The Complete Genome of Probiotic Lactobacillus sakei Derived from Plateau Yak Feces
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Probiotics Strain
2.3. Genome DNA Extraction
2.4. Library Construction and Sequencing
2.5. Sequencing Data and Quality Control
2.6. Genome Assembly, Annotation, and Phylogenetic Analysis
2.7. Functional Analysis
2.8. Plasmid Analysis
2.9. Genome and Plasmid Comparison
3. Results
3.1. Raw Data Deposit
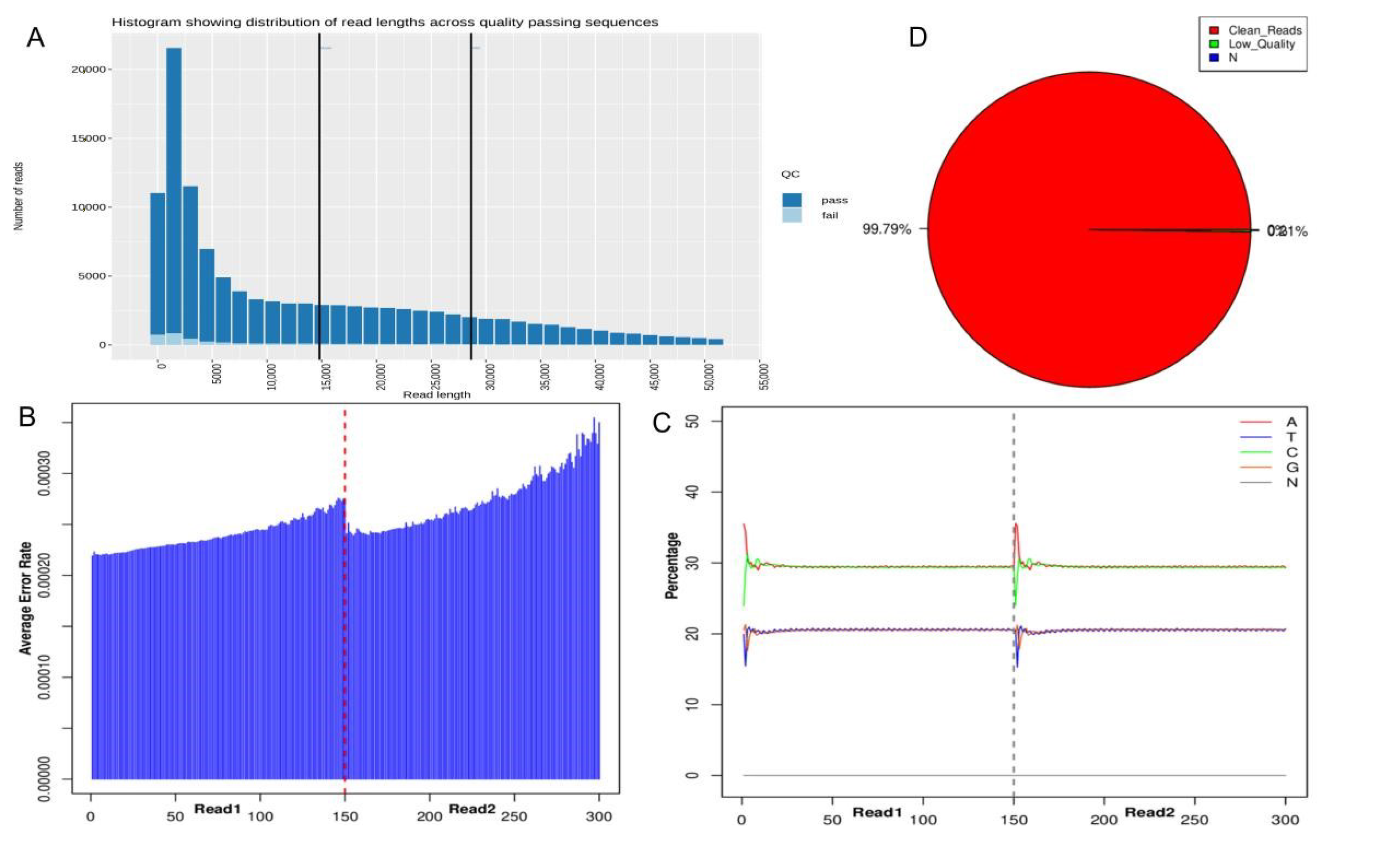
3.2. Sequencing Information and Quality Control Results
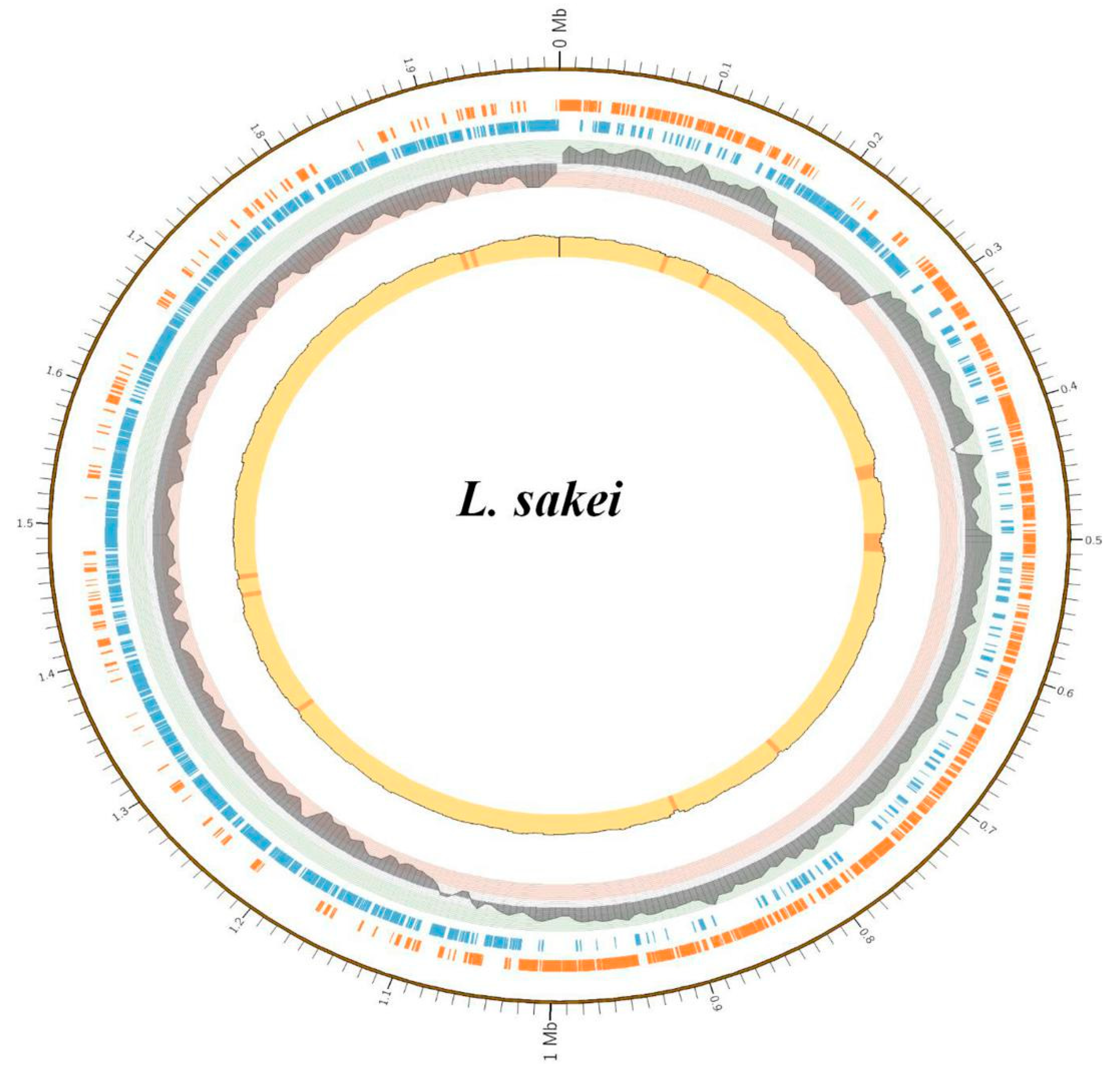
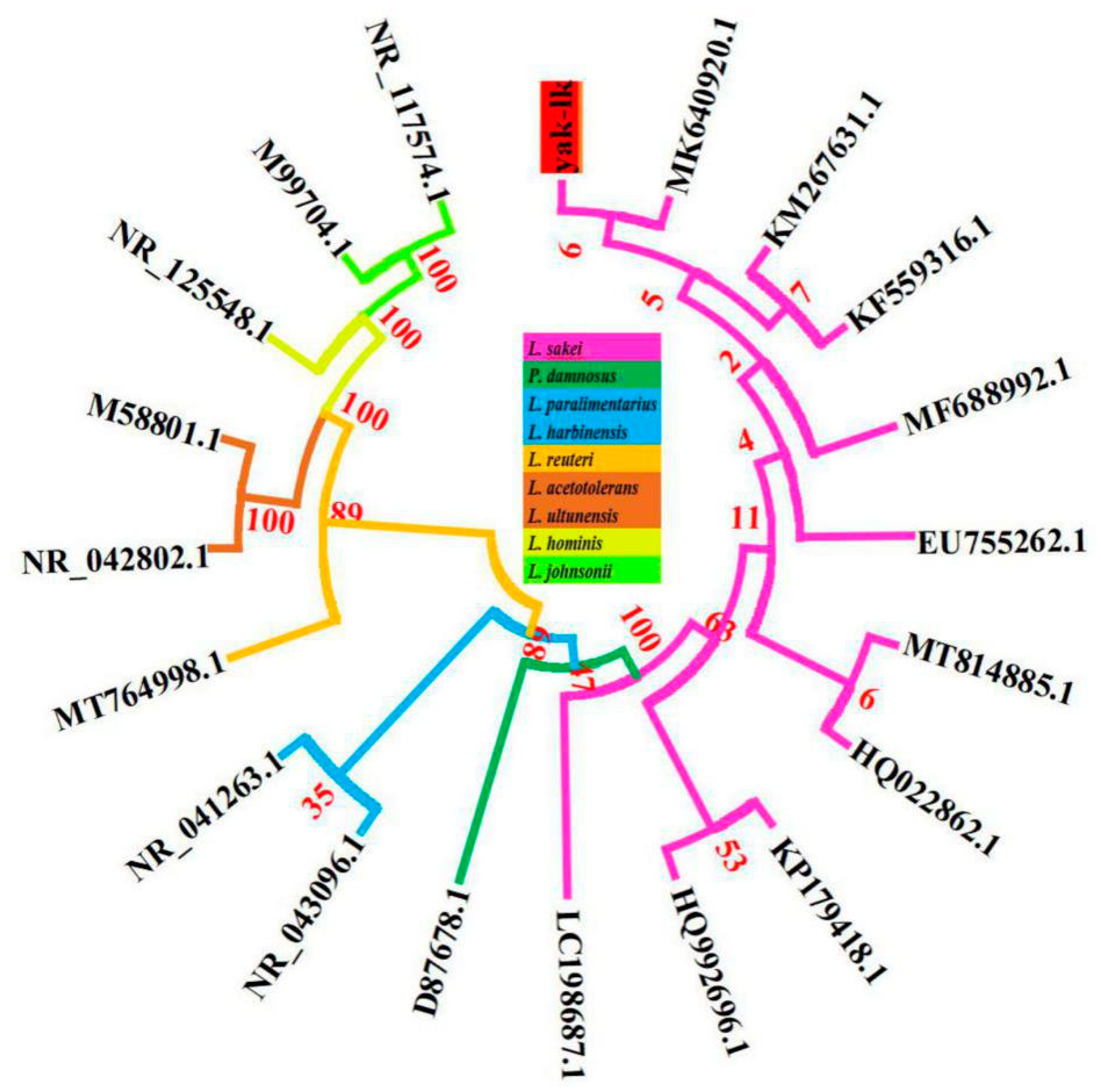
3.3. Genome Assembly, Annotation and Phylogenetic Tree
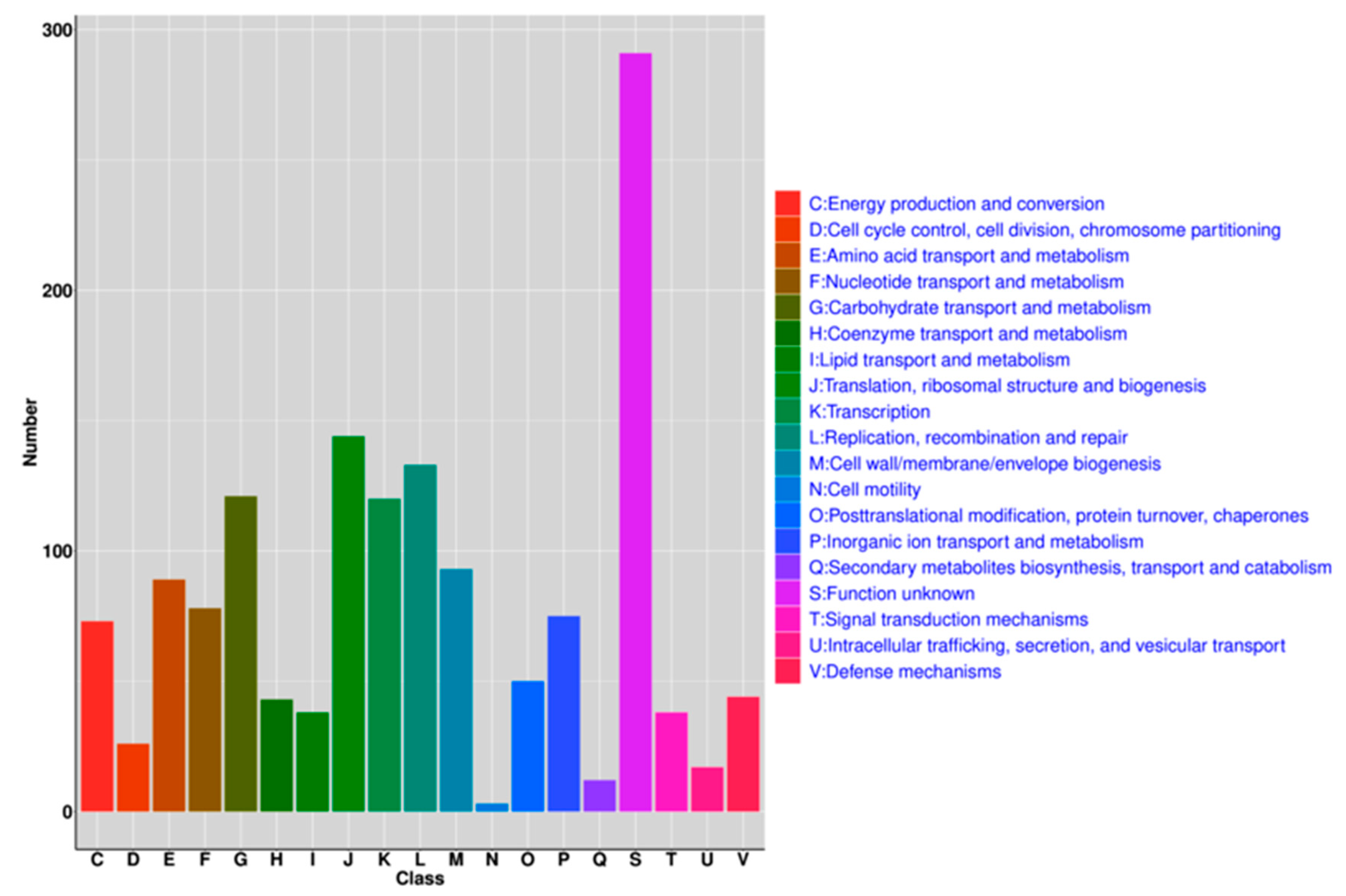
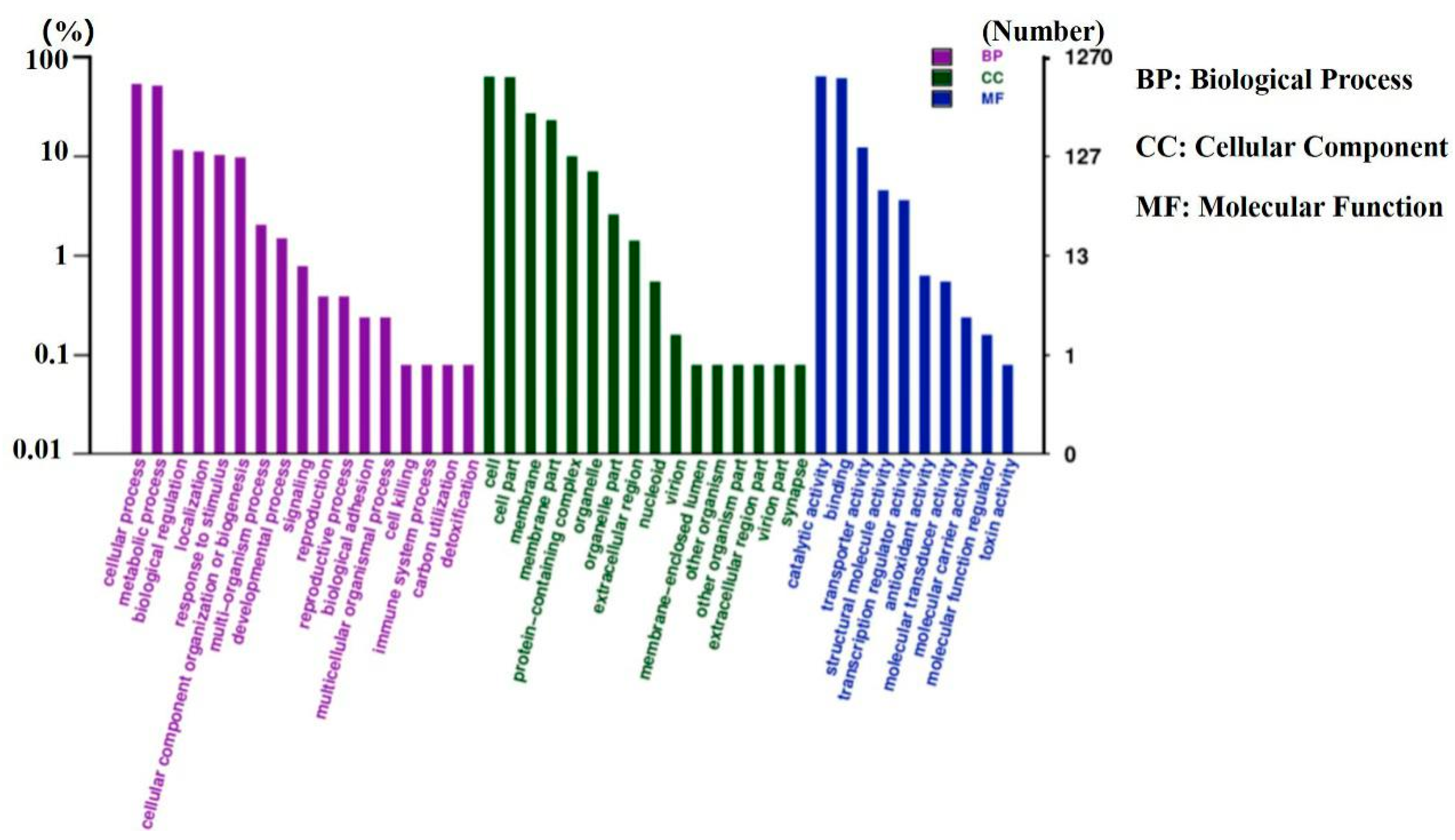
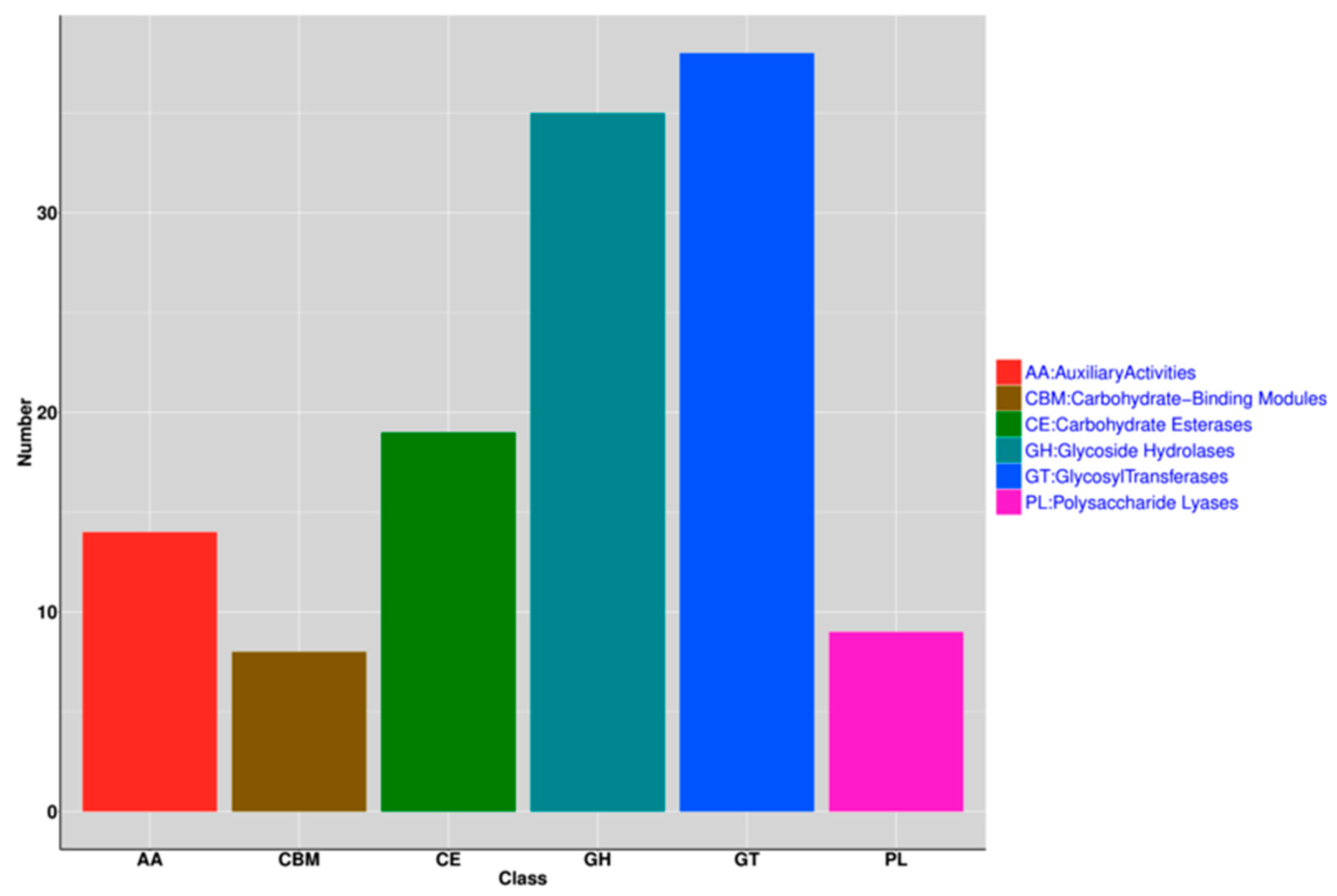
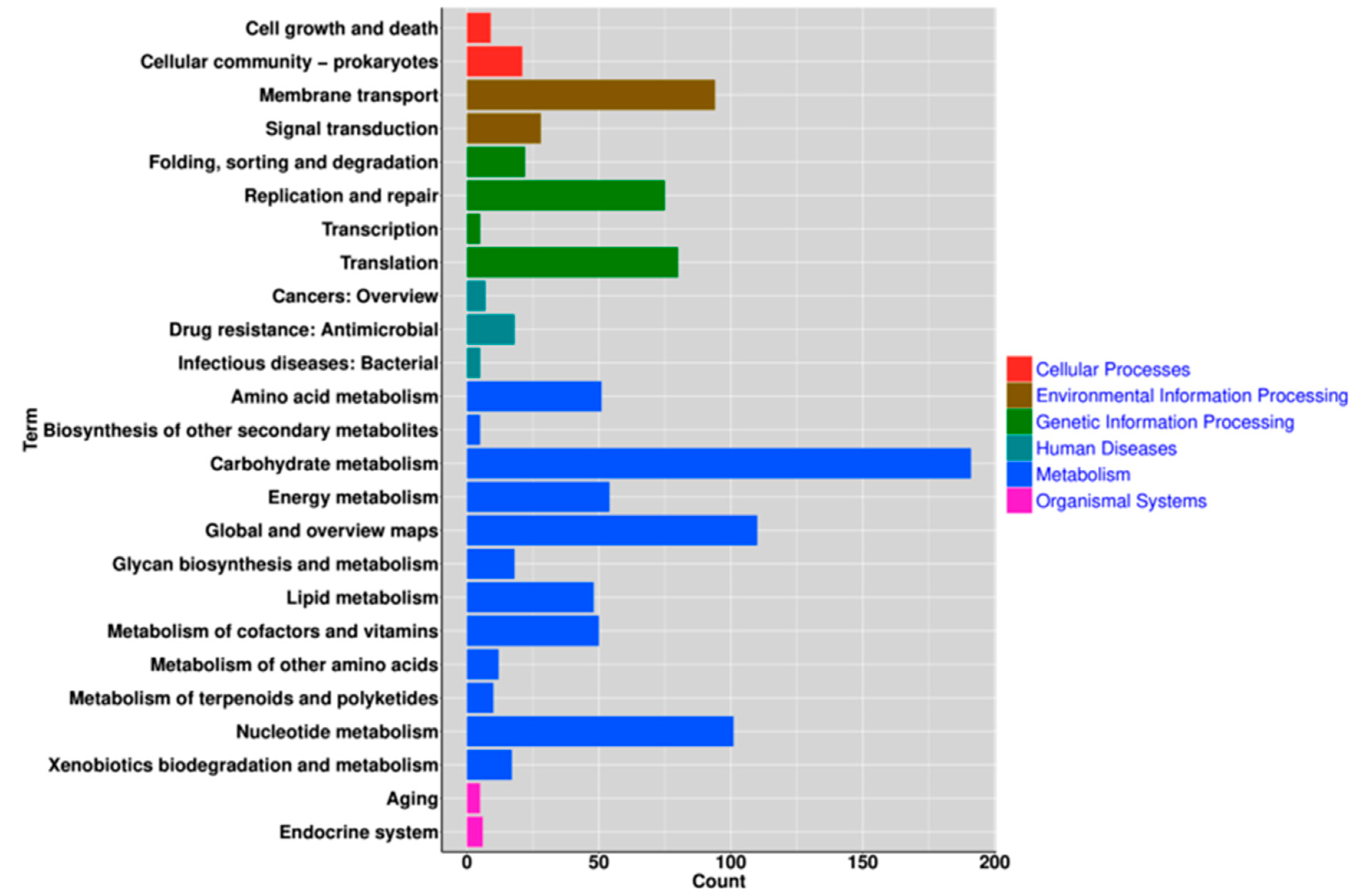
3.4. Functional Analysis
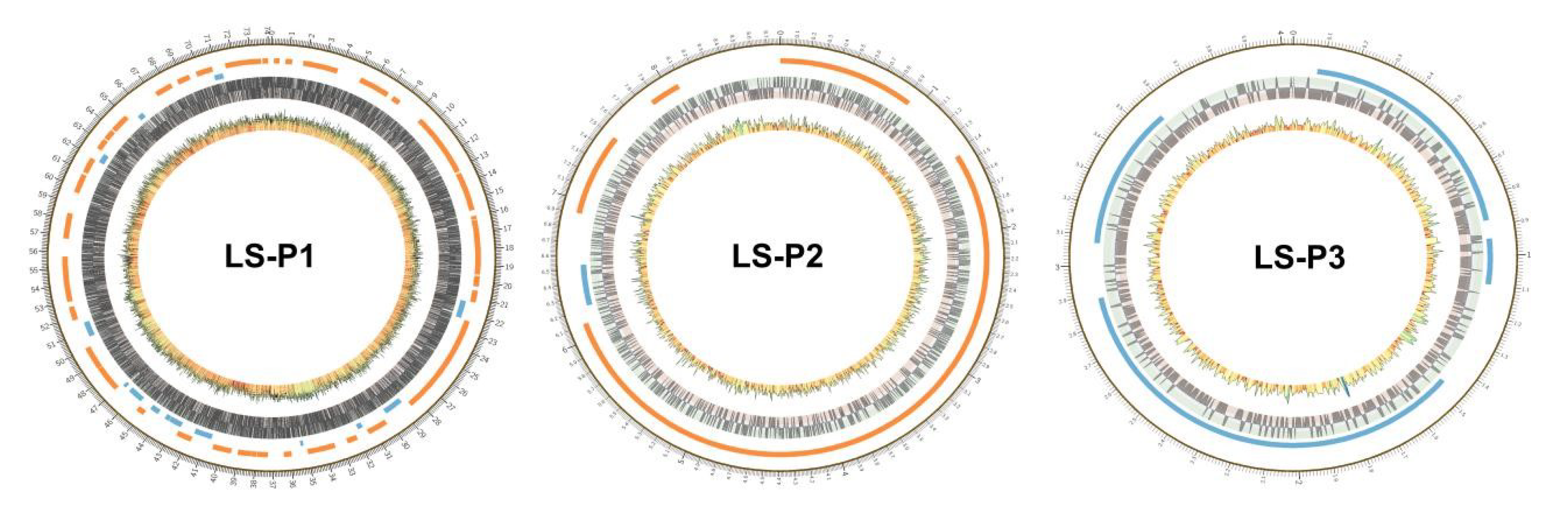
3.5. Plasmid Analysis
3.6. Genome and Plasmid Comparison Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Sanchez, B.; Delgado, S.; Blanco-Miguez, A.; Lourenco, A.; Gueimonde, M.; Margolles, A. Probiotics, gut microbiota, and their influence on host health and disease. Mol. Nutr. Food Res. 2017, 61. [Google Scholar] [CrossRef] [PubMed]
- Hill, C.; Guarner, F.; Reid, G.; Gibson, G.R.; Merenstein, D.J.; Pot, B.; Morelli, L.; Canani, R.B.; Flint, H.J.; Salminen, S.; et al. Expert consensus document. The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat. Rev. Gastroenterol. Hepatol. 2014, 11, 506–514. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.H.; Chun, B.H.; Baek, J.H.; Roh, S.W.; Lee, S.H.; Jeon, C.O. Genomic and metabolic features of Lactobacillus sakei as revealed by its pan-genome and the metatranscriptome of kimchi fermentation. Food Microbiol. 2020, 86, 103341. [Google Scholar] [CrossRef]
- Verplaetse, E.; Andre-Leroux, G.; Duhutrel, P.; Coeuret, G.; Chaillou, S.; Nielsen-Leroux, C.; Champomier-Verges, M.-C. Heme Uptake in Lactobacillus sakei Evidenced by a New Energy Coupling Factor (ECF)-Like Transport System. Appl. Environ. Microbiol. 2020, 86. [Google Scholar] [CrossRef]
- Liu, J.; Wang, Y.; Li, A.; Iqbal, M.; Zhang, L.; Pan, H.; Liu, Z.; Li, J. Probiotic potential and safety assessment of Lactobacillus isolated from yaks. Microbial. Pathog. 2020, 145, 104213. [Google Scholar] [CrossRef]
- Lee, S.-Y.; Jeong, J.-J.; Kim, K.-A.; Kim, D.-H. Lactobacillus sakei OK67 ameliorates collagen-induced arthritis in mice by inhibiting NF-κB activation and restoring Th17/Treg cell balance. J. Funct. Foods 2015, 18, 501–511. [Google Scholar] [CrossRef]
- Rather, I.A.; Bajpai, V.K.; Huh, Y.S.; Han, Y.K.; Bhat, E.A.; Lim, J.; Paek, W.K.; Park, Y.H. Probiotic Lactobacillus sakei proBio-65 Extract Ameliorates the Severity of Imiquimod Induced Psoriasis-Like Skin Inflammation in a Mouse Model. Front. Microbiol. 2018, 9, 1021. [Google Scholar] [CrossRef]
- Seo, S.; Shin, J.-S.; Lee, W.-S.; Rhee, Y.K.; Cho, C.-W.; Hong, H.-D.; Lee, K.-T. Anti-colitis effect of Lactobacillus sakei K040706 via suppression of inflammatory responses in the dextran sulfate sodium-induced colitis mice model. J. Funct. Foods 2017, 29, 256–268. [Google Scholar] [CrossRef]
- Jung, J.Y.; Shin, J.S.; Lee, S.G.; Rhee, Y.K.; Cho, C.W.; Hong, H.D.; Lee, K.T. Lactobacillus sakei K040706 evokes immunostimulatory effects on macrophages through TLR 2-mediated activation. Int. Immunopharmacol. 2015, 28, 88–96. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.M.; Jeong, J.J.; Woo, K.H.; Han, M.J.; Kim, D.H. Lactobacillus sakei OK67 ameliorates high-fat diet-induced blood glucose intolerance and obesity in mice by inhibiting gut microbiota lipopolysaccharide production and inducing colon tight junction protein expression. Nutr. Res. 2016, 36, 337–348. [Google Scholar] [CrossRef] [PubMed]
- Woo, S.; Kim, J.; Lee, Y.; Kim, N.; Hahn, Y. Effect of Lactobacillus sakei supplementation in children with atopic eczema-dermatitis syndrome. Ann. Allergy Asthma Immunol. 2010, 104, 343–348. [Google Scholar] [CrossRef] [PubMed]
- Li, K.; Gao, J.; Shahzad, M.; Han, Z.; Nabi, F.; Liu, M.; Zhang, D.; Li, J. Seroprevalence of Toxoplasma gondii infection in yaks (Bos grunniens) on the Qinghai-Tibetan Plateau of China. Vet. Parasitol. 2014, 205, 354–356. [Google Scholar] [CrossRef] [PubMed]
- Li, K.; Li, Z.; Zeng, Z.; Li, A.; Mehmood, K.; Shahzad, M.; Gao, K.; Li, J. Prevalence and molecular characterization of Cryptosporidium spp. in yaks (Bos grunniens) in Naqu, China. Microb. Pathog. 2020, 144, 104190. [Google Scholar] [CrossRef] [PubMed]
- Qiu, Q.; Zhang, G.; Ma, T.; Qian, W.; Wang, J.; Ye, Z.; Cao, C.; Hu, Q.; Kim, J.; Larkin, D.M.; et al. The yak genome and adaptation to life at high altitude. Nat. Genet. 2012, 44, 946–949. [Google Scholar] [CrossRef] [PubMed]
- Fan, Q.S.; Wanapat, M.; Yan, T.H.; Hou, F.J. Altitude influences microbial diversity and herbage fermentation in the rumen of yaks. BMC Microbiol. 2020, 20, 370. [Google Scholar] [CrossRef]
- Wu, D.W.; Vinitchaikul, P.; Deng, M.Y.; Zhang, G.R.; Sun, L.Y.; Wang, H.X.; Gou, X.; Mao, H.M.; Yang, S.L. Exploration of the effects of altitude change on bacteria and fungi in the rumen of yak (Bos grunniens). Arch. Microbiol. 2020. [Google Scholar] [CrossRef]
- Cryan, J.F.; Dinan, T.G. Mind-altering micro-organisms: The impact of the gut microbiota on brain and behaviour. Nat. Rev. Neurosci. 2012, 13, 701–712. [Google Scholar] [CrossRef]
- Kwun, S.Y.; Yoon, J.A.; Park, E.H.; Kim, M.D. Complete genome sequence data of Lactobacillus sakei MBEL1397 isolated from kimchi. Data Brief 2020, 31, 105740. [Google Scholar] [CrossRef]
- Eisenbach, L.; Geissler, A.J.; Ehrmann, M.A.; Vogel, R.F. Comparative genomics of Lactobacillus sakei supports the development of starter strain combinations. Microbiol. Res. 2019, 221, 1–9. [Google Scholar] [CrossRef]
- Kato, S.; Oikawa, T. Genome Sequence of Lactobacillus sakei LK-145 Isolated from a Japanese Sake Cellar as a High Producer of d-Amino Acids. Genome Announc. 2017, 5. [Google Scholar] [CrossRef]
- Jans, C.; Lagler, S.; Lacroix, C.; Meile, L.; Stevens, M.J.A. Complete Genome Sequences of Lactobacillus curvatus KG6, L. curvatus MRS6, and Lactobacillus sakei FAM18311, Isolated from Fermented Meat Products. Genome Announc. 2017, 5. [Google Scholar] [CrossRef] [PubMed]
- Tritt, A.; Eisen, J.A.; Facciotti, M.T.; Darling, A.E. An integrated pipeline for de novo assembly of microbial genomes. PLoS ONE 2012, 7, e42304. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef] [PubMed]
- Chin, C.S.; Peluso, P.; Sedlazeck, F.J.; Nattestad, M.; Concepcion, G.T.; Clum, A.; Dunn, C.; O’Malley, R.; Figueroa-Balderas, R.; Morales-Cruz, A.; et al. Phased diploid genome assembly with single-molecule real-time sequencing. Nat. Methods 2016, 13, 1050–1054. [Google Scholar] [CrossRef] [PubMed]
- Koren, S.; Walenz, B.P.; Berlin, K.; Miller, J.R.; Bergman, N.H.; Phillippy, A.M. Canu: Scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 2017, 27, 722–736. [Google Scholar] [CrossRef] [PubMed]
- Marcais, G.; Delcher, A.L.; Phillippy, A.M.; Coston, R.; Salzberg, S.L.; Zimin, A. MUMmer4: A fast and versatile genome alignment system. PLoS Comput. Biol. 2018, 14, e1005944. [Google Scholar] [CrossRef] [PubMed]
- Walker, B.J.; Abeel, T.; Shea, T.; Priest, M.; Abouelliel, A.; Sakthikumar, S.; Cuomo, C.A.; Zeng, Q.; Wortman, J.; Young, S.K.; et al. Pilon: An integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 2014, 9, e112963. [Google Scholar] [CrossRef] [PubMed]
- Besemer, J.; Lomsadze, A.; Borodovsky, M. GeneMarkS: A self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions. Nucleic Acids Res. 2001, 29, 2607–2618. [Google Scholar] [CrossRef]
- Lowe, T.M.; Eddy, S.R. tRNAscan-SE: A program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997, 25, 955–964. [Google Scholar] [CrossRef]
- Burge, S.W.; Daub, J.; Eberhardt, R.; Tate, J.; Barquist, L.; Nawrocki, E.P.; Eddy, S.R.; Gardner, P.P.; Bateman, A. Rfam 11.0: 10 years of RNA families. Nucleic Acids Res. 2013, 41, D226–D232. [Google Scholar] [CrossRef]
- Moriya, Y.; Itoh, M.; Okuda, S.; Yoshizawa, A.C.; Kanehisa, M. KAAS: An automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 2007, 35, W182–W185. [Google Scholar] [CrossRef] [PubMed]
- Stothard, P.; Wishart, D.S. Circular genome visualization and exploration using CGView. Bioinformatics 2005, 21, 537–539. [Google Scholar] [CrossRef] [PubMed]
- Petersen, T.N.; Brunak, S.; von Heijne, G.; Nielsen, H. SignalP 4.0: Discriminating signal peptides from transmembrane regions. Nat. Methods 2011, 8, 785–786. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Yu, P.; Luo, J.; Jiang, Y. Secreted protein prediction system combining CJ-SPHMM, TMHMM, and PSORT. Mamm. Genome 2003, 14, 859–865. [Google Scholar] [CrossRef] [PubMed]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 2015, 12, 59–60. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Bateman, A.; Clements, J.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Heger, A.; Hetherington, K.; Holm, L.; Mistry, J.; et al. Pfam: The protein families database. Nucleic Acids Res. 2014, 42, D222–D230. [Google Scholar] [CrossRef] [PubMed]
- Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef]
- Kant, R.; Blom, J.; Palva, A.; Siezen, R.J.; de Vos, W.M. Comparative genomics of Lactobacillus. Microb. Biotechnol. 2011, 4, 323–332. [Google Scholar] [CrossRef]
- Loux, V.; Coeuret, G.; Zagorec, M. Complete and Draft Genome Sequences of Nine Lactobacillus sakei Strains Selected from the Three Known Phylogenetic Lineages and Their Main Clonal Complexes. Genome Announc. 2018, 6, e00082-00018. [Google Scholar] [CrossRef]
- Chaillou, S.; Champomier-Verges, M.C.; Cornet, M.; Crutz-Le Coq, A.M.; Dudez, A.M.; Martin, V.; Beaufils, S.; Darbon-Rongere, E.; Bossy, R.; Loux, V.; et al. The complete genome sequence of the meat-borne lactic acid bacterium Lactobacillus sakei 23K. Nat. Biotechnol. 2005, 23, 1527–1533. [Google Scholar] [CrossRef]
- Liu, X.; Zeng, J.; Huang, K.; Wang, J. Structure of the mannose transporter of the bacterial phosphotransferase system. Cell Res. 2019, 29, 680–682. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.; Seo, H.S.; Jeong, J.; Yoon, H. Understanding the multifaceted roles of the phosphoenolpyruvate: Phosphotransferase system in regulation of Salmonella virulence using a mutant defective in ptsI and crr expression. Microbiol. Res. 2019, 223–225, 63–71. [Google Scholar] [CrossRef] [PubMed]
- Kotrba, P.; Inui, M.; Yukawa, H. Bacterial phosphotransferase system (PTS) in carbohydrate uptake and control of carbon metabolism. J. Biosci. Bioeng. 2001, 92, 502–517. [Google Scholar] [CrossRef]
- Laarman, A.; Milder, F.; Strijp, J.V.; Rooijakkers, S. Complement inhibition by gram-positive pathogens: Molecular mechanisms and therapeutic implications. J. Mol. Med. 2010, 88, 115–120. [Google Scholar] [CrossRef] [PubMed]
- Ye, K.; Li, P.; Gu, Q. Complete genome sequence analysis of a strain Lactobacillus pentosus ZFM94 and its probiotic characteristics. Genomics 2020, 112, 3142–3149. [Google Scholar] [CrossRef] [PubMed]
| Reads Type | Bases (bp) | Reads Number | Reads Mean Length (bp) | Reads N50 (bp) | Longest Reads (bp) | Q-Score |
|---|---|---|---|---|---|---|
| Raw | 1,722,592,614 | 117,399 | 14,672.97 | 28,556 | 179,164 | |
| Filtered | 1,671,594,841 | 113,051 | 14,786.2 | 28,642 | 179,164 | ≥7 |
| Total Reads | Clean Reads | Percentage (%) | Clean Bases | GC Content (%) | Q20 (%) | Q30 (%) |
|---|---|---|---|---|---|---|
| 7,089,390 | 7,074,292 | 99.79% | 1,061,058,974 | 41.0% | 97.95% | 93.42% |
| Category | Property |
|---|---|
| Total gene length (bp) | 1,776,396 |
| Average gene length (bp) | 875 |
| GC content in the gene region | 41 |
| Gene density(genes/Mb) | 1045 |
| Gene/Genome (%) | 89 |
| Intergenetic region length | 212,724 |
| Protein coding genes | 1943 |
| rRNA genes | 21 |
| tRNA genes | 65 |
| Gene density(genes/Mb) | 1045 |
| Gene/Genome(%) | 89 |
| Intergenetic region length | 212,724 |
| Sequencing depth | 533 |
| Accession No. | Size (Mbp) | GC (%) | Genes No. | Plasmids No. | Origin |
|---|---|---|---|---|---|
| CP064817 | 1.99 | 41.1 | 1943 | 3 | Yak feces |
| NZ_CP046037.1 | 2.02 | 41.1 | 1904 | 2 | Kimchi |
| NZ_CP025839.1 | 2.10 | 41.1 | 1964 | 0 | Kimchi |
| NZ_CP048116.1 | 1.99 | 41.0 | 1842 | 0 | Kimchi |
| NZ_CP025136.1 | 2.03 | 41.2 | 1936 | 2 | Kimchi |
| NZ_CP025206.1 | 2.04 | 41.0 | 1916 | 2 | Kimchi |
| NZ_CP020806.1 | 2.08 | 41.2 | 1975 | 2 | Kimchi |
| NZ_CP025203.1 | 2.04 | 41.2 | 1913 | 2 | Kimchi |
| JRFY01000024.1 | 2.19 | 40.6 | 1869 | ~ | Kimchi |
| NZ_LT960781.1 | 2.10 | 41.1 | 2024 | 2 | Sausage |
| NZ_LT960790.1 | 2.01 | 41.3 | 1886 | 2 | Sausage |
| NZ_LT907930.1 | 1.82 | 41.0 | 3532 | 0 | Sausage |
| NZ_LT907929.1 | 1.88 | 41.1 | 1752 | 0 | Sausage |
| NZ_ASTI01000039.1 | 2.03 | 40.9 | 1919 | 1 | Sausage |
| NZ_CP059697.1 | 2.06 | 41.0 | 2002 | 1 | Fermented vegetables |
| NZ_CP032652.1 | 2.02 | 41.2 | 1908 | 1 | Fermented vegetables |
| NZ_CP032640.1 | 2.02 | 41.2 | 1907 | 1 | Fermented vegetables |
| NZ_MKDM00000000.1 | 2.02 | 40.9 | 1934 | ~ | Meat |
| NZ_MKDO00000000.1 | 2.03 | 40.9 | 1938 | ~ | Meat |
| NZ_MKDC00000000.1 | 1.94 | 41.1 | 1892 | ~ | Meat |
| NZ_MKDN00000000.1 | 1.91 | 41.0 | 1838 | ~ | Meat |
| NZ_AZFG01000049.1 | 1.99 | 41.0 | 1923 | ~ | Meat |
| NZ_LT907933.1 | 1.98 | 41.0 | 1864 | 0 | Horse meat |
| NZ_LT960788.1 | 2.04 | 41.1 | 1936 | 1 | Beef carpaccio |
| NZ_LT960784.1 | 2.06 | 41.1 | 1928 | 3 | Beef carpaccio |
| NZ_CP022709.1 | 2.08 | 41.2 | 1955 | 2 | Water |
| NZ_LSFF00000000.1 | 2.01 | 41.0 | 1929 | ~ | Mukeunji |
| NZ_CP032633.1 | 2.02 | 41.2 | 1909 | 1 | Raw cow milk |
| NZ_CP032635.1 | 2.02 | 41.2 | 1909 | 1 | Raw cow milk |
| NZ_AP017931.1 | 1.99 | 41.2 | 1898 | 3 | Microbial mat material |
| NZ_CP020459.1 | 2.06 | 41.0 | 1921 | 2 | Food |
| NZ_LT960777.1 | 1.96 | 41.2 | 1813 | 3 | Human feces |
| NZ_CABMJT000000000.1 | 1.96 | 41.2 | 1813 | ~ | Human gut |
| NZ_OVTU00000000.1 | 2.06 | 40.8 | 1986 | ~ | Potatoes |
| NZ_OKRC00000000.1 | 2.03 | 40.9 | 1972 | ~ | Potatoes |
| NZ_CABFKU000000000.1 | 2.00 | 41.0 | 1923 | ~ | Human nasopharynx |
| NZ_SCIF00000000.1 | 1.98 | 41.0 | 1915 | ~ | Bovine |
| NZ_QOSE00000000.1 | 1.94 | 41.1 | 1859 | 3 | Sauerkraut |
| NZ_MKGH01000040.1 | 1.95 | 41.0 | 1878 | ~ | Cacao bean fermentation |
| NZ_MKGG01000009.1 | 1.91 | 41.0 | 1847 | ~ | Cacao bean fermentation |
| NZ_BJVN01000001.1 | 1.90 | 41.1 | 1822 | 1 | Moto starter of sake |
| NZ_AP017929.1 | 1.94 | 41.2 | 1834 | 1 | Moto starter of sake |
| NZ_AZDN01000001.1 | 1.91 | 41.0 | 1818 | ~ | Moto starter of sake |
| NZ_MKDI01000019.1 | 1.93 | 41.0 | 1849 | ~ | ~ |
| NZ_BKAA01000001.1 | 1.97 | 41.0 | 1920 | ~ | ~ |
| NC_007576.1/CR936503.1 | 1.88 | 41.3 | 1777 | 0 | ~ |
| BALW01000001.1 | 1.91 | 41.0 | ~ | ~ | ~ |
| NZ_PUFE01000027.1 | 1.94 | 41.1 | 1838 | ~ | ~ |
| NZ_LT907931.1 | 1.90 | 41.1 | 1782 | 0 | ~ |
| NZ_MKGB01000012.1 | 1.99 | 41.0 | 1900 | ~ | ~ |
| NZ_MKDL01000023.1 | 1.97 | 40.9 | 1890 | ~ | ~ |
| NZ_MKDJ00000000.1 | 1.92 | 41.1 | 1848 | ~ | ~ |
| NZ_MKDD00000000.1 | 2.01 | 41.0 | 1926 | ~ | ~ |
| NZ_BJLN00000000.1 | 1.94 | 41.1 | 1855 | ~ | ~ |
| NZ_MKDE00000000.1 | 1.95 | 40.9 | 1885 | ~ | ~ |
| Accession No. | Size (bp) | GC (%) | Genes No. | Origin |
|---|---|---|---|---|
| MW265923 | 74,183 | 40.0 | 75 | Yak feces |
| MW265924 | 8784 | 35.0 | 7 | Yak feces |
| MW265925 | 4034 | 35.0 | 6 | Yak feces |
| NZ_ASTI01000039.1 | 20,510 | 37.6 | 17 | Sausage |
| NZ_LT960791.1 | 33,463 | 41.2 | 28 | Sausage |
| NZ_LT960792.1 | 13,205 | 43.5 | 14 | Sausage |
| NZ_LT960782.1 | 46,347 | 39.2 | 43 | Sausage |
| NZ_LT960783.1 | 1526 | 40.2 | 2 | Sausage |
| NZ_CP020460.1 | 84,581 | 34.5 | 80 | Food |
| NZ_CP020461.1 | 26,098 | 40.4 | 21 | Food |
| NZ_CP022710.1 | 75,468 | 41.5 | 57 | Water |
| NZ_CP022711.1 | 11,504 | 34.6 | 10 | Water |
| NZ_AP017932.1 | 33,266 | 40.0 | 32 | Microbial mat material |
| NZ_AP017933.1 | 6196 | 35.9 | 6 | Microbial mat material |
| NZ_AP017934.1 | 4315 | 35.3 | 5 | Microbial mat material |
| NZ_LT960778.1 | 31,701 | 37.3 | 27 | Human feces |
| NZ_LT960779.1 | 12,663 | 43.4 | 13 | Human feces |
| NZ_LT960780.1 | 11,068 | 39.3 | 9 | Human feces |
| NZ_LT960789.1 | 47,094 | 40.7 | 38 | Beef carpaccio |
| NZ_LT960786.1 | 11,996 | 44.5 | 13 | Beef carpaccio |
| NZ_LT960787.1 | 11,156 | 39.5 | 8 | Beef carpaccio |
| NZ_LT960785.1 | 57,338 | 38.8 | 50 | Beef carpaccio |
| NZ_CP046038.1 | 53,648 | 39.5 | 53 | Kimchi |
| NZ_CP046039.1 | 20,312 | 38.5 | 21 | Kimchi |
| NZ_CP020807.1 | 13,019 | 41.2 | 9 | Kimchi |
| NZ_CP020808.1 | 13,083 | 38.1 | 10 | Kimchi |
| NZ_CP025137.1 | 42,297 | 41.4 | 32 | Kimchi |
| NZ_CP025138.1 | 14,883 | 36.3 | 20 | Kimchi |
| NZ_CP025207.1 | 45,431 | 40.4 | 42 | Kimchi |
| NZ_CP025208.1 | 44,839 | 38.5 | 38 | Kimchi |
| NZ_CP025204.1 | 75,290 | 41.1 | 66 | Kimchi |
| NZ_CP025205.1 | 14,876 | 36.3 | 20 | Kimchi |
| NZ_CM010625.1 | 11,246 | 39.7 | 8 | Sauerkraut |
| NZ_CM010626.1 | 6593 | 35.9 | 8 | Sauerkraut |
| NZ_CM010627.1 | 2499 | 33.8 | 3 | Sauerkraut |
| NZ_CP032636.1 | 51,286 | 41.9 | 40 | Raw cow milk |
| NZ_CP032634.1 | 51,286 | 41.9 | 42 | Raw cow milk |
| NZ_CP032641.1 | 51,284 | 41.9 | 39 | Fermented vegetables |
| NZ_CP032653.1 | 51,286 | 41.9 | 39 | Fermented vegetables |
| NZ_AP017930.1 | 6214 | 36.0 | 7 | Moto starter of sake |
| NZ_CP059698.1 | 56,165 | 41.1 | 48 | ~ |
| NC_004942.1 | 12,959 | 38.1 | 13 | ~ |
| EF605268.1 | 5031 | 36.7 | 3 | ~ |
| EU555173.1 | 1790 | 34.0 | 2 | ~ |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, K.; Liu, J.; Zeng, Z.; Kulyar, M.F.-e.-A.; Wang, Y.; Li, A.; Bhutta, Z.A.; Aqib, A.I.; Shahzad, M.; Li, J.; et al. The Complete Genome of Probiotic Lactobacillus sakei Derived from Plateau Yak Feces. Genes 2020, 11, 1527. https://doi.org/10.3390/genes11121527
Li K, Liu J, Zeng Z, Kulyar MF-e-A, Wang Y, Li A, Bhutta ZA, Aqib AI, Shahzad M, Li J, et al. The Complete Genome of Probiotic Lactobacillus sakei Derived from Plateau Yak Feces. Genes. 2020; 11(12):1527. https://doi.org/10.3390/genes11121527
Chicago/Turabian StyleLi, Kun, Juanjuan Liu, Zhibo Zeng, Muhammad Fakhar-e-Alam Kulyar, Yaping Wang, Aoyun Li, Zeeshan Ahmad Bhutta, Amjad Islam Aqib, Muhammad Shahzad, Jiakui Li, and et al. 2020. "The Complete Genome of Probiotic Lactobacillus sakei Derived from Plateau Yak Feces" Genes 11, no. 12: 1527. https://doi.org/10.3390/genes11121527
APA StyleLi, K., Liu, J., Zeng, Z., Kulyar, M. F.-e.-A., Wang, Y., Li, A., Bhutta, Z. A., Aqib, A. I., Shahzad, M., Li, J., & Qi, D. (2020). The Complete Genome of Probiotic Lactobacillus sakei Derived from Plateau Yak Feces. Genes, 11(12), 1527. https://doi.org/10.3390/genes11121527







