Molecular Characterization of Velogenic Newcastle Disease Virus (Sub-Genotype VII.1.1) from Wild Birds, with Assessment of Its Pathogenicity in Susceptible Chickens
Abstract
Simple Summary
Abstract
1. Introduction
2. Material and Methods
2.1. Ethical Considerations
2.2. Study Area, Sample Collection, and Sample Preparation
2.3. Standard NDV
2.4. Molecular Characterization of NDV
2.4.1. Viral RNA Extraction
2.4.2. Real-Time Reverse Transcriptase Polymerase Chain Reaction
2.4.3. Reverse-Transcriptase Polymerase Chain Reaction
2.4.4. Sequencing and Phylogenetic Analysis of the Selected Samples
2.5. Pathogenicity Testing of Wild Bird Sub-Genotype VII.1.1 NDV Strains in Susceptible Chickens
2.5.1. Susceptible Chickens
2.5.2. NDV Strains
2.5.3. Virus Propagation in Embryonated Chicken Eggs
2.5.4. Virus Titration
2.5.5. Experimental Design
2.5.6. Histopathological Examination
3. Results
3.1. Clinical Signs and Post-Mortem Lesions
3.2. Molecular Characterization of NDV
3.2.1. Real-Time Reverse Transcriptase Polymerase Chain Reaction
3.2.2. Reverse Transcriptase Polymerase Chain Reaction
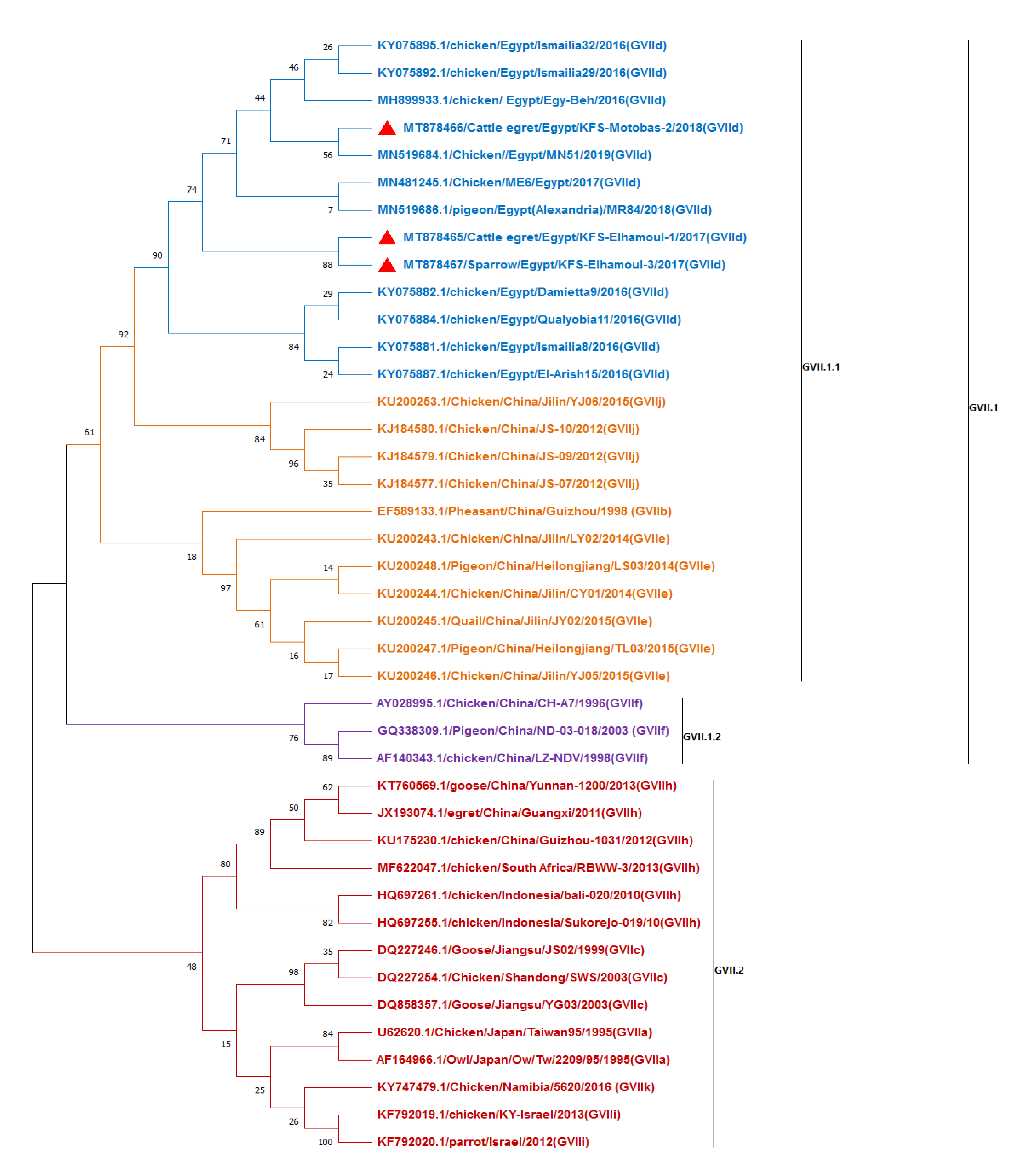
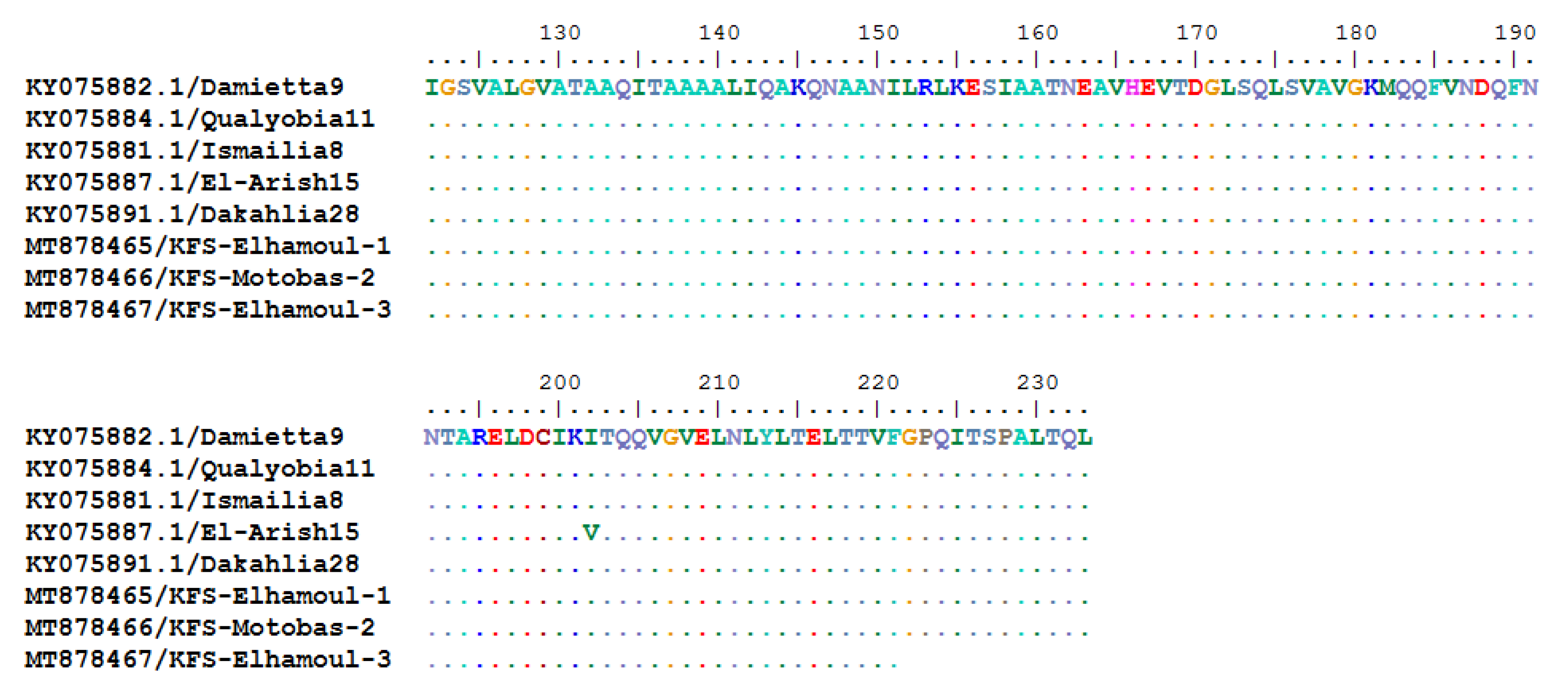
3.2.3. Sequencing and Phylogenetic Analysis of NDV F Gene Fragment
3.3. Pathogenicity Testing of Wild Birds’ NDV Sub-Genotype VII.1.1 Strains in Susceptible Chickens
3.3.1. Virus Propagation in Embryonated Chicken Eggs
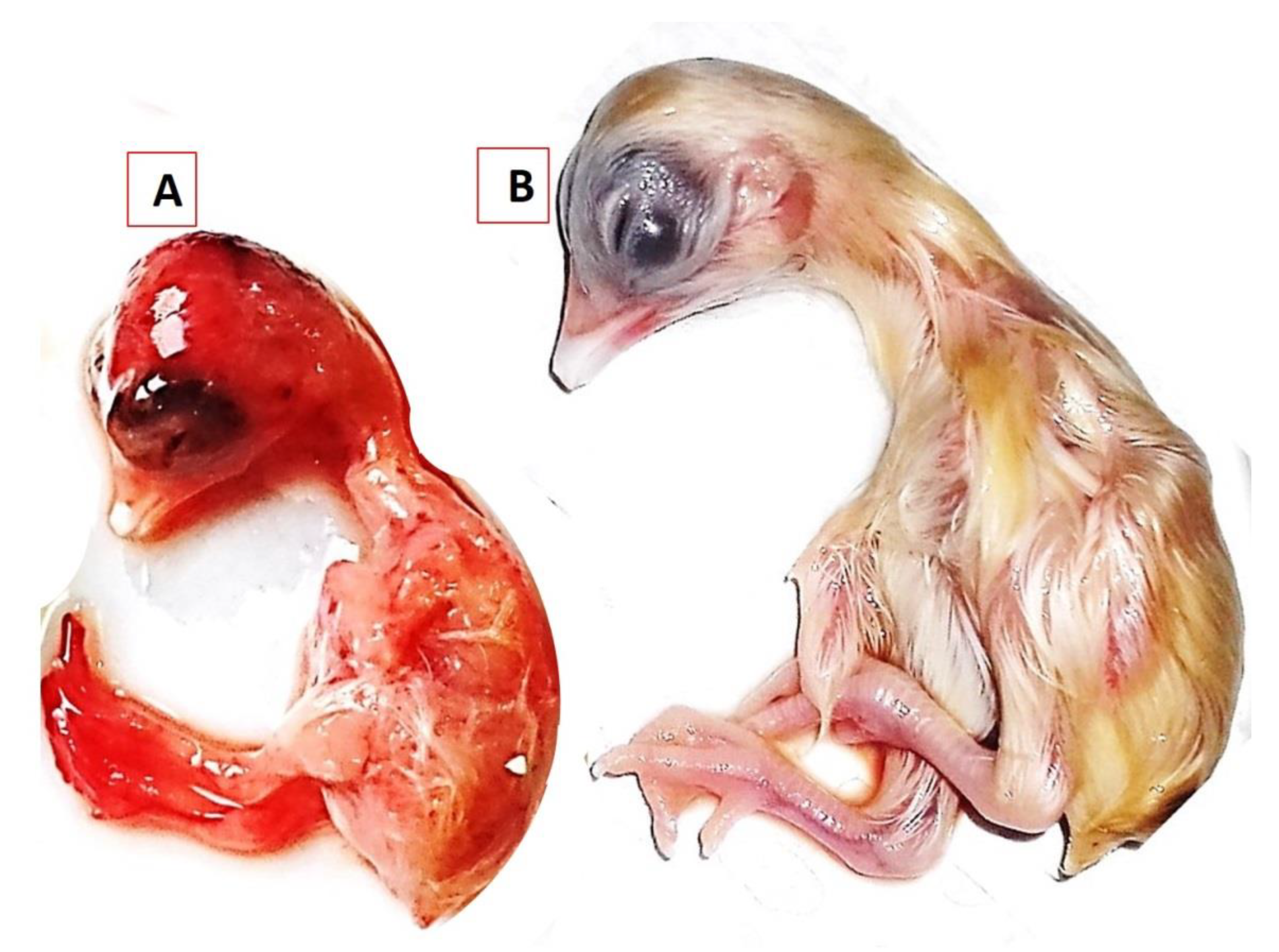
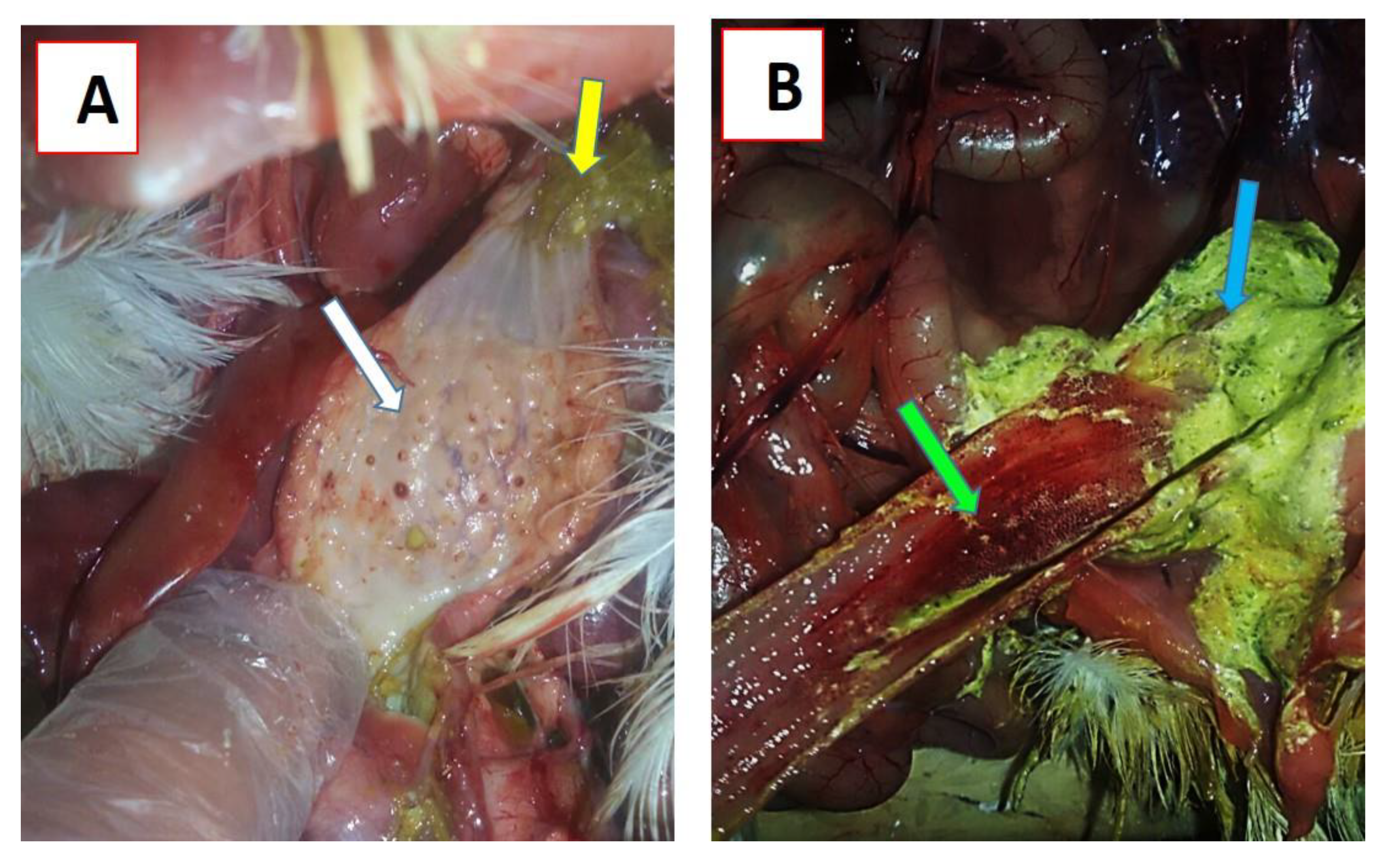
3.3.2. Clinical Signs and Post-Mortem Changes in Experimentally Infected Chickens
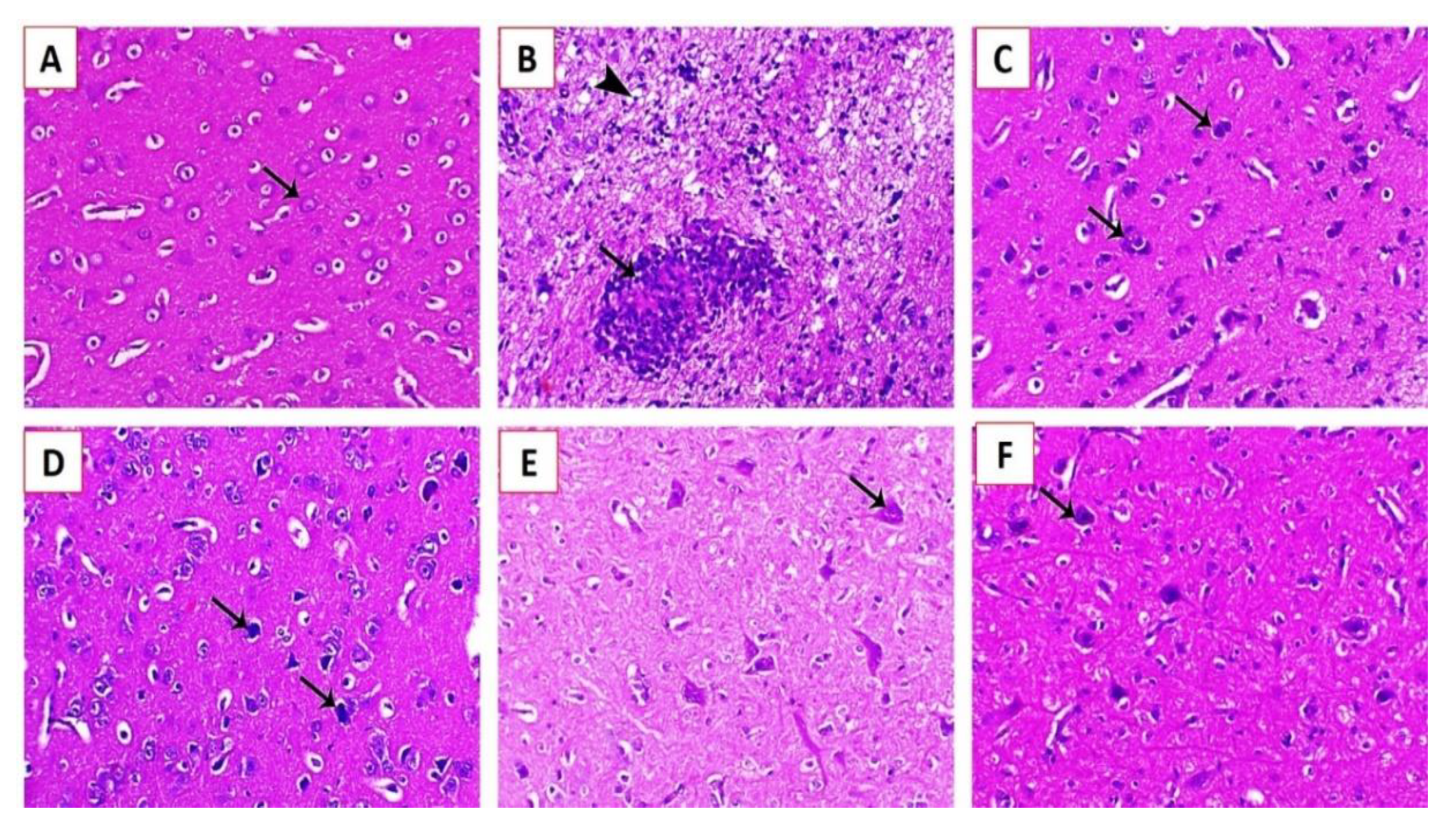
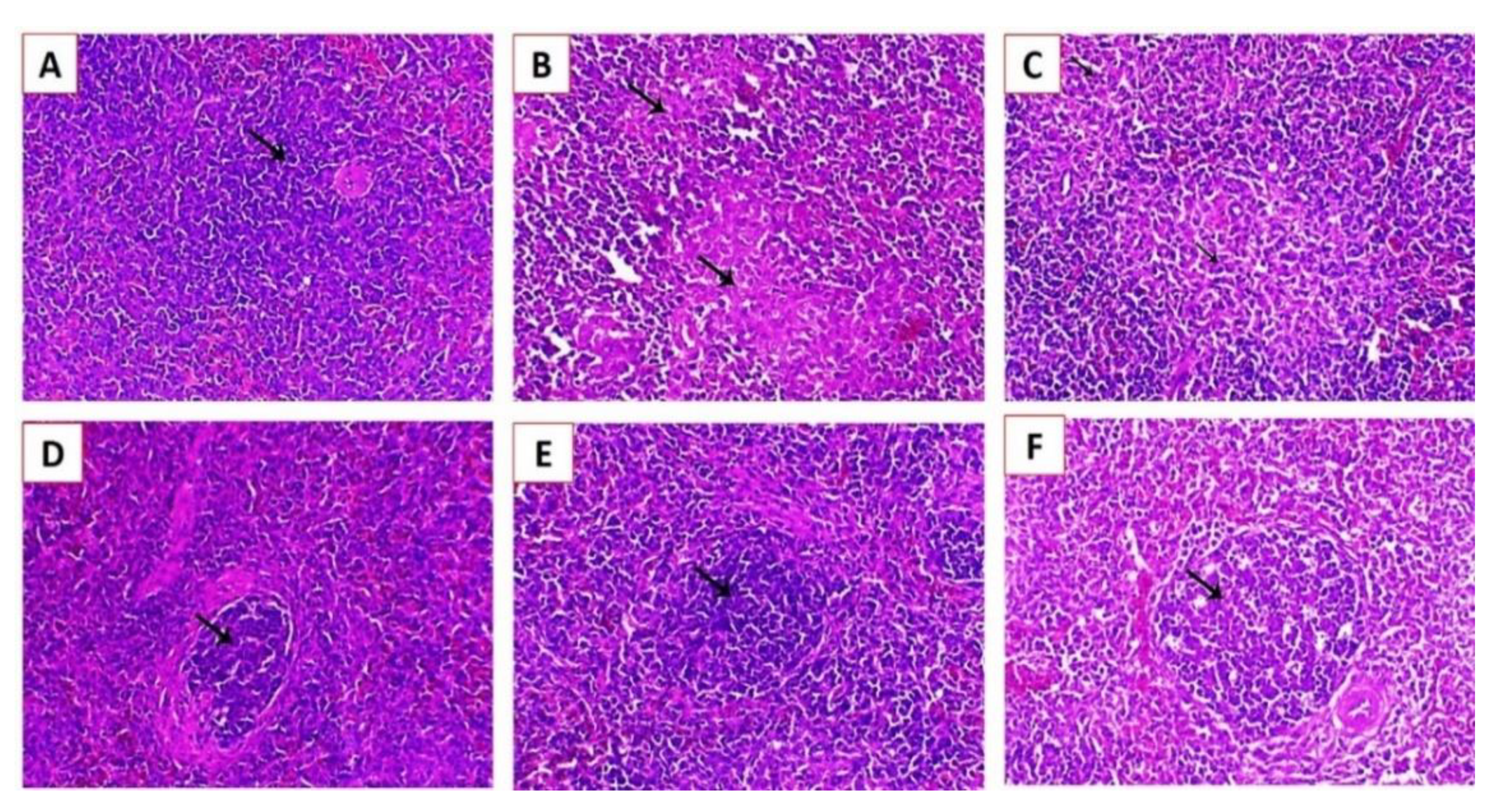
3.3.3. Histopathological Examination
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Alexander, D.J.; Aldous, E.W.; Fuller, C.M. The long view: A selective review of 40 years of Newcastle disease research. Avian Pathol. 2012, 41, 329–335. [Google Scholar] [CrossRef] [PubMed]
- OIE. World Animal Health Information Database (WAHIS Interface)—Version 1. Available online: http://www.oie.int/wahis_2/public/wahid.php/Wahidhome/Home (accessed on 12 December 2020).
- Anonymous. World Livestock Disease Atlas: A Quantitative Analysis of GlobalAnimal Health Data A (2006–2009); The World Bank for Reconstruction and Development, The World Bank ad TAFS Forum: Washington, DC, USA, 2011. [Google Scholar]
- Miller, P.J.; Koch, G. Newcastle disease. Dis. Poult. 2013, 13, 89–138. [Google Scholar]
- Bello, M.B.; Yusoff, K.M.; Ideris, A.; Hair-Bejo, M.; Peeters, B.P.H.; Jibril, A.H.; Tambuwal, F.M.; Omar, A.R. Genotype Diversity of Newcastle Disease Virus in Nigeria: Disease Control Challenges and Future Outlook. Adv. Virol. 2018, 2018. [Google Scholar] [CrossRef]
- Absalon, A.E.; Cortes-Espinosa, D.V.; Lucio, E.; Miller, P.J.; Afonso, C.L. Epidemiology, control, and prevention of Newcastle disease in endemic regions: Latin America. Trop. Anim. Health Prod. 2019, 51, 1033–1048. [Google Scholar] [CrossRef]
- Kapczynski, D.R.; Afonso, C.L.; Miller, P.J. Immune responses of poultry to Newcastle disease virus. Dev. Comp. Immunol. 2013, 41, 447–453. [Google Scholar] [CrossRef]
- OIE. Newcastle disease. In OIE Manual of Standards for Diagnostic Tests and Vaccines; Chapter 2.3.14; NB: Version adopted by the World Assembly of Delegates of the OIE; OIE: Paris, France, 2012. [Google Scholar]
- ICTV. International Committee on Taxonomy of Viruses Virus Taxonomy: 2018b Release. 2019. Available online: https://talk.ictvonline.org/taxonomy/ (accessed on 15 December 2020).
- Moura, V.; Susta, L.; Cardenas-Garcia, S.; Stanton, J.; Miller, P.; Afonso, C.; Brown, C. Neuropathogenic capacity of lentogenic, mesogenic, and velogenic Newcastle disease virus strains in day-old chickens. Vet. Pathol. 2016, 53, 53–64. [Google Scholar] [CrossRef]
- Dimitrov, K.M.; Abolnik, C.; Afonso, C.L.; Albina, E.; Bahl, J.; Berg, M.; Briand, F.-X.; Brown, I.H.; Choi, K.-S.; Chvala, I. Updated unified phylogenetic classification system and revised nomenclature for Newcastle disease virus. Infect. Genet. Evol. 2019, 74. [Google Scholar] [CrossRef]
- Xue, C.; Cong, Y.; Yin, R.; Sun, Y.; Ding, C.; Yu, S.; Liu, X.; Hu, S.; Qian, J.; Yuan, Q. Genetic diversity of the genotype VII Newcastle disease virus: Identification of a novel VIIj sub-genotype. Virus genes 2017, 53, 63–70. [Google Scholar] [CrossRef] [PubMed]
- Dimitrov, K.; Dong-Hun, L.; Williams-Coplin, D.; Olivier, T.; Miller, P.; Afonso, C. Newcastle Disease Viruses Causing Recent Outbreaks Worldwide Show Unexpectedly High Genetic Similarity with Historical Virulent Isolates from the 1940’s. J. Clin. Microbiol. 2016, 54. [Google Scholar] [CrossRef]
- Snoeck, C.; Owoade, A.; Couacy-Hymann, E.; Alkali, B.; Okwen, M.; Adeyanju, T.; Komoyo, G.F.; Nakouné, E.; Faou, A.; Muller, C. High Genetic Diversity of Newcastle Disease Virus in Poultry in West and Central Africa: Cocirculation of Genotype XIV and Newly Defined Genotypes XVII and XVIII. J. Clin. Microbiol. 2013, 51. [Google Scholar] [CrossRef] [PubMed]
- Diel, D.G.; da Silva, L.H.; Liu, H.; Wang, Z.; Miller, P.J.; Afonso, C.L. Genetic diversity of avian paramyxovirus type 1: Proposal for a unified nomenclature and classification system of Newcastle disease virus genotypes. Infect. Genet. Evol. 2012, 12, 1770–1779. [Google Scholar] [CrossRef]
- Abolnik, C.; Mubamba, C.; Wandrag, D.B.; Horner, R.; Gummow, B.; Dautu, G.; Bisschop, S.P. Tracing the origins of genotype VII h Newcastle disease in Southern Africa. Transbound. Emerg. Dis. 2018, 65, e393–e403. [Google Scholar] [CrossRef]
- Fuller, C.; Löndt, B.; Dimitrov, K.; Lewis, N.; van Boheemen, S.; Fouchier, R.; Coven, F.; Goujgoulova, G.; Haddas, R.; Brown, I. An epizootiological report of the re-emergence and spread of a lineage of virulent Newcastle disease virus into Eastern Europe. Transbound. Emerg. Dis. 2017, 64, 1001–1007. [Google Scholar] [CrossRef]
- Kammon, A.; Monne, I.; Asheg, A.; Cattoli, G. Molecular detection and characterisation of avian paramyxovirus type 1 in backyard chickens and pigeons in Alzintan city of Libya. Open Vet. J. 2018, 8, 401–405. [Google Scholar] [CrossRef] [PubMed]
- Mapaco, L.P.; Monjane, I.V.; Nhamusso, A.E.; Viljoen, G.J.; Dundon, W.G.; Achá, S.J. Phylogenetic analysis of Newcastle disease viruses isolated from commercial poultry in Mozambique (2011–2016). Virus Genes 2016, 52, 748–753. [Google Scholar] [CrossRef] [PubMed]
- Jindal, N.; Chander, Y.; Chockalingam, A.K.; De Abin, M.; Redig, P.T.; Goyal, S.M. Phylogenetic analysis of Newcastle disease viruses isolated from waterfowl in the upper midwest region of the United States. Virol. J. 2009, 6, 1–9. [Google Scholar] [CrossRef]
- Miller, P.J.; Haddas, R.; Simanov, L.; Lublin, A.; Rehmani, S.F.; Wajid, A.; Bibi, T.; Khan, T.A.; Yaqub, T.; Setiyaningsih, S. Identification of new sub-genotypes of virulent Newcastle disease virus with potential panzootic features. Infect. Genet. Evol. 2015, 29, 216–229. [Google Scholar] [CrossRef]
- Molini, U.; Aikukutu, G.; Khaiseb, S.; Cattoli, G.; Dundon, W.G. First genetic characterization of Newcastle disease viruses from Namibia: Identification of a novel VIIk subgenotype. Arch. Virol. 2017, 162, 2427–2431. [Google Scholar] [CrossRef] [PubMed]
- Radwan, M.M.; Darwish, S.F.; El-Sabagh, I.M.; El-Sanousi, A.A.; Shalaby, M.A. Isolation and molecular characterization of Newcastle disease virus genotypes II and VIId in Egypt between 2011 and 2012. Virus Genes 2013, 47, 311–316. [Google Scholar] [CrossRef] [PubMed]
- Brown, V.R.; Bevins, S.N. A review of virulent Newcastle disease viruses in the United States and the role of wild birds in viral persistence and spread. Vet. Res. 2017, 48, 68. [Google Scholar] [CrossRef] [PubMed]
- Sabban, M.; Zied, A.A.; Basyouni, A.; Nadiem, S.; Barhouma, N.; Habashi, Y. Susceptibility and possible role of doves in transmission of Newcastle disease in Egypt. Zent. Vet. Reihe B 1982, 29, 193–198. [Google Scholar] [CrossRef] [PubMed]
- Mayo, M.A. Virus taxonomy—Houston 2002. Arch. Virol. 2002, 147, 1071–1076. [Google Scholar] [CrossRef] [PubMed]
- Perttula, L. Epidemiology and Characterization of Newcastle Disease in Smallholder Poultry in Mozambique; Examensarbete (Sveriges Lantbruksuniversitet, Fakulteten för Veterinärmedicin och Husdjursvetenskap, Veterinärprogrammet): Uppsala, Sweden, 2010. [Google Scholar]
- Aziz-ul-Rahman, M.H.; Shabbir, M.Z. Suppl-2, M3: Adaptation of Newcastle Disease Virus (NDV) in Feral Birds and their Potential Role in Interspecies Transmission. Open Virol. J. 2018, 12, 52–68. [Google Scholar] [CrossRef]
- Seal, B.; Wise, M.; Pedersen, J.; Senne, D.; Alvarez, R.; Scott, M.; King, D.; Yu, Q.; Kapczynski, D. Genomic sequences of low-virulence avian paramyxovirus-1 (Newcastle disease virus) isolates obtained from live-bird markets in North America not related to commonly utilized commercial vaccine strains. Vet. Microbiol. 2005, 106, 7–16. [Google Scholar] [CrossRef]
- Jørgensen, H.P.; Handberg, K.; Ahrens, P.; Therkildsen, O.; Manvell, R.; Alexander, D. Strains of avian paramyxovirus type 1 of low pathogenicity for chickens isolated from poultry and wild birds in Denmark. Vet. Record 2004, 154, 497–500. [Google Scholar] [CrossRef] [PubMed]
- Dimitrov, K.; Ramey, A.; Qiu, X.; Bahl, J.; Afonso, C. Temporal, geographic, and host distribution of avian paramyxovirus 1 (Newcastle disease virus). Infect. Genet. Evol. 2016, 39, 22–34. [Google Scholar] [CrossRef]
- Mosad, S.M.; El-Gohary, F.A.; Ali, H.S.; El-Sharkawy, H.; Elmahallawy, E.K. Pathological and Molecular Characterization of H5 Avian Influenza Virus in Poultry Flocks from Egypt over a Ten-Year Period (2009–2019). Animals (Basel) 2020, 10, 1010. [Google Scholar] [CrossRef] [PubMed]
- Ramey, A.M.; Goraichuk, I.V.; Hicks, J.T.; Dimitrov, K.M.; Poulson, R.L.; Stallknecht, D.E.; Bahl, J.; Afonso, C.L. Assessment of contemporary genetic diversity and inter-taxa/inter-region exchange of avian paramyxovirus serotype 1 in wild birds sampled in North America. Virol. J. 2017, 14, 43. [Google Scholar] [CrossRef] [PubMed]
- Fagbohun, O.; Oluwayelu, D.; Owoade, A.; Olayemi, F. Survey for antibodies to Newcastle disease virus in cattle egrets, pigeons and Nigerians laughing doves. Afr. J. Biomed. Res. 2000, 3, 193–194. [Google Scholar]
- Hicks, J.T.; Dimitrov, K.M.; Afonso, C.L.; Ramey, A.M.; Bahl, J. Global phylodynamic analysis of avian paramyxovirus-1 provides evidence of inter-host transmission and intercontinental spatial diffusion. BMC Evol. Biol. 2019, 19, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Turan, N.; Ozsemir, C.; Yilmaz, A.; Cizmecigil, U.Y.; Aydin, O.; Bamac, O.E.; Gurel, A.; Kutukcu, A.; Ozsemir, K.; Tali, H.E.; et al. Identification of Newcastle disease virus subgenotype VII.2 in wild birds in Turkey. BMC Vet. Res. 2020, 16, 277. [Google Scholar] [CrossRef]
- Maiti, S.K.; Tiwary, R.; Vasan, P.; Dutta, A. Xylazine, diazepam and midazolam premedicated ketamine anaesthesia in white Leghorn cockerels for typhlectomy. J. S. Afr. Vet. Assoc. 2006, 77, 12–18. [Google Scholar] [CrossRef] [PubMed]
- Grimes, S.E. A Basic Laboratory Manual for the Small-Scale Production and Testing of I-2 Newcastle Disease Vaccine; FAO Regional Office for Asia and the Pacific (RAP): Bangkok, Thailand, 2002; p. 22. [Google Scholar]
- Awad, M.; Mosad, S.; El-Kenawy, A. Molecular differentiation between velogenic isolates and lentogenic LaSota strain of Newcastle disease virus. Mansoura Vet. Med. J. 2020, 21, 193–198. [Google Scholar] [CrossRef]
- Wise, M.G.; Suarez, D.L.; Seal, B.S.; Pedersen, J.C.; Senne, D.A.; King, D.J.; Kapczynski, D.R.; Spackman, E. Development of a real-time reverse-transcription PCR for detection of newcastle disease virus RNA in clinical samples. J. Clin. Microbiol. 2004, 42, 329–338. [Google Scholar] [CrossRef] [PubMed]
- Selim, K.M.; Selim, A.; Arafa, A.; Hussein, H.A.; Elsanousi, A.A. Molecular characterization of full fusion protein (F) of Newcastle disease virus genotype VIId isolated from Egypt during 2012-2016. Vet. World 2018, 11, 930–938. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Hall, T. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. In Nucleic Acids Symposium; Oxford Academic Press: Cary, NC, USA, 1999; Volume 41, pp. 95–98. [Google Scholar]
- Ali, H.H. Study serologic status of newcastle disease in broilers chikens by haemagglutination inhibition test in Suliamania province. Glob. J. Bio-Sci. Biotechnol. 2015, 4, 364–369. [Google Scholar]
- Reed, L.J.; Muench, H. A simple method of estimating fifty per cent endpoints. Am. J. Epidemiol. 1938, 27, 493–497. [Google Scholar] [CrossRef]
- Ahmed, A.; Kandeil, A.; Kutkat, M.; Abdel-Moez, A. Advancement in Vaccination of Broiler Chickens with Genotype-Matched Vaccines to Currently Epidemic Newcastle Disease Virus Genotype VII in Egypt. J. World’s Poult. Res. 2019, 9, 117–123. [Google Scholar] [CrossRef]
- Downie, T. Theory and practice of histological techniques edited by JD Bancroft & a. Stevens, Churchill Livingstone, Edinburgh, 740 pages, £ 55.00. Histopathology 1990, 17, 386. [Google Scholar]
- Abdisa, T.; Tagesu, T. Review on Newcastle Disease in Poultry and its Public Health Importance. J. Anim. Poult. Sci. 2017, 6, 29–39. [Google Scholar] [CrossRef]
- Silva, J.; Mota, R.; Vilela, S.; Doretto Júnior, L.; Pinheiro Júnior, J.; Silva, L. Newcastle disease virus infection in Sparrows (Passer domesticus, Linneaus, 1758) captured in poultry farms of the Agreste region of the State of Pernambuco. Braz. J. Poult. Sci. 2006, 8, 125–129. [Google Scholar] [CrossRef][Green Version]
- Munir, T.; Aslam, A.; Zahid, B.; Ahmed, I.; Imran, M.; Ijaz, M. Potential of commonly resident wild birds towards newcastle disease virus transmission. Pak. Vet. J. 2015, 35, 106–107. [Google Scholar]
- Lindh, E.; Ek-Kommonen, C.; Vaananen, V.M.; Alasaari, J.; Vaheri, A.; Vapalahti, O.; Huovilainen, A. Molecular epidemiology of outbreak-associated and wild-waterfowl-derived newcastle disease virus strains in Finland, including a novel class I genotype. J. Clin. Microbiol. 2012, 50, 3664–3673. [Google Scholar] [CrossRef] [PubMed]
- Aldous, E.W.; Mynn, J.K.; Banks, J.; Alexander, D.J. A molecular epidemiological study of avian paramyxovirus type 1 (Newcastle disease virus) isolates by phylogenetic analysis of a partial nucleotide sequence of the fusion protein gene. Avian Pathol. 2003, 32, 239–256. [Google Scholar] [CrossRef] [PubMed]
- Blaxland, J. Newcastle disease in shags and cormorants and its significance as a factor in the spread of this disease among domestic poultry. Vet. Rec. 1951, 63, 731–733. [Google Scholar]
- Dhama, K.; Mahendran, M.; Tomar, S. Pathogens Transmitted by Migratory Birds: Threat Perceptions to Poultry Health and Production. Int. J. Poult. Sci. 2008, 7, 516–525. [Google Scholar] [CrossRef]
- Rehman, Z.U.; Meng, C.; Sun, Y.; Mahrose, K.M.; Umar, S.; Ding, C.; Munir, M. Pathobiology of Avian avulavirus 1: Special focus on waterfowl. Vet. Res. 2018, 49, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Elmberg, J.; Berg, C.; Lerner, H.; Waldenström, J.; Hessel, R. Potential disease transmission from wild geese and swans to livestock, poultry and humans: A review of the scientific literature from a One Health perspective. Infect. Ecol. Epidemiol. 2017, 7. [Google Scholar] [CrossRef]
- Saif, B.G.J.; Fadly, L.R.M.; Swayne, D. (Eds.) Diseases of Poultry; Iowa State University Press: Ames, IA, USA, 2005; Volume 4, pp. 63–99. [Google Scholar]
- Alexander, D.J. Ecology and epidemiology of Newcastle disease. In Avian Influenza and Newcastle Disease; Springer: Berlin/Heidelberg, Germany, 2009; pp. 19–26. [Google Scholar]
- Alexander, D. Newcastle disease and other avian paramyxoviruses. Rev. Sci. Tech. Off. Int. Epizoo. 2000, 19, 443–455. [Google Scholar] [CrossRef]
- Alexander, D.; Banks, J.; Collins, M.; Manvell, R.; Frost, K.; Speidel, E.; Aldous, E. Antigenic and genetic characerisation of Newcastle disease viruses isolated from outbreaks in domestic fowl and turkeys in Great Britain during 1997. Vet. Rec. 1999, 145, 417–421. [Google Scholar] [CrossRef] [PubMed]
- Guirguis, W.I. The Role Played by white Egrets in Transmission of Newcastle Disease Virus (Velogenicviscerotropic Strains). Master’s Thesis, Cairo Univsity Library, Cairo, Egypt, 1983. [Google Scholar]
- Xu, H.; Lohr, J.; Greiner, M. The selection of ELISA cut-off points for testing antibody to Newcastle disease by two-graph receiver operating characteristic (TG-ROC) analysis. J. Immunol. Methods 1997, 208, 61–64. [Google Scholar] [CrossRef]
- Zhang, L.; Liu, N.; Ma, X.; Jiang, L. The transcriptional control machinery as well as the cell wall integrity and its regulation are involved in the detoxification of the organic solvent dimethyl sulfoxide in Saccharomyces cerevisiae. FEMS Yeast Res. 2013, 13, 200–218. [Google Scholar] [CrossRef]
- Alexander, D.; Gough, R.; Saif, Y.; Barnes, H.; Glisson, J.; Fadly, A.; McDougald, L.; Swayne, D. Newcastle Disease, Other Avian Paramyxoviruses, and Pneumovirus Infections. Disease of Poultry, 11th ed.; Iowa State Press: Ames, IA, USA, 2003. [Google Scholar]
- Mazumder, A.; Khatun, S.; Nooruzzaman, M.; Chowdhury, E.; Das, P.; Islam, M. Isolation and identification of Newcastle disease viruses from field outbreaks in chickens and pigeons. Bangladesh Vet. 2012, 29, 41–48. [Google Scholar] [CrossRef]
- Williams, R.; Boshoff, C.H.; Verwoerd, D.; Schoeman, M.; van Wyk, A.; Gerdes, T.H.; Roos, K. Detection of antibodies to Newcastle disease virus in ostriches (Struthio camelus) by an indirect ELISA. Avian Dis. 1997, 864–869. [Google Scholar] [CrossRef]
- Suarez, D.L.; Miller, P.J.; Koch, G.; Mundt, E.; Rautenschlein, S. Newcastle disease, other avian paramyxoviruses, and avian metapneumovirus infections. Dis. Poult. 2020, 109–166. [Google Scholar] [CrossRef]
- Aldous, E.; Alexander, D. Detection and differentiation of Newcastle disease virus (avian paramyxovirus type 1). Avian Pathol. 2001, 30, 117–128. [Google Scholar] [CrossRef]
- Beguas, R.; Umali, D. Genetic characterization of Newcastle Disease virus from broiler flocks in selected areas in Central Luzon, Philippines. Philipp. J. Vet. Med. 2018, 55, 25–34. [Google Scholar]
- El Naggar, R.F.; Rohaim, M.A.; Bazid, A.H.; Ahmed, K.A.; Hussein, H.A.; Munir, M. Biological characterization of wild-bird-origin avian avulavirus 1 and efficacy of currently applied vaccines against potential infection in commercial poultry. Arch. Virol. 2018, 163, 2743–2755. [Google Scholar] [CrossRef]
- Schelling, E.; Thur, B.; Griot, C.; Audige, L. Epidemiological study of Newcastle disease in backyard poultry and wild bird populations in Switzerland. Avian Pathol. 1999, 28, 263–272. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Renfu, Y.; Qian, J.; Sun, Y.; Wang, C.; Ding, C.; Yu, S.; Hu, Z.; Liu, X.; Cong, Y.; et al. Identification and pathotypical analysis of a novel VIk sub-genotype Newcastle disease virus obtained from pigeon in China. Virus Res. 2017, 238. [Google Scholar] [CrossRef]
- Kim, L.M.; King, D.J.; Curry, P.E.; Suarez, D.L.; Swayne, D.E.; Stallknecht, D.E.; Slemons, R.D.; Pedersen, J.C.; Senne, D.A.; Winker, K.; et al. Phylogenetic diversity among low-virulence newcastle disease viruses from waterfowl and shorebirds and comparison of genotype distributions to those of poultry-origin isolates. J. Virol. 2007, 81, 12641–12653. [Google Scholar] [CrossRef]
- Thomazelli, L.; de Araujo, J.; Ferreira, C.d.S.; Hurtado, R.; Oliveira, D.; Ometto, T.; Golono, M.; Sanfilippo, L.; Demetrio, C.; Figueiredo, M. Molecular surveillance of the Newcastle disease virus in domestic and wild birds on the North Eastern Coast and Amazon biome of Brazil. Braz. J. Poult. Sci. 2012, 14, 1–7. [Google Scholar] [CrossRef]
- Nunes, C.F.; Fonseca, F.; Leite, A.T.M.; Silva Filho, R.P.d.; Finger, P.F.; Castro, C.C.; Fischer, G.; Vargas, G.; Hübner, S.d.O. Investigation on Newcastle disease virus (NDV), infectious bursal disease virus (IBDV) and avian poxvirus (APV) in Magellanic Penguins in Southern Region of Brazil. Braz. Arch. Biol. Technol. 2012, 55, 537–542. [Google Scholar] [CrossRef]
- Daodu, O.B.; Aiyedun, J.O.; Kadir, R.A.; Ambali, H.M.; Oludairo, O.O.; Olorunshola, I.D.; Daodu, O.C.; Baba, S.S. Awareness and antibody detection of Newcastle disease virus in a neglected society in Nigeria. Vet. World 2019, 12, 112–118. [Google Scholar] [CrossRef] [PubMed]
- van Seventer, J.M.; Hochberg, N.S. Principles of Infectious Diseases: Transmission, Diagnosis, Prevention, and Control. Int. Encycl. Public Health 2017, 22–39. [Google Scholar] [CrossRef]
- Mariappan, A.K.; Munusamy, P.; Kumar, D.; Latheef, S.K.; Singh, S.D.; Singh, R.; Dhama, K. Pathological and molecular investigation of velogenic viscerotropic Newcastle disease outbreak in a vaccinated chicken flocks. Virusdisease 2018, 29, 180–191. [Google Scholar] [CrossRef]
- Bhaiyat, M.; Ochiai, K.; Itakura, C.; Islam, M.; Kida, H. Brain lesions in young broiler chickens naturally infected with a mesogenic strain of Newcastle disease virus. Avian Pathol. 1994, 23, 693–708. [Google Scholar] [CrossRef] [PubMed]

| Chicken Farm No. | Chicken Flock Size | Type of Poultry Farms | Farm Location | Collection Date (Month/Year) | Number of Collected Wild Birds |
|---|---|---|---|---|---|
| 1 | 3000 | Broiler | El-Hamoul | 10/2017 | Five cattle egrets and five house sparrows were collected from the vicinity of each farm. |
| 2 | 2000 | Broiler | El-Hamoul | 12/2017 | |
| 3 | 4000 | Layer | Kafrelsheikh | 3/2018 | |
| 4 | 2500 | Broiler | Balteem | 5/2018 | |
| 5 | 5000 | Broiler | Motobas | 6/2018 | |
| 6 | 3200 | Layer | El-Hamoul | 11/2018 | |
| 7 | 4500 | Broiler | Kafrelsheikh | 1/2019 | |
| 8 | 5000 | Broiler | Motobas | 3/2019 | |
| 9 | 2700 | Broiler | Kafrelsheikh | 5/2019 | |
| 10 | 5000 | Layer | Balteem | 10/2019 |
| Primer | Sequence (5’–3’) Direction | Product Size (bp) | Reference |
|---|---|---|---|
| F+4839 | TCCGGAGGATACAAGGGTCT | 101 | [41] |
| F+4894 (VFP-1) | [FAM]AAGCGTTTCTGTCTCCTTCCTCCA[TAMRA] | ||
| F-4939 | AGCTGTTGCAACCCCAAG | ||
| NDV-F330 forward | AGGAAGGAGACAAAAACGTTTTATAGG | 370 | [42] |
| NDV-R700 reverse | TCAGCTGAGTTAATGCAGGGGAGG |
| Farm No. | Wild Bird No. | Clinical Signs | Post-mortem Changes | RRT-PCR (Positive/Total) | RT-PCR (Positive/Total) | Sequencing |
|---|---|---|---|---|---|---|
| 1 | 5 cattle egrets | Whitish-green diarrhea (1/5) | Enteritis (1/5) | 3/5 | 2/5 | KFS-Elhamoul-1 strain |
| 5 house sparrows | No signs | No PM lesions | 2/5 | 2/5 | ||
| 2 | 5 cattle egrets | No signs | No PM lesions | 1/5 | 1/5 | |
| 5 house sparrows | Whitish-green diarrhea (1/5) | Enteritis (1/5) | 2/5 | 1/5 | KFS-Elhamoul-3 strain | |
| 3 | 5 cattle egrets | Ruffled feathers (1/5) | Air sac cloudiness (1/5) | 3/5 | 3/5 | |
| 5 house sparrows | No signs | No PM lesions | 1/5 | 1/5 | ||
| 4 | 5 cattle egrets | No signs | No PM lesions | 2/5 | 1/5 | |
| 5 house sparrows | Ruffled feathers (2/5) | No PM lesions | 1/5 | 1/5 | ||
| 5 | 5 cattle egrets | Whitish-green diarrhea (1/5) | Enteritis (1/5) | 2/5 | 2/5 | KFS-Motobas-2 strain |
| 5 house sparrows | No signs | No PM lesions | 1/5 | 1/5 | ||
| 6 | 5 cattle egrets | Ruffled feathers (1/5) | No PM lesions | 2/5 | 2/5 | |
| 5 house sparrows | No signs | No PM lesions | 1/5 | 0/5 | ||
| 7 | 5 cattle egrets | No signs | No PM lesions | 3/5 | 2/5 | |
| 5 house sparrows | Whitish-green diarrhea (1/5) | Enteritis (1/5) | 2/5 | 1/5 | ||
| 8 | 5 cattle egrets | Whitish-green diarrhea (1/5) | Enteritis (1/5) | 3/5 | 3/5 | |
| 5 house sparrows | No signs | No PM lesions | 2/5 | 2/5 | ||
| 9 | 5 cattle egrets | Ruffled feathers (2/5) | No PM lesions | 2/5 | 2/5 | |
| 5 house sparrows | Ruffled feathers | No PM lesions | 1/5 | 1/5 | ||
| 10 | 5 cattle egrets | No signs | No PM lesions | 1/5 | 0/5 | |
| 5 house sparrows | No signs | No PM lesions | 0/5 | 0/5 |
| Group | Clinical Signs | Number of Affected Birds (Days Post-Inoculation) | Total % Mortality | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 3rd | 4th | 5th | 6th | 7th | 8th | 9th | 10th | |||
| Depression and ruffled feathers | 12/40 | 30/34 | 26/30 | 23/25 | 21/24 | 8/14 | 6/12 | 3/9 | ||
| Decreased feed intake | 12/40 | 25/34 | 26/30 | 23/25 | 21/24 | 8/14 | 6/12 | 3/9 | ||
| G1 | Greenish diarrhea | 0/40 | 15/34 | 25/30 | 21/25 | 20/24 | 7/14 | 5/12 | 2/9 | |
| Swollen head | 0/40 | 17/34 | 23/30 | 20/25 | 20/24 | 7/14 | 5/12 | 2/9 | ||
| Respiratory signs | 0/40 | 17/34 | 23/30 | 20/25 | 20/24 | 7/14 | 5/12 | 2/9 | ||
| Nervous signs | 0/40 | 0/34 | 9/30 | 18/25 | 17/24 | 10/14 | 9/12 | 6/9 | ||
| Mortality | 3/40 | 4/34 | 5/30 | 1/25 | 7/24 | 2/14 | 3/12 | 3/9 | 32/40 (80%) | |
| Depression and ruffled feathers | 10/40 | 30/34 | 28/31 | 25/27 | 19/23 | 13/17 | 10/14 | 7/11 | ||
| Decreased feed intake | 10/40 | 27/34 | 29/31 | 25/27 | 19/23 | 13/17 | 10/14 | 7/11 | ||
| G2 | Greenish diarrhea | 0/40 | 17/34 | 22/31 | 21/27 | 15/23 | 9/17 | 6/14 | 3/11 | |
| Swollen head | 0/40 | 12/34 | 17/31 | 16/27 | 16/23 | 10/17 | 7/14 | 4/11 | ||
| Respiratory signs | 0/40 | 12/34 | 17/31 | 16/27 | 16/23 | 10/17 | 7/14 | 4/11 | ||
| Nervous signs | 0/40 | 0/34 | 8/31 | 19/27 | 16/23 | 12/17 | 9/14 | 6/11 | ||
| Mortality | 3/40 | 3/34 | 4/31 | 4/27 | 3/23 | 3/17 | 3/14 | 4/11 | 27/40 (67.5%) | |
| Depression and ruffled feathers | 9/40 | 26/35 | 28/31 | 26/28 | 22/24 | 15/17 | 10/12 | 8/10 | ||
| G3 | Decreased feed intake | 9/40 | 29/35 | 29/31 | 25/28 | 22/24 | 15/17 | 10/12 | 8/10 | |
| Greenish diarrhea | 0/40 | 17/35 | 21/31 | 22/28 | 18/24 | 11/17 | 7/12 | 5/10 | ||
| Swollen head | 0/40 | 16/35 | 19/31 | 19/28 | 15/24 | 9/17 | 5/12 | 3/10 | ||
| Respiratory signs | 0/40 | 16/35 | 19/31 | 19/28 | 15/24 | 9/17 | 5/12 | 3/10 | ||
| Nervous signs | 0/40 | 0/35 | 7/31 | 12/28 | 13/24 | 10/17 | 5/12 | 6/10 | ||
| Mortality | 2/40 | 4/35 | 3/31 | 4/28 | 4/24 | 5/17 | 2/12 | 1/10 | 25/40 (62.5%) | |
| G4 | Clinical signs | 0/40 | 0/40 | 0/40 | 0/40 | 0/40 | 0/40 | 0/40 | 0/40 | |
| Mortality | 0/40 | 0/40 | 0/40 | 0/40 | 0/40 | 0/40 | 0/40 | 0/40 | 0/40 (0%) | |
| Group | Post-Mortem Changes | Lesions in Dead Birds (Days Post-Inoculation) | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 3rd | 4th | 5th | 6th | 7th | 8th | 9th | 10th | ||||
| No. of examined birds | Dead | 3 | 4 | 5 | 1 | 7 | 2 | 3 | 3 | ||
| Euthanized | 3 | 0 | 0 | 0 | 3 | 0 | 0 | 3 | |||
| Hemorrhagic tracheitis | 0/6 | 3/4 | 3/5 | 0/1 | 9/10 | 2/2 | 2/3 | 6/6 | |||
| G1 | Lung congestion | 0/6 | 3/4 | 3/5 | 0/1 | 8/10 | 1/2 | 2/3 | 6/6 | ||
| Hemorrhagic cecal tonsils | 1/6 | 2/4 | 3/5 | 0/1 | 6/10 | 1/2 | 3/3 | 5/6 | |||
| Enlarged spleen | 1/6 | 4/4 | 5/5 | 1/1 | 9/10 | 2/2 | 3/3 | 5/6 | |||
| Hemorrhages on the periventricular gland tips | 0/6 | 3/4 | 3/5 | 1/1 | 10/10 | 2/2 | 2/3 | 6/6 | |||
| Greenish proventricular and intestinal contents | 0/6 | 3/4 | 5/5 | 1/1 | 8/10 | 2/2 | 3/3 | 6/6 | |||
| Hemorrhagic enteritis | 0/6 | 3/4 | 3/5 | 1/1 | 6/10 | 1/2 | 2/3 | 5/6 | |||
| No. of examined birds | Dead | 3 | 3 | 4 | 4 | 3 | 3 | 3 | 4 | ||
| Euthanized | 3 | 0 | 0 | 0 | 3 | 0 | 0 | 3 | |||
| G2 | Hemorrhagic tracheitis | 0/6 | 2/3 | 2/4 | 3/4 | 4/6 | 2/3 | 1/3 | 6/7 | ||
| Lung congestion | 0/6 | 3/3 | 3/4 | 2/4 | 4/6 | 1/3 | 0/3 | 5/7 | |||
| Hemorrhagic cecal tonsils | 2/6 | 2/3 | 1/4 | 3/4 | 3/6 | 1/3 | 2/3 | 3/7 | |||
| Enlarged spleen | 4/6 | 3/3 | 4/4 | 4/4 | 6/6 | 3/3 | 3/3 | 6/7 | |||
| Hemorrhages on the periventricular gland tips | 0/6 | 3/3 | 4/4 | 3/4 | 3/6 | 2/3 | 1/3 | 7/7 | |||
| Greenish proventricular | 0/6 | 3/3 | 4/4 | 4/4 | 4/6 | 3/3 | 3/3 | 7/7 | |||
| Hemorrhagic enteritis | 0/6 | 2/3 | 2/4 | 4/4 | 3/6 | 1/3 | 1/3 | 5/7 | |||
| No. of examined birds | Dead | 2 | 4 | 3 | 4 | 4 | 5 | 2 | 1 | ||
| Euthanized | 3 | 0 | 0 | 0 | 3 | 0 | 0 | 3 | |||
| G3 | Hemorrhagic tracheitis | 0/5 | 3/4 | 1/3 | 1/4 | 5/7 | 2/5 | 0/2 | 3/4 | ||
| Lung congestion | 0/5 | 2/4 | 2/3 | 2/4 | 6/7 | 3/5 | 1/2 | 2/4 | |||
| Hemorrhagic cecal tonsils | 0/5 | 3/4 | 2/3 | 3/4 | 5/7 | 4/5 | 0/2 | 0/4 | |||
| Enlarged spleen | 2/5 | 4/4 | 3/3 | 3/4 | 7/7 | 4/5 | 1/2 | 4/4 | |||
| Hemorrhages on the periventricular gland tips | 0/5 | 2/4 | 3/3 | 2/4 | 7/7 | 2/5 | 2/2 | 3/4 | |||
| Greenish proventricular and intestinal contents | 0/5 | 3/4 | 3/3 | 4/4 | 7/7 | 5/5 | 2/2 | 4/4 | |||
| Hemorrhagic enteritis | 0/5 | 1/4 | 2/3 | 1/4 | 5/7 | 3/5 | 0/2 | 1/4 | |||
| G4 | No. of examined birds | Dead | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | |
| Euthanized | 3 | 0 | 0 | 0 | 3 | 0 | 0 | 3 | |||
| PM lesions | 0/3 | 0/0 | 0/0 | 0/0 | 0/3 | 0/0 | 0/0 | 0/3 | |||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abd Elfatah, K.S.; Elabasy, M.A.; El-khyate, F.; Elmahallawy, E.K.; Mosad, S.M.; El-Gohary, F.A.; Abdo, W.; Al-Brakati, A.; Seadawy, M.G.; Tahoon, A.E.; et al. Molecular Characterization of Velogenic Newcastle Disease Virus (Sub-Genotype VII.1.1) from Wild Birds, with Assessment of Its Pathogenicity in Susceptible Chickens. Animals 2021, 11, 505. https://doi.org/10.3390/ani11020505
Abd Elfatah KS, Elabasy MA, El-khyate F, Elmahallawy EK, Mosad SM, El-Gohary FA, Abdo W, Al-Brakati A, Seadawy MG, Tahoon AE, et al. Molecular Characterization of Velogenic Newcastle Disease Virus (Sub-Genotype VII.1.1) from Wild Birds, with Assessment of Its Pathogenicity in Susceptible Chickens. Animals. 2021; 11(2):505. https://doi.org/10.3390/ani11020505
Chicago/Turabian StyleAbd Elfatah, Khaled Saad, Moshira Abas Elabasy, Faris El-khyate, Ehab Kotb Elmahallawy, Samah M. Mosad, Fatma A. El-Gohary, Walied Abdo, Ashraf Al-Brakati, Mohamed G. Seadawy, Abd Elnaby Tahoon, and et al. 2021. "Molecular Characterization of Velogenic Newcastle Disease Virus (Sub-Genotype VII.1.1) from Wild Birds, with Assessment of Its Pathogenicity in Susceptible Chickens" Animals 11, no. 2: 505. https://doi.org/10.3390/ani11020505
APA StyleAbd Elfatah, K. S., Elabasy, M. A., El-khyate, F., Elmahallawy, E. K., Mosad, S. M., El-Gohary, F. A., Abdo, W., Al-Brakati, A., Seadawy, M. G., Tahoon, A. E., & El-Gohary, A. E. (2021). Molecular Characterization of Velogenic Newcastle Disease Virus (Sub-Genotype VII.1.1) from Wild Birds, with Assessment of Its Pathogenicity in Susceptible Chickens. Animals, 11(2), 505. https://doi.org/10.3390/ani11020505







