Potential Added Value of PET/CT Radiomics for Survival Prognostication beyond AJCC 8th Edition Staging in Oropharyngeal Squamous Cell Carcinoma
Abstract
1. Introduction
2. Results
2.1. Cohort Characteristics
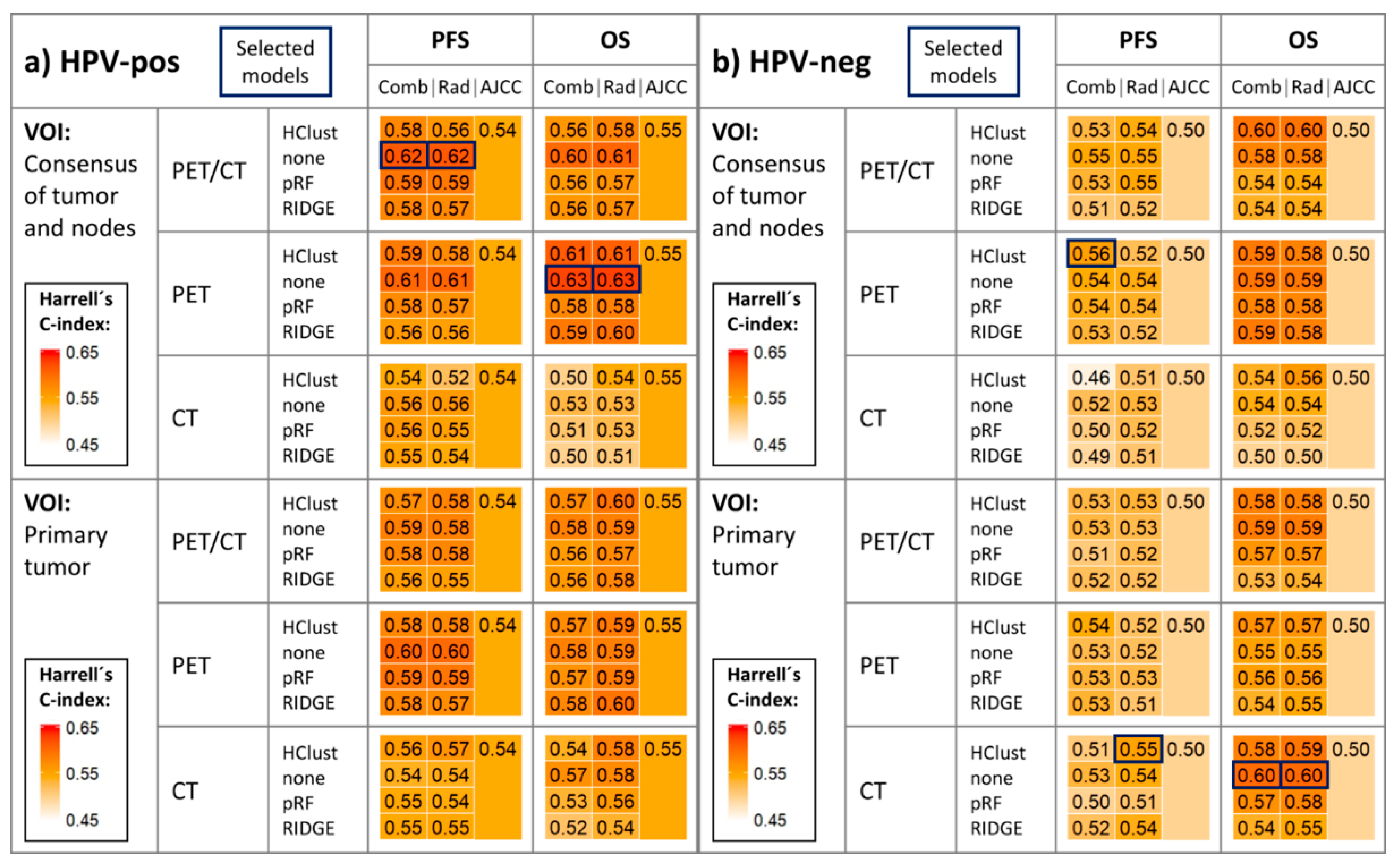
2.2. Survival Model Performance
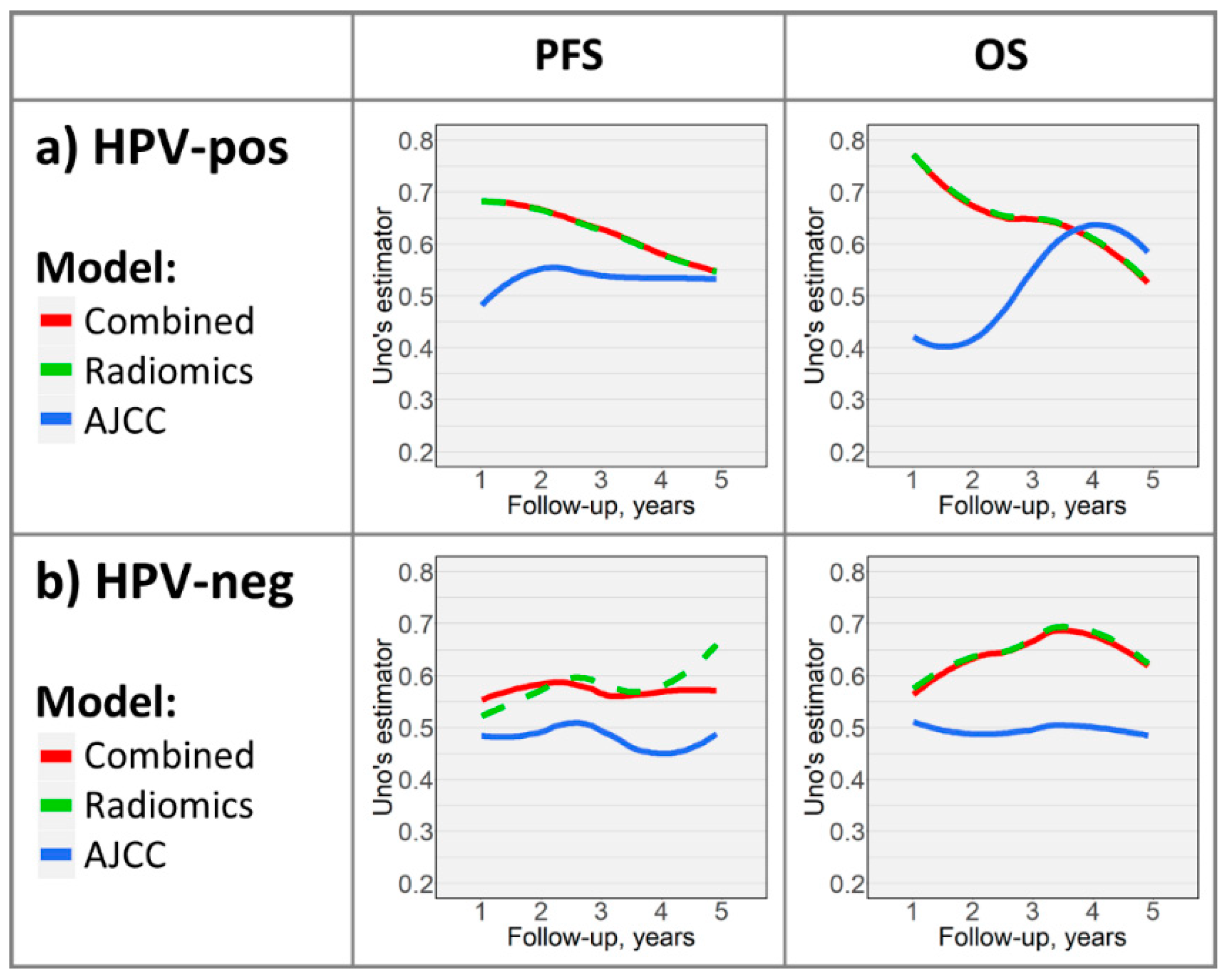
2.3. Time-Dependent Survival Model Evaluation
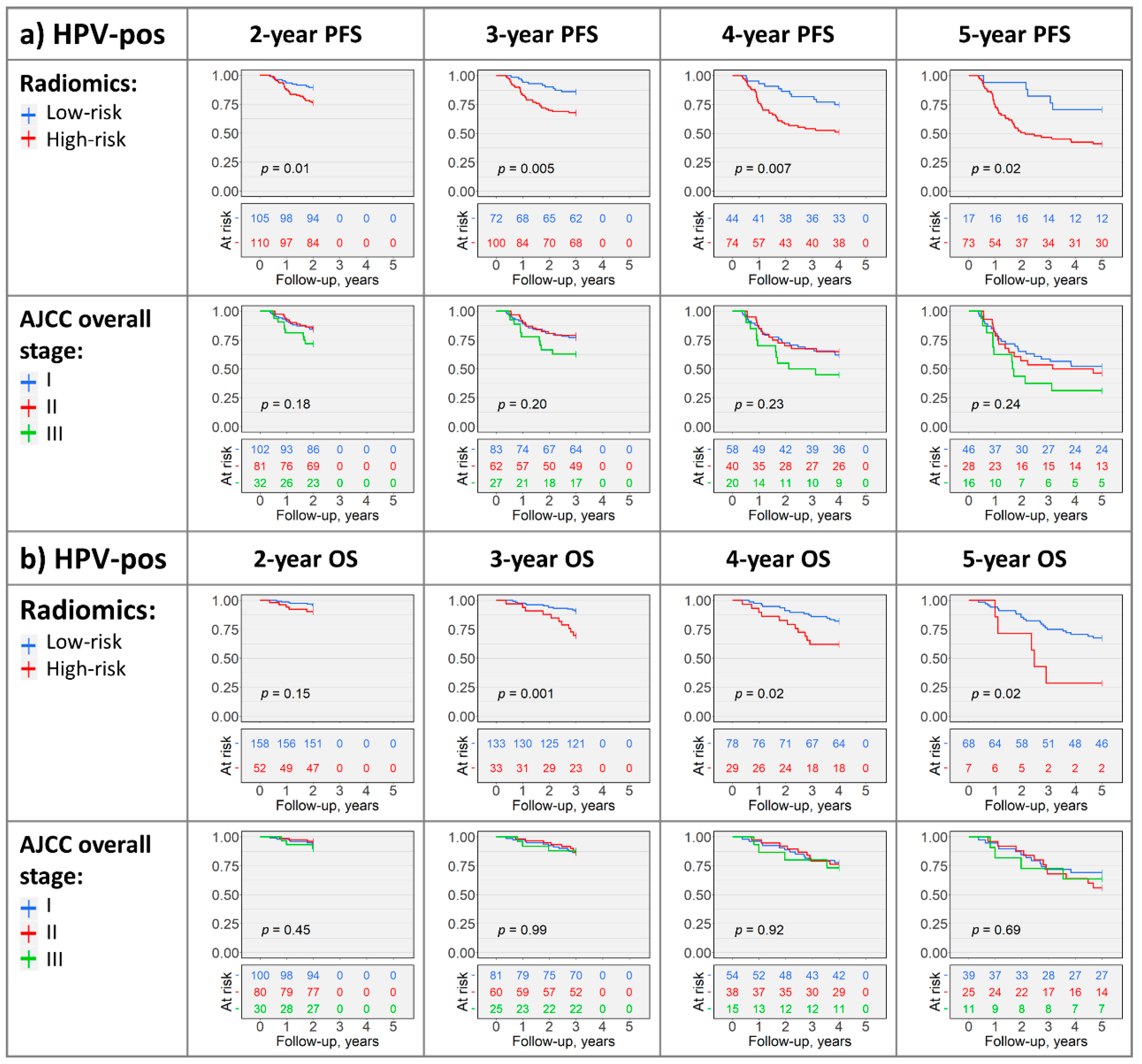
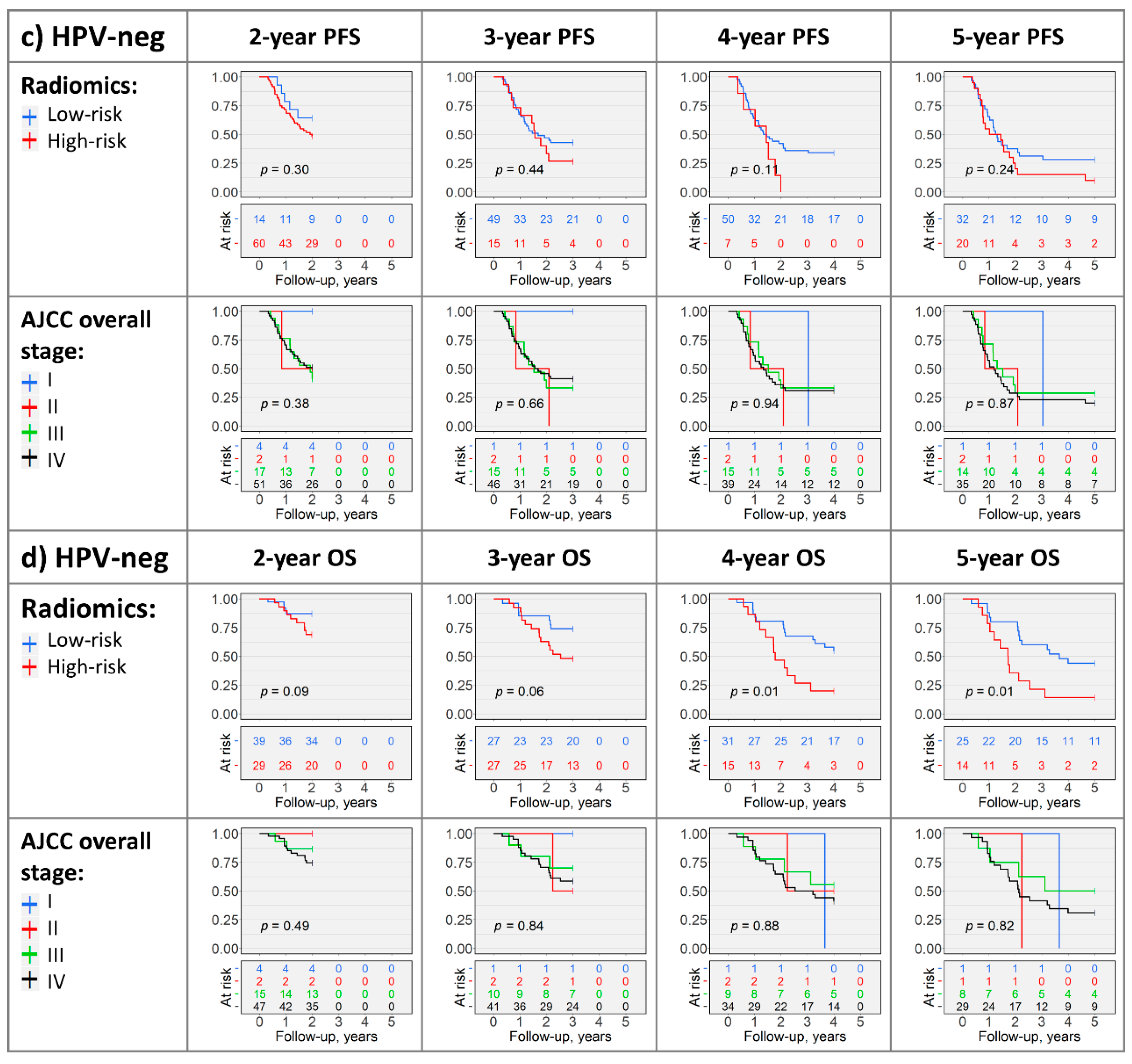
2.4. Kaplan–Meier Analysis
3. Discussion
4. Materials and Methods
4.1. Data Acquisition
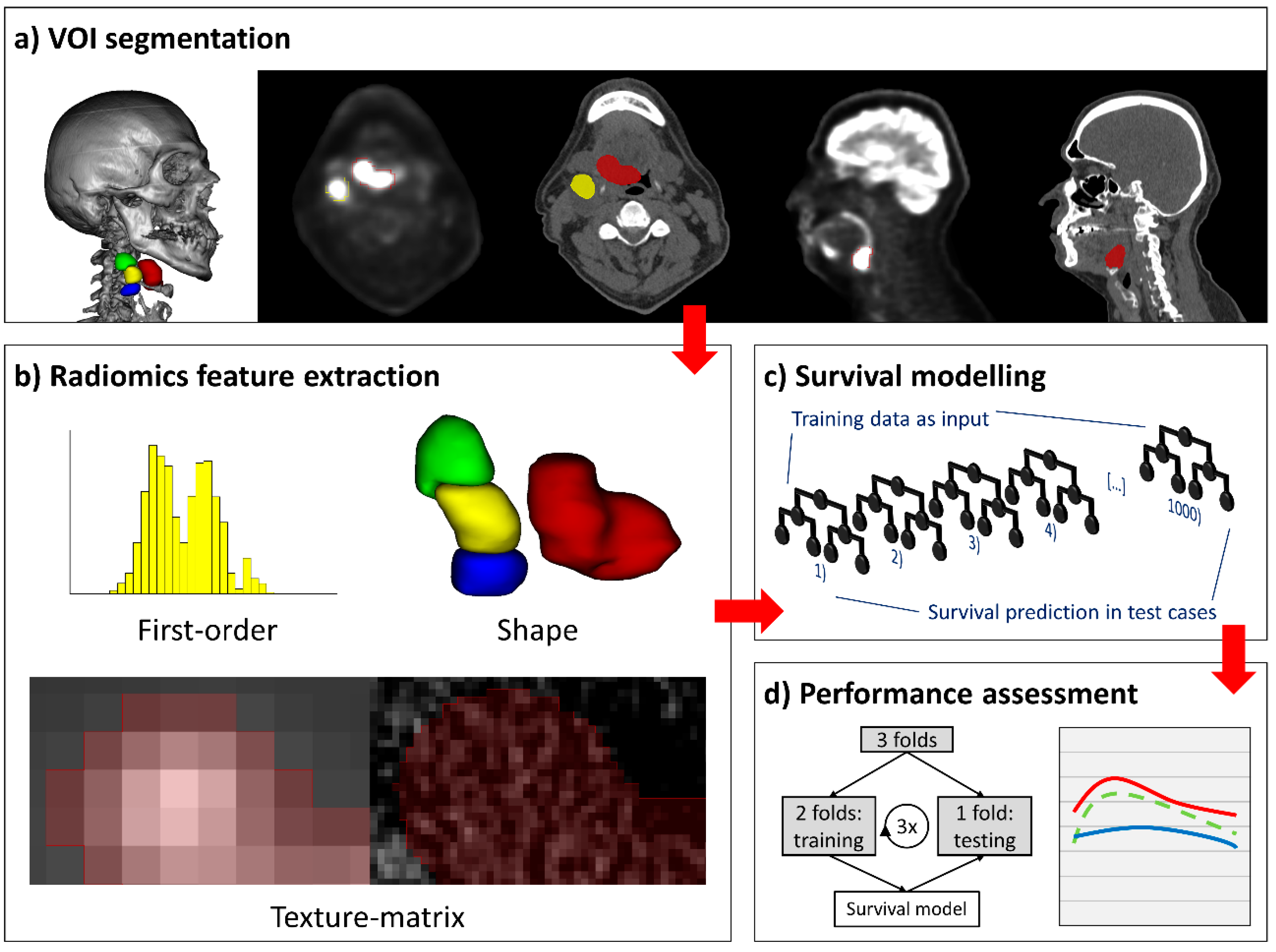
4.2. Lesion Segmentation
4.3. Radiomics Feature Extraction
4.4. Survival Study Arms and Cohorts
4.5. Survival Modelling
4.6. Cross-Validation and Performance Evaluation of Survival Models
4.7. Risk-Stratification and Kaplan–Meier Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| AJCC | American Joint Committee on Cancer |
| AUC | area under the receiver operating characteristic curve |
| CT | computed tomography |
| HPV | human papillomavirus |
| IQR | interquartile range |
| OPSCC | oropharyngeal squamous cell carcinoma |
| OS | overall survival |
| PET | [18F]fluorodeoxyglucose positron emission tomography |
| PFS | progression-free survival |
| RCF | random classification forest |
| RSF | random survival forest |
| SD | standard deviation |
| TCIA | The Cancer Imaging Archive |
| UICC | Union for International Cancer Control |
| VOI | volume of interest |
References
- Gillison, M.L.; Chaturvedi, A.K.; Anderson, W.F.; Fakhry, C. Epidemiology of Human Papillomavirus-Positive Head and Neck Squamous Cell Carcinoma. J. Clin. Oncol. 2015, 33, 3235–3242. [Google Scholar] [CrossRef] [PubMed]
- Gupta, B.; Johnson, N.W.; Kumar, N. Global Epidemiology of Head and Neck Cancers: A Continuing Challenge. Oncology 2016, 91, 13–23. [Google Scholar] [CrossRef] [PubMed]
- Mourad, M.; Jetmore, T.; Jategaonkar, A.A.; Moubayed, S.; Moshier, E.; Urken, M.L. Epidemiological Trends of Head and Neck Cancer in the United States: A SEER Population Study. J. Oral. Maxillofac. Surg. 2017, 75, 2562–2572. [Google Scholar] [CrossRef] [PubMed]
- Mehanna, H.; Beech, T.; Nicholson, T.; El-Hariry, I.; McConkey, C.; Paleri, V.; Roberts, S. Prevalence of human papillomavirus in oropharyngeal and nonoropharyngeal head and neck cancer--systematic review and meta-analysis of trends by time and region. Head Neck 2013, 35, 747–755. [Google Scholar] [CrossRef] [PubMed]
- Benson, E.; Li, R.; Eisele, D.; Fakhry, C. The clinical impact of HPV tumor status upon head and neck squamous cell carcinomas. Oral. Oncol. 2014, 50, 565–574. [Google Scholar] [CrossRef] [PubMed]
- Taberna, M.; Mena, M.; Pavon, M.A.; Alemany, L.; Gillison, M.L.; Mesia, R. Human papillomavirus-related oropharyngeal cancer. Ann. Oncol. 2017, 28, 2386–2398. [Google Scholar] [CrossRef]
- Ang, K.K.; Harris, J.; Wheeler, R.; Weber, R.; Rosenthal, D.I.; Nguyen-Tan, P.F.; Westra, W.H.; Chung, C.H.; Jordan, R.C.; Lu, C.; et al. Human papillomavirus and survival of patients with oropharyngeal cancer. N. Engl. J. Med. 2010, 363, 24–35. [Google Scholar] [CrossRef]
- Fakhry, C.; Westra, W.H.; Li, S.; Cmelak, A.; Ridge, J.A.; Pinto, H.; Forastiere, A.; Gillison, M.L. Improved survival of patients with human papillomavirus-positive head and neck squamous cell carcinoma in a prospective clinical trial. J. Natl. Cancer Inst. 2008, 100, 261–269. [Google Scholar] [CrossRef]
- AJCC Cancer Staging Manual, 8th ed.; Amin, M., Edge, S., Greene, F., Byrd, D., Brookland, R., Washington, M., Gershenwald, J., Compton, C., Hess, K.E.A., Eds.; Springer International Publishing: Cham, Switzerland, 2017. [Google Scholar]
- Union for International Cancer Control. TNM Classification of Malignant Tumours, 8th ed.; Brierley, J.D., Gospodarowicz, M.K., Wittekind, C., Eds.; Wiley-Blackwell: Hoboken, NJ, USA, 2016. [Google Scholar]
- Lydiatt, W.M.; Patel, S.G.; O’Sullivan, B.; Brandwein, M.S.; Ridge, J.A.; Migliacci, J.C.; Loomis, A.M.; Shah, J.P. Head and Neck cancers-major changes in the American Joint Committee on cancer eighth edition cancer staging manual. CA Cancer J. Clin. 2017, 67, 122–137. [Google Scholar] [CrossRef]
- O’Sullivan, B.; Huang, S.H.; Su, J.; Garden, A.S.; Sturgis, E.M.; Dahlstrom, K.; Lee, N.; Riaz, N.; Pei, X.; Koyfman, S.A.; et al. Development and validation of a staging system for HPV-related oropharyngeal cancer by the International Collaboration on Oropharyngeal cancer Network for Staging (ICON-S): A multicentre cohort study. Lancet Oncol. 2016, 17, 440–451. [Google Scholar] [CrossRef]
- Gillies, R.J.; Kinahan, P.E.; Hricak, H. Radiomics: Images Are More than Pictures, They Are Data. Radiology 2016, 278, 563–577. [Google Scholar] [CrossRef] [PubMed]
- Forghani, R.; Savadjiev, P.; Chatterjee, A.; Muthukrishnan, N.; Reinhold, C.; Forghani, B. Radiomics and Artificial Intelligence for Biomarker and Prediction Model Development in Oncology. Comput. Struct. Biotechnol. J. 2019, 17, 995–1008. [Google Scholar] [CrossRef] [PubMed]
- Guha, A.; Connor, S.; Anjari, M.; Naik, H.; Siddiqui, M.; Cook, G.; Goh, V. Radiomic analysis for response assessment in advanced head and neck cancers, a distant dream or an inevitable reality? A systematic review of the current level of evidence. Br. J. Radiol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Giraud, P.; Giraud, P.; Gasnier, A.; El Ayachy, R.; Kreps, S.; Foy, J.P.; Durdux, C.; Huguet, F.; Burgun, A.; Bibault, J.E. Radiomics and Machine Learning for Radiotherapy in Head and Neck Cancers. Front. Oncol. 2019, 9, 174. [Google Scholar] [CrossRef] [PubMed]
- Haider, S.P.; Burtness, B.; Yarbrough, W.G.; Payabvash, S. Applications of radiomics in precision diagnosis, prognostication and treatment planning of head and neck squamous cell carcinomas. Cancers Head Neck 2020, 5, 6. [Google Scholar] [CrossRef]
- Yip, S.S.; Aerts, H.J. Applications and limitations of radiomics. Phys. Med. Biol. 2016, 61, R150. [Google Scholar] [CrossRef]
- Haider, S.P.; Mahajan, A.; Zeevi, T.; Baumeister, P.; Reichel, C.; Sharaf, K.; Forghani, R.; Kucukkaya, A.S.; Kann, B.H.; Judson, B.L.; et al. PET/CT radiomics signature of human papilloma virus association in oropharyngeal squamous cell carcinoma. Eur. J. Nucl. Med. Mol. Imaging 2020. [Google Scholar] [CrossRef]
- Chiesa-Estomba, C.M.; Echaniz, O.; Larruscain, E.; Gonzalez-Garcia, J.A.; Sistiaga-Suarez, J.A.; Grana, M. Radiomics and Texture Analysis in Laryngeal Cancer. Looking for New Frontiers in Precision Medicine through Imaging Analysis. Cancers (Basel) 2019, 11, 1409. [Google Scholar] [CrossRef]
- Wong, A.J.; Kanwar, A.; Mohamed, A.S.; Fuller, C.D. Radiomics in head and neck cancer: From exploration to application. Transl. Cancer Res. 2016, 5, 371–382. [Google Scholar] [CrossRef]
- Cheng, N.M.; Fang, Y.H.; Chang, J.T.; Huang, C.G.; Tsan, D.L.; Ng, S.H.; Wang, H.M.; Lin, C.Y.; Liao, C.T.; Yen, T.C. Textural features of pretreatment 18F-FDG PET/CT images: Prognostic significance in patients with advanced T-stage oropharyngeal squamous cell carcinoma. J. Nucl. Med. 2013, 54, 1703–1709. [Google Scholar] [CrossRef]
- Cheng, N.M.; Fang, Y.H.; Lee, L.Y.; Chang, J.T.; Tsan, D.L.; Ng, S.H.; Wang, H.M.; Liao, C.T.; Yang, L.Y.; Hsu, C.H.; et al. Zone-size nonuniformity of 18F-FDG PET regional textural features predicts survival in patients with oropharyngeal cancer. Eur. J. Nucl. Med. Mol. Imaging 2015, 42, 419–428. [Google Scholar] [CrossRef] [PubMed]
- Folkert, M.R.; Setton, J.; Apte, A.P.; Grkovski, M.; Young, R.J.; Schoder, H.; Thorstad, W.L.; Lee, N.Y.; Deasy, J.O.; Oh, J.H. Predictive modeling of outcomes following definitive chemoradiotherapy for oropharyngeal cancer based on FDG-PET image characteristics. Phys. Med. Biol. 2017, 62, 5327–5343. [Google Scholar] [CrossRef] [PubMed]
- Ger, R.B.; Zhou, S.; Elgohari, B.; Elhalawani, H.; Mackin, D.M.; Meier, J.G.; Nguyen, C.M.; Anderson, B.M.; Gay, C.; Ning, J.; et al. Radiomics features of the primary tumor fail to improve prediction of overall survival in large cohorts of CT- and PET-imaged head and neck cancer patients. PLoS ONE 2019, 14, e0222509. [Google Scholar] [CrossRef] [PubMed]
- Leijenaar, R.T.; Carvalho, S.; Hoebers, F.J.; Aerts, H.J.; van Elmpt, W.J.; Huang, S.H.; Chan, B.; Waldron, J.N.; O’Sullivan, B.; Lambin, P. External validation of a prognostic CT-based radiomic signature in oropharyngeal squamous cell carcinoma. Acta Oncol. 2015, 54, 1423–1429. [Google Scholar] [CrossRef] [PubMed]
- M. D. Anderson Cancer Center Head and Neck Quantitative Imaging Working Group. Investigation of radiomic signatures for local recurrence using primary tumor texture analysis in oropharyngeal head and neck cancer patients. Sci. Rep. 2018, 8, 1524. [Google Scholar] [CrossRef]
- Zdilar, L.; Vock, D.M.; Marai, G.E.; Fuller, C.D.; Mohamed, A.S.R.; Elhalawani, H.; Elgohari, B.A.; Tiras, C.; Miller, A.; Canahuate, G. Evaluating the Effect of Right-Censored End Point Transformation for Radiomic Feature Selection of Data From Patients With Oropharyngeal Cancer. JCO Clin. Cancer Inform. 2018, 2, 1–19. [Google Scholar] [CrossRef]
- Mascitti, M.; Tempesta, A.; Togni, L.; Capodiferro, S.; Troiano, G.; Rubini, C.; Maiorano, E.; Santarelli, A.; Favia, G.; Limongelli, L. Histological Features and Survival in Young Patients with HPV Negative Oral Squamous Cell Carcinoma. Oral. Dis. 2020. [Google Scholar] [CrossRef]
- Ishwaran, H.; Kogalur, U.B.; Blackstone, E.H.; Lauer, M.S. Random survival forests. Ann. Appl. Stat. 2008, 2, 841–860. [Google Scholar] [CrossRef]
- Schmid, M.; Wright, M.; Ziegler, A. On the use of Harrell’s C for clinical risk prediction via random survival forests. Expert Syst. Appl. 2016, 63, 450–459. [Google Scholar] [CrossRef]
- Wright, M.N.; Ziegler, A. ranger: A Fast Implementation of Random Forests for High Dimensional Data in C plus plus and R. J. Stat. Softw. 2017, 77, 1–17. [Google Scholar] [CrossRef]
- Beesley, L.J.; Hawkins, P.G.; Amlani, L.M.; Bellile, E.L.; Casper, K.A.; Chinn, S.B.; Eisbruch, A.; Mierzwa, M.L.; Spector, M.E.; Wolf, G.T.; et al. Individualized survival prediction for patients with oropharyngeal cancer in the human papillomavirus era. Cancer 2019, 125, 68–78. [Google Scholar] [CrossRef] [PubMed]
- Nauta, I.H.; Rietbergen, M.M.; van Bokhoven, A.; Bloemena, E.; Lissenberg-Witte, B.I.; Heideman, D.A.M.; Baatenburg de Jong, R.J.; Brakenhoff, R.H.; Leemans, C.R. Evaluation of the eighth TNM classification on p16-positive oropharyngeal squamous cell carcinomas in the Netherlands and the importance of additional HPV DNA testing. Ann. Oncol. 2018, 29, 1273–1279. [Google Scholar] [CrossRef] [PubMed]
- Deschuymer, S.; Dok, R.; Laenen, A.; Hauben, E.; Nuyts, S. Patient Selection in Human Papillomavirus Related Oropharyngeal Cancer: The Added Value of Prognostic Models in the New TNM 8th Edition Era. Front. Oncol. 2018, 8, 273. [Google Scholar] [CrossRef] [PubMed]
- Clark, K.; Vendt, B.; Smith, K.; Freymann, J.; Kirby, J.; Koppel, P.; Moore, S.; Phillips, S.; Maffitt, D.; Pringle, M.; et al. The Cancer Imaging Archive (TCIA): Maintaining and operating a public information repository. J. Digit. Imaging 2013, 26, 1045–1057. [Google Scholar] [CrossRef]
- Vallières, M.; Kay-Rivest, E.; Perrin, L.J.; Liem, X.; Furstoss, C.; Khaouam, N.; Nguyen-Tan, P.F.; Wang, C.; Sultanem, K. Data from Head-Neck-PET-CT. Cancer Imaging Arch. 2017. [Google Scholar] [CrossRef]
- Vallieres, M.; Kay-Rivest, E.; Perrin, L.J.; Liem, X.; Furstoss, C.; Aerts, H.; Khaouam, N.; Nguyen-Tan, P.F.; Wang, C.S.; Sultanem, K.; et al. Radiomics strategies for risk assessment of tumour failure in head-and-neck cancer. Sci. Rep. 2017, 7, 10117. [Google Scholar] [CrossRef]
- Grossberg, A.; Mohamed, A.; Elhalawani, H.; Bennett, W.; Smith, K.; Nolan, T.; Chamchod, S.; Kanto, r.M.; Browne, T.; Hutcheson, K.; et al. Data from Head and Neck Cancer CT Atlas. Cancer Imaging Arch. 2017. [Google Scholar] [CrossRef]
- Grossberg, A.J.; Mohamed, A.S.R.; Elhalawani, H.; Bennett, W.C.; Smith, K.E.; Nolan, T.S.; Williams, B.; Chamchod, S.; Heukelom, J.; Kantor, M.E.; et al. Imaging and clinical data archive for head and neck squamous cell carcinoma patients treated with radiotherapy. Sci. Data 2018, 5, 180173. [Google Scholar] [CrossRef]
- Ger, R.B.; Craft, D.F.; Mackin, D.S.; Zhou, S.; Layman, R.R.; Jones, A.K.; Elhalawani, H.; Fuller, C.D.; Howell, R.M.; Li, H.; et al. Practical guidelines for handling head and neck computed tomography artifacts for quantitative image analysis. Comput. Med. Imaging Graph. 2018, 69, 134–139. [Google Scholar] [CrossRef]
- Lewis, J.S., Jr.; Beadle, B.; Bishop, J.A.; Chernock, R.D.; Colasacco, C.; Lacchetti, C.; Moncur, J.T.; Rocco, J.W.; Schwartz, M.R.; Seethala, R.R.; et al. Human Papillomavirus Testing in Head and Neck Carcinomas: Guideline From the College of American Pathologists. Arch. Pathol. Lab. Med. 2018, 142, 559–597. [Google Scholar] [CrossRef]
- Kikinis, R.; Pieper, S.D.; Vosburgh, K.G. 3D Slicer: A Platform for Subject-Specific Image Analysis, Visualization, and Clinical Support. In Intraoperative Imaging and Image-Guided Therapy; Jolesz, F.A., Ed.; Springer: New York, NY, USA, 2014; pp. 277–289. [Google Scholar]
- Fedorov, A.; Beichel, R.; Kalpathy-Cramer, J.; Finet, J.; Fillion-Robin, J.C.; Pujol, S.; Bauer, C.; Jennings, D.; Fennessy, F.; Sonka, M.; et al. 3D Slicer as an image computing platform for the Quantitative Imaging Network. Magn. Reson. Imaging 2012, 30, 1323–1341. [Google Scholar] [CrossRef] [PubMed]
- Van Griethuysen, J.J.M.; Fedorov, A.; Parmar, C.; Hosny, A.; Aucoin, N.; Narayan, V.; Beets-Tan, R.G.H.; Fillion-Robin, J.C.; Pieper, S.; Aerts, H. Computational Radiomics System to Decode the Radiographic Phenotype. Cancer Res. 2017, 77, e104–e107. [Google Scholar] [CrossRef] [PubMed]
- Pyradiomics-community. Pyradiomics Documentation Release 2.1.2. Available online: https://readthedocs.org/projects/pyradiomics/downloads/pdf/2.1.2/ (accessed on 15 December 2019).
- Traverso, A.; Wee, L.; Dekker, A.; Gillies, R. Repeatability and Reproducibility of Radiomic Features: A Systematic Review. Int. J. Radiat. Oncol. Biol. Phys. 2018, 102, 1143–1158. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.; Lv, W.; Jiang, J.; Ma, J.; Feng, Q.; Rahmim, A.; Chen, W. Robustness of Radiomic Features in [(11)C]Choline and [(18)F]FDG PET/CT Imaging of Nasopharyngeal Carcinoma: Impact of Segmentation and Discretization. Mol. Imaging Biol. 2016, 18, 935–945. [Google Scholar] [CrossRef]
- Leijenaar, R.T.; Carvalho, S.; Velazquez, E.R.; van Elmpt, W.J.; Parmar, C.; Hoekstra, O.S.; Hoekstra, C.J.; Boellaard, R.; Dekker, A.L.; Gillies, R.J.; et al. Stability of FDG-PET Radiomics features: An integrated analysis of test-retest and inter-observer variability. Acta Oncol. 2013, 52, 1391–1397. [Google Scholar] [CrossRef] [PubMed]
- Doumou, G.; Siddique, M.; Tsoumpas, C.; Goh, V.; Cook, G.J. The precision of textural analysis in (18)F-FDG-PET scans of oesophageal cancer. Eur. Radiol. 2015, 25, 2805–2812. [Google Scholar] [CrossRef]
- Aerts, H.J.; Velazquez, E.R.; Leijenaar, R.T.; Parmar, C.; Grossmann, P.; Carvalho, S.; Bussink, J.; Monshouwer, R.; Haibe-Kains, B.; Rietveld, D.; et al. Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nat. Commun. 2014, 5, 4006. [Google Scholar] [CrossRef]
- Kalpathy-Cramer, J.; Mamomov, A.; Zhao, B.; Lu, L.; Cherezov, D.; Napel, S.; Echegaray, S.; Rubin, D.; McNitt-Gray, M.; Lo, P.; et al. Radiomics of Lung Nodules: A Multi-Institutional Study of Robustness and Agreement of Quantitative Imaging Features. Tomography 2016, 2, 430–437. [Google Scholar] [CrossRef] [PubMed]
- Yu, K.; Zhang, Y.; Yu, Y.; Huang, C.; Liu, R.; Li, T.; Yang, L.; Morris, J.S.; Baladandayuthapani, V.; Zhu, H. Radiomic analysis in prediction of Human Papilloma Virus status. Clin. Transl. Radiat. Oncol. 2017, 7, 49–54. [Google Scholar] [CrossRef]
- R Development Core Team. R: A language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2019. [Google Scholar]
- Harrell, F.E., Jr.; Califf, R.M.; Pryor, D.B.; Lee, K.L.; Rosati, R.A. Evaluating the yield of medical tests. JAMA 1982, 247, 2543–2546. [Google Scholar] [CrossRef]
- Brentnall, A.R.; Cuzick, J. Use of the concordance index for predictors of censored survival data. Stat. Methods Med. Res. 2018, 27, 2359–2373. [Google Scholar] [CrossRef] [PubMed]
- Bouckaert, R.R.; Frank, E. Evaluating the Replicability of Significance Tests for Comparing Learning Algorithms; Springer: Berlin/Heidelberg, Germany, 2004; pp. 3–12. [Google Scholar]
- Nadeau, C.; Bengio, Y. Inference for the Generalization Error. Mach. Learn. 2003, 52, 239–281. [Google Scholar] [CrossRef]
- Uno, H.; Cai, T.X.; Tian, L.; Wei, L.J. Evaluating prediction rules for t-year survivors with censored regression models. J. Am. Stat. Assoc. 2007, 102, 527–537. [Google Scholar] [CrossRef]
- Schmid, M.; Kestler, H.A.; Potapov, S. On the validity of time-dependent AUC estimators. Brief Bioinform. 2015, 16, 153–168. [Google Scholar] [CrossRef] [PubMed]
- Harrell, F.E.J.; Dupont, C. Hmisc: Harrell Miscellaneous, Version 4.3-0; 2019. Available online: https://cran.r-project.org/web/packages/Hmisc/index.html (accessed on 2 July 2020).
- Potapov, S.; Adler, W.; Schmid, M. survAUC: Estimators of Prediction Accuracy for Time-to-Event Data, Version 1.0-5; 2012. Available online: http://cran.rproject.org/web/packages/survAUC/index.html (accessed on 2 July 2020).
- Wickham, H. ggplot2. In Elegant Graphics for Data Analysis, 1st ed.; Springer: New York, NY, USA, 2009. [Google Scholar] [CrossRef]
- Cleveland, W.S.; Grosse, E.; Shyu, W.M. Local regression models. In Statistical Models in S; Chambers, J.M., Hastie, T.J., Eds.; The Wadsworth & Brooks/Cole Mathematics; Springer: Cham, Switzerland, 1992. [Google Scholar]
| Survival Endpoint | Progression-Free Survival | Overall Survival |
|---|---|---|
| Number of patients1—n | 311 | 306 |
| Included lymph nodes—n | 475 | 462 |
| Events—n (%) | 94 (30.2%) | 58 (19.0%) |
| Follow-up (days)—median (IQR) | 1170 (798–1645) | 1197 (818–1656) |
| Data source—n (%) | ||
| Yale | 201 (64.6%) | 200 (65.4%) |
| TCIA | 110 (35.4%) | 106 (34.6%) |
| Sex—n (%) | ||
| Male | 253 (81.4%) | 249 (81.4%) |
| Female | 58 (18.6%) | 57 (18.6%) |
| Age (years)—mean (SD) | 60.61 (9.24) | 60.60 (9.28) |
| HPV status2—n (%) | ||
| Positive | 235 (75.6%) | 233 (76.1%) |
| Negative | 76 (24.4%) | 73 (23.9%) |
| Smoking—n (%) | ||
| Never-smoker | 76 (24.4%) | 76 (24.8%) |
| Smoker | 143 (46.0%) | 142 (46.4%) |
| Pack-years—median (IQR) | 20 (10–40) | 20 (10–40) |
| Pack-years unknown—n | 20 | 20 |
| Unknown | 92 (29.6%) | 88 (28.8 %) |
| T stage3—n (%) | ||
| T1 | 43 (13.8%) | 42 (13.7%) |
| T2 | 120 (38.6%) | 120 (39.2%) |
| T3 | 99 (31.8%) | 97 (31.7%) |
| T4 | 49 (15.8%) | 47 (15.4%) |
| N stage3—n (%) | ||
| N0 | 60 (19.3%) | 59 (19.3%) |
| N1 | 149 (47.9%) | 149 (48.7%) |
| N2 | 97 (31.2%) | 94 (30.7%) |
| N3 | 5 (1.6%) | 4 (1.3 %) |
| Overall stage3—n (%) | ||
| I | 117 (37.6%) | 117 (38.2%) |
| II | 91 (29.3%) | 91 (29.7%) |
| III | 50 (16.1%) | 47 (15.4%) |
| IV | 53 (17.0%) | 51 (16.7%) |
| Included lymph nodes/patient—range | 0–8 | 0–8 |
| Primary treatment—n (%) | ||
| CCRT or CBRT | 208 (66.9%) | 204 (66.7%) |
| RT alone | 28 (9.0%) | 27 (8.8%) |
| Surgery | ||
| Without adjuvant therapy | 13 (4.2%) | 13 (4.2%) |
| With adjuvant RT, CCRT, or CBRT | 62 (19.9%) | 62 (20.3%) |
| PET4—mean (SD) | ||
| Slice thickness (mm) | 3.40 (0.38) | 3.39 (0.38) |
| In-plane pixel spacing (mm) | 4.30 (0.91) | 4.30 (0.92) |
| In-plane image matrix (n × n) | 147.16 (58.88) × idem | 147.32 (59.34) × idem |
| CT4—mean (SD) | ||
| Slice thickness (mm) | 3.12 (0.55) | 3.10 (0.54) |
| In-plane pixel spacing (mm) | 1.12 (0.18) | 1.12 (0.18) |
| In-plane image matrix (n × n) | 512 × 512 | 512 × 512 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Haider, S.P.; Zeevi, T.; Baumeister, P.; Reichel, C.; Sharaf, K.; Forghani, R.; Kann, B.H.; Judson, B.L.; Prasad, M.L.; Burtness, B.; et al. Potential Added Value of PET/CT Radiomics for Survival Prognostication beyond AJCC 8th Edition Staging in Oropharyngeal Squamous Cell Carcinoma. Cancers 2020, 12, 1778. https://doi.org/10.3390/cancers12071778
Haider SP, Zeevi T, Baumeister P, Reichel C, Sharaf K, Forghani R, Kann BH, Judson BL, Prasad ML, Burtness B, et al. Potential Added Value of PET/CT Radiomics for Survival Prognostication beyond AJCC 8th Edition Staging in Oropharyngeal Squamous Cell Carcinoma. Cancers. 2020; 12(7):1778. https://doi.org/10.3390/cancers12071778
Chicago/Turabian StyleHaider, Stefan P., Tal Zeevi, Philipp Baumeister, Christoph Reichel, Kariem Sharaf, Reza Forghani, Benjamin H. Kann, Benjamin L. Judson, Manju L. Prasad, Barbara Burtness, and et al. 2020. "Potential Added Value of PET/CT Radiomics for Survival Prognostication beyond AJCC 8th Edition Staging in Oropharyngeal Squamous Cell Carcinoma" Cancers 12, no. 7: 1778. https://doi.org/10.3390/cancers12071778
APA StyleHaider, S. P., Zeevi, T., Baumeister, P., Reichel, C., Sharaf, K., Forghani, R., Kann, B. H., Judson, B. L., Prasad, M. L., Burtness, B., Mahajan, A., & Payabvash, S. (2020). Potential Added Value of PET/CT Radiomics for Survival Prognostication beyond AJCC 8th Edition Staging in Oropharyngeal Squamous Cell Carcinoma. Cancers, 12(7), 1778. https://doi.org/10.3390/cancers12071778






