloidogenic polypeptides into unstructured, off-pathway oligomers. Nat. Struct. Mol. Biol. 2008, 15, 558–566. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Zhang, Y.; Quan, Z.; Wong, W.; Guo, J.; Zhang, R.; Yang, Q.; Dai, R.; McGeer, P.L.; Qing, H. Epigallocatechin gallate (EGCG) inhibits alpha-synuclein aggregation: A potential agent for Parkinson’s disease. Neurochem. Res. 2016, 41, 2788–2796. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Liang, Q.; Sun, Q.; Chen, C.; Xu, L.; Ding, Y.; Zhou, P. (−)-Epigallocatechin-3-gallate (EGCG) inhibits fibrillation, disaggregates amyloid fibrils of α-synuclein, and protects PC12 cells against α-synuclein-induced toxicity. RSC Adv. 2017, 7, 32508–32517. [Google Scholar] [CrossRef]
- Roy, S.; Bhat, R. Suppression, disaggregation, and modulation of γ-Synuclein fibrillation pathway by green tea polyphenol EGCG. Protein Sci. 2019, 28, 382–402. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.-H.; Lin, Y.; Cox, S.J.; Kinoshita, M.; Sahoo, B.R.; Ivanova, M.; Ramamoorthy, A. Zinc boosts EGCG’s hIAPP amyloid inhibition both in solution and membrane. BBA Proteins Proteom. 2019, 1867, 529–536. [Google Scholar] [CrossRef]
- Xu, Z.-X.; Ma, G.-L.; Zhang, Q.; Chen, C.-H.; He, Y.-M.; Xu, L.-H.; Zhou, G.-R.; Li, Z.-H.; Yang, H.-J.; Zhou, P. Inhibitory mechanism of epigallocatechin gallate on fibrillation and aggregation of amidated human islet amyloid polypeptide. Chem. Phys. Chem. 2017, 18, 1611–1619. [Google Scholar] [CrossRef]
- Ehrnhoefer, D.E.; Duennwald, M.; Markovic, P.; Wacker, J.L.; Engemann, S.; Roark, M.; Legleiter, J.; Marsh, J.L.; Thompson, L.M.; Lindquist, S.; et al. Green tea (−)-epigallocatechin-gallate modulates early events in huntingtin misfolding and reduces toxicity in Huntington’s disease models. Hum. Mol. Genet. 2006, 15, 2743–2751. [Google Scholar] [CrossRef]
- Wobst, H.J.; Sharma, A.; Diamond, M.I.; Wanker, E.E.; Bieschke, J. The green tea polyphenol (−)-epigallocatechin gallate prevents the aggregation of tau protein into toxic oligomers at substoichiometric ratios. FEBS Lett. 2014, 589, 77–83. [Google Scholar] [CrossRef]
- Palhano, F.L.; Lee, J.; Grimster, N.P.; Kelly, J.W. Toward the molecular mechanism(s) by which EGCG treatment remodels mature amyloid fibrils. J. Am. Chem. Soc. 2013, 135, 7503–7510. [Google Scholar] [CrossRef] [PubMed]
- Najarzadeh, Z.; Pedersen, J.N.; Christiansen, G.; Shojaosadati, S.A.; Pedersen, J.S.; Otzen, D.E. Bacterial amphiphiles as amyloid inducers_ effect of rhamnolipid and lipopolysaccharide on FapC fibrillation. BBA Proteins Proteom. 2019, 1867, 140263. [Google Scholar] [CrossRef]
- Bamunuarachchi, N.I.; Khan, F.; Kim, Y.M. Inhibition of virulence factors and biofilm formation of Acinetobacter baumannii by naturally-derived and synthetic drugs. Curr. Drug Targets 2021, 22, 734–759. [Google Scholar] [CrossRef] [PubMed]
- O’May, C.; Ciobanu, A.; Lam, H.; Tufenkji, N. Tannin derived materials can block swarming motility and enhance biofilm formation in Pseudomonas aeruginosa. Biofouling 2012, 28, 1063–1076. [Google Scholar] [CrossRef] [PubMed]
- Bikels-Goshen, T.; Landau, E.; Saguy, S.; Shapira, R. Staphylococcal strains adapted to epigallocathechin gallate (EGCG) show reduced susceptibility to vancomycin, oxacillin and ampicillin, increased heat tolerance, and altered cell morphology. Int. J. Food Microbiol. 2010, 138, 26–31. [Google Scholar] [CrossRef]
- Miller, A.L.; Bessho, S.; Grando, K.; Tükel, C. Microbiome or infections: Amyloid-containing biofilms as a trigger for complex human diseases. Front. Immunol. 2021, 12, 638867. [Google Scholar] [CrossRef]
| Amyloid Type | Locus | Amyloid Subunits | Bacteria | |
|---|---|---|---|---|
| Intrinsic amyloids | ||||
| Curli | csgDEFG; csgBAC | CsgA, CsgB | E. coli, Salmonella | [40] |
| Fap | fapABCDEF | FapC, FapB, | P. aeruginosa | [41] |
| MTP | mtp | MTP | Mycobacterium tuberculosis | [42] |
| Chaplins/Rodlins | chpA, chpD, rdlA, rdlB; chpC, chpH; chpF, chpG; chpB, chpE | ChpD-H, RdlB | Streptomyces coelicolor | [43,44] |
| Facultative amyloids | ||||
| PSMs | psmα1–4; psmβ1–2; hld | PSMα, PSMβ, δ-toxin | S. aureus, CNS 1 | [17,45] |
| TasA | tapA, sipW, tasA | TasA | B. subtilis, B. cereus | [46,47] |
| Bap | bap | BapB-domain | S. aureus, CNS 1 | [48] |
| Esp | esp | N-terminal domain | E. faecalis | [49] |
| P1 | p1 | AgII-C123 region | S. mutans | [50] |
| WapA | wapA | WapA | S. mutans | [51] |
| SMU_63C | smu_63c | SMU_63C | S. mutans | [51] |
| Type | Amyloid Inhibitors | Anti-Biofilm Effect | Bacterial Amyloid Target | Eukaryotic Amyloid Target | Reference |
|---|---|---|---|---|---|
| Peptides | ANK6 | S. Typhimurium | CsgA | Aβ | [87] |
| DB3DB3 | S. Typhimurium | CsgA | Aβ | [87] | |
| AP90 | S. aureus | PSMα1 | ND | [88] | |
| AFhPs | S. mutans, S. sanguis, S. aureus, E. coli | ND | ND | [89] | |
| P1 | S. mutans | ND | ND | [89] | |
| Proteins | TTR | E. coli | CsgA | Aβ, HepF-N | [89] |
| B. subtilis | ND | ||||
| Antibodies | 3H3 | S. Typhimurium | Curli | Aβ, TTR, Tau | [90] |
| Molecular tweezers | CLR01 | S. aureus | PSMα1 | Aβ, α-syn, Tau | [91] |
| CLR05 | S. aureus | PSMα1 | Aβ, α-syn, Tau | [91] | |
| Curlicides | FN075 | E. coli | Curli | Aβ | [92] |
| BibC6 | E. coli | Curli | ND | [92] | |
| VA028 | E. coli | Curli | ND | [92] | |
| Bioactive compounds | AA-861 | B. subtilis | TasA | New1 | [93] |
| S. mutans | P1 WapA | [92] | |||
| Parthenolide | B. subtilis | TasA | New1 | [93] | |
| Polyphenols | EGCG | E. coli | CsgA CsgB | Aβ, α-syn, Tau | [94] |
| Pseudomonas sp. | FapC | [95,96] | |||
| S. mutans | P1 WapA SMU_63c | [89] | |||
| S. aureus | PSMα1PSMα4 | [97] | |||
| PGG | Pseudomonas sp. | FapC | Aβ | [95] | |
| Tannic acid | S. mutans | P1 WapA | Prion PrP, Aβ | [51] | |
| Luteolin | E. coli | CsgA | Aβ, α-syn | [98] | |
| Morin | E. coli | CsgA | Aβ, α-syn | [98] | |
| Myricetin | E. coli,S. aureus | CsgA Bap | Aβ, α-syn | [98,99] | |
| Quercetin | E. coli | CsgA Bap | Aβ, α-syn | [98,99] | |
| Phloretin | E. coli | CsgA CsgB | Aβ, αSA53T | [98] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Matilla-Cuenca, L.; Toledo-Arana, A.; Valle, J. Anti-Biofilm Molecules Targeting Functional Amyloids. Antibiotics 2021, 10, 795. https://doi.org/10.3390/antibiotics10070795
Matilla-Cuenca L, Toledo-Arana A, Valle J. Anti-Biofilm Molecules Targeting Functional Amyloids. Antibiotics. 2021; 10(7):795. https://doi.org/10.3390/antibiotics10070795
Chicago/Turabian StyleMatilla-Cuenca, Leticia, Alejandro Toledo-Arana, and Jaione Valle. 2021. "Anti-Biofilm Molecules Targeting Functional Amyloids" Antibiotics 10, no. 7: 795. https://doi.org/10.3390/antibiotics10070795
APA StyleMatilla-Cuenca, L., Toledo-Arana, A., & Valle, J. (2021). Anti-Biofilm Molecules Targeting Functional Amyloids. Antibiotics, 10(7), 795. https://doi.org/10.3390/antibiotics10070795






