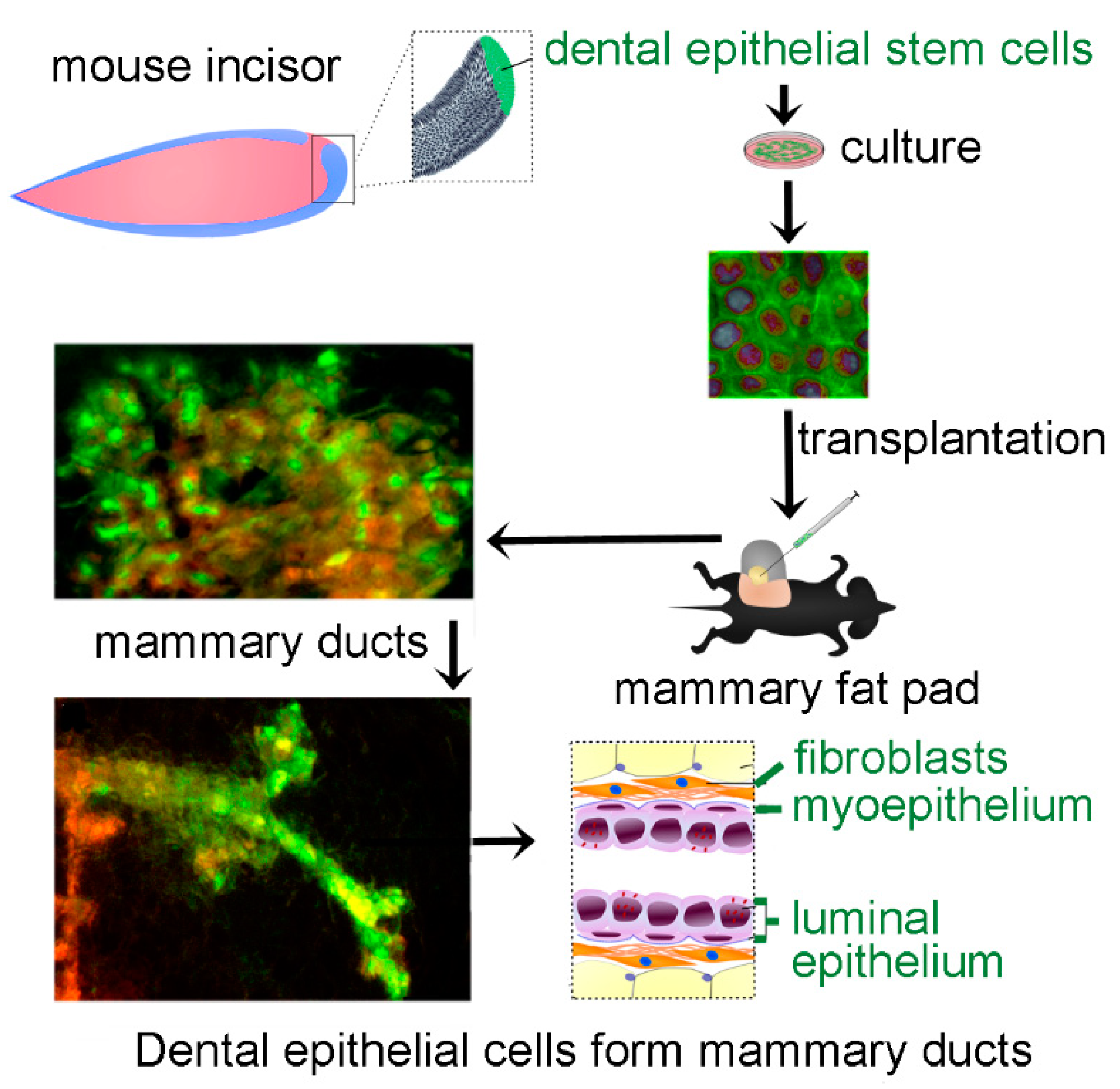
Dental Epithelial Stem Cells as a Source for Mammary Gland Regeneration and Milk Producing Cells In Vivo
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Isolation and Lentiviral Infection
2.2. Animals and Surgical Procedures
2.3. Immunofluorescence and Immunohistochemistry
2.4. RNA Extraction and Real Time Quantitative Polymerase Chain Reaction (RT-qPCRs)
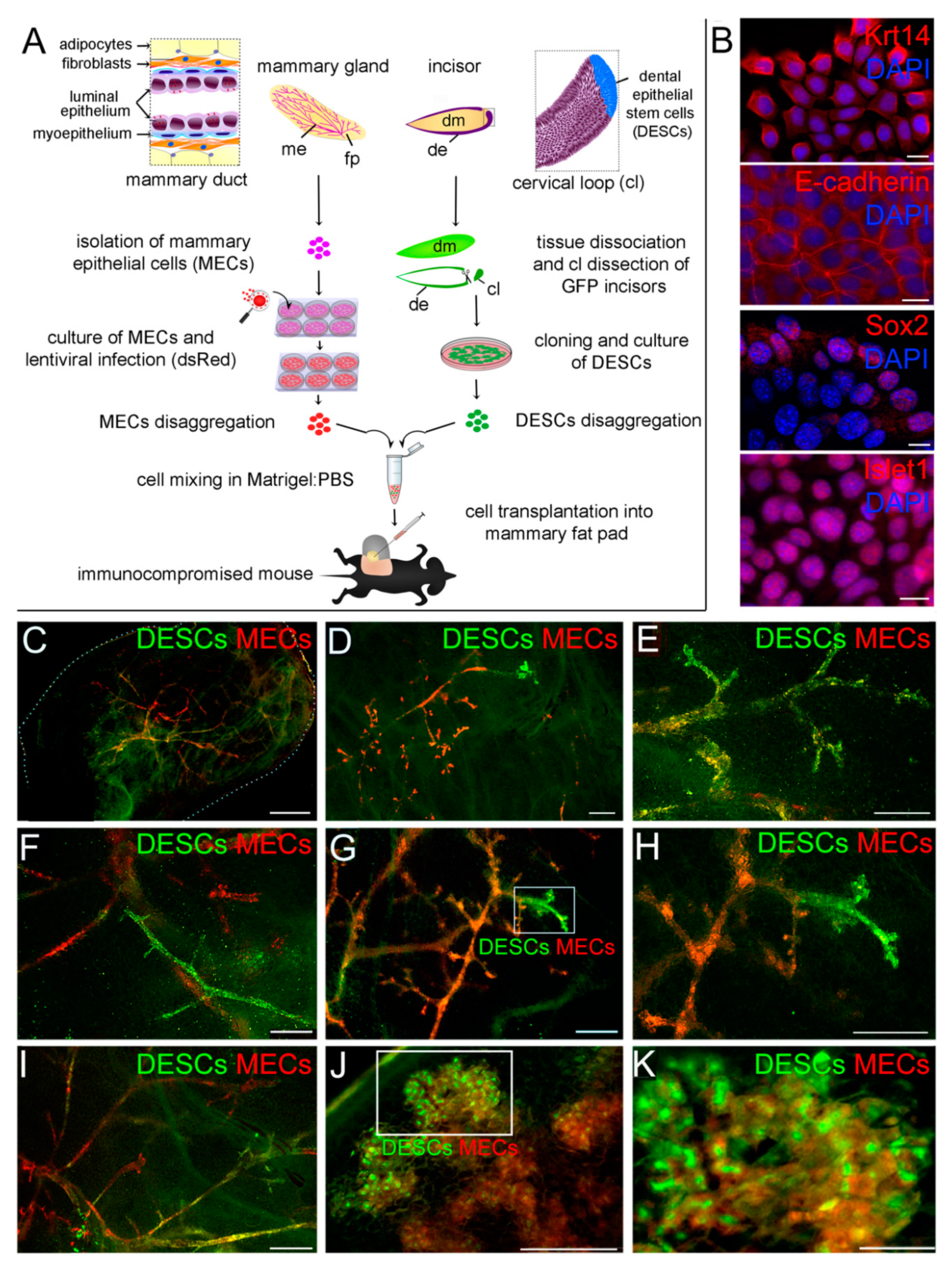
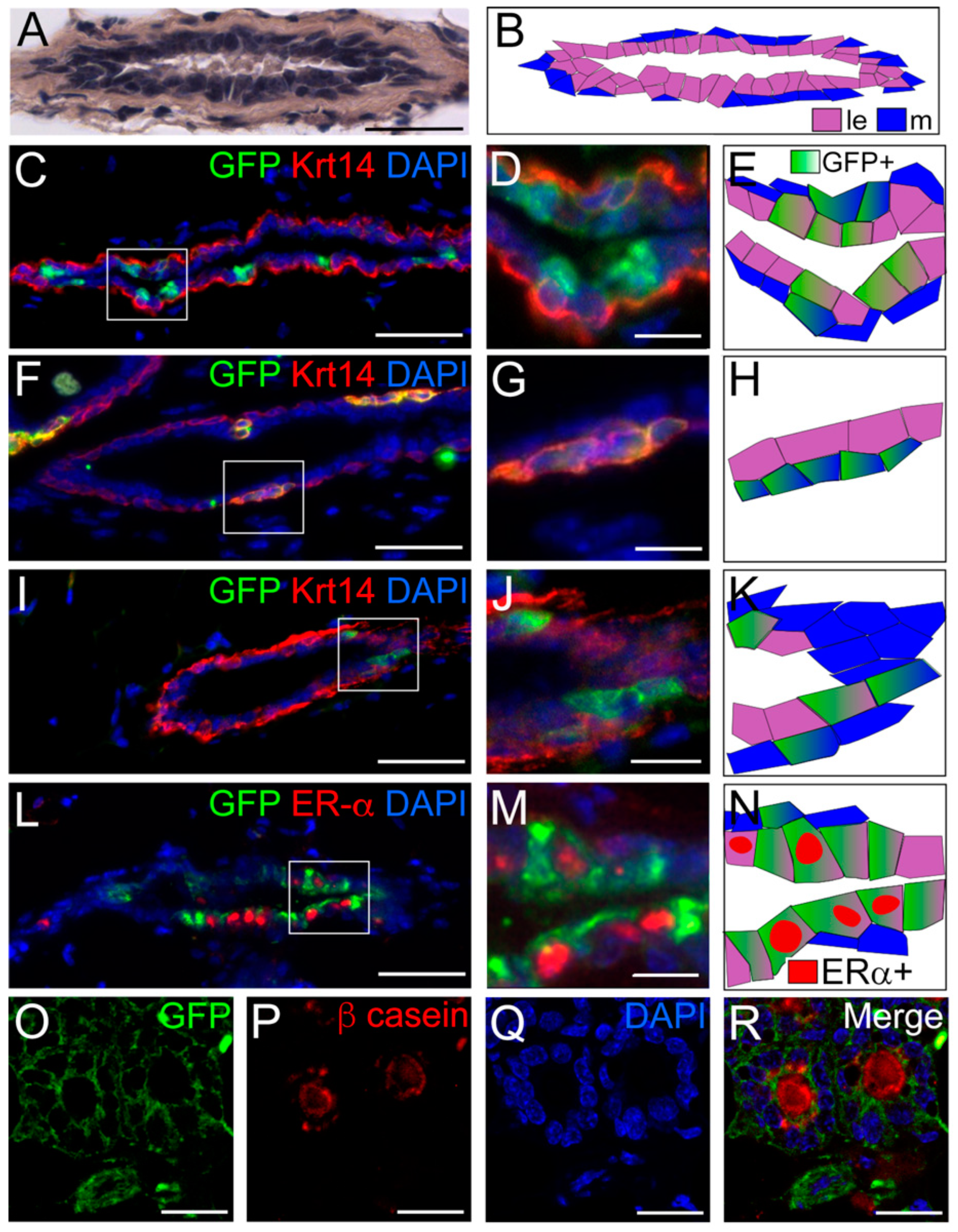
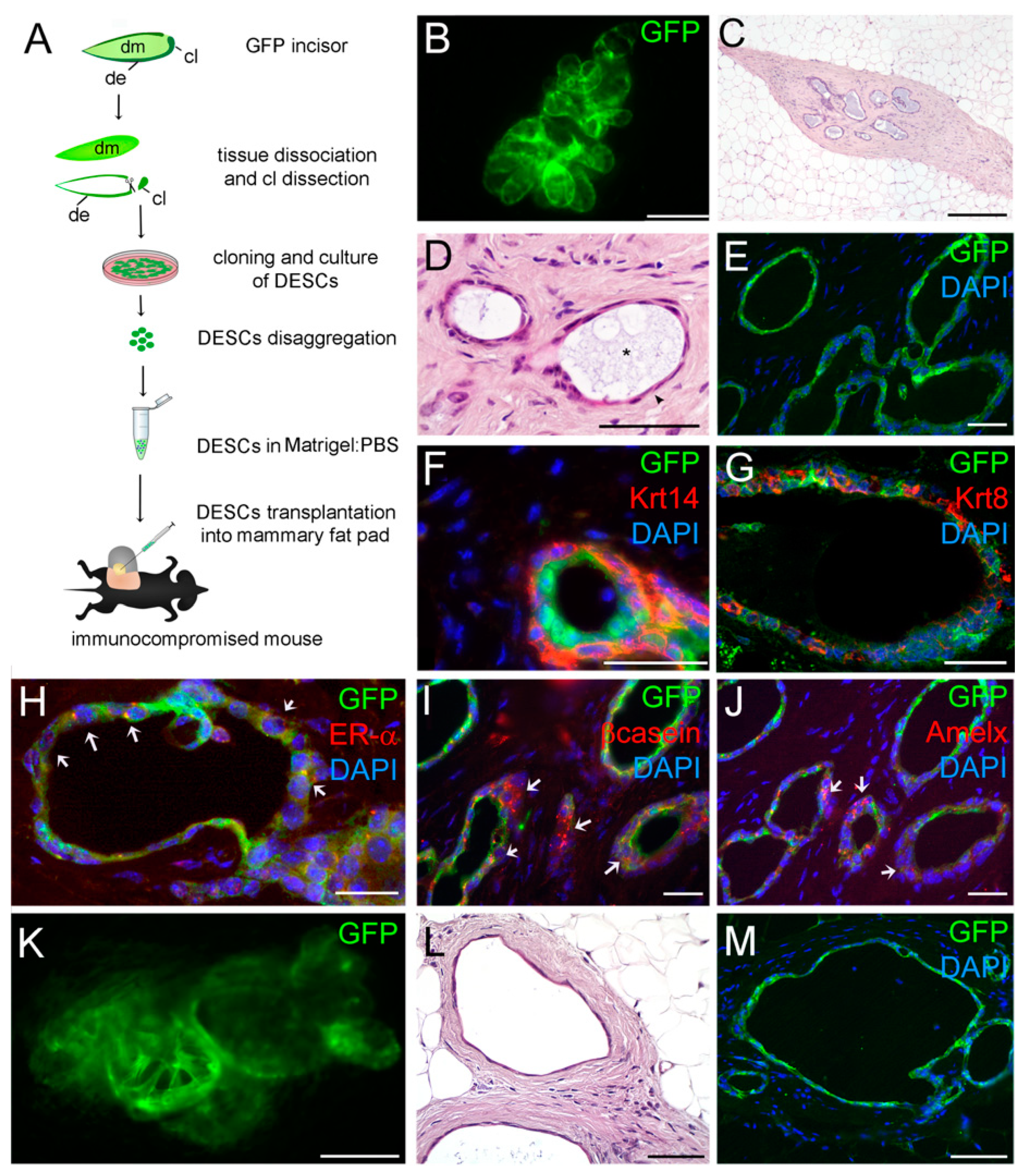
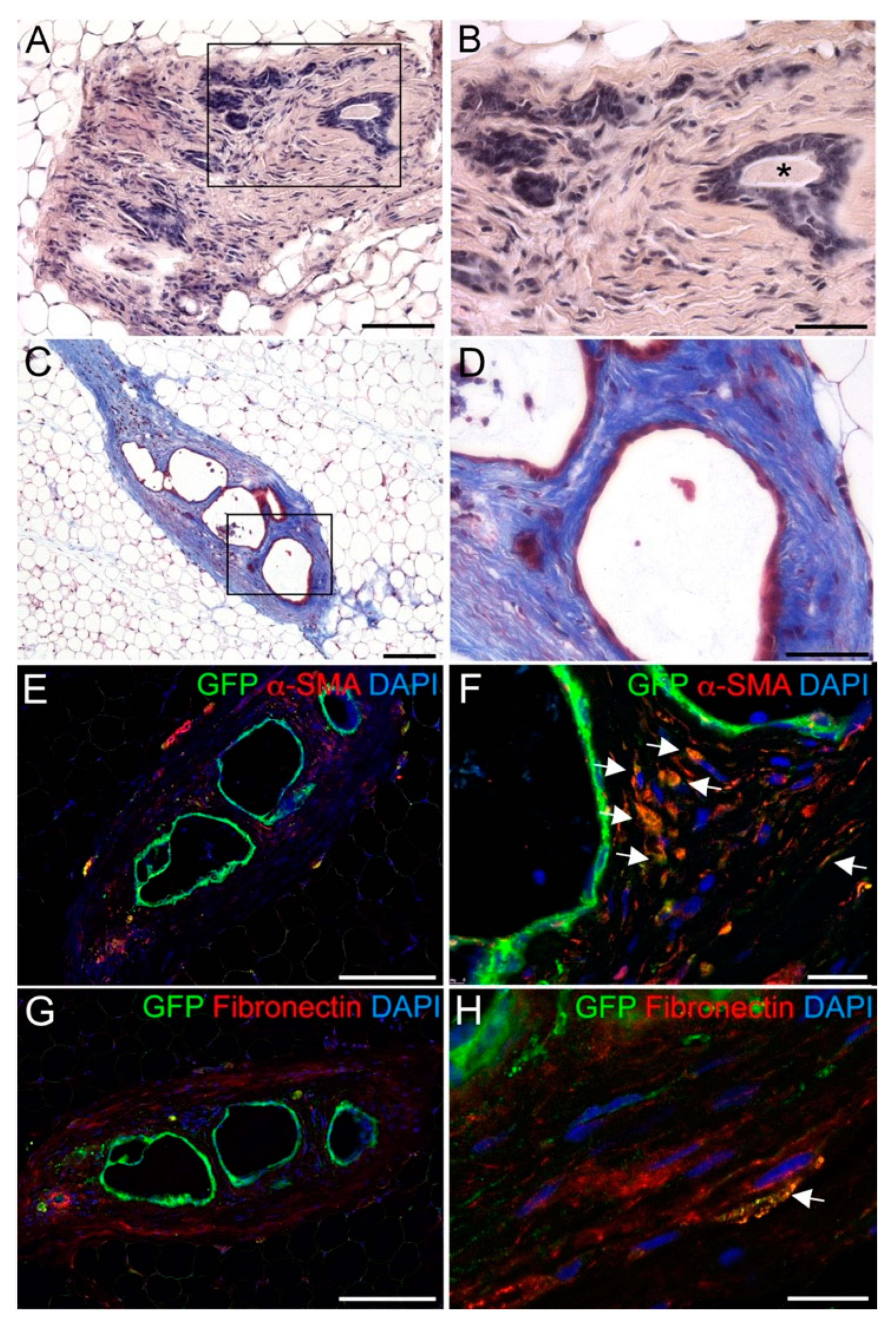
3. Results
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| DESCs | Dental epithelial stem cells |
| EGF | Epidermal growth factor |
| FGF | Fibroblasts growth factor |
| GFP | Green fluorescent protein |
| MECs | Mammary epithelial cells |
References
- Jimenez-Rojo, L.; Granchi, Z.; Graf, D.; Mitsiadis, T.A. Stem Cell Fate Determination during Development and Regeneration of Ectodermal Organs. Front. Physiol. 2012, 3, 107. [Google Scholar] [CrossRef] [PubMed]
- Biggs, L.C.; Mikkola, M.L. Early inductive events in ectodermal appendage morphogenesis. Semin. Cell Dev. Biol. 2014, 25, 11–21. [Google Scholar] [CrossRef] [PubMed]
- Yoshizaki, K.; Hu, L.; Nguyen, T.; Sakai, K.; He, B.; Fong, C.; Yamada, Y.; Bikle, D.D.; Oda, Y. Ablation of coactivator Med1 switches the cell fate of dental epithelia to that generating hair. PLoS ONE 2014, 9, e99991. [Google Scholar] [CrossRef] [PubMed]
- Kratochwil, K. Organ specificity in mesenchymal induction demonstrated in the embryonic development of the mammary gland of the mouse. Dev. Biol. 1969, 20, 46–71. [Google Scholar] [CrossRef]
- Lumsden, A.G. Spatial organization of the epithelium and the role of neural crest cells in the initiation of the mammalian tooth germ. Development 1988, 103, 155–169. [Google Scholar]
- Blanpain, C.; Fuchs, E. Stem cell plasticity. Plasticity of epithelial stem cells in tissue regeneration. Science 2014, 344, 1242281. [Google Scholar] [CrossRef]
- Wabik, A.; Jones, P.H. Switching roles: The functional plasticity of adult tissue stem cells. EMBO J. 2015, 34, 1164–1179. [Google Scholar] [CrossRef]
- Seidel, K.; Ahn, C.P.; Lyons, D.; Nee, A.; Ting, K.; Brownell, I.; Cao, T.; Carano, R.A.; Curran, T.; Schober, M.; et al. Hedgehog signaling regulates the generation of ameloblast progenitors in the continuously growing mouse incisor. Development 2010, 137, 3753–3761. [Google Scholar] [CrossRef]
- Juuri, E.; Saito, K.; Ahtiainen, L.; Seidel, K.; Tummers, M.; Hochedlinger, K.; Klein, O.D.; Thesleff, I.; Michon, F. Sox2+ stem cells contribute to all epithelial lineages of the tooth via Sfrp5+ progenitors. Dev. Cell 2012, 23, 317–328. [Google Scholar] [CrossRef]
- Biehs, B.; Hu, J.K.; Strauli, N.B.; Sangiorgi, E.; Jung, H.; Heber, R.P.; Ho, S.; Goodwin, A.F.; Dasen, J.S.; Capecchi, M.R.; et al. BMI1 represses Ink4a/Arf and Hox genes to regulate stem cells in the rodent incisor. Nat. Cell Biol. 2013, 15, 846–852. [Google Scholar] [CrossRef]
- Shackleton, M.; Vaillant, F.; Simpson, K.J.; Stingl, J.; Smyth, G.K.; Asselin-Labat, M.L.; Wu, L.; Lindeman, G.J.; Visvader, J.E. Generation of a functional mammary gland from a single stem cell. Nature 2006, 439, 84–88. [Google Scholar] [CrossRef] [PubMed]
- Boulanger, C.A.; Smith, G.H. Reprogramming cell fates in the mammary microenvironment. Cell Cycle 2009, 8, 1127–1132. [Google Scholar] [CrossRef] [PubMed]
- Deome, K.B.; Faulkin, L.J.; Bern, H.A., Jr.; Blair, P.B. Development of mammary tumors from hyperplastic alveolar nodules transplanted into gland-free mammary fat pads of female C3H mice. Cancer Res. 1959, 19, 515–520. [Google Scholar] [PubMed]
- Asselin-Labat, M.L.; Vaillant, F.; Sheridan, J.M.; Pal, B.; Wu, D.; Simpson, E.R.; Yasuda, H.; Smyth, G.K.; Martin, T.J.; Lindeman, G.J.; et al. Control of mammary stem cell function by steroid hormone signalling. Nature 2010, 465, 798–802. [Google Scholar] [CrossRef]
- Wang, D.; Cai, C.; Dong, X.; Yu, Q.C.; Zhang, X.O.; Yang, L.; Zeng, Y.A. Identification of multipotent mammary stem cells by protein C receptor expression. Nature 2015, 517, 81–84. [Google Scholar] [CrossRef]
- Boulanger, C.A.; Mack, D.L.; Booth, B.W.; Smith, G.H. Interaction with the mammary microenvironment redirects spermatogenic cell fate in vivo. Proc. Natl. Acad. Sci. USA 2007, 104, 3871–3876. [Google Scholar] [CrossRef]
- Booth, B.W.; Mack, D.L.; Androutsellis-Theotokis, A.; McKay, R.D.; Boulanger, C.A.; Smith, G.H. The mammary microenvironment alters the differentiation repertoire of neural stem cells. Proc. Natl. Acad. Sci. USA 2008, 105, 14891–14896. [Google Scholar] [CrossRef]
- Boulanger, C.A.; Bruno, R.D.; Rosu-Myles, M.; Smith, G.H. The mouse mammary microenvironment redirects mesoderm-derived bone marrow cells to a mammary epithelial progenitor cell fate. Stem Cells Dev. 2012, 21, 948–954. [Google Scholar] [CrossRef]
- Smalley, M.J. Isolation, culture and analysis of mouse mammary epithelial cells. Methods Mol. Biol. 2010, 633, 139–170. [Google Scholar]
- Mitsiadis, T.A.; Lardelli, M.; Lendahl, U.; Thesleff, I. Expression of Notch 1, 2 and 3 is regulated by epithelial-mesenchymal interactions and retinoic acid in the developing mouse tooth and associated with determination of ameloblast cell fate. J. Cell Biol. 1995, 130, 407–418. [Google Scholar] [CrossRef]
- Gusterson, B.A.; Ross, D.T.; Heath, V.J.; Stein, T. Basal cytokeratins and their relationship to the cellular origin and functional classification of breast cancer. Breast Cancer Res. BCR 2005, 7, 143–148. [Google Scholar] [CrossRef] [PubMed]
- Rodilla, V.; Dasti, A.; Huyghe, M.; Lafkas, D.; Laurent, C.; Reyal, F.; Fre, S. Luminal progenitors restrict their lineage potential during mammary gland development. PLoS Biol. 2015, 13, e1002069. [Google Scholar] [CrossRef] [PubMed]
- Visvader, J.E.; Stingl, J. Mammary stem cells and the differentiation hierarchy: Current status and perspectives. Genes Dev. 2014, 28, 1143–1158. [Google Scholar] [CrossRef] [PubMed]
- Hennighausen, L.; Robinson, G.W. Information networks in the mammary gland. Nat. Rev. Mol. Cell Biol. 2005, 6, 715–725. [Google Scholar] [CrossRef]
- Barron, M.J.; Brookes, S.J.; Kirkham, J.; Shore, R.C.; Hunt, C.; Mironov, A.; Kingswell, N.J.; Maycock, J.; Shuttleworth, C.A.; Dixon, M.J. A mutation in the mouse Amelx tri-tyrosyl domain results in impaired secretion of amelogenin and phenocopies human X-linked amelogenesis imperfecta. Hum. Mol. Genet. 2010, 19, 1230–1247. [Google Scholar] [CrossRef]
- Mitsiadis, T.A.; Graf, D. Cell fate determination during tooth development and regeneration. Birth Defects Res. C Embryol. Today 2009, 87, 199–211. [Google Scholar] [CrossRef]
- Inman, J.L.; Robertson, C.; Mott, J.D.; Bissell, M.J. Mammary gland development: Cell fate specification, stem cells and the microenvironment. Development 2015, 142, 1028–1042. [Google Scholar] [CrossRef]
- Mues, G.; Bonds, J.; Xiang, L.; Vieira, A.R.; Seymen, F.; Klein, O.; D’Souza, R.N. The WNT10A gene in ectodermal dysplasias and selective tooth agenesis. Am. J. Med. Genet. A 2014, 164, 2455–2460. [Google Scholar] [CrossRef]
- Hens, J.R.; Wysolmerski, J.J. Key stages of mammary gland development: Molecular mechanisms involved in the formation of the embryonic mammary gland. Breast Cancer Res. 2005, 7, 220–224. [Google Scholar] [CrossRef]
- McBryan, J.; Howlin, J. Pubertal Mammary Gland Development: Elucidation of In Vivo Morphogenesis Using Murine Models. Methods Mol. Biol. 2017, 1501, 77–114. [Google Scholar]
- Pond, A.C.; Bin, X.; Batts, T.; Roarty, K.; Hilsenbeck, S.; Rosen, J.M. Fibroblast growth factor receptor signaling is essential for normal mammary gland development and stem cell function. Stem Cells 2013, 31, 178–189. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Martinez, D.; Koledova, Z.; Qiao, G.; Streuli, C.H.; Lu, P. FGF ligands of the postnatal mammary stroma regulate distinct aspects of epithelial morphogenesis. Development 2014, 141, 3352–3362. [Google Scholar] [CrossRef] [PubMed]
- Kettunen, P.; Laurikkala, J.; Itaranta, P.; Vainio, S.; Itoh, N.; Thesleff, I. Associations of FGF-3 and FGF-10 with signaling networks regulating tooth morphogenesis. Dev. Dyn. 2000, 219, 322–332. [Google Scholar] [PubMed]
- Harada, H.; Kettunen, P.; Jung, H.S.; Mustonen, T.; Wang, Y.A.; Thesleff, I. Localization of putative stem cells in dental epithelium and their association with Notch and FGF signaling. J. Cell Biol 1999, 147, 105–120. [Google Scholar] [CrossRef] [PubMed]
- Harada, H.; Toyono, T.; Toyoshima, K.; Yamasaki, M.; Itoh, N.; Kato, S.; Sekine, K.; Ohuchi, H. FGF10 maintains stem cell compartment in developing mouse incisors. Development 2002, 129, 1533–1541. [Google Scholar]
- Mitsiadis, T.A.; Barrandon, O.; Rochat, A.; Barrandon, Y.; De Bari, C. Stem cell niches in mammals. Exp. Cell Res. 2007, 313, 3377–3385. [Google Scholar]
- Couse, J.F.; Korach, K.S. Estrogen receptor null mice: What have we learned and where will they lead us? Endocr. Rev. 1999, 20, 358–417. [Google Scholar] [CrossRef]
- Feng, Y.; Manka, D.; Wagner, K.U.; Khan, S.A. Estrogen receptor-α expression in the mammary epithelium is required for ductal and alveolar morphogenesis in mice. Proc. Natl. Acad. Sci. USA 2007, 104, 14718–14723. [Google Scholar] [CrossRef]
- Mallepell, S.; Krust, A.; Chambon, P.; Brisken, C. Paracrine signaling through the epithelial estrogen receptor α is required for proliferation and morphogenesis in the mammary gland. Proc. Natl. Acad. Sci. USA 2006, 103, 2196–2201. [Google Scholar] [CrossRef]
- Diegelmann, R.F.; Evans, M.C. Wound healing: An overview of acute, fibrotic and delayed healing. Front. Biosci. A J. Virtual Libr. 2004, 9, 283–289. [Google Scholar] [CrossRef]
- Wynn, T.A.; Ramalingam, T.R. Mechanisms of fibrosis: Therapeutic translation for fibrotic disease. Nature Med. 2012, 18, 1028–1040. [Google Scholar] [CrossRef] [PubMed]
- Brownfield, D.G.; Venugopalan, G.; Lo, A.; Mori, H.; Tanner, K.; Fletcher, D.A.; Bissell, M.J. Patterned collagen fibers orient branching mammary epithelium through distinct signaling modules. Curr. Biol. 2013, 23, 703–709. [Google Scholar] [CrossRef] [PubMed]
- Schedin, P.; Keely, P.J. Mammary gland ECM remodeling, stiffness, and mechanosignaling in normal development and tumor progression. Cold Spring Harb. Perspect. Biol. 2011, 3, a003228. [Google Scholar] [CrossRef] [PubMed]
- Silberstein, G.B.; Flanders, K.C.; Roberts, A.B.; Daniel, C.W. Regulation of mammary morphogenesis: Evidence for extracellular matrix-mediated inhibition of ductal budding by transforming growth factor-β 1. Dev. Biol 1992, 152, 354–362. [Google Scholar] [CrossRef]
- Nelson, C.M.; Vanduijn, M.M.; Inman, J.L.; Fletcher, D.A.; Bissell, M.J. Tissue geometry determines sites of mammary branching morphogenesis in organotypic cultures. Science 2006, 314, 298–300. [Google Scholar] [CrossRef] [PubMed]
- Ewan, K.B.; Oketch-Rabah, H.A.; Ravani, S.A.; Shyamala, G.; Moses, H.L.; Barcellos-Hoff, M.H. Proliferation of estrogen receptor-α-positive mammary epithelial cells is restrained by transforming growth factor-β1 in adult mice. Am. J. Pathol 2005, 167, 409–417. [Google Scholar] [CrossRef]
- Moses, H.; Barcellos-Hoff, M.H. TGF-β biology in mammary development and breast cancer. Cold Spring Harb. Perspect. Biol 2011, 3, a003277. [Google Scholar] [CrossRef]
- Iezzi, I.; Pagella, P.; Mattioli-Belmonte, M.; Mitsiadis, T.A. The effects of ageing on dental pulp stem cells, the tooth longevity elixir. Eur. Cell Mater. 2019, 26, 175–185. [Google Scholar] [CrossRef]
- Sharir, A.; Marangoni, P.; Zilionis, R.; Wan, M.; Wald, T.; Hu, J.K.; Kawaguchi, K.; Castillo-Azofeifa, D.; Epstein, L.; Harrington, K.; et al. A large pool of actively cycling progenitors orchestrates self-renewal and injury repair of an ectodermal appendage. Nat. Cell Biol. 2019, 21, 1102–1112. [Google Scholar] [CrossRef]
| No. of Dental Epithelial Stem Cells (DESCs) | No. of Mammary Epithelial Cells (MECs) | No. of Inoculated Fat Pads | No. of Intakes | No. of Outgrowths |
|---|---|---|---|---|
| 50,000 | 50,000 | 7 | 6 | 6 |
| 100,000 | 50,000 | 4 | 3 | 3 |
| 100,000 | 0 | 6 | 6 | 6 |
| 50,000 * | 50,000 * | 5 * | 5 * | 5 * |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jimenez-Rojo, L.; Pagella, P.; Harada, H.; Mitsiadis, T.A. Dental Epithelial Stem Cells as a Source for Mammary Gland Regeneration and Milk Producing Cells In Vivo. Cells 2019, 8, 1302. https://doi.org/10.3390/cells8101302
Jimenez-Rojo L, Pagella P, Harada H, Mitsiadis TA. Dental Epithelial Stem Cells as a Source for Mammary Gland Regeneration and Milk Producing Cells In Vivo. Cells. 2019; 8(10):1302. https://doi.org/10.3390/cells8101302
Chicago/Turabian StyleJimenez-Rojo, Lucia, Pierfrancesco Pagella, Hidemitsu Harada, and Thimios A. Mitsiadis. 2019. "Dental Epithelial Stem Cells as a Source for Mammary Gland Regeneration and Milk Producing Cells In Vivo" Cells 8, no. 10: 1302. https://doi.org/10.3390/cells8101302
APA StyleJimenez-Rojo, L., Pagella, P., Harada, H., & Mitsiadis, T. A. (2019). Dental Epithelial Stem Cells as a Source for Mammary Gland Regeneration and Milk Producing Cells In Vivo. Cells, 8(10), 1302. https://doi.org/10.3390/cells8101302







