Probiotic Lactobacillus spp. Act Against Helicobacter pylori-induced Inflammation
Abstract
1. Introduction
2. Materials and Methods
2.1. Chemicals and Antibodies
2.2. Cell Culture
2.3. Identification of Lactobacillus Strains that Possess Probiotic Activities
2.4. Bacterial Strains and Culture
2.5. Determination of Anti-H. pylori Activity by Lactobacillus Strains
2.6. Assays for H. pylori Adhesion and Invasion to AGS Cells
2.7. NF-κB Reporter Luciferase Assay
2.8. Measurement of Interleukin (IL)-8 Production
2.9. H. pylori CagA Translocation and Phosphorylation Assay
2.10. Animal Study and Immunohistochemistry Analysis
2.11. Stool Collection
2.12. Analysis of Gut Specific Bacteria using Quantitative Real-time PCR (qRT-PCR)
2.13. Statistical Analysis
3. Results
3.1. Lactobacillus spp. Inhibits MDR-H. Pylori Growth
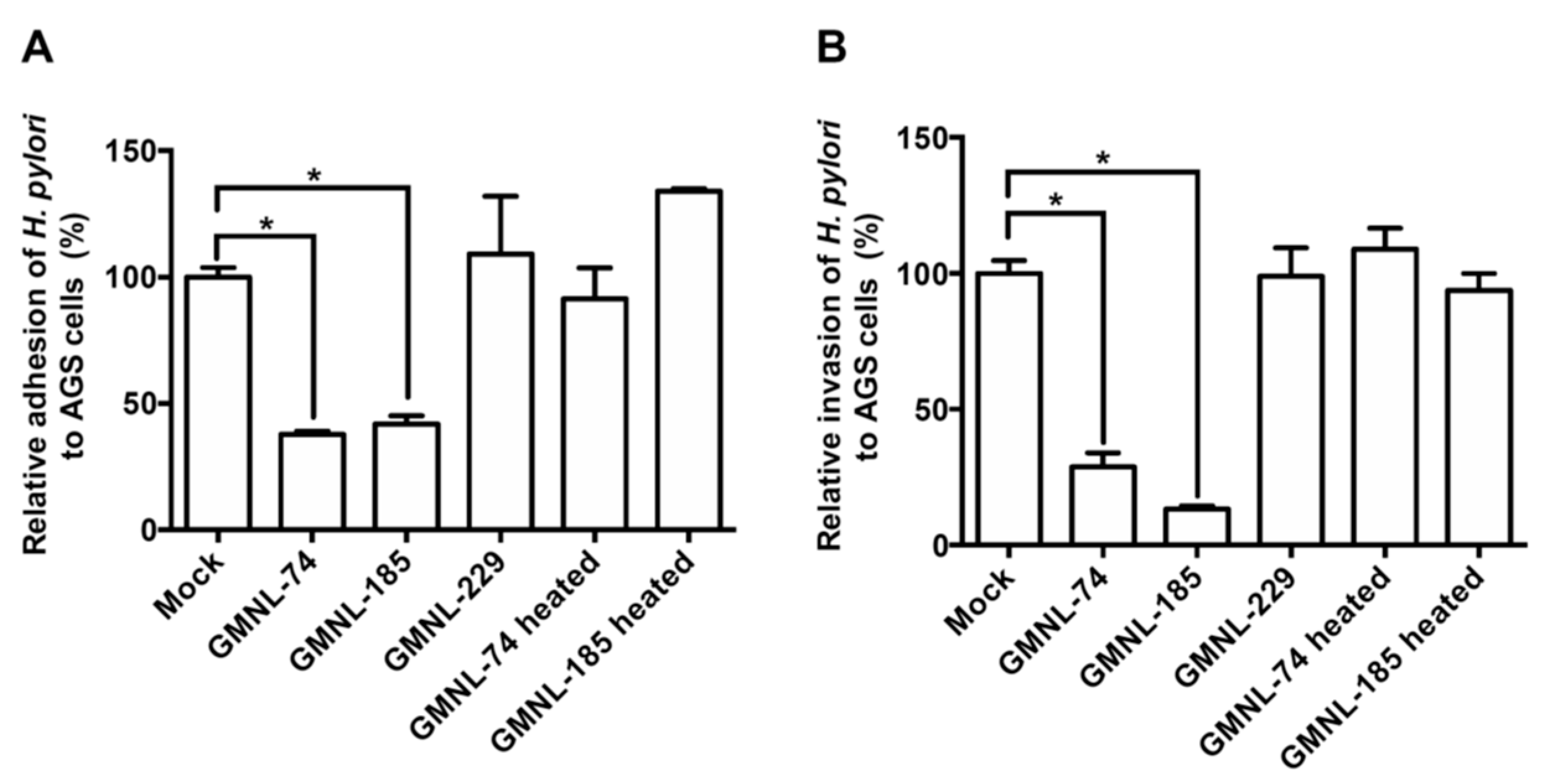
3.2. Lactobacillus spp. Suppresses H. pylori Adhesion and Invasion Activities
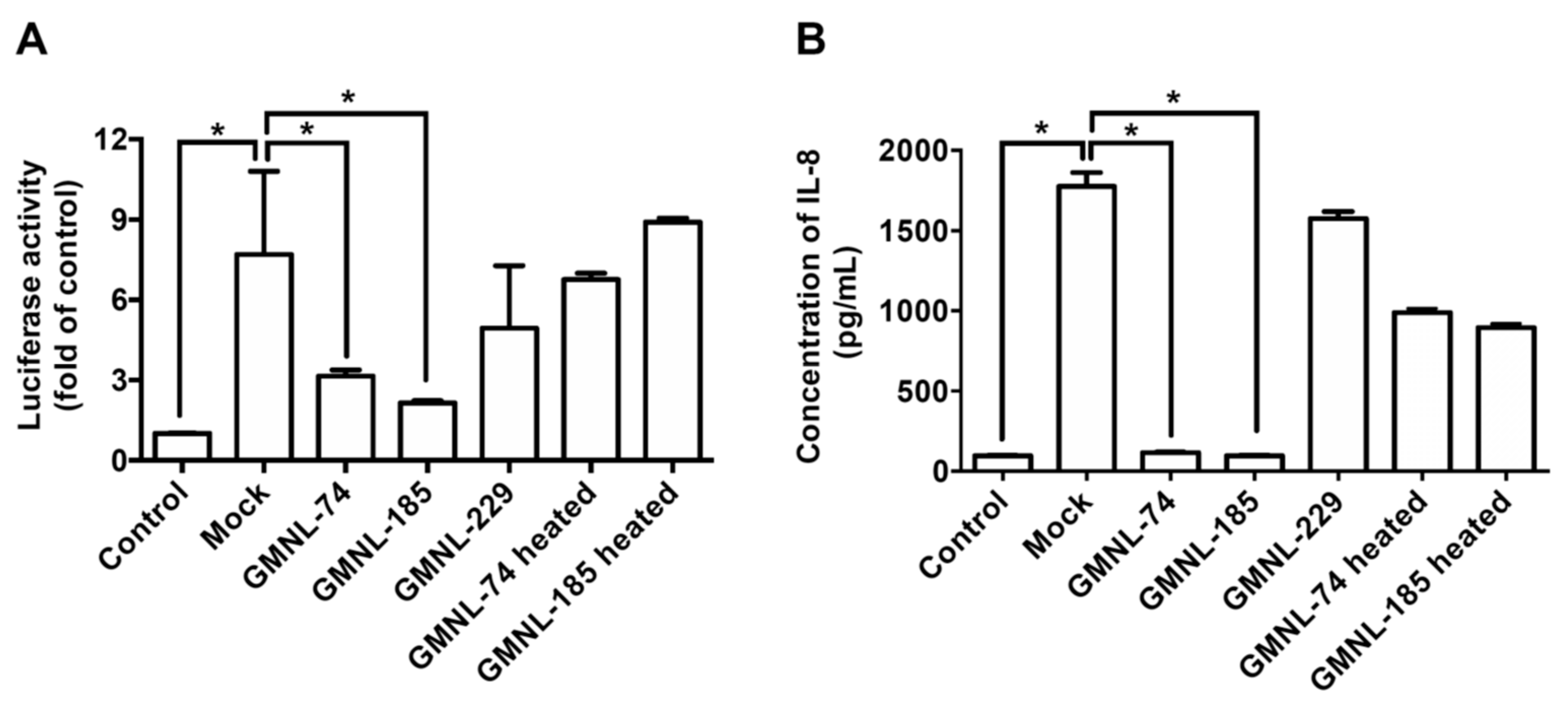
3.3. Lactobacillus spp. Ameliorates H. pylori-induced Inflammation
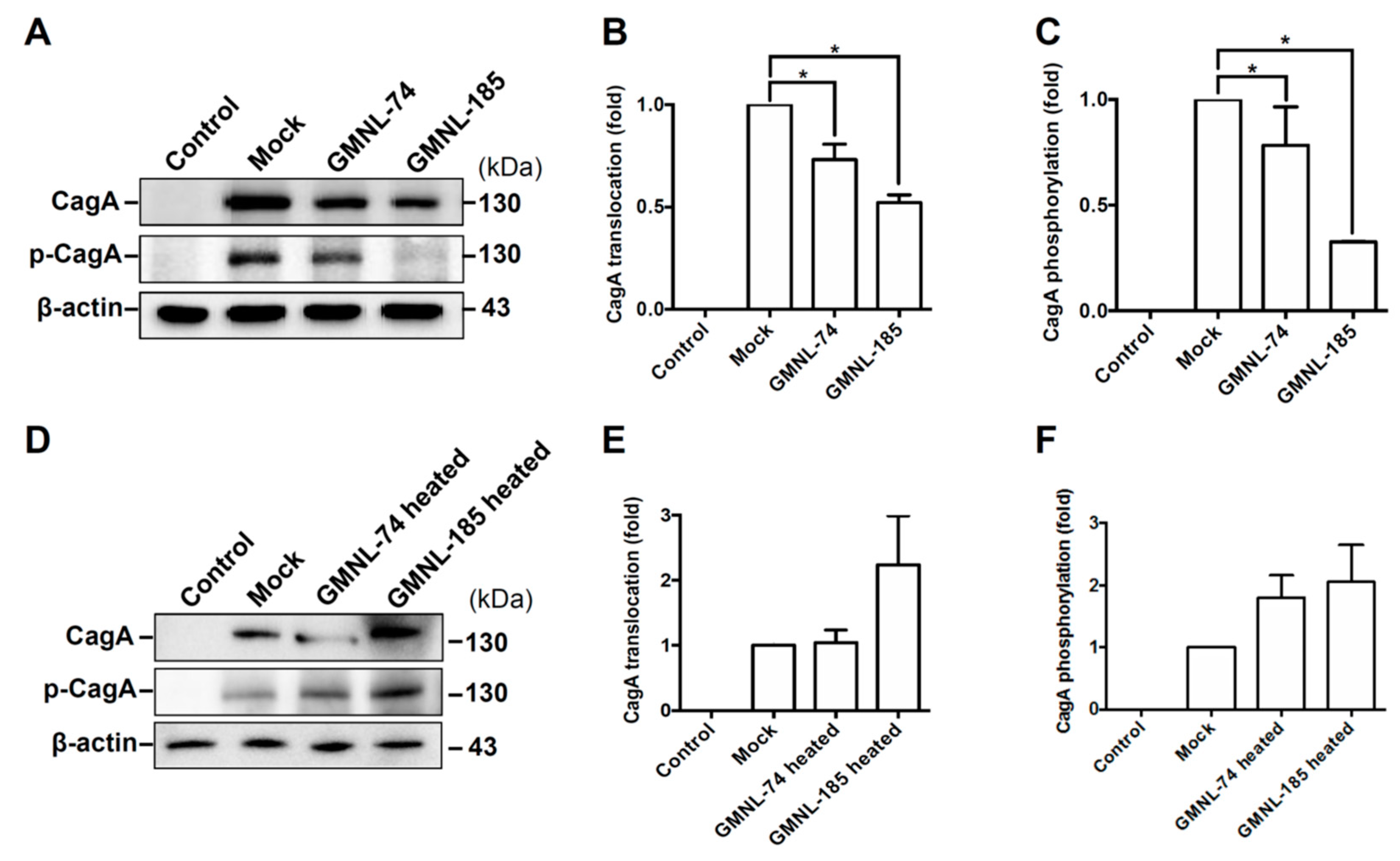
3.4. Lactobacillus spp. Attenuates H. pylori CagA Translocation and Phosphorylation
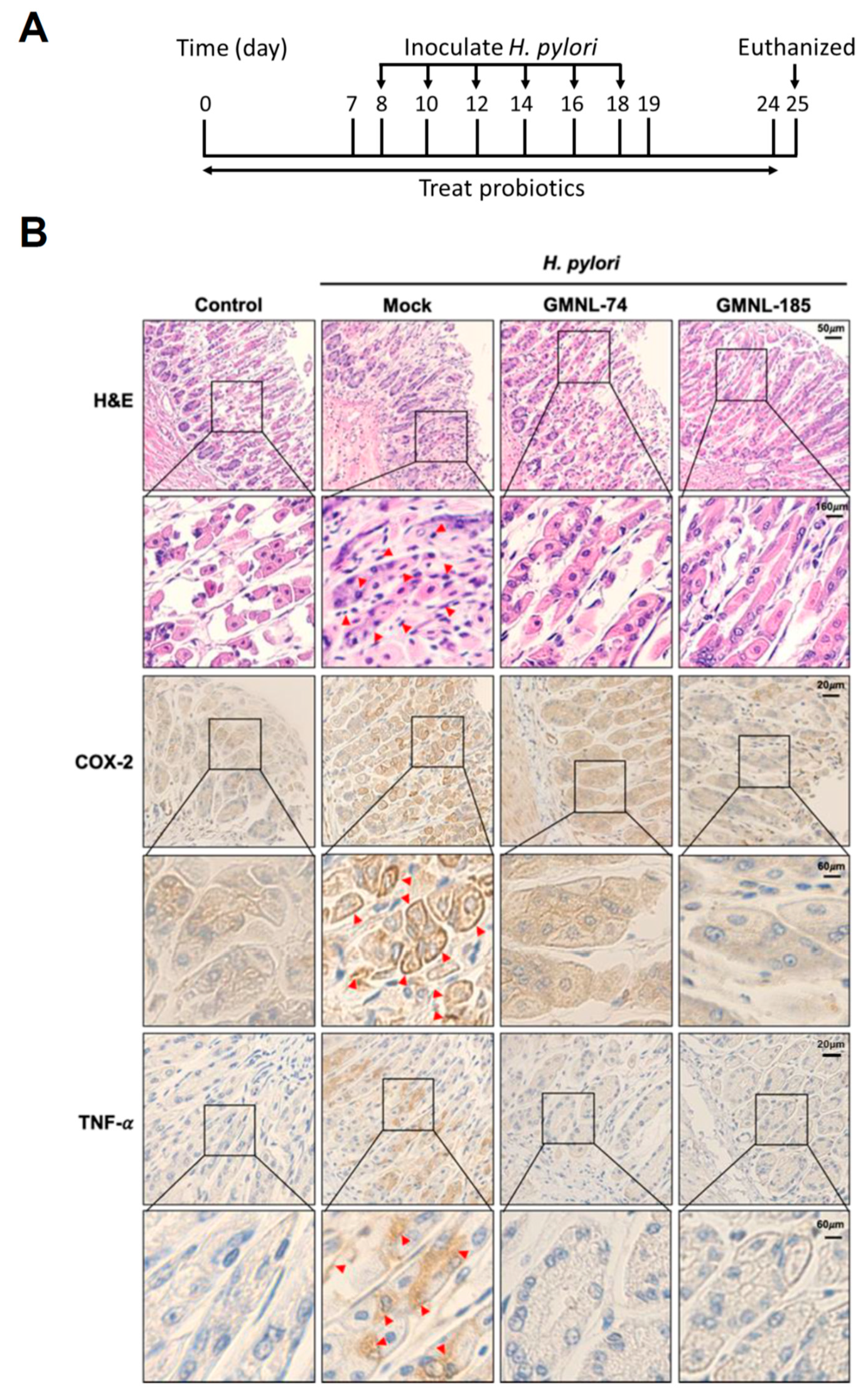
3.5. Lactobacillus spp. Suppresses Gastric Inflammation in Mice Infected with H. pylori
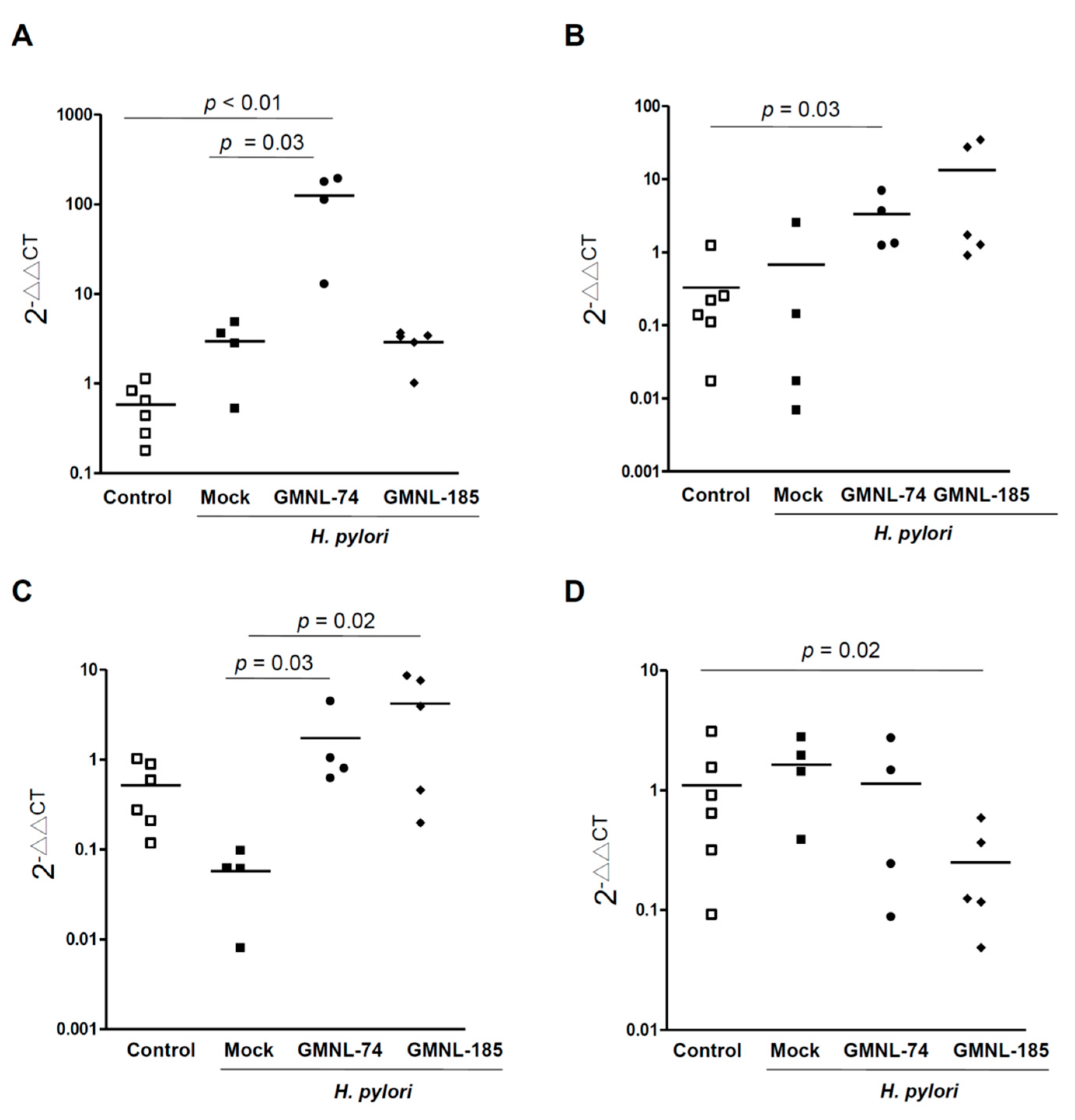
3.6. Changes in the Specific Microbiota Members by Treatment with Lactobacillus
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hooi, J.K.Y.; Lai, W.Y.; Ng, W.K.; Suen, M.M.Y.; Underwood, F.E.; Tanyingoh, D.; Malfertheiner, P.; Graham, D.Y.; Wong, V.W.S.; Wu, J.C.Y.; et al. Global Prevalence of Helicobacter pylori Infection: Systematic Review and Meta-Analysis. Gastroenterology 2017, 153, 420–429. [Google Scholar] [CrossRef]
- Eusebi, L.H.; Zagari, R.M.; Bazzoli, F. Epidemiology of Helicobacter pylori infection. Helicobacter 2014, 19 (Suppl. 1), 1–5. [Google Scholar] [CrossRef]
- Venerito, M.; Vasapolli, R.; Rokkas, T.; Delchier, J.C.; Malfertheiner, P. Helicobacter pylori, gastric cancer and other gastrointestinal malignancies. Helicobacter 2017, 22 (Suppl. 1), e12413. [Google Scholar] [CrossRef] [PubMed]
- Marshall, B. Helicobacter pylori: 20 years on. Clin. Med. (Lond.) 2002, 2, 147–152. [Google Scholar] [CrossRef] [PubMed]
- Polk, D.B.; Peek, R.M., Jr. Helicobacter pylori: Gastric cancer and beyond. Nat. Rev. Cancer 2010, 10, 403–414. [Google Scholar] [CrossRef] [PubMed]
- Covacci, A.; Rappuoli, R. Tyrosine-phosphorylated bacterial proteins: Trojan horses for the host cell. J. Exp. Med. 2000, 191, 587–592. [Google Scholar] [CrossRef] [PubMed]
- Tegtmeyer, N.; Neddermann, M.; Asche, C.I.; Backert, S. Subversion of host kinases: A key network in cellular signaling hijacked by Helicobacter pylori CagA. Mol. Microbiol. 2017, 105, 358–372. [Google Scholar] [CrossRef]
- Brandt, S.; Kwok, T.; Hartig, R.; Konig, W.; Backert, S. NF-kappaB activation and potentiation of proinflammatory responses by the Helicobacter pylori CagA protein. Proc. Natl. Acad. Sci. USA 2005, 102, 9300–9305. [Google Scholar] [CrossRef]
- Van Doorn, L.J.; Figueiredo, C.; Sanna, R.; Plaisier, A.; Schneeberger, P.; de Boer, W.; Quint, W. Clinical relevance of the cagA, vacA, and iceA status of Helicobacter pylori. Gastroenterology 1998, 115, 58–66. [Google Scholar] [CrossRef]
- Kim, S.H.; Woo, H.; Park, M.; Rhee, K.J.; Moon, C.; Lee, D.; Seo, W.D.; Kim, J.B. Cyanidin 3-O-glucoside reduces Helicobacter pylori VacA-induced cell death of gastric KATO III cells through inhibition of the SecA pathway. Int. J. Med. Sci. 2014, 11, 742–747. [Google Scholar] [CrossRef]
- Cover, T.L.; Blanke, S.R. Helicobacter pylori VacA, a paradigm for toxin multifunctionality. Nat. Rev. Microbiol. 2005, 3, 320–332. [Google Scholar] [CrossRef] [PubMed]
- Backert, S.; Neddermann, M.; Maubach, G.; Naumann, M. Pathogenesis of Helicobacter pylori infection. Helicobacter 2016, 21 (Suppl. 1), 19–25. [Google Scholar] [CrossRef]
- Malfertheiner, P.; Megraud, F.; O’Morain, C.A.; Gisbert, J.P.; Kuipers, E.J.; Axon, A.T.; Bazzoli, F.; Gasbarrini, A.; Atherton, J.; Graham, D.Y.; et al. Management of Helicobacter pylori infection-the Maastricht V/Florence Consensus Report. Gut 2017, 66, 6–30. [Google Scholar] [CrossRef] [PubMed]
- O’Connor, A.; Lamarque, D.; Gisbert, J.P.; O’Morain, C. Treatment of Helicobacter pylori infection 2017. Helicobacter 2017, 22 (Suppl. 1). [Google Scholar] [CrossRef]
- O’Morain, N.R.; Dore, M.P.; O’Connor, A.J.P.; Gisbert, J.P.; O’Morain, C.A. Treatment of Helicobacter pylori infection in 2018. Helicobacter 2018, 23 (Suppl. 1), e12519. [Google Scholar] [CrossRef] [PubMed]
- Alba, C.; Blanco, A.; Alarcon, T. Antibiotic resistance in Helicobacter pylori. Curr. Opin. Infect. Dis. 2017, 30, 489–497. [Google Scholar] [CrossRef] [PubMed]
- Nenci, A.; Becker, C.; Wullaert, A.; Gareus, R.; van Loo, G.; Danese, S.; Huth, M.; Nikolaev, A.; Neufert, C.; Madison, B.; et al. Epithelial NEMO links innate immunity to chronic intestinal inflammation. Nature 2007, 446, 557–561. [Google Scholar] [CrossRef] [PubMed]
- Are, A.; Aronsson, L.; Wang, S.; Greicius, G.; Lee, Y.K.; Gustafsson, J.A.; Pettersson, S.; Arulampalam, V. Enterococcus faecalis from newborn babies regulate endogenous PPARgamma activity and IL-10 levels in colonic epithelial cells. Proc. Natl. Acad. Sci. USA 2008, 105, 1943–1948. [Google Scholar] [CrossRef] [PubMed]
- Patel, R.M.; Lin, P.W. Developmental biology of gut-probiotic interaction. Gut Microbes 2010, 1, 186–195. [Google Scholar] [CrossRef]
- Servin, A.L. Antagonistic activities of lactobacilli and bifidobacteria against microbial pathogens. FEMS Microbiol. Rev. 2004, 28, 405–440. [Google Scholar] [CrossRef] [PubMed]
- De Keersmaecker, S.C.; Verhoeven, T.L.; Desair, J.; Marchal, K.; Vanderleyden, J.; Nagy, I. Strong antimicrobial activity of Lactobacillus rhamnosus GG against Salmonella typhimurium is due to accumulation of lactic acid. FEMS Microbiol. Lett. 2006, 259, 89–96. [Google Scholar] [CrossRef] [PubMed]
- Prasad, J.; Gill, H.; Smart, J.; Gopal, P.K. Selection and characterisation of Lactobacillus and Bifidobacterium strains for use as probiotics. Int. Dairy J. 1998, 8, 993–1002. [Google Scholar] [CrossRef]
- Presti, I.; D’Orazio, G.; Labra, M.; La Ferla, B.; Mezzasalma, V.; Bizzaro, G.; Giardina, S.; Michelotti, A.; Tursi, F.; Vassallo, M.; et al. Evaluation of the probiotic properties of new Lactobacillus and Bifidobacterium strains and their in vitro effect. Appl. Microbiol. Biotech. 2015, 99, 5613–5626. [Google Scholar] [CrossRef] [PubMed]
- Turkova, K.; Mavric, A.; Narat, M.; Rittich, B.; Spanova, A.; Rogelj, I.; Matijasic, B.B. Evaluation of Lactobacillus strains for selected probiotic properties. Folia Microbiol. 2013, 58, 261–267. [Google Scholar] [CrossRef] [PubMed]
- Lievin-Le Moal, V.; Servin, A.L. Anti-infective activities of lactobacillus strains in the human intestinal microbiota: From probiotics to gastrointestinal anti-infectious biotherapeutic agents. Clin. Microbiol. Rev. 2014, 27, 167–199. [Google Scholar] [CrossRef]
- Ribeiro, L.A.; Azevedo, V.; Le Loir, Y.; Oliveira, S.C.; Dieye, Y.; Piard, J.C.; Gruss, A.; Langella, P. Production and targeting of the Brucella abortus antigen L7/L12 in Lactococcus lactis: A first step towards food-grade live vaccines against brucellosis. Appl. Environ. Microbiol. 2002, 68, 910–916. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.H.; Roussel, Y.; Wilks, M.; Tabaqchali, S. Expression of Helicobacter pylori urease subunit B gene in Lactococcus lactis MG1363 and its use as a vaccine delivery system against H. pylori infection in mice. Vaccine 2001, 19, 3927–3935. [Google Scholar] [CrossRef]
- Delgado, S.; Leite, A.M.; Ruas-Madiedo, P.; Mayo, B. Probiotic and technological properties of Lactobacillus spp. strains from the human stomach in the search for potential candidates against gastric microbial dysbiosis. Front. Microbiol. 2014, 5, 766. [Google Scholar] [CrossRef]
- Fong, Y.C.; Maa, M.C.; Tsai, F.J.; Chen, W.C.; Lin, J.G.; Jeng, L.B.; Yang, R.S.; Fu, W.M.; Tang, C.H. Osteoblast-derived TGF-beta1 stimulates IL-8 release through AP-1 and NF-kappaB in human cancer cells. J. Bone Miner. Res. 2008, 23, 961–970. [Google Scholar] [CrossRef]
- Lu, D.Y.; Chen, H.C.; Yang, M.S.; Hsu, Y.M.; Lin, H.J.; Tang, C.H.; Lee, C.H.; Lai, C.K.; Lin, C.J.; Shyu, W.C.; et al. Ceramide and Toll-like receptor 4 are mobilized into membrane rafts in response to Helicobacter pylori infection in gastric epithelial cells. Infect. Immun. 2012, 80, 1823–1833. [Google Scholar] [CrossRef]
- Lai, C.H.; Fang, S.H.; Rao, Y.K.; Geethangili, M.; Tang, C.H.; Lin, Y.J.; Hung, C.H.; Wang, W.C.; Tzeng, Y.M. Inhibition of Helicobacter pylori-induced inflammation in human gastric epithelial AGS cells by Phyllanthus urinaria extracts. J. Ethnopharmacol. 2008, 118, 522–526. [Google Scholar] [CrossRef] [PubMed]
- Lai, C.H.; Kuo, C.H.; Chen, P.Y.; Poon, S.K.; Chang, C.S.; Wang, W.C. Association of antibiotic resistance and higher internalization activity in resistant Helicobacter pylori isolates. J. Antimicrob. Chemother. 2006, 57, 466–471. [Google Scholar] [CrossRef]
- Liao, P.H.; Kuo, W.W.; Hsieh, D.J.; Yeh, Y.L.; Day, C.H.; Chen, Y.H.; Chang, S.H.; Padma, V.V.; Chen, Y.H.; Huang, C.Y. Heat-killed Lactobacillus Reuteri GMNL-263 Prevents Epididymal Fat Accumulation and Cardiac Injury in High-Calorie Diet-Fed Rats. Int. J. Med. Sci. 2016, 13, 569–577. [Google Scholar] [CrossRef] [PubMed]
- Geethangili, M.; Fang, S.H.; Lai, C.H.; Rao, Y.K.; Lien, H.M.; Tzeng, Y.M. Inhibitory effect of Antrodia camphorata constituents on the Helicobacter pylori-associated gastric inflammation. Food Chem. 2010, 119, 149–153. [Google Scholar] [CrossRef]
- Lin, H.J.; Hsu, F.Y.; Chen, W.W.; Lee, C.H.; Lin, Y.J.; Chen, Y.Y.; Chen, C.J.; Huang, M.Z.; Kao, M.C.; Chen, Y.A.; et al. Helicobacter pylori Activates HMGB1 Expression and Recruits RAGE into Lipid Rafts to Promote Inflammation in Gastric Epithelial Cells. Front. Immunol. 2016, 7, 341. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.J.; Cheng, W.C.; Cheng, H.H.; Lai, C.H.; Wang, W.C. Helicobacter pylori cholesteryl glucosides interfere with host membrane phase and affect type IV secretion system function during infection in AGS cells. Mol. Microbiol. 2012, 83, 67–84. [Google Scholar] [CrossRef]
- Lai, C.H.; Chang, Y.C.; Du, S.Y.; Wang, H.J.; Kuo, C.H.; Fang, S.H.; Fu, H.W.; Lin, H.H.; Chiang, A.S.; Wang, W.C. Cholesterol depletion reduces Helicobacter pylori CagA translocation and CagA-induced responses in AGS cells. Infect. Immun. 2008, 76, 3293–3303. [Google Scholar] [CrossRef]
- Lai, C.K.; Lu, Y.L.; Hsieh, J.T.; Tsai, S.C.; Feng, C.L.; Tsai, Y.S.; Tsai, P.C.; Su, H.L.; Lin, Y.H.; Lai, C.H. Development of chitosan/heparin nanoparticle-encapsulated cytolethal distending toxin for gastric cancer therapy. Nanomedicine 2014, 9, 803–817. [Google Scholar] [CrossRef]
- Brzozowski, T.; Konturek, P.C.; Kwiecien, S.; Konturek, S.J.; Pajdo, R.; Drozdowicz, D.; Ptak, A.; Pawlik, M.; Stachura, J.; Pawlik, W.W.; et al. Triple eradication therapy counteracts functional impairment associated with Helicobacter pylori infection in Mongolian gerbils. J. Physiol. Pharmacol. 2003, 54, 33–51. [Google Scholar]
- Marshall, B.J.; Warren, J.R.; Francis, G.J.; Langton, S.R.; Goodwin, C.S.; Blincow, E.D. Rapid urease test in the management of Campylobacter pyloridis-associated gastritis. Am. J. Gastroenterol. 1987, 82, 200–210. [Google Scholar]
- Amieva, M.R.; El-Omar, E.M. Host-bacterial interactions in Helicobacter pylori infection. Gastroenterology 2008, 134, 306–323. [Google Scholar] [CrossRef] [PubMed]
- Wilson, K.T.; Ramanujam, K.S.; Mobley, H.L.; Musselman, R.F.; James, S.P.; Meltzer, S.J. Helicobacter pylori stimulates inducible nitric oxide synthase expression and activity in a murine macrophage cell line. Gastroenterology 1996, 111, 1524–1533. [Google Scholar] [CrossRef]
- Martinez, R.C.; Bedani, R.; Saad, S.M. Scientific evidence for health effects attributed to the consumption of probiotics and prebiotics: An update for current perspectives and future challenges. Br. J. Nutr. 2015, 114, 1993–2015. [Google Scholar] [CrossRef] [PubMed]
- Espinoza, J.L.; Matsumoto, A.; Tanaka, H.; Matsumura, I. Gastric microbiota: An emerging player in Helicobacter pylori-induced gastric malignancies. Cancer Lett. 2018, 414, 147–152. [Google Scholar] [CrossRef] [PubMed]
- Megraud, F.; Lamouliatte, H. Review article: The treatment of refractory Helicobacter pylori infection. Aliment. Pharmacol. Ther. 2003, 17, 1333–1343. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, A.; Edlund, C.; Nord, C.E. Effect of antimicrobial agents on the ecological balance of human microflora. Lancet Infect. Dis. 2001, 1, 101–114. [Google Scholar] [CrossRef]
- Adamsson, I.; Nord, C.E.; Lundquist, P.; Sjostedt, S.; Edlund, C. Comparative effects of omeprazole, amoxycillin plus metronidazole versus omeprazole, clarithromycin plus metronidazole on the oral, gastric and intestinal microflora in Helicobacter pylori-infected patients. J. Antimicrob. Chemother. 1999, 44, 629–640. [Google Scholar] [CrossRef] [PubMed]
- Buhling, A.; Radun, D.; Muller, W.A.; Malfertheiner, P. Influence of anti-Helicobacter triple-therapy with metronidazole, omeprazole and clarithromycin on intestinal microflora. Aliment. Pharmacol. Ther. 2001, 15, 1445–1452. [Google Scholar] [CrossRef]
- Francavilla, R.; Polimeno, L.; Demichina, A.; Maurogiovanni, G.; Principi, B.; Scaccianoce, G.; Ierardi, E.; Russo, F.; Riezzo, G.; Di Leo, A.; et al. Lactobacillus reuteri strain combination in Helicobacter pylori infection: A randomized, double-blind, placebo-controlled study. J. Clin. Gastroenterol. 2014, 48, 407–413. [Google Scholar] [CrossRef]
- Lai, C.H.; Huang, J.C.; Cheng, H.H.; Wu, M.C.; Huang, M.Z.; Hsu, H.Y.; Chen, Y.A.; Hsu, C.Y.; Pan, Y.J.; Chu, Y.T.; et al. Helicobacter pylori cholesterol glucosylation modulates autophagy for increasing intracellular survival in macrophages. Cell. Microbiol. 2018, e12947. [Google Scholar] [CrossRef]
- Lin, C.J.; Liao, W.C.; Chen, Y.A.; Lin, H.J.; Feng, C.L.; Lin, C.L.; Lin, Y.J.; Kao, M.C.; Huang, M.Z.; Lai, C.H.; et al. Statin Therapy Is Associated with Reduced Risk of Peptic Ulcer Disease in the Taiwanese Population. Front. Pharmacol. 2017, 8, 210. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.J.; Liao, W.C.; Lin, H.J.; Hsu, Y.M.; Lin, C.L.; Chen, Y.A.; Feng, C.L.; Chen, C.J.; Kao, M.C.; Lai, C.H.; et al. Statins Attenuate Helicobacter pylori CagA Translocation and Reduce Incidence of Gastric Cancer: In Vitro and Population-Based Case-Control Studies. PLoS ONE 2016, 11, e0146432. [Google Scholar] [CrossRef] [PubMed]
- London, L.E.; Kumar, A.H.; Wall, R.; Casey, P.G.; O’Sullivan, O.; Shanahan, F.; Hill, C.; Cotter, P.D.; Fitzgerald, G.F.; Ross, R.P.; et al. Exopolysaccharide-producing probiotic Lactobacilli reduce serum cholesterol and modify enteric microbiota in ApoE-deficient mice. J. Nutr. 2014, 144, 1956–1962. [Google Scholar] [CrossRef] [PubMed]
- Michael, D.R.; Davies, T.S.; Moss, J.W.E.; Calvente, D.L.; Ramji, D.P.; Marchesi, J.R.; Pechlivanis, A.; Plummer, S.F.; Hughes, T.R. The anti-cholesterolaemic effect of a consortium of probiotics: An acute study in C57BL/6J mice. Sci. Rep. 2017, 7, 2883. [Google Scholar] [CrossRef] [PubMed]
| Bacterial Species | Primer | Nucleotide Sequence (5’-3’) |
|---|---|---|
| Total bacteria | Forward | GTGSTGCAYGGYTGTCGTCA |
| Reverse | ACGTCRTCCMCACCTTCCTC | |
| Bifidobacterium | Forward | CGCGTCYGGTGTGAAAG |
| Reverse | CCCCACATCCAGCATCCA | |
| Akkermansia muciniphila | Forward | CAGCACGTGAAGGTGGGGAC |
| Reverse | CCTTGCGGTTGGCTTCAGAT | |
| Escherichia coli | Forward | CATGCCGCGTGTATGAAGAA |
| Reverse | CGGGTAACGTCAATGAGCAAA | |
| Clostridium cluster I | Forward | TACCHRAGGAGGAAGCCAC |
| Reverse | GTTCTTCCTAATCTCTACGCAT | |
| Bacteroides-Provetella | Forward | AAGGTCCCCCACATTGG |
| Reverse | CCGCGGCKGCTGGCAC | |
| Proteobacteria | Forward | TGGTGTAGGGGTAAAATCCG |
| Reverse | AGGTAAGGTTCTTCGYGTATC | |
| Actinobacteria | Forward | TGTAGCGGTGGAATGCGC |
| Reverse | AATTAAGCCACATGCTCCGCT | |
| Fusobacteria | Forward | AAGCGCGTCTAGGTGGTTATGT |
| Reverse | TGTAGTTCCGCTTACCTCTCCAG | |
| Enterococcus | Forward | CCCTTATTGTTAGTTGCCATCATT |
| Reverse | ACTCGTTGTACTTCCCATTGT | |
| Firmicutes | Forward | GCGTGAGTGAAGAAGT |
| Reverse | CTACGCTCCCTTTACAC | |
| Bacteroidetes | Forward | CTGAACCAGCCAAGTAGCG |
| Reverse | CCGCAAACTTTCACAACTGACTTA |
| Inhibition Zone (mm) ‡ | |||
|---|---|---|---|
| Reference Strain | MDR-H. pylori † | ||
| Treatment | 26695 | v633 | v1354 |
| GMNL-74 | 12.3 ± 0.5 | 8.3 ± 0.5 | 8.7 ± 0.5 |
| GMNL-185 | 11.3 ± 0.5 | 7.7 ± 0.5 | 9.0 ± 0.8 |
| GMNL-229 | 0 | 0 | 0 |
| GMNL-814 | 0 | 0 | 0 |
| MRS broth | 0 | 0 | 0 |
| Clarithromycin | 21.5 ± 0.3 | 0 | 0 |
| Metronidazole | 18.3 ± 0.4 | 0 | 0 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, Y.-H.; Tsai, W.-H.; Wu, H.-Y.; Chen, C.-Y.; Yeh, W.-L.; Chen, Y.-H.; Hsu, H.-Y.; Chen, W.-W.; Chen, Y.-W.; Chang, W.-W.; et al. Probiotic Lactobacillus spp. Act Against Helicobacter pylori-induced Inflammation. J. Clin. Med. 2019, 8, 90. https://doi.org/10.3390/jcm8010090
Chen Y-H, Tsai W-H, Wu H-Y, Chen C-Y, Yeh W-L, Chen Y-H, Hsu H-Y, Chen W-W, Chen Y-W, Chang W-W, et al. Probiotic Lactobacillus spp. Act Against Helicobacter pylori-induced Inflammation. Journal of Clinical Medicine. 2019; 8(1):90. https://doi.org/10.3390/jcm8010090
Chicago/Turabian StyleChen, Yi-Hsing, Wan-Hua Tsai, Hui-Yu Wu, Chun-Ya Chen, Wen-Ling Yeh, Ya-Hui Chen, Hui-Ying Hsu, Wei-Wei Chen, Yu-Wen Chen, Wen-Wei Chang, and et al. 2019. "Probiotic Lactobacillus spp. Act Against Helicobacter pylori-induced Inflammation" Journal of Clinical Medicine 8, no. 1: 90. https://doi.org/10.3390/jcm8010090
APA StyleChen, Y.-H., Tsai, W.-H., Wu, H.-Y., Chen, C.-Y., Yeh, W.-L., Chen, Y.-H., Hsu, H.-Y., Chen, W.-W., Chen, Y.-W., Chang, W.-W., Lin, T.-L., Lai, H.-C., Lin, Y.-H., & Lai, C.-H. (2019). Probiotic Lactobacillus spp. Act Against Helicobacter pylori-induced Inflammation. Journal of Clinical Medicine, 8(1), 90. https://doi.org/10.3390/jcm8010090






