Association Mapping of Fertility Restorer Gene for CMS PET1 in Sunflower
Abstract
1. Introduction
2. Materials and Methods
3. Results and Discussion
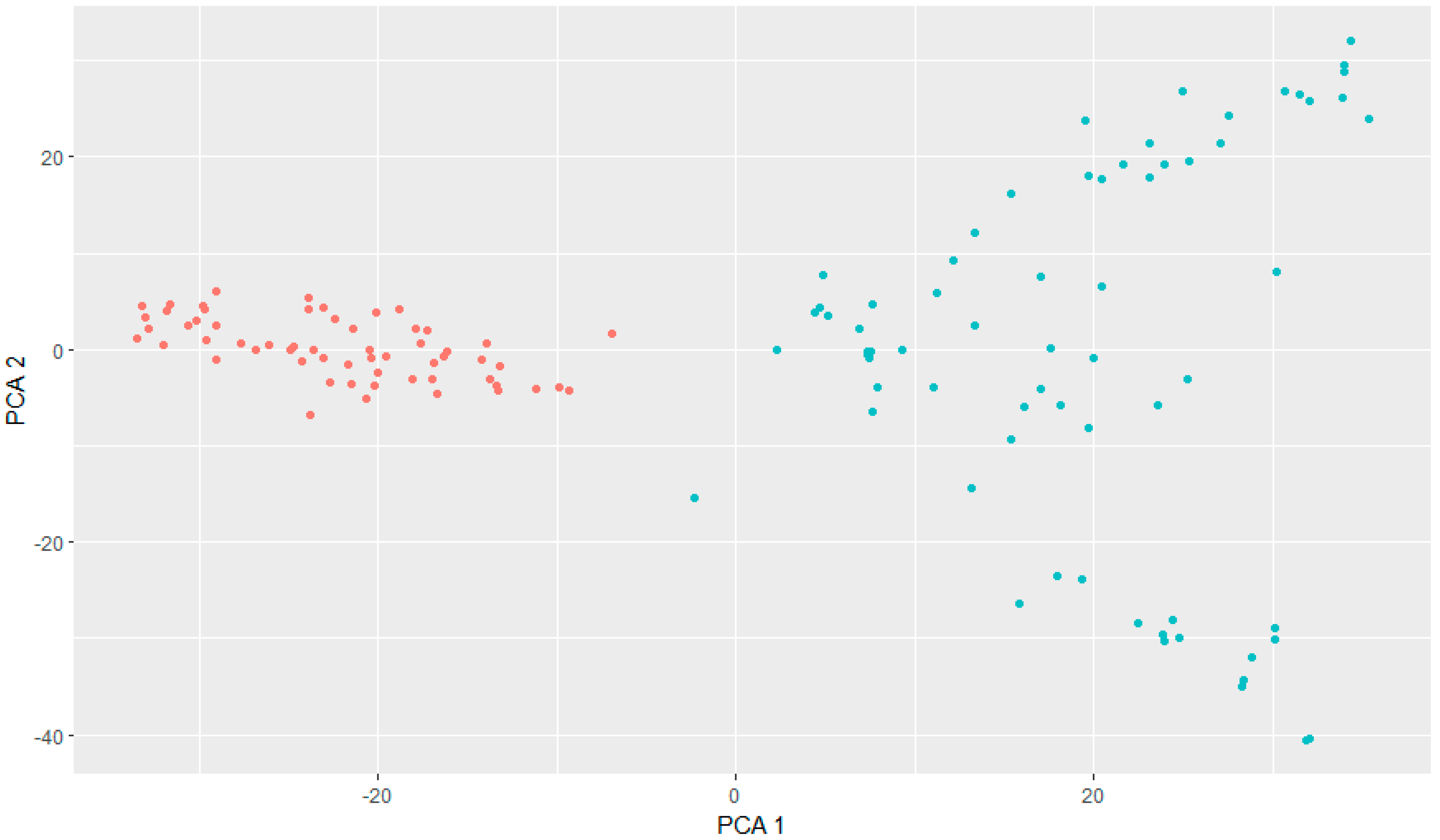
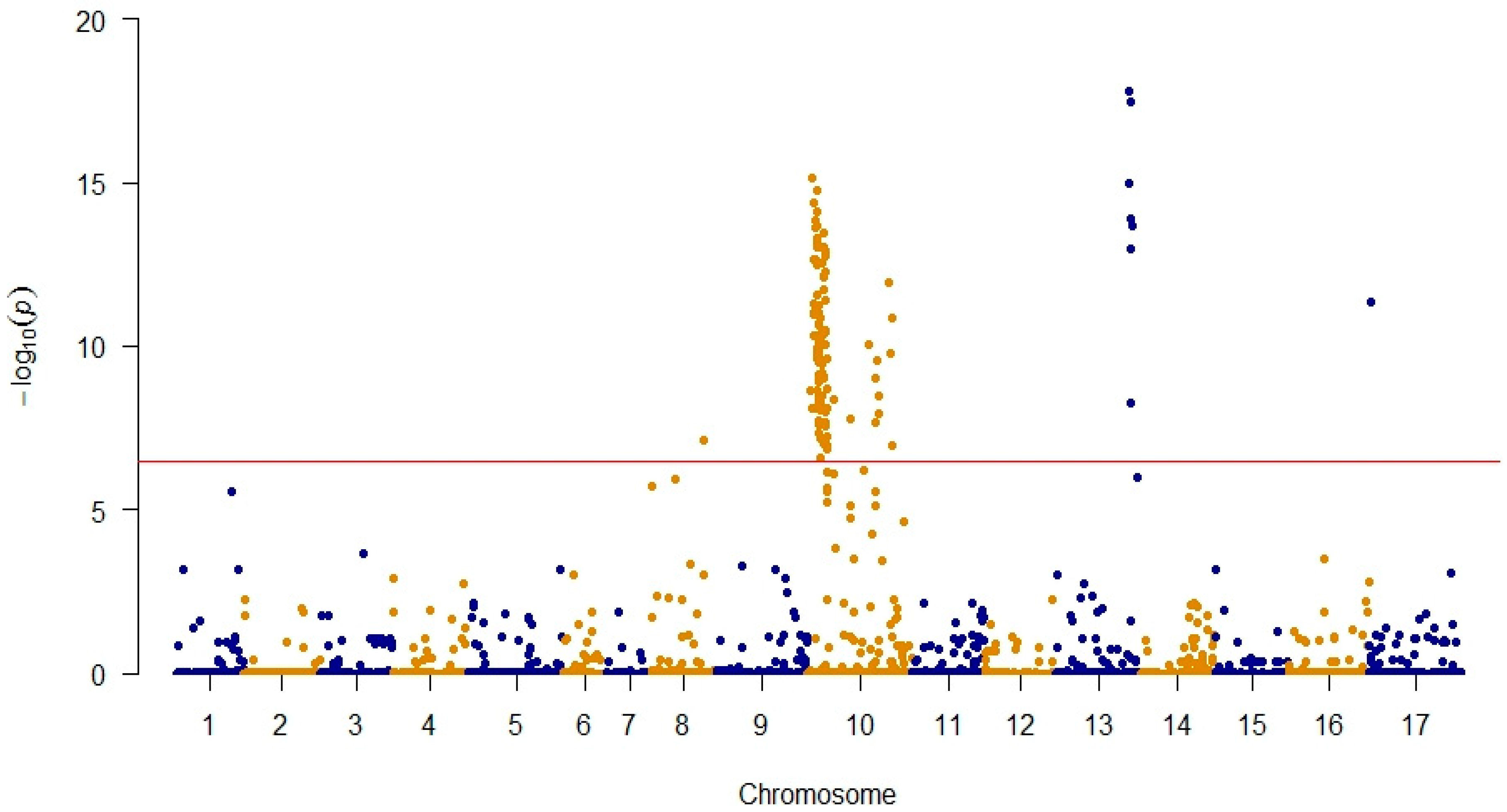
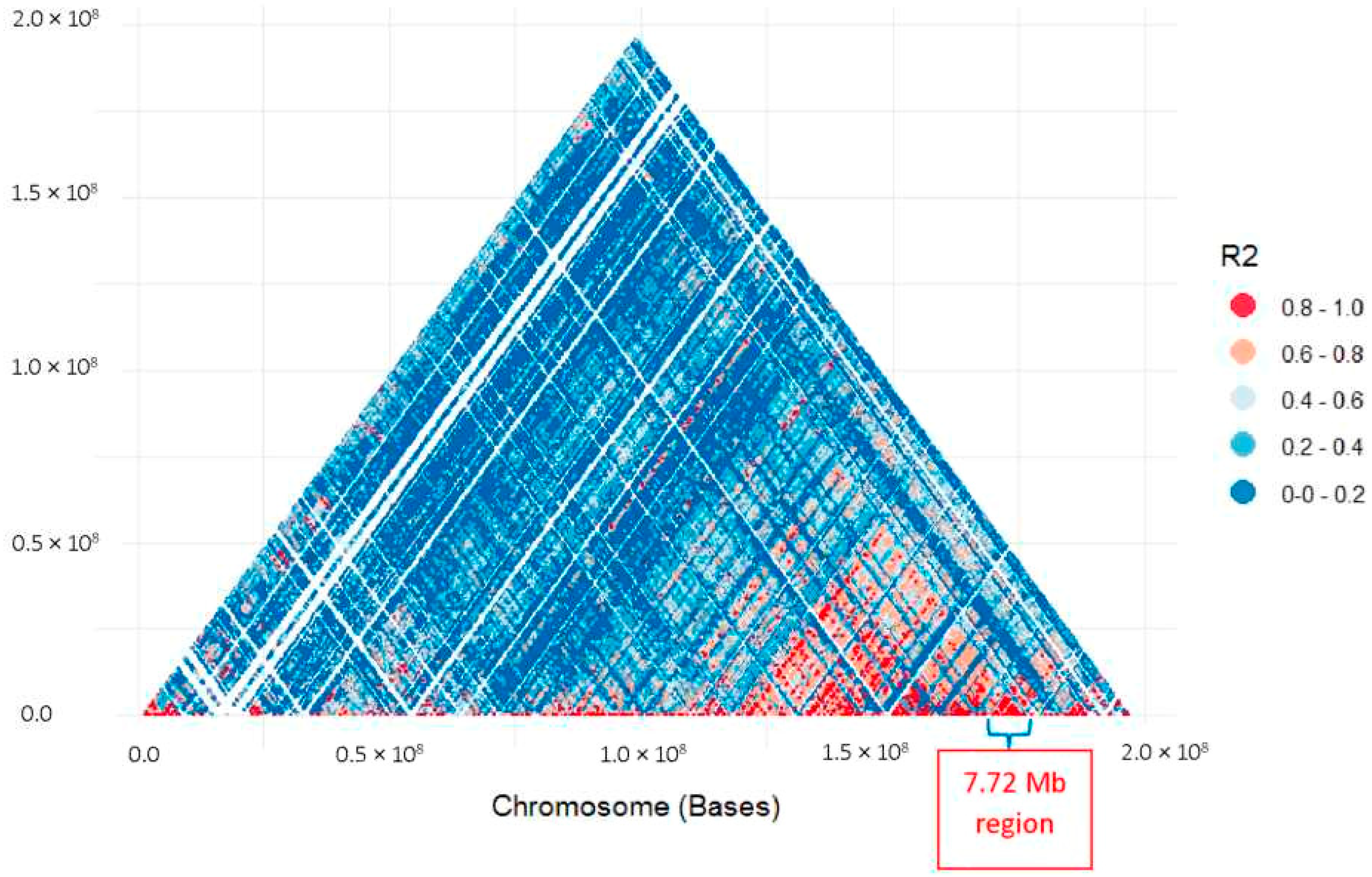
3.1. Genotyping and GWAS Analysis
3.2. Identification of Rf1 Candidate Genes
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Rhoades, M.M. Cytoplasmic inheritance of male sterility in Zea mays. Science 1931, 73, 340–341. [Google Scholar] [CrossRef] [PubMed]
- Schnable, P. The molecular basis of cytoplasmic male sterility and fertility restoration. Trends Plant Sci. 1998, 3, 175–180. [Google Scholar] [CrossRef]
- Ivanov, M.K.; Dymshits, G.M. Cytoplasmic male sterility and restoration of pollen fertility in higher plants. Russ. J. Genet. 2007, 43, 354–368. [Google Scholar] [CrossRef]
- Anisimova, I.N.; Gavrilova, V.A.; Rozhkova, V.T.; Timofeeva, G.I.; Tikhonova, M.A. Molecular markers in identification of sunflower pollen fertility restorer genes. Russ. Agric. Sci. 2009, 35, 367–370. [Google Scholar] [CrossRef]
- Leclercq, P. Une sterilite male cytoplasmique chez le tournesol. Ann. Amel. Plantes 1969, 19, 99–106. [Google Scholar]
- Horn, R.; Köhler, R.H.; Zetsche, K. A mitochondrial 16 kDa protein is associated with cytoplasmic male sterility in sunflower. Plant Mol. Biol. 1991, 17, 29–36. [Google Scholar] [CrossRef] [PubMed]
- Hans Köhler, R.; Horn, R.; Lössl, A.; Zetsche, K. Cytoplasmic male sterility in sunflower is correlated with the co-transcription of a new open reading frame with the atpA gene. MGG Mol. Gen. Genet. 1991, 227, 369–376. [Google Scholar] [CrossRef]
- Miller, J.F.; Fick, G.N. Genetics of sunflower. In Sunflower Technology and Production; American Society of Agronomy, Crop Science Society of America, Soil Science Society of America: Madison, WI, USA, 1997; pp. 441–495. ISBN 978-0-89118-135-4. [Google Scholar]
- Anaschenko, A.V.; Duca, M.V. Studies of sunflower (Helianthus annuus L.) CMS–Rf genetic system: II. Male fertility restoration in hybrids based on CMSP. Russ. J. Genet. 1985, 21, 1999–2004. [Google Scholar]
- Serieys, H. Identification, study and utilization in breeding programs of new CMS sources. Helia 1996, 19, 144–158. [Google Scholar]
- Gentzbittel, L.; Vear, F.; Zhang, Y.-X.; Bervillé, A.; Nicolas, P. Development of a consensus linkage RFLP map of cultivated sunflower (Helianthus annuus L.). Theor. Appl. Genet. 1995, 90, 1079–1086. [Google Scholar] [CrossRef]
- Jan, C.C.; Vick, B.A.; Miller, J.F.; Kahler, A.L.; Butler, E.T. Construction of an RFLP linkage map for cultivated sunflower. Theor. Appl. Genet. 1998, 96, 15–22. [Google Scholar] [CrossRef]
- Kusterer, B.; Horn, R.; Friedt, W. Molecular mapping of the fertility restoration locus Rf1 in sunflower and development of diagnostic markers for the restorer gene. Euphytica 2005, 143, 35–42. [Google Scholar] [CrossRef]
- Tang, S.; Yu, J.-K.; Slabaugh, B.; Shintani, K.; Knapp, J. Simple sequence repeat map of the sunflower genome. Theor. Angew. Genet. 2002, 105, 1124–1136. [Google Scholar] [CrossRef]
- Horn, R.; Kusterer, B.; Lazarescu, E.; Prüfe, M.; Friedt, W. Molecular mapping of the Rf1 gene restoring pollen fertility in PET1-based F1 hybrids in sunflower (Helianthus annuus L.). Theor. Appl. Genet. 2003, 106, 599–606. [Google Scholar] [CrossRef] [PubMed]
- Markin, N.; Usatov, A.; Makarenko, M.; Azarin, K.; Gorbachenko, O.; Kolokolova, N.; Usatenko, T.; Markina, O.; Gavrilova, V. Study of informative DNA markers of the Rf1 gene in sunflower for breeding practice. Czech J. Genet. Plant Breed. 2017, 53, 69–75. [Google Scholar] [CrossRef]
- Anisimova, I.N.; Gavrilova, V.A.; Rozhkova, V.T.; Port, A.I.; Timofeeva, G.I.; Duka, M.V. Genetic diversity of sources of sunflower pollen fertility restorer genes. Russ. Agric. Sci. 2011, 37, 192–196. [Google Scholar] [CrossRef]
- Sykes, T.; Yates, S.; Nagy, I.; Asp, T.; Small, I.; Studer, B. In-silico identification of candidate genes for fertility restoration in cytoplasmic male sterile perennial ryegrass (Lolium perenne L.). Genome Biol. Evol. 2016, 9, 351–362. [Google Scholar]
- Horn, R.; Hamrit, S. Gene cloning and characterization. In Genetics, Genomics and Breeding of Sunflower; CRC Press: Boca Raton, FL, USA, 2010; pp. 173–219. ISBN 978-1-138-11513-2. [Google Scholar]
- Yue, B.; Vick, B.A.; Cai, X.; Hu, J. Genetic mapping for the Rf1 (fertility restoration) gene in sunflower (Helianthus annuus L.) by SSR and TRAP markers. Plant Breed. 2010, 129, 24–28. [Google Scholar] [CrossRef]
- Anisimova, I.N.; Alpatieva, N.V.; Rozhkova, V.T.; Kuznetsova, E.B.; Pinaev, A.G.; Gavrilova, V.A. Polymorphism among RFL-PPR homologs in sunflower (Helianthus annuus L.) lines with varying ability for the suppression of the cytoplasmic male sterility phenotype. Russ. J. Genet. 2014, 50, 712–721. [Google Scholar] [CrossRef]
- Cui, X.; Wise, R.P.; Schnable, P.S. The rf2 nuclear restorer gene of male-sterile T-cytoplasm maize. Science 1996, 272, 1334–1336. [Google Scholar] [CrossRef]
- Itabashi, E.; Iwata, N.; Fujii, S.; Kazama, T.; Toriyama, K. The fertility restorer gene, Rf2, for Lead Rice-type cytoplasmic male sterility of rice encodes a mitochondrial glycine-rich protein: Rf2 for CMS rice encodes a glycine-rich protein. Plant J. 2011, 65, 359–367. [Google Scholar] [CrossRef] [PubMed]
- Melonek, J.; Stone, J.D.; Small, I. Evolutionary plasticity of restorer-of-fertility-like proteins in rice. Sci. Rep. 2016, 6, 35152. [Google Scholar] [CrossRef] [PubMed]
- Lurin, C. Genome-wide analysis of Arabidopsis pentatricopeptide repeat proteins reveals their essential role in organelle biogenesis. Plant Cell 2004, 16, 2089–2103. [Google Scholar] [CrossRef] [PubMed]
- Cheng, S.; Gutmann, B.; Zhong, X.; Ye, Y.; Fisher, M.F.; Bai, F.; Castleden, I.; Song, Y.; Song, B.; Huang, J.; et al. Redefining the structural motifs that determine RNA binding and RNA editing by pentatricopeptide repeat proteins in land plants. Plant J. Cell Mol. Biol. 2016, 85, 532–547. [Google Scholar] [CrossRef] [PubMed]
- Fujii, S.; Bond, C.S.; Small, I.D. Selection patterns on restorer-like genes reveal a conflict between nuclear and mitochondrial genomes throughout angiosperm evolution. Proc. Natl. Acad. Sci. USA 2011, 108, 1723–1728. [Google Scholar] [CrossRef] [PubMed]
- Kaur, P.; Verma, M. Insights into PPR Gene Family in Cajanus cajan and other legume species. J. Data Min. Genom. Proteom. 2016, 7. [Google Scholar] [CrossRef]
- Dahan, J.; Mireau, H. The Rf and Rf-like PPR in higher plants, a fast-evolving subclass of PPR genes. RNA Biol. 2013, 10, 1469–1476. [Google Scholar] [CrossRef]
- Gaborieau, L.; Brown, G.G.; Mireau, H. The propensity of pentatricopeptide repeat genes to evolve into restorers of cytoplasmic male sterility. Front. Plant Sci. 2016, 7, 1816. [Google Scholar] [CrossRef]
- Rieseberg, L.H.; Blackman, B.K. Speciation genes in plants. Ann. Bot. 2010, 106, 439–455. [Google Scholar] [CrossRef]
- Zhigunov, A.V.; Ulianich, P.S.; Lebedeva, M.V.; Chang, P.L.; Nuzhdin, S.V.; Potokina, E.K. Development of F1 hybrid population and the high-density linkage map for European aspen (Populus tremula L.) using RADseq technology. BMC Plant Biol. 2017, 17, 180. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Bradbury, P.J.; Zhang, Z.; Kroon, D.E.; Casstevens, T.M.; Ramdoss, Y.; Buckler, E.S. TASSEL: Software for association mapping of complex traits in diverse samples. Bioinform. Oxf. Engl. 2007, 23, 2633–2635. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef] [PubMed]
- Badouin, H.; Gouzy, J.; Grassa, C.J.; Murat, F.; Staton, S.E.; Cottret, L.; Lelandais-Brière, C.; Owens, G.L.; Carrère, S.; Mayjonade, B.; et al. The sunflower genome provides insights into oil metabolism, flowering and Asterid evolution. Nature 2017, 546, 148–152. [Google Scholar] [CrossRef]
- Wickham, H. ggplot2; Springer: New York, NY, USA, 2009; ISBN 978-0-387-98140-6. [Google Scholar]
- Zhang, Z.; Ersoz, E.; Lai, C.-Q.; Todhunter, R.J.; Tiwari, H.K.; Gore, M.A.; Bradbury, P.J.; Yu, J.; Arnett, D.K.; Ordovas, J.M.; et al. Mixed linear model approach adapted for genome-wide association studies. Nat. Genet. 2010, 42, 355–360. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.C.; Chow, C.C.; Tellier, L.C.; Vattikuti, S.; Purcell, S.M.; Lee, J.J. Second-generation PLINK: Rising to the challenge of larger and richer datasets. GigaScience 2015, 4, 7. [Google Scholar] [CrossRef] [PubMed]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.R.; Bender, D.; Maller, J.; Sklar, P.; de Bakker, P.I.W.; Daly, M.J.; et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef]
- Turner, S.D. QQman: An R package for visualizing GWAS results using QQ and manhattan plots. BioRxiv 2014. [Google Scholar] [CrossRef]
- Tang, S.; Leon, A.; Bridges, W.C.; Knapp, S.J. Quantitative trait loci for genetically correlated seed traits are tightly linked to branching and pericarp pigment loci in sunflower. Crop Sci. 2006, 46, 721. [Google Scholar] [CrossRef]
- Nambeesan, S.U.; Mandel, J.R.; Bowers, J.E.; Marek, L.F.; Ebert, D.; Corbi, J.; Rieseberg, L.H.; Knapp, S.J.; Burke, J.M. Association mapping in sunflower (Helianthus annuus L.) reveals independent control of apical vs. basal branching. BMC Plant Biol. 2015, 15, 84. [Google Scholar] [CrossRef]
- Yu, J.-K.; Tang, S.; Slabaugh, M.B.; Heesacker, A.; Cole, G.; Herring, M.; Soper, J.; Han, F.; Chu, W.-C.; Webb, D.M.; et al. Towards a saturated molecular genetic linkage map for cultivated sunflower. Crop Sci. 2003, 43, 367. [Google Scholar] [CrossRef]
- Kusterer, B.; Rozynek, B.; Brahm, L.; Prüfe, M.; Tzigos, S.; Horn, R.; Friedt, W. Construction of a genetic map and localization of major traits in sunflower (Helianthus annuus L.). Helia 2004, 27, 15–23. [Google Scholar] [CrossRef]
- Hernandez Mora, J.R.; Rivals, E.; Mireau, H.; Budar, F. Sequence analysis of two alleles reveals that intra-and intergenic recombination played a role in the evolution of the radish fertility restorer (Rfo). BMC Plant Biol. 2010, 10, 35. [Google Scholar] [CrossRef] [PubMed]
- Kazama, T.; Toriyama, K. A fertility restorer gene, Rf4, widely used for hybrid rice breeding encodes a pentatricopeptide repeat protein. Rice 2014, 7, 28. [Google Scholar] [CrossRef] [PubMed]
- Madugula, P.; Uttam, A.G.; Tonapi, V.A.; Ragimasalawada, M. Fine mapping of Rf2, a major locus controlling pollen fertility restoration in sorghum A1 cytoplasm, encodes a PPR gene and its validation through expression analysis. Plant Breed. 2018, 137, 148–161. [Google Scholar] [CrossRef]
- Bentolila, S.; Alfonso, A.A.; Hanson, M.R. A pentatricopeptide repeat-containing gene restores fertility to cytoplasmic male-sterile plants. Proc. Natl. Acad. Sci. USA 2002, 99, 10887–10892. [Google Scholar] [CrossRef]
- Schmitz-Linneweber, C.; Small, I. Pentatricopeptide repeat proteins: A socket set for organelle gene expression. Trends Plant Sci. 2008, 13, 663–670. [Google Scholar] [CrossRef]
- Aubourg, S.; Boudet, N.; Kreis, M.; Lecharny, A. In Arabidopsis thaliana, 1% of the genome codes for a novel protein family unique to plants. Plant Mol. Biol. 2000, 42, 603–613. [Google Scholar] [CrossRef]
- Small, I.D.; Peeters, N. The PPR motif—A TPR-related motif prevalent in plant organellar proteins. Trends Biochem. Sci. 2000, 25, 46–47. [Google Scholar] [CrossRef]
- Wei, K.; Han, P. Pentatricopeptide repeat proteins in maize. Mol. Breed. 2016, 36. [Google Scholar] [CrossRef]
- Manna, S. An overview of pentatricopeptide repeat proteins and their applications. Biochimie 2015, 113, 93–99. [Google Scholar] [CrossRef] [PubMed]
- Small, I.D.; Rackham, O.; Filipovska, A. Organelle transcriptomes: Products of a deconstructed genome. Curr. Opin. Microbiol. 2013, 16, 652–658. [Google Scholar] [CrossRef] [PubMed]
- Barkan, A.; Small, I. Pentatricopeptide repeat proteins in plants. Annu. Rev. Plant Biol. 2014, 65, 415–442. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Wang, D.; Feng, J.; Seiler, G.J.; Cai, X.; Jan, C.-C. Diversifying sunflower germplasm by integration and mapping of a novel male fertility restoration gene. Genetics 2013, 193, 727–737. [Google Scholar] [CrossRef] [PubMed]

| Marker | Position | p-Value |
|---|---|---|
| S13_170494693 | 170494693 | 1.01 × 10−15 |
| S13_171053833 | 171053833 | 1.53 × 10−18 |
| S13_173268042 | 173268042 | 3.46 × 10−18 |
| S13_173832391 | 173832391 | 5.69 × 10−9 |
| S13_174474103 | 174474103 | 1.22 × 10−14 |
| S13_174474122 | 174474122 | 1.22 × 10−14 |
| S13_174809087 | 174809087 | 1.10 × 10−13 |
| S13_178217103 | 178217103 | 2.03 × 10−14 |
| Gene | Start | End | Strand | Product | Gene Bank Accession Number of Translated Protein | Hits for All PROSITE (Release 2018_11) Motifs |
|---|---|---|---|---|---|---|
| HanXRQChr13g0418841 | 170850155 | 170852002 | + | Putative pentatricopeptide repeat | OTG02960 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0418861 | 170908019 | 170909110 | + | Putative pentatricopeptide repeat | OTG02962 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0419621 | 173473487 | 173475525 | − | Probable pentatricopeptide repeat-containing protein At2g41080 | OTG03034 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0419631 | 173484455 | 173500401 | + | Putative pentatricopeptide repeat | OTG03035 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0419931 | 174209661 | 174217234 | − | Putative pentatricopeptide repeat | OTG03064 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0420121 | 174799667 | 174801481 | + | Probable pentatricopeptide repeat (PPR) superfamily protein | OTG03081 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0420241 | 174944047 | 174945506 | + | Putative pentatricopeptide repeat | OTG03093 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0420261 | 174962084 | 174962512 | + | Putative pentatricopeptide repeat | OTG03095 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0420351 | 175219425 | 175219886 | − | Putative pentatricopeptide repeat | OTG03099 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0420811 | 176970038 | 176972308 | + | Probable pentatricopeptide repeat (PPR) superfamily protein | OTG03141 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0421081 | 178216563 | 178219635 | − | Probable putative pentatricopeptide repeat-containing protein At4g17915 | OTG03166 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0418851 | 170877322 | 170879307 | + | Putative tetratricopeptide-like helical domain | OTG02961 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0419881 | 174159006 | 174160682 | − | Putative tetratricopeptide-like helical domain | OTG03060 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0420271 | 175002640 | 175003793 | + | Putative tetratricopeptide-like helical domain | OTG03096 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0420281 | 175016437 | 175018065 | + | Putative tetratricopeptide-like helical domain | OTG03097 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0420301 | 175055952 | 175057826 | + | Putative tetratricopeptide-like helical domain | OTG03098 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0420371 | 175253986 | 175294219 | − | Putative tetratricopeptide-like helical domain | OTG03101 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0420861 | 177597409 | 177599211 | + | Putative tetratricopeptide-like helical domain | OTG03145 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0420881 | 177609240 | 177611054 | + | Putative tetratricopeptide-like helical domain | OTG03147 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0421271 | 178655189 | 178657150 | − | Probable tetratricopeptide repeat (TPR)-like superfamily protein | OTG03183 | PS51375 Pentatricopeptide (PPR) repeat |
| HanXRQChr13g0419821 | 174082899 | 174091500 | − | Probable aldehyde dehydrogenase 5F1 | OTG03054 | NA |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Goryunov, D.V.; Anisimova, I.N.; Gavrilova, V.A.; Chernova, A.I.; Sotnikova, E.A.; Martynova, E.U.; Boldyrev, S.V.; Ayupova, A.F.; Gubaev, R.F.; Mazin, P.V.; et al. Association Mapping of Fertility Restorer Gene for CMS PET1 in Sunflower. Agronomy 2019, 9, 49. https://doi.org/10.3390/agronomy9020049
Goryunov DV, Anisimova IN, Gavrilova VA, Chernova AI, Sotnikova EA, Martynova EU, Boldyrev SV, Ayupova AF, Gubaev RF, Mazin PV, et al. Association Mapping of Fertility Restorer Gene for CMS PET1 in Sunflower. Agronomy. 2019; 9(2):49. https://doi.org/10.3390/agronomy9020049
Chicago/Turabian StyleGoryunov, Denis V., Irina N. Anisimova, Vera A. Gavrilova, Alina I. Chernova, Evgeniia A. Sotnikova, Elena U. Martynova, Stepan V. Boldyrev, Asiya F. Ayupova, Rim F. Gubaev, Pavel V. Mazin, and et al. 2019. "Association Mapping of Fertility Restorer Gene for CMS PET1 in Sunflower" Agronomy 9, no. 2: 49. https://doi.org/10.3390/agronomy9020049
APA StyleGoryunov, D. V., Anisimova, I. N., Gavrilova, V. A., Chernova, A. I., Sotnikova, E. A., Martynova, E. U., Boldyrev, S. V., Ayupova, A. F., Gubaev, R. F., Mazin, P. V., Gurchenko, E. A., Shumskiy, A. A., Petrova, D. A., Garkusha, S. V., Mukhina, Z. M., Benko, N. I., Demurin, Y. N., Khaitovich, P. E., & Goryunova, S. V. (2019). Association Mapping of Fertility Restorer Gene for CMS PET1 in Sunflower. Agronomy, 9(2), 49. https://doi.org/10.3390/agronomy9020049






