Exploring the Impact of the Biofloc Rearing System and an Oral WSSV Challenge on the Intestinal Bacteriome of Litopenaeus vannamei
Abstract
1. Introduction
2. Materials and Methods
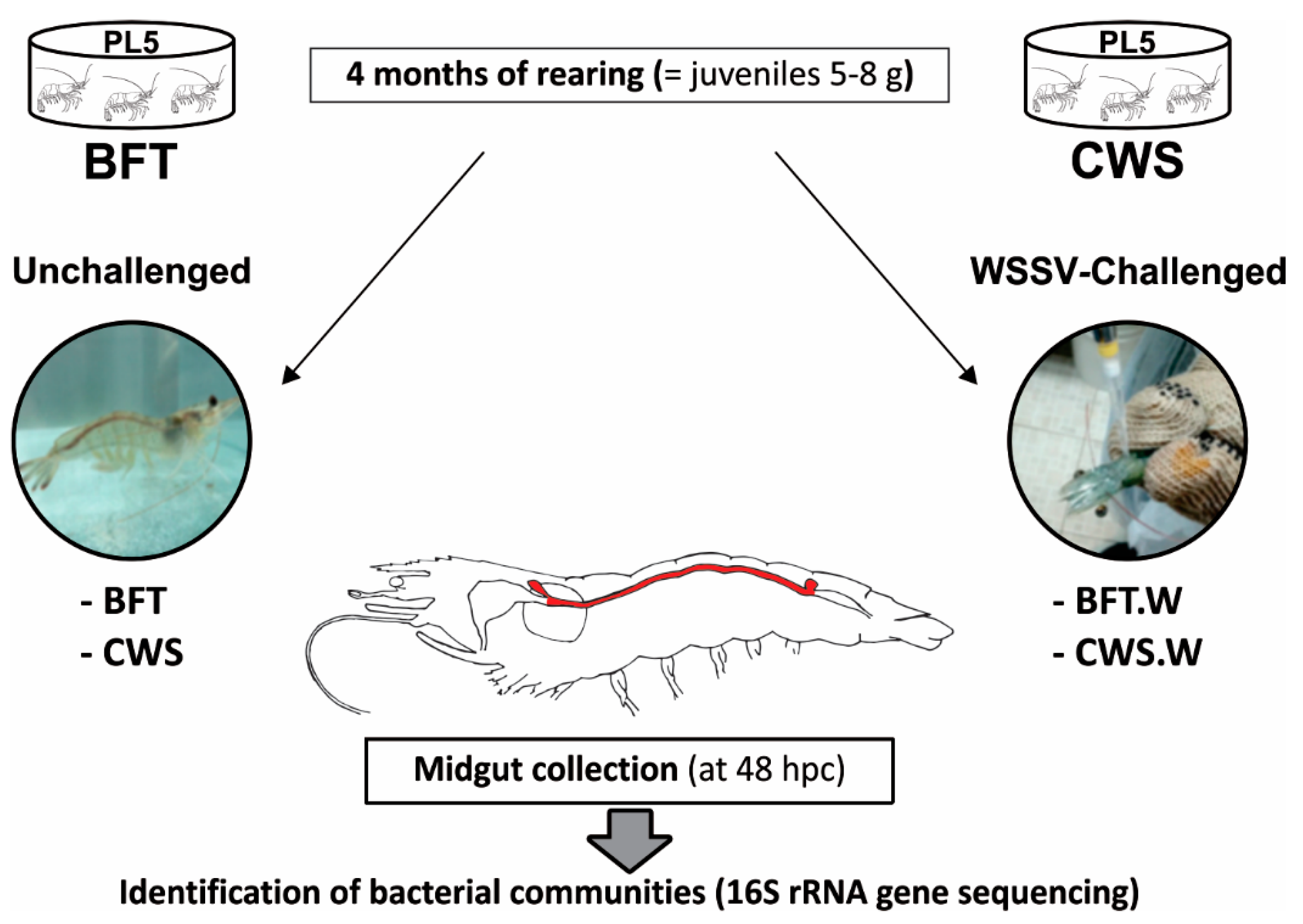
2.1. Animals and Experimental Design
2.2. WSSV Per Os Challenge and Midgut Collection
2.3. Genomic DNA (gDNA) Extraction
2.4. 16S rRNA Gene Library Preparation and High Throughput Sequencing
2.5. Sequence Data Analysis
3. Results and Discussion
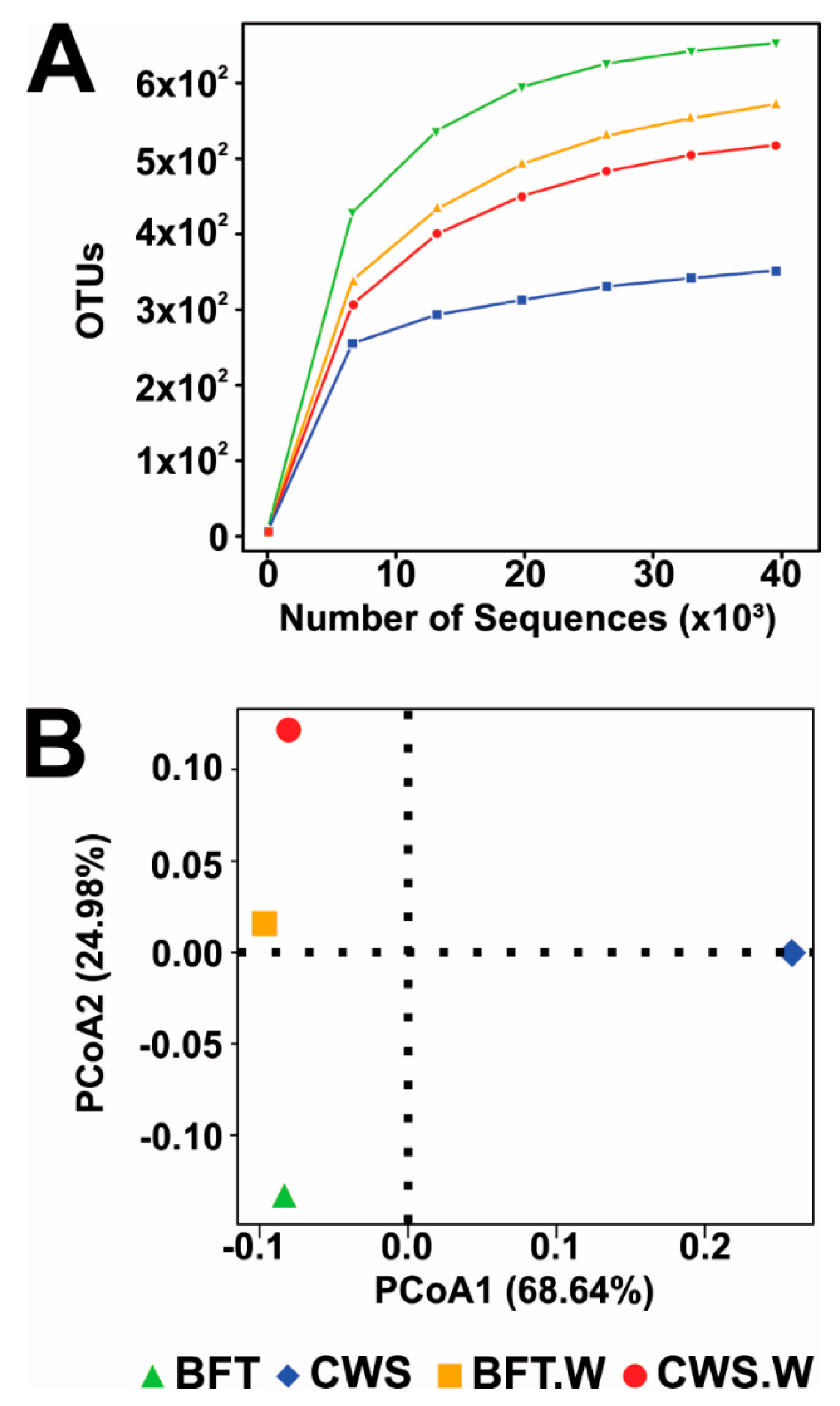
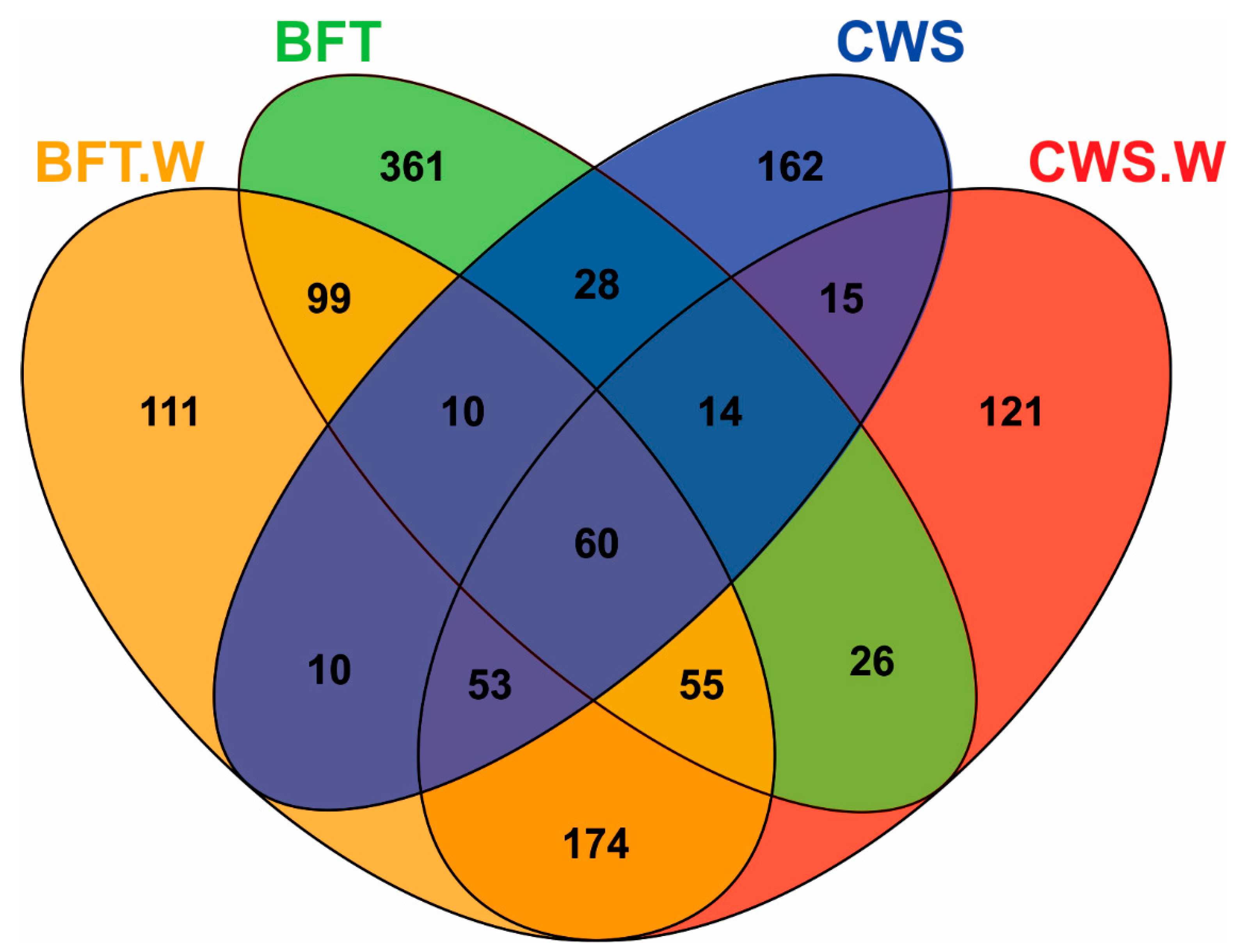
3.1. Overview of the Illumina Sequencing, Diversity, and Richness of Bacteria
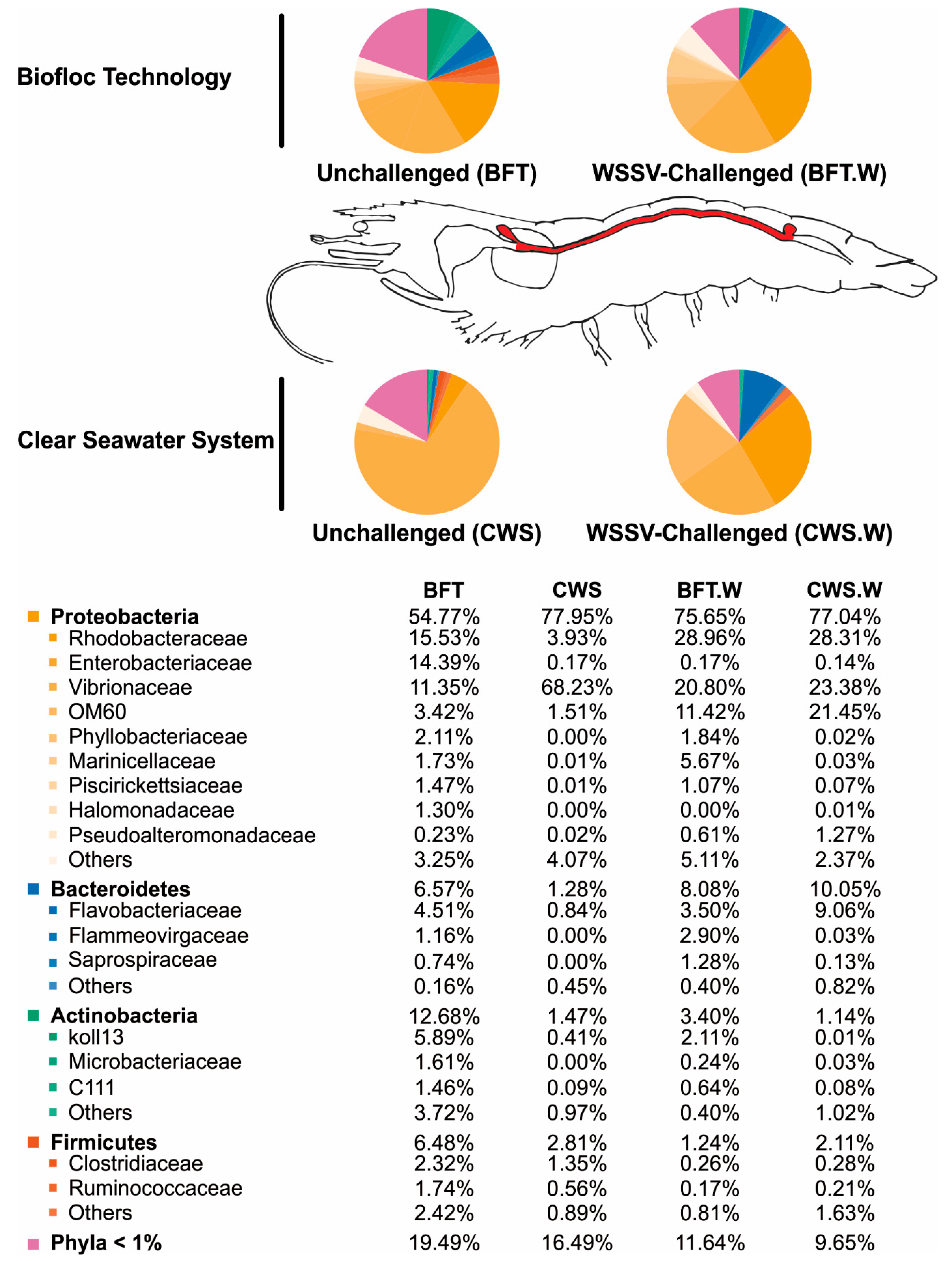
3.2. Influence of Rearing Conditions on the Bacterial Communities of Shrimp Midgut
3.3. Shrimp Intestinal Microbiota Plasticity in Response to a Viral Challenge
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Lightner, D.V.; Redman, R.M.; Pantoja, C.R.; Tang, K.F.J.; Noble, B.L.; Schofield, P.; Mohney, L.L.; Nunan, L.M.; Navarro, S.A. Historic emergence, impact and current status of shrimp pathogens in the Americas. J. Invertebr. Pathol. 2012, 110, 174–183. [Google Scholar] [CrossRef] [PubMed]
- Thitamadee, S.; Prachumwat, A.; Srisala, J.; Jaroenlak, P.; Salachan, P.V.; Sritunyalucksana, K.; Flegel, T.W.; Itsathitphaisarn, O. Review of current disease threats for cultivated penaeid shrimp in Asia. Aquaculture 2016, 452, 69–87. [Google Scholar] [CrossRef]
- Sánchez-Paz, A. White spot syndrome virus: An overview on an emergent concern. Vet. Res. 2010, 41. [Google Scholar] [CrossRef] [PubMed]
- Tran, L.; Nunan, L.; Redman, R.M.; Mohney, L.L.; Pantoja, C.R.; Fitzsimmons, K.; Lightner, D.V. Determination of the infectious nature of the agent of acute hepatopancreatic necrosis syndrome affecting penaeid shrimp. Dis. Aquat. Organ. 2013, 105, 45–55. [Google Scholar] [CrossRef] [PubMed]
- Kondo, H.; Van, P.T.; Dang, L.T. Draft Genome Sequence of Non-Vibrio parahaemolyticus Acute Diseased Shrimp in Vietnam. Genome Announc. 2015, 3, e00978-15. [Google Scholar] [CrossRef] [PubMed]
- Browdy, C.L.; Bratvold, D.; Stokesand, A.D.; McIntosh, R.P. Perspectives on the application of closed shrimp culture systems. In The New Wave. Proceedings of the Special Session on Sustainable Shrimp Farming; Browdy, C.L., Jory, D.E., Eds.; World Aquaculture Society: Valley Centers, CA, USA, 2001; pp. 20–34. [Google Scholar]
- Ahmad, I.; Babitha Rani, A.M.; Verma, A.K.; Maqsood, M. Biofloc technology: An emerging avenue in aquatic animal healthcare and nutrition. Aquac. Int. 2017, 25, 1215–1226. [Google Scholar] [CrossRef]
- Emerenciano, M.G.C.; Martínez-Córdova, L.R.; Martínez-Porchas, M.; Miranda-Baeza, A. Biofloc Technology (BFT): A Tool for Water Quality Management in Aquaculture. In Water Quality; InTech: London, UK, 2017; pp. 91–109. [Google Scholar]
- Bossier, P.; Ekasari, J. Biofloc technology application in aquaculture to support sustainable development goals. Microb. Biotechnol. 2017, 10, 1012–1016. [Google Scholar] [CrossRef] [PubMed]
- Moss, S.M.; Moss, D.R.; Arce, S.M.; Lightner, D.V.; Lotz, J.M. The role of selective breeding and biosecurity in the prevention of disease in penaeid shrimp aquaculture. J. Invertebr. Pathol. 2012, 110, 247–250. [Google Scholar] [CrossRef] [PubMed]
- Avnimelech, Y. Carbon/nitrogen ratio as a control element in aquaculture systems. Aquaculture 1999, 176, 227–235. [Google Scholar] [CrossRef]
- Burford, M.A.; Thompson, P.J.; McIntosh, R.P.; Bauman, R.H.; Pearson, D.C. The contribution of flocculated material to shrimp (Litopenaeus vannamei) nutrition in a high-intensity, zero-exchange system. Aquaculture 2004, 232, 525–537. [Google Scholar] [CrossRef]
- Kim, M.S.; Min, E.Y.; Kim, J.H.; Koo, J.K.; Kang, J.C. Growth performance and immunological and antioxidant status of Chinese shrimp, Fennerpenaeus chinensis reared in bio-floc culture system using probiotics. Fish Shellfish Immunol. 2015, 47, 141–146. [Google Scholar] [CrossRef] [PubMed]
- Ekasari, J.; Hanif Azhar, M.; Surawidjaja, E.H.; Nuryati, S.; De Schryver, P.; Bossier, P. Immune response and disease resistance of shrimp fed biofloc grown on different carbon sources. Fish Shellfish Immunol. 2014, 41, 332–339. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.K.; Pang, Z.; Seo, H.C.; Cho, Y.R.; Samocha, T.; Jang, I.K. Effect of bioflocs on growth and immune activity of Pacific white shrimp, Litopenaeus vannamei postlarvae. Aquac. Res. 2014, 45, 362–371. [Google Scholar] [CrossRef]
- Crab, R.; Lambert, A.; Defoirdt, T.; Bossier, P.; Verstraete, W. The application of bioflocs technology to protect brine shrimp (Artemia franciscana) from pathogenic Vibrio harveyi. J. Appl. Microbiol. 2010, 109, 1643–1649. [Google Scholar] [CrossRef] [PubMed]
- Wong, S.; Rawls, J.F. Intestinal microbiota composition in fishes is influenced by host ecology and environment. Mol. Ecol. 2012, 21, 3100–3102. [Google Scholar] [CrossRef] [PubMed]
- Montalban-Arques, A.; De Schryver, P.; Bossier, P.; Gorkiewicz, G.; Mulero, V.; Gatlin, D.M.; Galindo-Villegas, J. Selective manipulation of the gut microbiota improves immune status in vertebrates. Front. Immunol. 2015, 6, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Cornejo-Granados, F.; Lopez-Zavala, A.A.; Gallardo-Becerra, L.; Mendoza-Vargas, A.; Sánchez, F.; Vichido, R.; Brieba, L.G.; Viana, M.T.; Sotelo-Mundo, R.R.; Ochoa-Leyva, A. Microbiome of Pacific Whiteleg shrimp reveals differential bacterial community composition between Wild, Aquacultured and AHPND/EMS outbreak conditions. Sci. Rep. 2017, 7, 11783. [Google Scholar] [CrossRef] [PubMed]
- Cheung, M.K.; Yip, H.Y.; Nong, W.; Law, P.T.W.; Chu, K.H.; Kwan, H.S.; Hui, J.H.L. Rapid Change of Microbiota Diversity in the Gut but Not the Hepatopancreas during Gonadal Development of the New Shrimp Model Neocaridina denticulata. Mar. Biotechnol. 2015, 17, 811–819. [Google Scholar] [CrossRef] [PubMed]
- Tzuc, J.T.; Escalante, D.R.; Rojas Herrera, R.; Gaxiola Cortés, G.; Ortiz, M.L.A. Microbiota from Litopenaeus vannamei: Digestive tract microbial community of Pacific white shrimp (Litopenaeus vannamei). Springerplus 2014, 3, 280. [Google Scholar] [CrossRef] [PubMed]
- Cardona, E.; Gueguen, Y.; Magré, K.; Lorgeoux, B.; Piquemal, D.; Pierrat, F.; Noguier, F.; Saulnier, D. Bacterial community characterization of water and intestine of the shrimp Litopenaeus stylirostris in a biofloc system. BMC Microbiol. 2016, 16, 157. [Google Scholar] [CrossRef] [PubMed]
- Schveitzer, R.; Arantes, R.; Costódio, P.F.S.; do Espírito Santo, C.M.; Arana, L.V.; Seiffert, W.Q.; Andreatta, E.R. Effect of different biofloc levels on microbial activity, water quality and performance of Litopenaeus vannamei in a tank system operated with no water exchange. Aquac. Eng. 2013, 56, 59–70. [Google Scholar] [CrossRef]
- Lo, C.F.; Leu, J.H.; Ho, C.H.; Chen, C.H.; Peng, S.E.; Chen, Y.T.; Chou, C.M.; Yeh, P.Y.; Huang, C.J.; Chou, H.Y.; et al. Detection of baculovirus associated with white spot syndrome (WSBV) in penaeid shrimps using polymerase chain reaction. Dis. Aquat. Organ. 1996, 25, 133–141. [Google Scholar] [CrossRef]
- Goncalves, P.; Guertler, C.; Bachère, E.; de Souza, C.R.B.; Rosa, R.D.; Perazzolo, L.M. Molecular signatures at imminent death: Hemocyte gene expression profiling of shrimp succumbing to viral and fungal infections. Dev. Comp. Immunol. 2014, 42, 294–301. [Google Scholar] [CrossRef] [PubMed]
- Silveira, A.S.; Matos, G.M.; Falchetti, M.; Ribeiro, F.S.; Bressan, A.; Bachère, E.; Perazzolo, L.M.; Rosa, R.D. An immune-related gene expression atlas of the shrimp digestive system in response to two major pathogens brings insights into the involvement of hemocytes in gut immunity. Dev. Comp. Immunol. 2018, 79, 44–50. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Lee, C.; Kim, J.; Hwang, S. Group-specific primer and probe sets to detect methanogenic communities using quantitative real-time polymerase chain reaction. Biotechnol. Bioeng. 2005, 89, 670–679. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. Correspondence QIIME allows analysis of high-throughput community sequencing data Intensity normalization improves color calling in SOLiD sequencing. Nat. Publ. Gr. 2010, 7, 335–336. [Google Scholar] [CrossRef]
- Bokulich, N.A.; Subramanian, S.; Faith, J.J.; Gevers, D.; Gordon, I.; Knight, R.; Mills, D.A.; Caporaso, J.G. Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat. Methods 2013, 10, 57–59. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef] [PubMed]
- DeSantis, T.Z.; Hugenholtz, P.; Larsen, N.; Rojas, M.; Brodie, E.L.; Keller, K.; Huber, T.; Dalevi, D.; Hu, P.; Andersen, G.L. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 2006, 72, 5069–5072. [Google Scholar] [CrossRef] [PubMed]
- Chao, A. Nonparametric Estimation of the Number of Classes in a Population Author. Scanadinavian J. Stat. 1984, 11, 265–270. [Google Scholar] [CrossRef]
- Chao, A.; Lee, S.-M. Estimating the Number of Classes via Sanple Coverage. J. Am. Stat. Assoc. 1992, 87, 210–217. [Google Scholar] [CrossRef]
- Shannon, C.E. A mathematical theory of communication. ACM SIGMOBILE Mobile Comput. Commun. Rev. 1948, 27, 379–423. [Google Scholar] [CrossRef]
- Simpson, E.H. Measurement of diversity. Nature 1949, 163, 688. [Google Scholar] [CrossRef]
- Martin, G.G.; Simcox, R.; Nguyen, A.; Chilingaryan, A. Peritrophic membrane of the penaeid shrimp Sicyonia ingentis: Structure, formation, and permeability. Biol. Bull. 2006, 211, 275–285. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Li, F.; Wang, B.; Xiang, J. Structure and partial protein profiles of the peritrophic membrane (PM) from the gut of the shrimp Litopenaeus vannamei. Fish Shellfish Immunol. 2012, 33, 1285–1291. [Google Scholar] [CrossRef] [PubMed]
- Saraiva, R.G.; Kang, S.; Simões, M.L.; Angleró-Rodríguez, Y.I.; Dimopoulos, G. Mosquito gut antiparasitic and antiviral immunity. Dev. Comp. Immunol. 2016, 64, 53–64. [Google Scholar] [CrossRef] [PubMed]
- Deriu, E.; Boxx, G.M.; He, X.; Pan, C.; Benavidez, S.D.; Cen, L.; Rozengurt, N.; Shi, W.; Cheng, G. Influenza Virus Affects Intestinal Microbiota and Secondary Salmonella Infection in the Gut through Type I Interferons. PLoS Pathog. 2016, 12, 1–26. [Google Scholar] [CrossRef] [PubMed]
- Zeng, S.; Huang, Z.; Hou, D.; Liu, J.; Weng, S.; He, J. Composition, diversity and function of intestinal microbiota in pacific white shrimp (Litopenaeus vannamei) at different culture stages. PeerJ 2017, 5, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Simões, N.; Jones, D.; Soto-rodríguez, S.; Roque, A. Las Bacterias en el Inicio de la Alimentación Exógena en Larvas de Camarones Peneidos: Efectos de la Calidad del Agua, Tasas de Ingestión y Rutas de Colonización del Tracto Digestivo. In Proceedings of the Avances en Nutrición Acuícola VI. Memorias del VI Simposium Internacional de Nutrición Acuícola, Cancún, Mexico, 3–6 September 2002; pp. 243–276. [Google Scholar]
- Koenig, J.E.; Spor, A.; Scalfone, N.; Fricker, A.D.; Stombaugh, J.; Knight, R.; Angenent, L.T.; Ley, R.E. Succession of microbial consortia in the developing infant gut microbiome. Proc. Natl. Acad. Sci. USA 2011, 108, 4578–4585. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Sun, Y.; Chen, K.; Yu, N.; Zhou, Z.; Chen, L.; Du, Z.; Li, E. Characterization of the intestinal microbiota in Pacific white shrimp, Litopenaeus vannamei, fed diets with different lipid sources. Aquaculture 2014, 434, 449–455. [Google Scholar] [CrossRef]
- Rungrassamee, W.; Klanchui, A.; Maibunkaew, S.; Karoonuthaisiri, N. Bacterial dynamics in intestines of the black tiger shrimp and the Pacific white shrimp during Vibrio harveyi exposure. J. Invertebr. Pathol. 2016, 133, 12–19. [Google Scholar] [CrossRef] [PubMed]
- Sha, Y.; Liu, M.; Wang, B.; Jiang, K.; Qi, C.; Wang, L. Bacterial population in intestines of Litopenaeus vannamei fed different probiotics or probiotic supernatant. J. Microbiol. Biotechnol. 2016, 26, 1736–1745. [Google Scholar] [CrossRef] [PubMed]
- Vargas-Albores, F.; Porchas-Cornejo, M.A.; Martínez-Porchas, M.; Villalpando-Canchola, E.; Gollas-Galván, T.; Martínez-Córdova, L.R. Bacterial biota of shrimp intestine is significantly modified by the use of a probiotic mixture: A high throughput sequencing approach. Helgol. Mar. Res. 2017, 71, 5. [Google Scholar] [CrossRef]
- Aguilera-Rivera, D.; Prieto-Davó, A.; Escalante, K.; Chávez, C.; Cuzon, G.; Gaxiola, G. Probiotic effect of FLOC on Vibrios in the pacific white shrimp Litopenaeus vannamei. Aquaculture 2014, 424–425, 215–219. [Google Scholar] [CrossRef]
- Thompson, F.L.; Iida, T.; Swings, J. Biodiversity of Vibrios. Microbiol. Mol. Biol. Rev. 2004, 68, 403–431. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.Y.; Ng, T.H.; Wu, J.H.; Chen, J.W.; Wang, H.C. Microbiome Dynamics in a Shrimp Grow-out Pond with Possible Outbreak of Acute Hepatopancreatic Necrosis Disease. Sci. Rep. 2017, 7, 1–12. [Google Scholar] [CrossRef] [PubMed]
- De Schryver, P.; Defoirdt, T.; Sorgeloos, P. Early Mortality Syndrome Outbreaks: A Microbial Management Issue in Shrimp Farming? PLoS Pathog. 2014, 10, 10–11. [Google Scholar] [CrossRef] [PubMed]
- Molina-Aja, A.; Garcia-Gasca, A.; Abreu-Grobois, A.; Bolan-Mejia, C.; Roque, A.; Gomez-Gil b, B. Plasmid profling and antibiotic resistance of Vibrio strains isolated from cultured penaeid shrimp. FEMS Microbiol. Lett. 2002, 213, 7–12. [Google Scholar] [CrossRef] [PubMed]
- Emerenciano, M.; Gaxiola, G.; Cuzon, G. Biofloc Technology (BFT): A Review for Aquaculture Application and Animal Food Industry. In Biomass Now-Cultivation and Utilization; InTech: London, UK, 2013; pp. 301–328. ISBN 978-953-51-1106-1. [Google Scholar]
- Defoirdt, T.; Boon, N.; Sorgeloos, P.; Verstraete, W.; Bossier, P. Alternatives to antibiotics to control bacterial infections: Luminescent vibriosis in aquaculture as an example. Trends Biotechnol. 2007, 25, 472–479. [Google Scholar] [CrossRef] [PubMed]
- Elifantz, H.; Horn, G.; Ayon, M.; Cohen, Y.; Minz, D. Rhodobacteraceae are the key members of the microbial community of the initial biofilm formed in Eastern Mediterranean coastal seawater. FEMS Microbiol. Ecol. 2013, 85, 348–357. [Google Scholar] [CrossRef] [PubMed]
- Avnimelech, Y. Bio-filters: The need for an new comprehensive approach. Aquac. Eng. 2006, 34, 172–178. [Google Scholar] [CrossRef]
- Hjelm, M.; Riaza, A.; Formoso, F.; Melchiorsen, J.; Gram, L. Seasonal incidence of autochthonous antagonistic Roseobacter spp. and Vibrionaceae strains in a turbot larva (Scophthalmus maximus) rearing system. Appl. Environ. Microbiol. 2004, 70, 7288–7294. [Google Scholar] [CrossRef] [PubMed]
- Woebken, D.; Fuchs, B.M.; Kuypers, M.M.M.; Amann, R. Potential interactions of particle-associated anammox bacteria with bacterial and archaeal partners in the Namibian upwelling system. Appl. Environ. Microbiol. 2007, 73, 4648–4657. [Google Scholar] [CrossRef] [PubMed]
- Kirchman, D.L. The ecology of Cytophaga-Flavobacteria in aquatic environments. FEMS Microbiol. Ecol. 2002, 39, 91–100. [Google Scholar] [CrossRef]
- Xiong, J.; Wang, K.; Wu, J.; Qiuqian, L.; Yang, K.; Qian, Y.; Zhang, D. Changes in intestinal bacterial communities are closely associated with shrimp disease severity. Appl. Microbiol. Biotechnol. 2015, 99, 6911–6919. [Google Scholar] [CrossRef] [PubMed]
- Burford, M.A.; Thompson, P.J.; McIntosh, R.P.; Bauman, R.H.; Pearson, D.C. Nutrient and microbial dynamics in high-intensity, zero-exchange shrimp ponds in Belize. Aquaculture 2003, 219, 393–411. [Google Scholar] [CrossRef]
- Thompson, J.; Gregory, S.; Plummer, S.; Shields, R.J.; Rowley, A.F. An in vitro and in vivo assessment of the potential of Vibrio spp. as probiotics for the Pacific White shrimp, Litopenaeus vannamei. J. Appl. Microbiol. 2010, 109, 1177–1187. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Lin, G.; Yan, T.; Zheng, Z.; Chen, B.; Sun, F. The cellular community in the intestine of the shrimp Penaeus penicillatus and its culture environments. Fish. Sci. 2014, 80, 1001–1007. [Google Scholar] [CrossRef]
- Xu, W.J.; Pan, L.Q. Enhancement of immune response and antioxidant status of Litopenaeus vannamei juvenile in biofloc-based culture tanks manipulating high C/N ratio of feed input. Aquaculture 2013, 412–413, 117–124. [Google Scholar] [CrossRef]
- Brown, E.M.; Sadarangani, M.; Finlay, B.B. The role of the immune system in governing host-microbe interactions in the intestine. Nat. Immunol. 2013, 14, 660–667. [Google Scholar] [CrossRef] [PubMed]
- Nyholm, S.V.; Graf, J. Knowing your friends: Invertebrate innate immunity fosters beneficial bacterial symbioses. Nat. Rev. Microbiol. 2012, 10, 815–827. [Google Scholar] [CrossRef] [PubMed]
- Lazado, C.C.; Caipang, C.M.A. Mucosal immunity and probiotics in fish. Fish Shellfish Immunol. 2014, 39, 78–89. [Google Scholar] [CrossRef] [PubMed]
- Ponprateep, S.; Tharntada, S.; Somboonwiwat, K.; Tassanakajon, A. Gene silencing reveals a crucial role for anti-lipopolysaccharide factors from Penaeus monodon in the protection against microbial infections. Fish Shellfish Immunol. 2012, 32, 26–34. [Google Scholar] [CrossRef] [PubMed]
- Ding, Z.F.; Cao, M.J.; Zhu, X.S.; Xu, G.H.; Wang, R.L. Changes in the gut microbiome of the Chinese mitten crab (Eriocheir sinensis) in response to White spot syndrome virus (WSSV) infection. J. Fish Dis. 2017, 40, 1561–1571. [Google Scholar] [CrossRef] [PubMed]
- Apprill, A. Marine Animal Microbiomes: Toward Understanding Host–Microbiome Interactions in a Changing Ocean. Front. Mar. Sci. 2017, 4, 1–9. [Google Scholar] [CrossRef]
- Willing, B.P.; Gill, N.; Finlay, B.B. The role of the immune system in regulating the microbiota. Gut Microbes 2010, 1, 213–223. [Google Scholar] [CrossRef] [PubMed]
- Xiong, J.; Dai, W.; Li, C. Advances, challenges, and directions in shrimp disease control: The guidelines from an ecological perspective. Appl. Microbiol. Biotechnol. 2016, 100, 6947–6954. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Yu, M.; Liu, J.; Qiao, Y.; Wang, L.; Li, Z.; Zhang, X.H.; Yu, M. Bacterial community associated with healthy and diseased Pacific white shrimp (Litopenaeus vannamei) larvae and rearing water across different growth stages. Front. Microbiol. 2017, 8, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Zorriehzahra, M.J.; Banaederakhshan, R. Early Mortality Syndrome (EMS) as new Emerging Threat in Shrimp Industry. Adv. Anim. Vet. Sci. 2015, 3, 64–72. [Google Scholar] [CrossRef]
| Groups | OTUs | H’ | 1-D | Chao1 | ACE | Good’s Coverage |
|---|---|---|---|---|---|---|
| BFT | 653 | 6.24 | 0.96 | 674.05 | 672.40 | 0.99 |
| CWS | 352 | 4.13 | 0.79 | 487.07 | 408.01 | 0.99 |
| BFT.W | 572 | 5.79 | 0.96 | 624.99 | 627.61 | 0.99 |
| CWS.W | 518 | 4.55 | 0.89 | 542.84 | 552.32 | 0.99 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pilotto, M.R.; Goncalves, A.N.A.; Vieira, F.N.; Seifert, W.Q.; Bachère, E.; Rosa, R.D.; Perazzolo, L.M. Exploring the Impact of the Biofloc Rearing System and an Oral WSSV Challenge on the Intestinal Bacteriome of Litopenaeus vannamei. Microorganisms 2018, 6, 83. https://doi.org/10.3390/microorganisms6030083
Pilotto MR, Goncalves ANA, Vieira FN, Seifert WQ, Bachère E, Rosa RD, Perazzolo LM. Exploring the Impact of the Biofloc Rearing System and an Oral WSSV Challenge on the Intestinal Bacteriome of Litopenaeus vannamei. Microorganisms. 2018; 6(3):83. https://doi.org/10.3390/microorganisms6030083
Chicago/Turabian StylePilotto, Mariana R., André N. A. Goncalves, Felipe N. Vieira, Walter Q. Seifert, Evelyne Bachère, Rafael D. Rosa, and Luciane M. Perazzolo. 2018. "Exploring the Impact of the Biofloc Rearing System and an Oral WSSV Challenge on the Intestinal Bacteriome of Litopenaeus vannamei" Microorganisms 6, no. 3: 83. https://doi.org/10.3390/microorganisms6030083
APA StylePilotto, M. R., Goncalves, A. N. A., Vieira, F. N., Seifert, W. Q., Bachère, E., Rosa, R. D., & Perazzolo, L. M. (2018). Exploring the Impact of the Biofloc Rearing System and an Oral WSSV Challenge on the Intestinal Bacteriome of Litopenaeus vannamei. Microorganisms, 6(3), 83. https://doi.org/10.3390/microorganisms6030083





