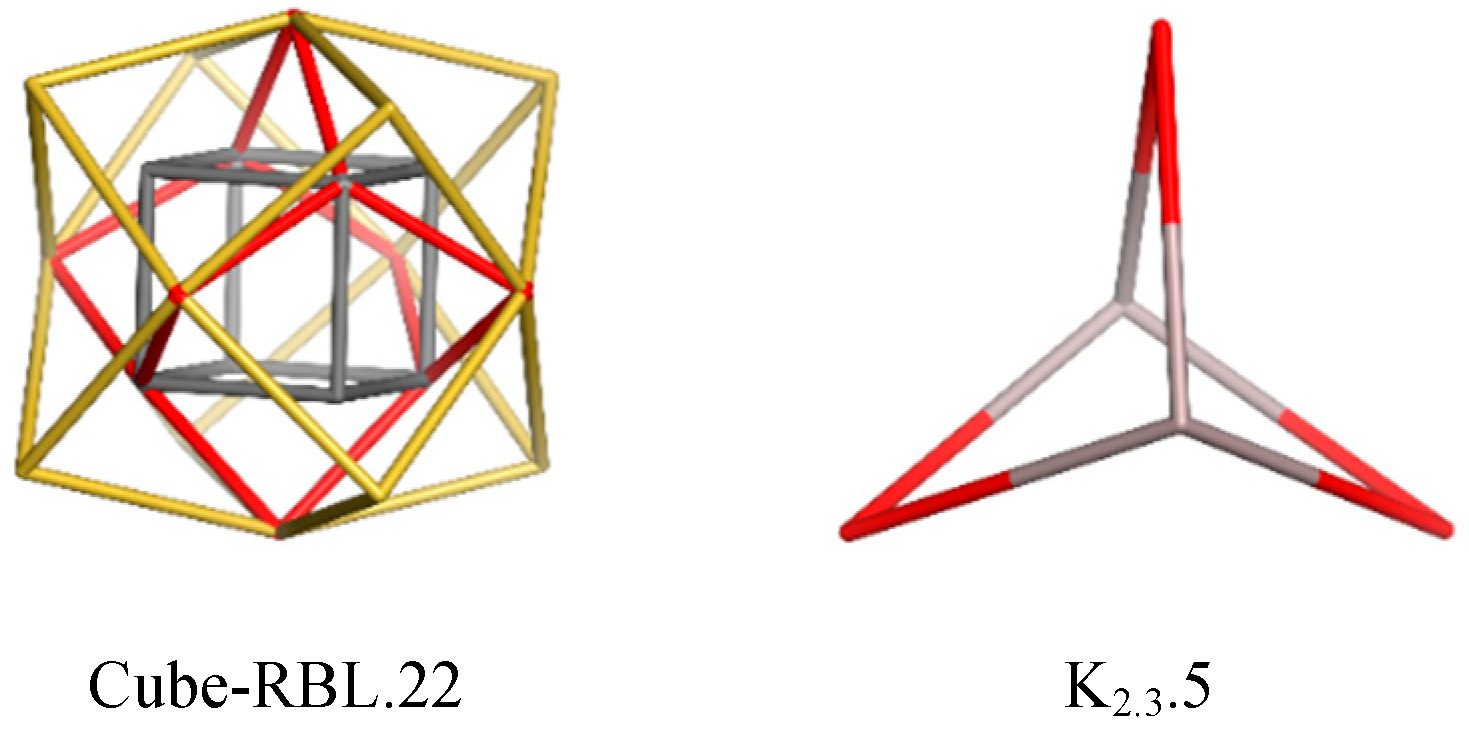
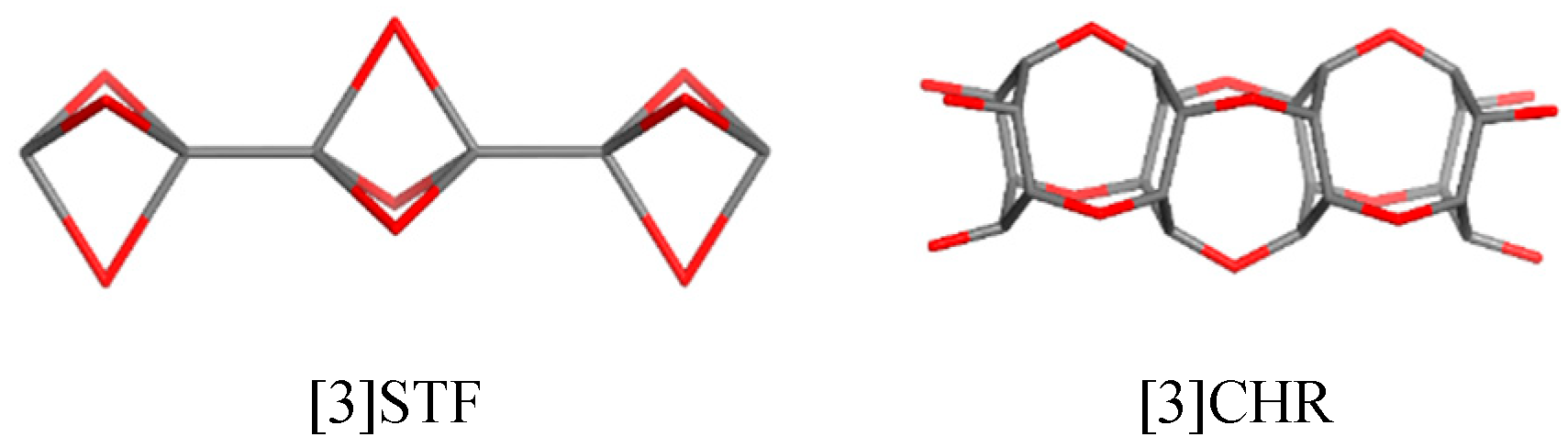
Docking Linear Ligands to Glucose Oxidase
Abstract
:1. Introduction
2. Materials and Methods
3. Results
3.1. The First Step of the Study
3.2. The Second Step of the Study
4. Discussion
5. Conclusions
Supplementary Materials
Funding
Acknowledgments
Conflicts of Interest
References
- Leskovac, V.; Trivić, S.; Wohlfahrt, G.; Kandrac, J.; Pericin, D. Glucose oxidase from Aspergillus niger: The mechanism of action with molecular oxygen, quinines and one electron acceptors. Int. J. Biochem. Cell Biol. 2005, 37, 731–750. [Google Scholar] [CrossRef] [PubMed]
- Yemul, O.; Imae, T. Synthesis and characterization of poly(ethyleneimine) dendrimers. Colloid Polym. Sci. 2008, 286, 747–752. [Google Scholar] [CrossRef]
- Rudolph, C.; Lausier, J.; Naundorf, S.; Müller, R.H.; Rosenecker, J. In vivo gene delivery to the lung using polyethylenimine and fractured polyamidoamine dendrimers. J. Gene Med. 2000, 2, 269–278. [Google Scholar] [CrossRef]
- Akinc, A.; Thomas, M.; Klibanov, A.M.; Langer, R. Exploring polyethylenimine-mediated DNA transfection and the proton sponge hypothesis. J. Gene Med. 2004, 7, 657–666. [Google Scholar] [CrossRef] [PubMed]
- Szefler, B.; Diudea, M.V.; Grudziński, I.P. Nature of polyethylene imine-glucose oxidase interactions. Studia UBB_Chemia 2016, 61, 249–260. [Google Scholar]
- Lungu, C.N.; Diudea, M.V.; Putz, M.V.; Grudziński, I.P. Linear and branched PEIs (Polyethylenimines) and their properties space. Int. J. Mol. Sci. 2016, 17, 555. [Google Scholar] [CrossRef] [PubMed]
- Szefler, B.; Diudea, M.V.; Putz, M.V.; Grudziński, I.P. Molecular Dynamic Studies of the Complex Polyethylenimine and Glucose Oxidase. Int. J. Mol. Sci. 2016, 17, 1796. [Google Scholar] [CrossRef]
- Lungu, C.N.; Diudea, M.V.; Putz, M.V.; Grudzinski, I.P. FAD molecular adaptability among surrounding amino acids and its catalytic role in glucose oxidase and related flavoproteins. Res. J. Life Sci. Bioinf. Pharm. Chem. Sci. 2017, 3. [Google Scholar] [CrossRef]
- Diudea, M.V. Rhombellanes—A new class of structures. Int. J. Chem. Model. 2017, 9, 91–96. [Google Scholar]
- Diudea, M.V. Cube-Rhombellane: From graph to molecule. Int. J. Chem. Model. 2017, 9, 97–103. [Google Scholar]
- Wiberg, K.B.; Walker, F.H. [1.1.1] Propellane. J. Am. Chem. Soc. 1982, 104, 5239–5240. [Google Scholar] [CrossRef]
- Kazynsky, P.; Michl, J. [n] Staffanes: A molecular-size tinkertoy construction set for nanotechnology. Preparation of end-functionalized telomers and a polymer of [1.1.1] propellane. J. Am. Chem. Soc. 1988, 110, 5225–5226. [Google Scholar] [CrossRef]
- Dilmaç, A.; Spuling, E.; Meijere, A.; Bräse, S. Propellanes—From a chemical curiosity to “explosive” materials and natural products. Angew. Chem. Int. Ed. 2017, 56, 5684–5718. [Google Scholar] [CrossRef] [PubMed]
- Diudea, M.V. Hypercube related polytopes. Iran. J. Math. Chem. 2018, 9, 1–8. [Google Scholar]
- Diudea, M.V. Rhombellanic crystals and quasicrystals. Iran. J. Math. Chem. 2018, 9, 167–178. [Google Scholar]
- Szefler, B.; Czeleń, P.; Diudea, M.V. Docking of indolizine derivatives on cube rhombellane functionalized homeomorphs. Studia Universitatis Babes-Bolyai Chemia 2018, 63, 7–18. [Google Scholar] [CrossRef]
- Diudea, M.V.; Medeleanu, M.; Khalaj, Z.; Ashrafi, A.R. Spongy Diamond. Iran. J. Math. Chem. 2019, 10, 1–9. [Google Scholar]
- Trott, O.; Olson, A.J. AutoDockVina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J. Comp. Chem. 2010, 31, 455–461. [Google Scholar]
- Bertrand, J.A.; Thieffine, S.; Vulpetti, A.; Cristiani, C.; Valsasina, B.; Knapp, S.; Kalisz, H.M.; Flocco, M. Structural characterization of the GSK-3beta active site using selective and non-selective ATP-mimetic inhibitors. J. Mol. Biol. 2003, 333, 393–407. [Google Scholar] [CrossRef]
- Shoichet, B.K.; Kuntz, I.D.; Bodian, D.L. Molecular docking using shape descriptors. J. Comput. Chem. 2004, 13, 380–397. [Google Scholar] [CrossRef]
- Dhananjayan, K.; Kalathil, K.; Sumathy, A.; Sivanandy, P. A computational study on binding affinity of bio-flavonoids on the crystal structure of 3-hydroxy-3-methyl-glutaryl-CoA reductase—An insilico molecular docking approach. Der Pharma Chemica 2014, 6, 378–387. [Google Scholar]
- Abagyan, R.; Totrov, M. High-throughput docking for lead generation. Curr. Opin. Chem. Biol. 2001, 5, 375–382. [Google Scholar] [CrossRef]
- Shen, M.; Zhou, S.; Li, Y.; Pan, P.; Zhang, L.; Hou, T. Discovery and optimization of triazine derivatives as ROCK1 inhibitors: Molecular docking, molecular dynamics simulations and free energy calculations. Mol. Biosyst. 2013, 9, 361–374. [Google Scholar] [CrossRef] [PubMed]
- Shen, M.; Yu, H.; Li, Y.; Li, P.; Pan, P.; Zhou, S.; Zhang, L.; Li, S.; Lee, S.M.Y.; Hou, T. Discovery of Rho-kinase inhibitors by docking-based virtual screening. Mol. Biosyst. 2013, 9, 1511–1521. [Google Scholar] [CrossRef] [PubMed]
- Jeffrey, G.A. An Introduction to Hydrogen Bonding; Oxford University Press: New York, NY, USA, 1997; Volume 12, p. 228. [Google Scholar]
© 2019 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Szefler, B. Docking Linear Ligands to Glucose Oxidase. Symmetry 2019, 11, 901. https://doi.org/10.3390/sym11070901
Szefler B. Docking Linear Ligands to Glucose Oxidase. Symmetry. 2019; 11(7):901. https://doi.org/10.3390/sym11070901
Chicago/Turabian StyleSzefler, Beata. 2019. "Docking Linear Ligands to Glucose Oxidase" Symmetry 11, no. 7: 901. https://doi.org/10.3390/sym11070901
APA StyleSzefler, B. (2019). Docking Linear Ligands to Glucose Oxidase. Symmetry, 11(7), 901. https://doi.org/10.3390/sym11070901