Antioxidant and Anti-Inflammatory Properties of Probiotic Candidate Strains Isolated during Fermentation of Agave (Agave angustifolia Haw)
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Sampling and Growth Conditions
2.2. Lysozyme, Low pH, and Bile Salt Resistance Tests
2.3. Adhesion and Antioxidant Assays
2.4. Assays in TNF-α-Activated HT-29 Cells
2.5. Analysis of LAB Strains against Oxidative Stress in HT-29 Cells
2.6. Animal Experiments
2.7. Macroscopic Scores
2.8. In Vivo Permeability Assay (FITC) and Myeloperoxidase Activity
2.9. Statistics
3. Results
3.1. Strain Identification
3.2. Lysozyme, pH, and Bile Salt Tolerance of the Isolated Bacteria
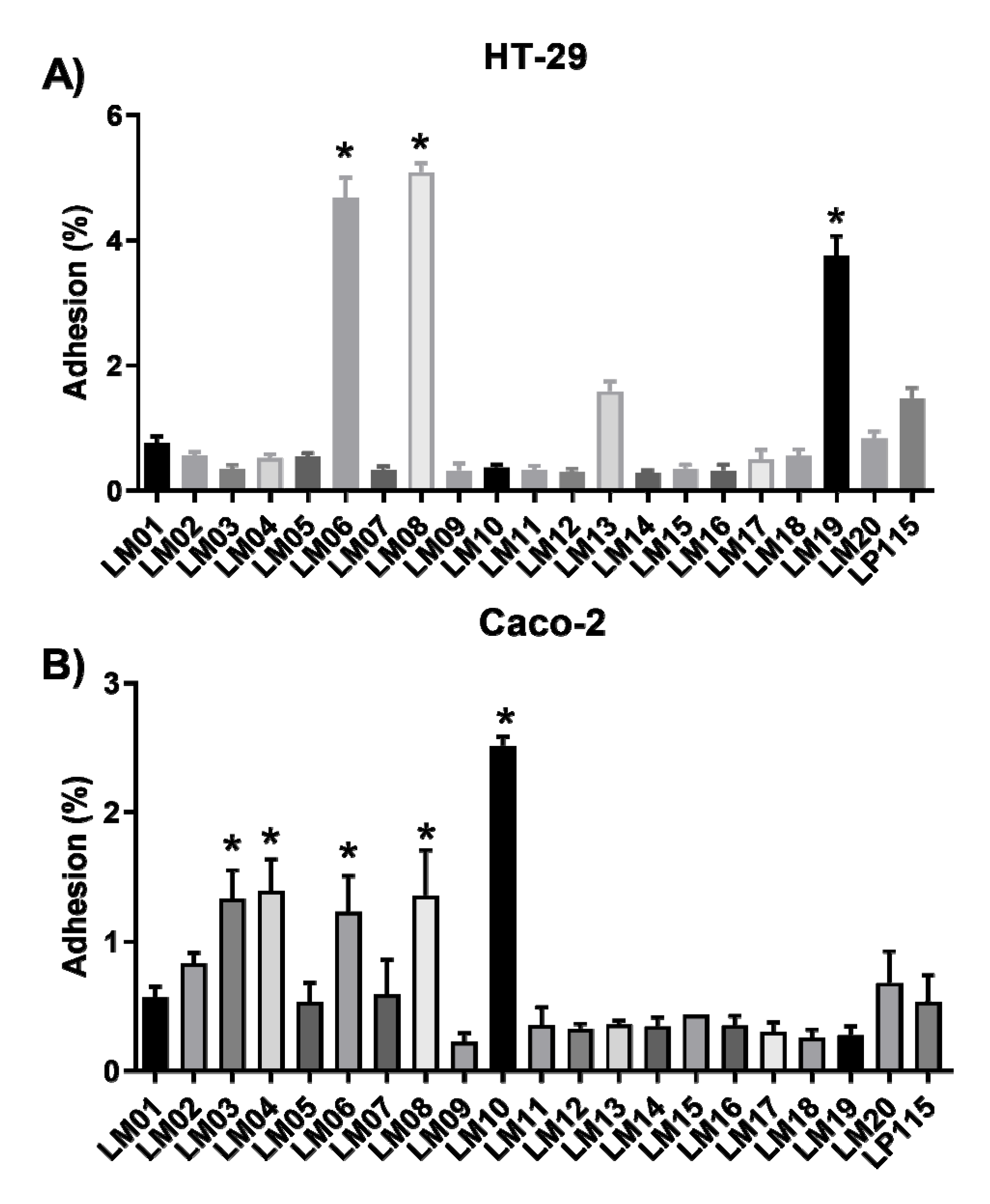
3.3. Cell Culture Methods and Biochemical Characterization of the Isolated Strains
3.4. Screening of Bacterial Strains in HT-29 Cells Stimulated with TNF-α
3.5. Role of LAB against Oxidative Stress
3.6. Anti-Inflammatory Effects of LAB Strains Isolated from Agave
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Hill, C.; Guarner, F.; Reid, G.; Gibson, G.R.; Merenstein, D.J.; Pot, B.; Morelli, L.; Canani, R.B.; Flint, H.J.; Salminen, S.; et al. Expert consensus document: The international scientific association for probiotics and prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat. Rev. Gastroenterol. Hepatol. 2014, 11, 506–514. [Google Scholar] [CrossRef]
- Choi, S.S.; Kim, Y.; Han, K.S.; You, S.; Oh, S.; Kim, S.H. Effects of Lactobacillus strains on cancer cell proliferation and oxidative stress in vitro. Lett. Appl. Microbiol. 2006, 42, 452–458. [Google Scholar] [CrossRef]
- George Kerry, R.; Patra, J.K.; Gouda, S.; Park, Y.; Shin, H.S.; Das, G. Benefaction of probiotics for human health: A review. J. Food Drug Anal. 2018, 26, 927–939. [Google Scholar] [CrossRef]
- Kaizu, H.; Sasaki, M.; Nakajima, H.; Suzuki, Y. Effect of antioxidative lactic acid bacteria on rats fed a diet deficient in vitamin E. J. Dairy Sci. 1993, 76, 2493–2499. [Google Scholar] [CrossRef]
- Kullisaar, G.; Songisepp, E.; Zilmer, M. Probiotics and oxidative stress. In Oxidative Stress Environmental Induction and Dietary Antioxidants; Lushchak, V., Ed.; InTech: Rijeka, Croatia, 2012; p. 402. ISBN 978-953-51-0553-4. [Google Scholar]
- Spyropoulos, B.G.; Misiakos, E.P.; Fotiadis, C.; Stoidis, C.N. Antioxidant properties of probiotics and their protective effects in the pathogenesis of radiation-induced enteritis and colitis. Dig. Dis. Sci. 2011, 56, 285–294. [Google Scholar] [CrossRef] [PubMed]
- Achuthan, A.A.; Duary, R.K.; Madathil, A.; Panwar, H.; Kumar, H.; Batish, V.K.; Grover, S. Antioxidative potential of lactobacilli isolated from the gut of Indian people. Mol. Biol. Rep. 2012, 39, 7887–7897. [Google Scholar] [CrossRef] [PubMed]
- Xing, J.; Wang, G.; Gu, Z.; Liu, X.; Zhang, Q.; Zhao, J.; Zhang, H.; Chen, Y.Q.; Chen, W. Cellular model to assess the antioxidant activity of lactobacilli. R. Soc. Chem. Adv. 2015, 5, 37626–37634. [Google Scholar] [CrossRef]
- Maresca, D.; Zotta, T.; Mauriello, G. Adaptation to aerobic environment of Lactobacillus johnsonii/gasseri strains. Front. Microbiol. 2018, 9, 1–11. [Google Scholar] [CrossRef]
- Mishra, V.; Shah, C.; Mokashe, N.; Chavan, R.; Yadav, H.; Prajapati, J. Probiotics as Potential Antioxidants: A Systematic Review. J. Agric. Food Chem. 2015, 63, 3615–3626. [Google Scholar] [CrossRef]
- Wu, R.; Wang, L.; Wang, J.; Li, H.; Menghe, B.; Wu, J.; Guo, M.; Zhang, H. Isolation and preliminary probiotic selection of lactobacilli from koumiss in Inner Mongolia. J. Basic Microbiol. 2009, 49, 318–326. [Google Scholar] [CrossRef]
- Lee, J.; Yun, H.S.; Cho, K.W.; Oh, S.; Kim, S.H.; Chun, T.; Kim, B.; Whang, K.Y. Evaluation of probiotic characteristics of newly isolated Lactobacillus spp.: Immune modulation and longevity. Int. J. Food Microbiol. 2011, 148, 80–86. [Google Scholar] [CrossRef] [PubMed]
- Torres-Maravilla, E.; Lenoir, M.; Mayorga-Reyes, L.; Allain, T.; Sokol, H.; Langella, P.; Sánchez-Pardo, M.E.; Bermúdez-Humarán, L.G. Identification of novel anti-inflammatory probiotic strains isolated from pulque. Appl. Microbiol. Biotechnol. 2015. [Google Scholar] [CrossRef] [PubMed]
- Durán-García, H.M.; González-Galván, E.J.; Matadamas-Ortíz, P. Mechanization process in the production of mezcal. J. Food Agric. Environ. 2007, 5, 32–35. [Google Scholar]
- Escalante-Minakata, P.; Blaschek, H.P.; Barba de la Rosa, A.P.; Santos, L.; De León-Rodríguez, A. Identification of yeast and bacteria involved in the mezcal fermentation of Agave salmiana. Lett. Appl. Microbiol. 2008, 46, 626–630. [Google Scholar] [CrossRef]
- Weisburg, W.G.; Barns, S.M.; Pelletier, D.A.; Lane, D.J. 16S Ribosomal DNA Amplification for Phylogenetic Study. J. Bacteriol. 1991, 173, 697–703. [Google Scholar] [CrossRef] [PubMed]
- Paroschi, T.P.; Breganó, J.W.; Simão, A.N.; Dichi, I.; Miglioranza, L.H. Effects of Sulfasalazine, Lactobacillus Plantarum (Lp-115) and Fish Oil in Experimental Colitis. J. Food Nutr. Disord. 2015, 1, 1–4. [Google Scholar]
- Zhou, Y.; Wang, J.Q.; Hu, C.H.; Ren, L.Q.; Wang, D.C.; Ye, B.C. Enhancement of bile resistance by maltodextrin supplementation in Lactobacillus plantarum Lp-115. J. Appl. Microbiol. 2019, 126, 1551–1557. [Google Scholar] [CrossRef]
- Vizoso Pinto, M.G.; Franz, C.M.A.P.; Schillinger, U.; Holzapfel, W.H. Lactobacillus spp. with in vitro probiotic properties from human faeces and traditional fermented products. Int. J. Food Microbiol. 2006, 109, 205–214. [Google Scholar] [CrossRef]
- Kechaou, N.; Chain, F.; Gratadoux, J.J.; Blugeon, S.; Bertho, N.; Chevalier, C.; Le Goffic, R.; Courau, S.; Molimard, P.; Chatel, J.M.; et al. Identification of one novel candidate probiotic Lactobacillus plantarum strain active against influenza virus infection in mice by a large-scale screening. Appl. Environ. Microbiol. 2013, 79, 1491–1499. [Google Scholar] [CrossRef]
- Muñoz-Provencio, D.; Pérez-Martínez, G.; Monedero, V. Characterization of a fibronectin-binding protein from Lactobacillus casei BL23. J. Appl. Microbiol. 2010, 108, 1050–1059. [Google Scholar] [CrossRef]
- Su, J.; Wang, T.; Li, Y.-Y.; Li, J.; Zhang, Y.; Wang, Y.; Wang, H.; Li, H. Antioxidant properties of wine lactic acid bacteria: Oenococcus oeni. Appl. Microbiol. Biotechnol. 2015, 99, 5189–5202. [Google Scholar] [CrossRef] [PubMed]
- Wang, A.N.; Yi, X.W.; Yu, H.F.; Dong, B.; Qiao, S.Y. Free radical scavenging activity of Lactobacillus fermentum in vitro and its antioxidative effect on growing-finishing pigs. J. Appl. Microbiol. 2009, 107, 1140–1148. [Google Scholar] [CrossRef]
- Linden, A.; Gu, M.; Maser, E.; Seibert, H. Peroxide-induced cell death and lipid peroxidation in C6 glioma cells. Toxicol. Vitr. 2008, 22, 1371–1376. [Google Scholar] [CrossRef] [PubMed]
- Dassarma, B.; Nandi, D.K.; Gangopadhyay, S.; Samanta, S. Hepatoprotective effect of food preservatives (butylated hydroxyanisole, butylated hydroxytoluene) on carbon tetrachloride-induced hepatotoxicity in rat. Toxicol. Rep. 2018, 5, 31–37. [Google Scholar] [CrossRef] [PubMed]
- Martín, R.; Chain, F.; Miquel, S.; Lu, J.; Gratadoux, J.J.; Sokol, H.; Verdu, E.F.; Bercik, P.; Bermúdez-Humarán, L.G.; Langella, P. The commensal bacterium faecalibacterium prausnitzii is protective in DNBS-induced chronic moderate and severe colitis models. Inflamm. Bowel Dis. 2014, 20, 417–430. [Google Scholar] [CrossRef] [PubMed]
- Martín, R.; Miquel, S.; Chain, F.; Natividad, J.M.; Jury, J.; Lu, J.; Sokol, H.; Theodorou, V.; Bercik, P.; Verdu, E.F.; et al. Faecalibacterium prausnitzii prevents physiological damages in a chronic low-grade inflammation murine model. BMC Microbiol. 2015, 15, 67. [Google Scholar] [CrossRef] [PubMed]
- Bradley, P.; Priebat, D.; Christensen, R. Measurement of cutaneous inflammation: Estimation of neutrophil content with an enzyme marker. J. Investig. Dermatol. 1982, 206–209. [Google Scholar] [CrossRef]
- Narváez-Zapata, J.A.; Rojas-Herrera, R.A.; Rodríguez-Luna, I.C.; Larralde-Corona, C.P. Culture-Independent Analysis of Lactic Acid Bacteria Diversity Associated with Mezcal Fermentation. Curr. Microbiol. 2010, 61, 444–450. [Google Scholar] [CrossRef]
- Douillard, F.P.; Ribbera, A.; Kant, R.; Pietilä, T.E.; Järvinen, H.M.; Messing, M.; Randazzo, C.L.; Paulin, L.; Laine, P.; Ritari, J.; et al. Comparative Genomic and Functional Analysis of 100 Lactobacillus rhamnosus Strains and Their Comparison with Strain GG. PLoS Genet. 2013, 9. [Google Scholar] [CrossRef]
- Samedi, L.; Charles, A.L. Isolation and characterization of potential probiotic Lactobacilli from leaves of food plants for possible additives in pellet feeding. Ann. Agric. Sci. 2019, 64, 55–62. [Google Scholar] [CrossRef]
- Kubota, H.; Senda, S.; Nomura, N.; Tokuda, H.; Uchiyama, H. Biofilm Formation by Lactic Acid Bacteria and Resistance to Environmental Stress. J. Biosci. Bioeng. 2008, 106, 381–386. [Google Scholar] [CrossRef]
- Corcoran, B.M.; Stanton, C.; Fitzgerald, G.F.; Ross, R.P. Survival of probiotic lactobacilli in acidic environments is enhanced in the presence of metabolizable sugars. Appl. Environ. Microbiol. 2005, 71, 3060–3067. [Google Scholar] [CrossRef]
- Dlamini, Z.C.; Langa, R.L.S.; Aiyegoro, O.A.; Okoh, A.I. Safety Evaluation and Colonisation Abilities of Four Lactic Acid Bacteria as Future Probiotics. Probiotics Antimicrob. Proteins 2019, 11, 397–402. [Google Scholar] [CrossRef] [PubMed]
- Pelletier, C.; Bouley, C.; Cayuela, C.; Bouttier, S.; Bourlioux, P.; Bellon-Fontaine, M. Cell surface characteristics of Lactobacillus casei subsp. casei, Lactobacillus paracasei subsp. paracasei, and Lactobacillus rhamnosus strains. Appl. Environ. Microbiol. 1997, 63, 1725–1731. [Google Scholar] [CrossRef]
- Yang, S.J.; Kim, K.T.; Kim, T.Y.; Paik, H.D. Probiotic properties and antioxidant activities of Pediococcus pentosaceus SC28 and Levilactobacillus brevis KU15151 in fermented black gamju. Foods 2020, 9, 1154. [Google Scholar] [CrossRef] [PubMed]
- Muñoz-Provencio, D.; Rodríguez-Díaz, J.; Collado, M.C.; Langella, P.; Bermúdez-Humarán, L.G.; Monedero, V. Functional Analysis of the Lactobacillus casei BL23 Sortases. Appl. Environ. Microbiol. 2012, 78, 8684–8693. [Google Scholar] [CrossRef] [PubMed]
- Benítez-Cabello, A.; Torres-Maravilla, E.; Bermúdez-Humarán, L.; Langella, P.; Martín, R.; Jiménez-Díaz, R.; Arroyo-López, F.N. Probiotic Properties of Lactobacillus Strains Isolated from Table Olive Biofilms. Probiotics Antimicrob. Proteins 2020, 12, 1071–1082. [Google Scholar] [CrossRef]
- Kim, H.; Jung, B.J.; Jung, J.H.; Kim, J.Y.; Chung, S.K.; Chung, D.K. Lactobacillus plantarum lipoteichoic acid alleviates TNF-α-induced inflammation in the HT-29 intestinal epithelial cell line. Mol. Cells 2012, 33, 479–486. [Google Scholar] [CrossRef]
- Jeffrey, M.P.; Strap, J.L.; Taggart, H.J.; Green-Johnson, J.M. Lactobacillus rhamnosus R0011 and Lactobacillus helveticus R0389 is mediated by secreted bioactive molecules. Front. Immunol. 2018, 9, 1–13. [Google Scholar] [CrossRef]
- Jeffrey, M.P.; MacPherson, C.W.; Mathieu, O.; Tompkins, T.A.; Green-Johnson, J.M. Secretome-Mediated Interactions with Intestinal Epithelial Cells: A Role for Secretome Components from Lactobacillus rhamnosus R0011 in the Attenuation of Salmonella enterica Serovar Typhimurium Secretome and TNF-α–Induced Proinflammatory Responses. J. Immunol. 2020, 204, 2523–2534. [Google Scholar] [CrossRef]
- Li, S.; Zhao, Y.; Zhang, L.; Zhang, X.; Huang, L.; Li, D.; Niu, C.; Yang, Z.; Wang, Q. Antioxidant activity of Lactobacillus plantarum strains isolated from traditional Chinese fermented foods. Food Chem. 2012, 135, 1914–1919. [Google Scholar] [CrossRef]
- Lin, M.Y.; Chang, F.J. Antioxidative effect of intestinal bacteria Bifidobacterium longum ATCC 15708 and Lactobacillus acidophilus ATCC 4356. Dig. Dis. Sci. 2000, 45, 1617–1622. [Google Scholar] [CrossRef]
- Stecchini, M.L.; Del Torre, M.; Munari, M. Determination of peroxy radical-scavenging of lactic acid bacteria. Int. J. Food Microbiol. 2001, 64, 183–188. [Google Scholar] [CrossRef]
- Sanders, J.W.; Leenhouts, K.J.; Haandrikman, A.J.; Venema, G.; Kok, J. Stress response in Lactococcus lactis: Cloning, expression analysis, and mutation of the lactococcal superoxide dismutase gene. J. Bacteriol. 1995, 177, 5254–5260. [Google Scholar] [CrossRef][Green Version]
- Kang, D.K.; Oh, H.K.; Ham, J.S.; Kim, J.G.; Yoon, C.H.; Ahn, Y.T.; Kim, H.U. Identification and characterization of hydrogen peroxide-generating Lactobacillus fermentum CS12-1. Asian-Australas. J. Anim. Sci. 2005, 18, 90–95. [Google Scholar] [CrossRef]
- Marsova, M.; Abilev, S.; Poluektova, E.; Danilenko, V. A bioluminescent test system reveals valuable antioxidant properties of lactobacillus strains from human microbiota. World J. Microbiol. Biotechnol. 2018, 34, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Kullisaar, T.; Zilmer, M.; Mikelsaar, M.; Vihalemm, T.; Annuk, H.; Kairane, C.; Kilk, A. Two antioxidative lactobacilli strains as promising probiotics. Int. J. Food Microbiol. 2002, 72, 215–224. [Google Scholar] [CrossRef]
- Morampudi, V.; Bhinder, G.; Wu, X.; Dai, C.; Sham, H.P.; Vallance, B.A.; Jacobson, K. DNBS/TNBS colitis models: Providing insights into inflammatory bowel disease and effects of dietary fat. J. Vis. Exp. 2014, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Yoda, K.; Miyazawa, K.; Hosoda, M.; Hiramatsu, M.; Yan, F.; He, F. Lactobacillus GG-fermented milk prevents DSS-induced colitis and regulates intestinal epithelial homeostasis through activation of epidermal growth factor receptor. Eur. J. Nutr. 2014, 53, 105–115. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Li, Y.; Wan, Y.; Hu, T.; Liu, L.; Yang, S.; Gong, Z.; Zeng, Q.; Wei, Y.; Yang, W.; et al. A novel postbiotic from lactobacillus rhamnosus GG with a beneficial effect on intestinal barrier function. Front. Microbiol. 2019, 10, 1–14. [Google Scholar] [CrossRef]
- Mao, Y.; Nobaek, S.; Kasravi, B.; Adawi, D.; Stenram, U.; Molin, G.; Jeppsson, B. The effects of Lactobacillus strains and oat fiber on methotrexate- induced enterocolitis in rats. Gastroenterology 1996, 111, 334–344. [Google Scholar] [CrossRef] [PubMed]
- Hansberry, D.R.; Shah, K.; Agarwal, P.; Agarwal, N. Fecal Myeloperoxidase as a Biomarker for Inflammatory Bowel Disease. Cureus 2017. [Google Scholar] [CrossRef] [PubMed]
- Collado, M.C.; Bäuerl, C.; Pérez-Martínez, G. Defining microbiota for developing new probiotics. Microb. Ecol. Health Dis. 2012, 23, 35–39. [Google Scholar] [CrossRef] [PubMed]




| Code | Microorganism | Region | 16S rRNA (%) |
|---|---|---|---|
| LM01 | Lactobacillus plantarum | Macuilxóchilt de Ártigas | 98.8 |
| LM02 | Lactobacillus plantarum | Macuilxóchilt de Ártigas | 99 |
| LM03 | Lactobacillus plantarum | Macuilxóchilt de Ártigas | 95.7 |
| LM04 | Lactococcus lactis ssp. lactis | Tlacolula de Matamoros | 99 |
| LM05 | Lactobacillus plantarum | Tlacolula de Matamoros | 97.4 |
| LM06 | Enterococcus faecium | Santiago Matatlán | 99 |
| LM07 | Lactobacillus rhamnosus | Tlacolula de Matamoros | 100 |
| LM08 | Lactobacillus plantarum | Tlacolula de Matamoros | 98.1 |
| LM09 | Lactobacillus plantarum | Macuilxóchilt de Ártigas | 97.5 |
| LM10 | Lactobacillus plantarum | Macuilxóchilt de Ártigas | 97.4 |
| LM11 | Lactobacillus plantarum | Tlacolula de Matamoros | 96.7 |
| LM12 | Lactobacillus plantarum | Santiago Matatlán | 96 |
| LM13 | Lactobacillus plantarum | Tlacolula de Matamoros | 97 |
| LM14 | Lactobacillus plantarum | Tlacolula de Matamoros | 97.6 |
| LM15 | Enterococcus faecium | Macuilxóchilt de Ártigas | 99 |
| LM16 | Lactobacillus rhamnosus | Tlacolula de Matamoros | 99 |
| LM17 | Lactobacillus plantarum | Tlacolula de Matamoros | 99.9 |
| LM18 | Lactobacillus plantarum | Tlacolula de Matamoros | 97 |
| LM19 | Lactobacillus plantarum | Santiago Matatlán | 99 |
| LM20 | Lactobacillus plantarum | Macuilxóchilt de Ártigas | 99 |
| Isolate | Lysozyme | pH 2.5 | 0.3% Bile Salt | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 180 | Survival (%) | 0 | 90 | 180 | Survival (%) | 0 | 90 | 180 | Survival (%) | |
| Log CFU/mL | Log CFU/mL | Log CFU/mL | |||||||||
| LM01 | 9.19 | 8.93 | 54.70 * | 9.19 | 9.09 | 8.77 | 38.10 | 9.19 | 9.10 | 8.92 | 53.68 |
| LM02 | 9.14 | 8.97 | 67.15 * | 9.31 | 8.56 | 8.33 | 10.46 | 9.31 | 9.23 | 8.70 | 24.84 |
| LM03 | 9.35 | 8.97 | 41.59 * | 9.35 | 9.04 | 8.74 | 24.70 | 9.35 | 9.11 | 8.92 | 37.50 |
| LM04 | 9.32 | 9.13 | 64.76 * | 9.33 | 9.02 | 8.92 | 38.58 | 9.33 | 9.23 | 9.00 | 45.99 |
| LM05 | 9.39 | 8.97 | 38.25 * | 9.35 | 9.03 | 8.82 | 28.91 | 9.35 | 9.21 | 9.02 | 46.31 |
| LM06 | 9.25 | 8.80 | 35.58 * | 9.39 | 8.89 | 8.56 | 14.52 | 9.39 | 9.28 | 8.72 | 21.24 |
| LM07 | 9.18 | 9.00 | 66.22 * | 9.29 | 8.71 | 8.44 | 13.95 | 9.29 | 9.21 | 8.75 | 28.57 |
| LM08 | 9.31 | 8.99 | 47.57 * | 9.45 | 9.18 | 8.97 | 33.10 | 9.45 | 9.28 | 8.94 | 31.19 |
| LM09 | 9.38 | 9.00 | 41.05 * | 9.36 | 9.08 | 8.83 | 29.82 | 9.36 | 9.24 | 8.88 | 33.04 |
| LM10 | 9.42 | 8.90 | 29.80 * | 9.35 | 9.13 | 9.00 | 44.35 | 9.35 | 9.27 | 8.95 | 39.58 |
| LM11 | 9.43 | 9.15 | 52.24 * | 9.21 | 8.76 | 8.66 | 27.64 | 9.21 | 9.15 | 8.97 | 57.32 * |
| LM12 | 9.25 | 8.95 | 50.19 * | 9.25 | 8.94 | 8.73 | 30.68 | 9.25 | 9.13 | 8.85 | 40.53 |
| LM13 | 9.15 | 8.85 | 49.77 * | 9.31 | 9.09 | 9.05 | 55.56 * | 9.31 | 9.14 | 9.06 | 55.88 * |
| LM14 | 9.19 | 8.82 | 42.31 * | 8.73 | 8.64 | 8.52 | 61.73 * | 8.73 | 8.62 | 8.52 | 61.73 * |
| LM15 | 9.27 | 9.16 | 77.06 * | 8.94 | 8.73 | 8.51 | 36.36 | 8.94 | 8.94 | 8.85 | 81.06 * |
| LM16 | 9.33 | 9.14 | 64.78 * | 8.86 | 8.60 | 8.50 | 43.52 | 8.86 | 8.64 | 8.51 | 44.44 |
| LM17 | 9.04 | 8.45 | 25.45 | 9.30 | 9.12 | 9.07 | 59.93 * | 9.3 | 9.22 | 8.93 | 43.43 |
| LM18 | 9.36 | 8.90 | 34.20 * | 9.26 | 9.12 | 8.84 | 38.15 | 9.26 | 8.99 | 8.85 | 39.63 |
| LM19 | 9.32 | 9.06 | 55.45 * | 8.96 | 8.93 | 8.80 | 68.84 * | 8.96 | 8.93 | 8.90 | 86.23 * |
| LM20 | 9.35 | 9.06 | 51.79 * | 9.06 | 9.00 | 8.94 | 75.86 * | 9.06 | 8.94 | 8.88 | 64.94 * |
| Lp115 | 9.44 | 8.88 | 27.74 | 8.97 | 8.92 | 8.56 | 38.30 | 8.97 | 8.78 | 8.67 | 49.65 |
| 2 mM Hydrogen Peroxide | |||||
|---|---|---|---|---|---|
| Group | MDA nmol/mg Protein | TAS mmol/mg Protein | SOD U/mg Protein | GPx U/mg Protein | CAT U/mg Protein |
| PBS | 5.20 ± 0.03 b,c | 0.16 ± 0.001 b,c | 18.45 ± 0.0003 b,c | 1.15 ± 0.08 b,c | 0.87 ± 0.11 b,c |
| BHT | 1.10 ± 0.02 a,c | 2.24 ± 0.03 a | 28.39 ± 0.001 a,c | 5.59 ± 0.05 a | 4.57 ± 0.06 a,c |
| Lp115 | 1.28 ± 0.07 a,b | 1.92 ± 0.65 a | 26.64 ± 0.49 a,b | 4.99 ± 0.06 a | 3.67 ± 0.12 a,b |
| LM07 | 2.11 ± 0.06 a,b,c | 1.40 ± 0.12 a,b,c | 22.36 ± 0.005 a,b,c | 4.38 ± 0.10 a,b,c | 3.36 ± 0.37 a,b |
| LM17 | 2.49 ± 0.11 a,b,c | 2.10 ± 0.08 a | 23.15 ± 0.003 a,b,c | 4.11 ± 0.11 a,b,c | 3.95 ± 0.21 a |
| LM19 | 2.73 ± 0.02 a,b,c | 1.73 ± 0.14 a,b | 23.42 ± 0.001 a,b,c | 4.39 ± 0.21 a,b,c | 3.25 ± 0.12 a,b |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hernández-Delgado, N.C.; Torres-Maravilla, E.; Mayorga-Reyes, L.; Martín, R.; Langella, P.; Pérez-Pastén-Borja, R.; Sánchez-Pardo, M.E.; Bermúdez-Humarán, L.G. Antioxidant and Anti-Inflammatory Properties of Probiotic Candidate Strains Isolated during Fermentation of Agave (Agave angustifolia Haw). Microorganisms 2021, 9, 1063. https://doi.org/10.3390/microorganisms9051063
Hernández-Delgado NC, Torres-Maravilla E, Mayorga-Reyes L, Martín R, Langella P, Pérez-Pastén-Borja R, Sánchez-Pardo ME, Bermúdez-Humarán LG. Antioxidant and Anti-Inflammatory Properties of Probiotic Candidate Strains Isolated during Fermentation of Agave (Agave angustifolia Haw). Microorganisms. 2021; 9(5):1063. https://doi.org/10.3390/microorganisms9051063
Chicago/Turabian StyleHernández-Delgado, Natalia C., Edgar Torres-Maravilla, Lino Mayorga-Reyes, Rebeca Martín, Philippe Langella, Ricardo Pérez-Pastén-Borja, María E. Sánchez-Pardo, and Luis G. Bermúdez-Humarán. 2021. "Antioxidant and Anti-Inflammatory Properties of Probiotic Candidate Strains Isolated during Fermentation of Agave (Agave angustifolia Haw)" Microorganisms 9, no. 5: 1063. https://doi.org/10.3390/microorganisms9051063
APA StyleHernández-Delgado, N. C., Torres-Maravilla, E., Mayorga-Reyes, L., Martín, R., Langella, P., Pérez-Pastén-Borja, R., Sánchez-Pardo, M. E., & Bermúdez-Humarán, L. G. (2021). Antioxidant and Anti-Inflammatory Properties of Probiotic Candidate Strains Isolated during Fermentation of Agave (Agave angustifolia Haw). Microorganisms, 9(5), 1063. https://doi.org/10.3390/microorganisms9051063







