Transcriptional Modulation by Idelalisib Synergizes with Bendamustine in Chronic Lymphocytic Leukemia
Abstract
1. Introduction
2. Results
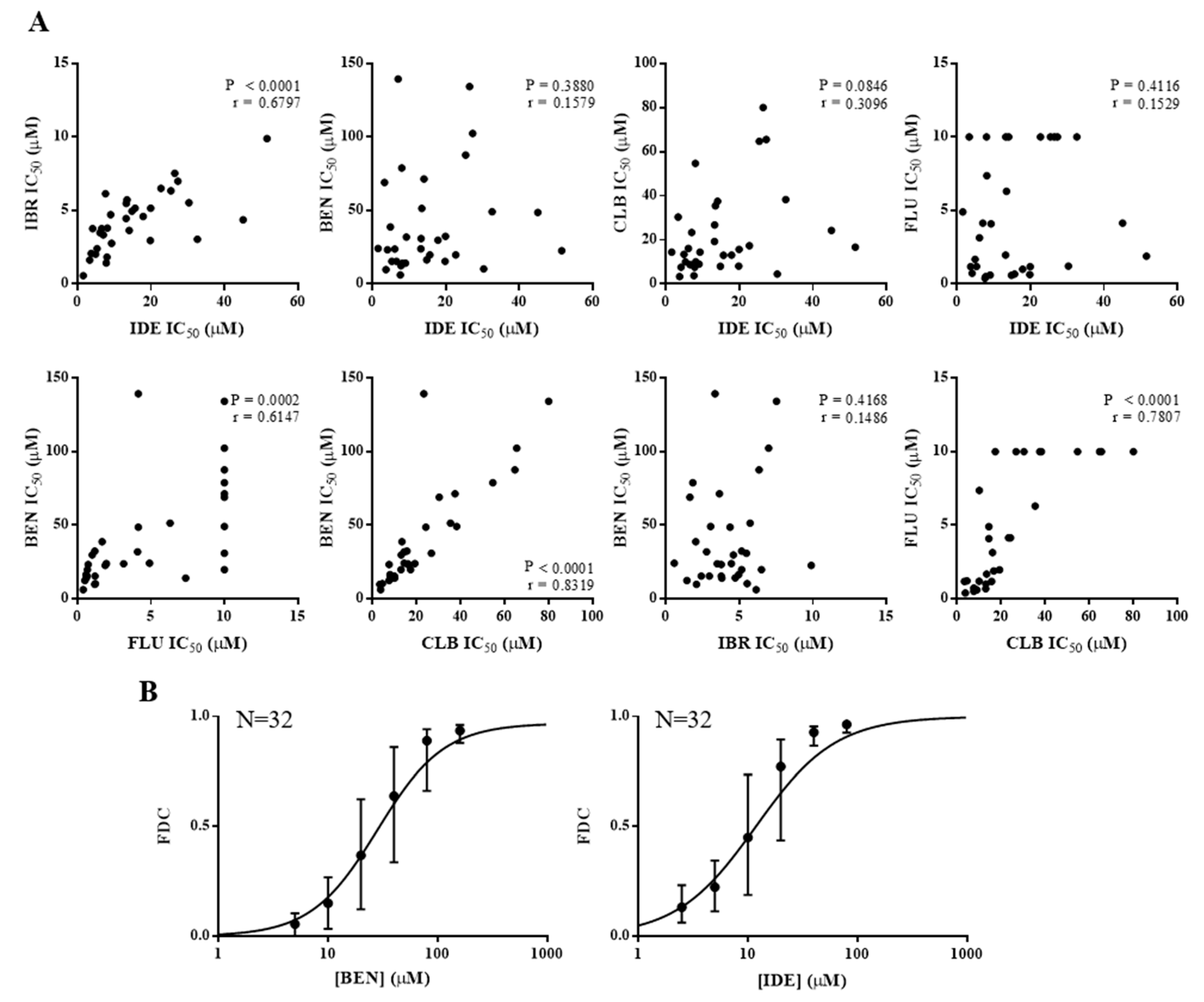
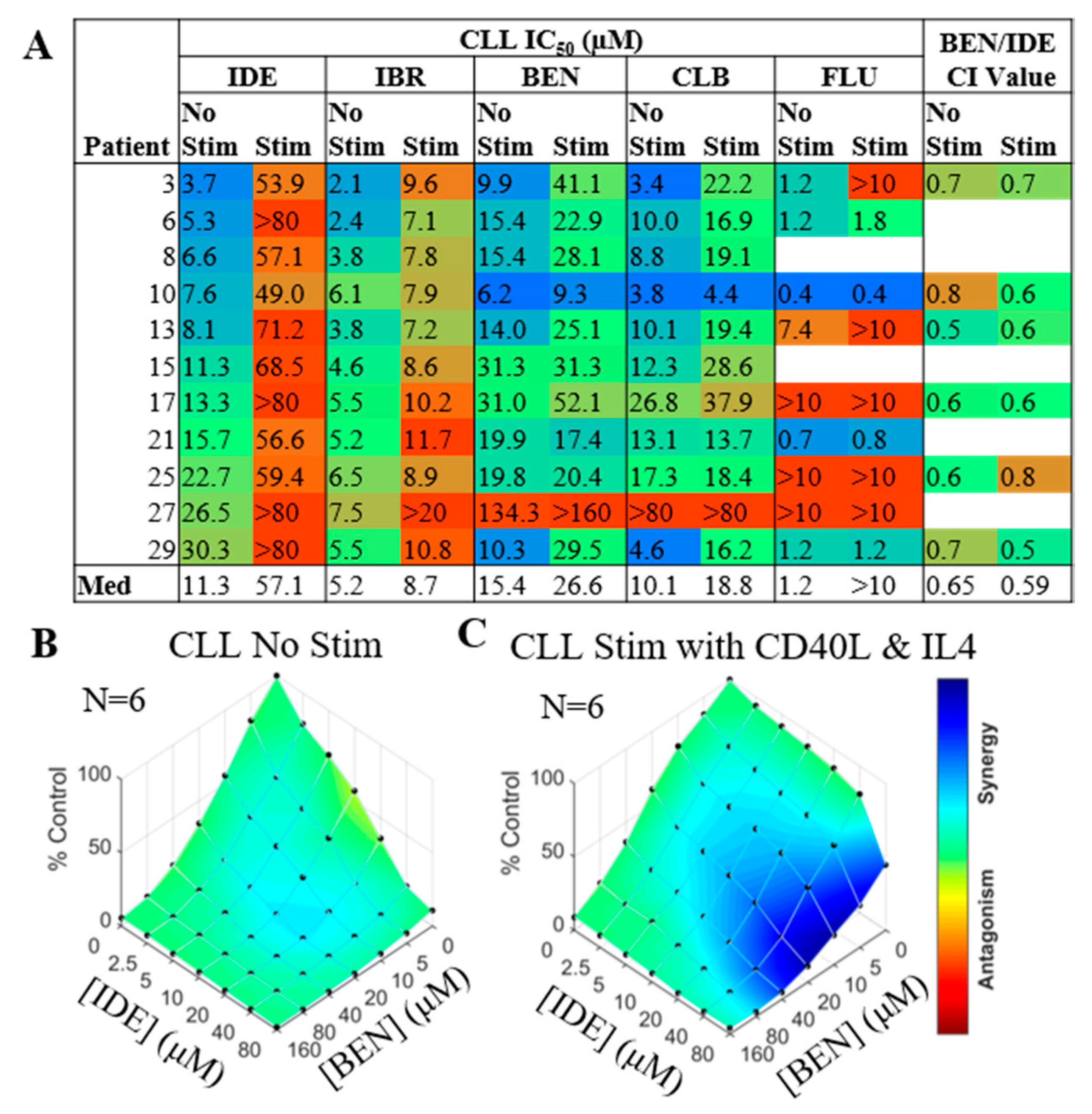
2.1. IDE Is Cross-Resistant with IBR but Not Chemotherapeutic Agents and Displays Synergy with Chemotherapeutics, Incubated Alone or with CD40/IL4
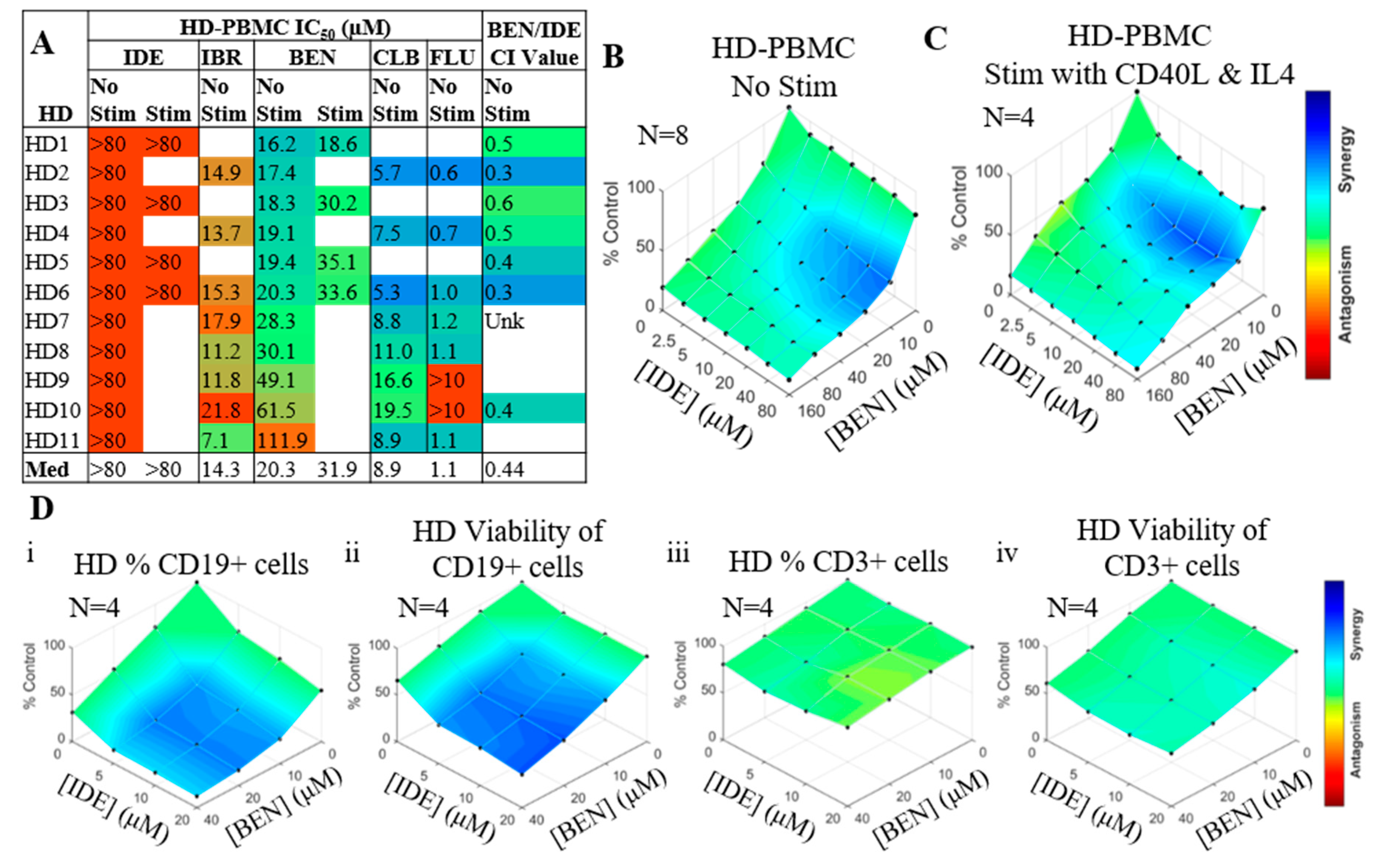
2.2. IDE and BEN Are Also Synergistic in Normal B Cells but Not T Cells
2.3. B Cells from Mice Lacking Functional PI3Kδ Display Reduced IDE Sensitivity and Synergy with BEN/IDE
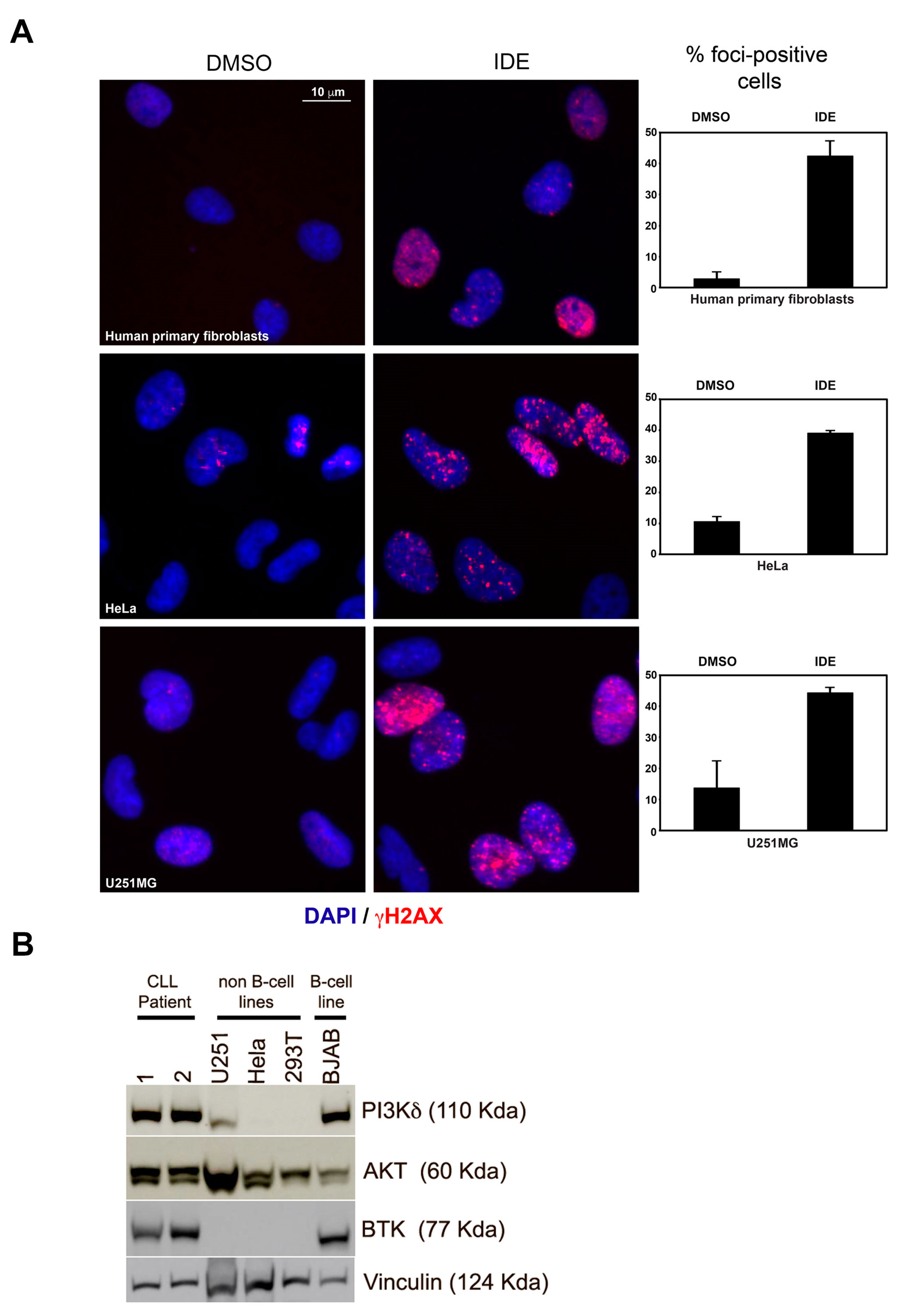
2.4. IDE Alone Induces γH2AX Formation but Not DNA Breaks, and Does Not Inhibit DNA Damage Repair in CLL Cells
2.5. IDE Induces γH2AX Formation Independent of PI3Kδ
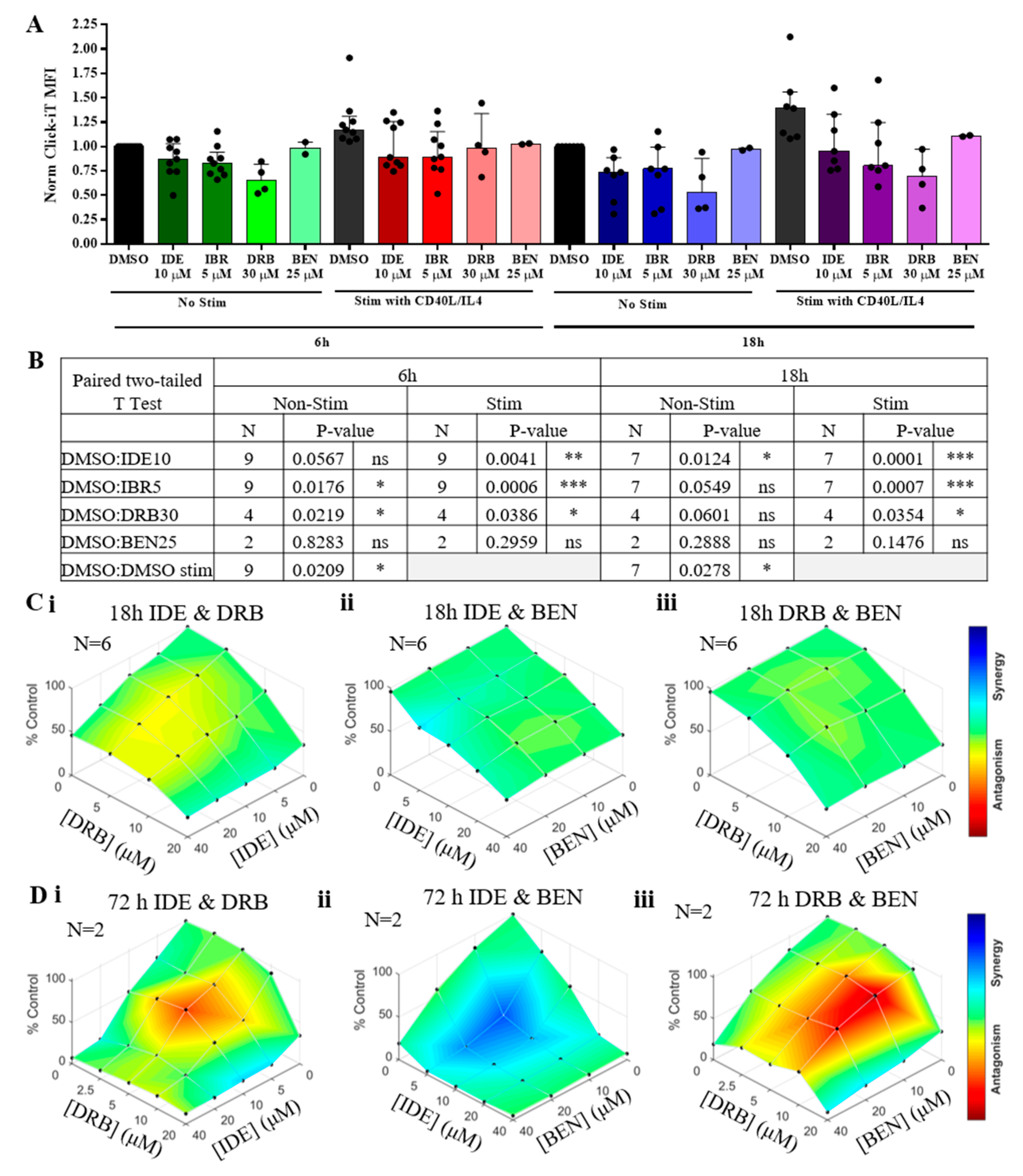
2.6. Synergy between BEN and IDE is Related to Changes in Transcription by IDE
3. Discussion
4. Materials and Methods
4.1. Patient Samples, Cell Lines, Culture Conditions
4.2. Drugs
4.3. Flow Cytometry and Drug Synergy
4.4. Animals
4.5. DNA Damage Analysis
4.6. Comet Assay
4.7. IR Recovery, Dose Response, and Synergy
4.8. Western Blots
4.9. Transcriptional Analysis
4.10. IGHV Mutational Analysis
4.11. FISH Analysis
4.12. Statistical and Synergy Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hallek, M.; Shanafelt, T.D.; Eichhorst, B. Chronic lymphocytic leukaemia. Lancet 2018, 391, 1524–1537. [Google Scholar] [CrossRef]
- Kost, S.E.F.; Bouchard, E.D.J.; Labossière, É.; Ye, X.; Queau, M.L.; Liang, W.S.; Banerji, V.; Gibson, S.B.; Katyal, S.; Johnston, J.B. Cross-resistance and synergy with bendamustine in chronic lymphocytic leukemia. Leuk. Res. 2016, 50, 63–71. [Google Scholar] [CrossRef] [PubMed]
- Leoni, L.M.; Bailey, B.; Reifert, J.; Bendall, H.H.; Zeller, R.W.; Corbeil, J.; Elliott, G.; Niemeyer, C.C. Bendamustine (Treanda) displays a distinct pattern of cytotoxicity and unique mechanistic features compared with other alkylating agents. Clin. Cancer Res. 2008, 14, 309–317. [Google Scholar] [CrossRef] [PubMed]
- El-Mabhouh, A.A.; Ayres, M.L.; Shpall, E.J.; Baladandayuthapani, V.; Keating, M.J.; Wierda, W.G.; Gandhi, V. Evaluation of bendamustine in combination with fludarabine in primary chronic lymphocytic leukemia cells. Blood 2014, 123, 3780–3789. [Google Scholar] [CrossRef] [PubMed]
- Burger, J.A.; Wiestner, A. Targeting B cell receptor signalling in cancer: Preclinical and clinical advances. Nat. Rev. Cancer 2018, 18, 148–167. [Google Scholar] [CrossRef] [PubMed]
- Okkenhaug, K.; Burger, J.A. PI3K signaling in normal B cells and chronic lymphocytic leukemia (CLL). Curr. Top. Microbiol. Immunol. 2016, 393, 123–142. [Google Scholar] [PubMed]
- Herman, S.E.M.; Gordon, A.L.; Wagner, A.J.; Heerema, N.A.; Zhao, W.; Flynn, J.M.; Jones, J.; Andritsos, L.; Puri, K.D.; Lannutti, B.J.; et al. Phosphatidylinositol 3-kinase-δ inhibitor CAL-101 shows promising preclinical activity in chronic lymphocytic leukemia by antagonizing intrinsic and extrinsic cellular survival signals. Blood 2010, 116, 2078–2088. [Google Scholar] [CrossRef]
- Lannutti, B.J.; Meadows, S.A.; Herman, S.E.M.; Kashishian, A.; Steiner, B.; Johnson, A.J.; Byrd, J.C.; Tyner, J.W.; Loriaux, M.M.; Deininger, M.; et al. CAL-101, a p110δ selective phosphatidylinositol-3-kinase inhibitor for the treatment of B-cell malignancies, inhibits PI3K signaling and cellular viability. Blood 2011, 117, 591–594. [Google Scholar] [CrossRef]
- Vanhaesebroeck, B.; Khwaja, A. PI3Kδ inhibition hits a sensitive spot in B cell malignancies. Cancer Cell 2014, 25, 269–271. [Google Scholar] [CrossRef]
- O’Brien, S.M.; Lamanna, N.; Kipps, T.J.; Flinn, I.; Zelenetz, A.D.; Burger, J.A.; Keating, M.; Mitra, S.; Holes, L.; Yu, A.S.; et al. A phase 2 study of idelalisib plus rituximab in treatment-naive older patients with chronic lymphocytic leukemia. Blood 2015, 126, 2686–2694. [Google Scholar] [CrossRef]
- Nair, K.S.; Cheson, B. The role of idelalisib in the treatment of relapsed and refractory chronic lymphocytic leukemia. Ther. Adv. Hematol. 2016, 7, 69–84. [Google Scholar] [CrossRef] [PubMed]
- Brown, J.R.; Byrd, J.C.; Coutre, S.E.; Benson, D.M.; Flinn, I.W.; Wagner-johnston, N.D.; Spurgeon, S.E.; Kahl, B.S.; Bello, C.; Webb, H.K.; et al. Idelalisib, an inhibitor of phosphatidylinositol 3-kinase p110 d, for relapsed/refractory chronic lymphocytic leukemia. Blood 2014, 123, 3390–3397. [Google Scholar] [CrossRef] [PubMed]
- Furman, R.R.; Sharman, J.P.; Coutre, S.E.; Cheson, B.D.; Pagel, J.M.; Hillmen, P.; Barrientos, J.C.; Zelenetz, A.D.; Kipps, T.J.; Flinn, I.; et al. Idelalisib and rituximab in relapsed chronic lymphocytic leukemia. N. Engl. J. Med. 2014, 370, 997–1007. [Google Scholar] [CrossRef] [PubMed]
- Sharman, J.P.; Coutre, S.E.; Furman, R.R.; Cheson, B.D.; Pagel, J.M.; Hillmen, P.; Barrientos, J.C.; Zelenetz, A.D.; Kipps, T.J.; Flinn, I.W.; et al. Final Results of a Randomized, Phase III Study of Rituximab With or Without Idelalisib Followed by Open-Label Idelalisib in Patients With Relapsed Chronic Lymphocytic Leukemia. J. Clin. Oncol. 2019, 37, 1391–1402. [Google Scholar] [CrossRef] [PubMed]
- De Weerdt, I.; Koopmans, S.M.; Kater, A.P.; Van Gelder, M. Incidence and management of toxicity associated with ibrutinib and idelalisib: A practical approach. Haematologica 2017, 102, 1629–1639. [Google Scholar] [CrossRef]
- Zelenetz, A.D.; Barrientos, J.C.; Brown, J.R.; Coiffier, B.; Delgado, J.; Egyed, M.; Ghia, P.; Illés, Á.; Jurczak, W.; Marlton, P.; et al. Idelalisib or placebo in combination with bendamustine and rituximab in patients with relapsed or refractory chronic lymphocytic leukaemia: Interim results from a phase 3, randomised, double-blind, placebo-controlled trial. Lancet Oncol. 2017, 18, 297–311. [Google Scholar] [CrossRef]
- Modi, P.; Balakrishnan, K.; Yang, Q.; Wierda, W.G. Idelalisib and bendamustine combination is synergistic and increases DNA damage response in chronic lymphocytic leukemia cells. Oncotarget 2017, 8, 16259–16274. [Google Scholar] [CrossRef][Green Version]
- Montraveta, A.; Lee-Vergés, E.; Roldán, J.; Jiménez, L.; Cabezas, S.; Clot, G.; Pinyol, M.; Xargay-Torrent, S.; Rosich, L.; Arimany-Nardí, C.; et al. CD69 expression potentially predicts response to bendamustine and its modulation by ibrutinib or idelalisib enhances cytotoxic effect in chronic lymphocytic leukemia. Oncotarget 2016, 7, 5507–5520. [Google Scholar] [CrossRef]
- Di Veroli, G.Y.; Fornari, C.; Wang, D.; Mollard, S.; Bramhall, J.L.; Richards, F.M.; Jodrell, D.I. Combenefit: An interactive platform for the analysis and visualization of drug combinations. Bioinformatics 2016, 32, 2866–2868. [Google Scholar] [CrossRef]
- Liston, D.R.; Davis, M. Clinically relevant concentrations of anticancer drugs: A guide for nonclinical studies. Clin. Cancer Res. 2017, 23, 3489–3498. [Google Scholar] [CrossRef]
- Ali, A.Y.A.Y.; Wu, X.; Eissa, N.; Hou, S.; Ghia, J.-E.J.E.; Murooka, T.T.T.T.; Banerji, V.; Johnston, J.B.J.B.; Lin, F.; Gibson, S.B.S.B.; et al. Distinct roles for phosphoinositide 3-kinases γ and δ in malignant B cell migration. Leukemia 2018, 32, 1958–1969. [Google Scholar] [CrossRef] [PubMed]
- Crassini, K.; Shen, Y.; Mulligan, S.; Best, O.G.; Crassini, K.; Shen, Y.; Mulligan, S.; Best, O.G. Modeling the chronic lymphocytic leukemia microenvironment in vitro. Leuk. Lymphoma 2017, 58, 266–279. [Google Scholar] [CrossRef] [PubMed]
- Mariotti, L.G.; Pirovano, G.; Savage, K.I.; Ghita, M.; Ottolenghi, A.; Prise, K.M.; Schettino, G.; Prise, M.; Schettino, G. Use of the γ-H2AX Assay to Investigate DNA Repair Dynamics Following Multiple Radiation Exposures. PLoS ONE 2013, 8, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Lezina, L.; Spriggs, R.V.; Beck, D.; Jones, C.; Dudek, K.M.; Bzura, A.; Jones, G.D.D.; Packham, G.; Willis, A.E.; Wagner, S.D. CD40L/IL-4-stimulated CLL demonstrates variation in translational regulation of DNA damage response genes including ATM. Blood Adv. 2018, 2, 1869–1881. [Google Scholar] [CrossRef] [PubMed]
- Bakkenist, C.J.; Kastan, M.B. DNA damage activates ATM through intermolecular autophosphorylation and dimer dissociation. Nature 2003, 421, 499–506. [Google Scholar] [CrossRef]
- Kingston, R.E.; Bunker, C.A.; Imbalzano, A.N. Repression and activation by multiprotein complexes that alter chromatin structure. Genes Dev. 1996, 10, 905–920. [Google Scholar] [CrossRef]
- Ding, W.; Shanafelt, T.D.; Lesnick, C.E.; Erlichman, C.; Leis, J.F.; Secreto, C.; Sassoon, T.R.; Call, T.G.; Bowen, D.A.; Conte, M.; et al. Akt inhibitor MK2206 selectively targets CLL B-cell receptor induced cytokines, mobilizes lymphocytes and synergizes with bendamustine to induce CLL apoptosis. Br. J. Haematol. 2014, 164, 146–150. [Google Scholar] [CrossRef]
- Yang, Q.; Chen, L.S.; Ha, M.J.; Do, K.A.; Neelapu, S.S.; Gandhi, V. Idelalisib impacts cell growth through inhibiting translation-regulatory mechanisms in mantle cell lymphoma. Clin. Cancer Res. 2017, 23, 181–192. [Google Scholar] [CrossRef]
- Landau, D.A.; Sun, C.; Rosebrock, D.; Herman, S.E.M.; Fein, J.; Sivina, M.; Underbayev, C.; Liu, D.; Hoellenriegel, J.; Ravichandran, S.; et al. The evolutionary landscape of chronic lymphocytic leukemia treated with ibrutinib targeted therapy. Nat. Commun. 2017, 8, 1–12. [Google Scholar] [CrossRef]
- Bojarczuk, K.; Sasi, B.K.; Gobessi, S.; Innocenti, I.; Pozzato, G.; Laurenti, L.; Efremov, D.G. BCR signaling inhibitors differ in their ability to overcome Mcl-1 – mediated resistance of CLL B cells to ABT-199. Blood 2016, 127, 3192–3201. [Google Scholar] [CrossRef]
- Turinetto, V.; Giachino, C. Survey and summary multiple facets of histone variant H2AX: A DNA double-strand-break marker with several biological functions. Nucleic Acids Res. 2015, 43, 2489–2498. [Google Scholar] [CrossRef] [PubMed]
- Stucki, M.; Clapperton, J.A.; Mohammad, D.; Yaffe, M.B.; Smerdon, S.J.; Jackson, S.P. MDC1 directly binds phosphorylated histone H2AX to regulate cellular responses to DNA double-strand breaks. Cell 2005, 123, 1213–1226. [Google Scholar] [CrossRef] [PubMed]
- Begleiter, A.; Verburg, L.; Israels, L.G.; Johnston, J.B. Factors influencing the inhibition of repair of irradiation-induced DNA damage by 2’-deoxycoformycin and deoxyadenosine. Cancer Chemother. Pharmacol. 1992, 30, 65–69. [Google Scholar] [CrossRef] [PubMed]
- Compagno, M.; Wang, Q.; Pighi, C.; Cheong, T.-C.C.; Meng, F.-L.L.; Poggio, T.; Yeap, L.-S.S.; Karaca, E.; Blasco, R.B.; Langellotto, F.; et al. Phosphatidylinositol 3-kinase δ blockade increases genomic instability in B cells. Nature 2017, 542, 489–493. [Google Scholar] [CrossRef]
- Holmes, K.B.; Sadreev, I.I.; Rawstron, A.C.; Munir, T.; Westhead, D.R.; Hillmen, P.; Lefevre, P.F. Ibrutinib induces chromatin reorganisation of chronic lymphocytic leukaemia cells. Oncogenesis 2019, 8, 32. [Google Scholar] [CrossRef]
- Singh, I.; Ozturk, N.; Cordero, J.; Mehta, A.; Hasan, D.; Cosentino, C.; Sebastian, C.; Krüger, M.; Looso, M.; Carraro, G.; et al. High mobility group protein-mediated transcription requires DNA damage marker γ-H2AX. Cell Res. 2015, 25, 837–850. [Google Scholar] [CrossRef]
- McManus, K.; Hendzel, M. ATM-dependent DNA Damage-independent Mitotic Phosphorylation of H2AX in Normally Growing Mammalian Cells. Mol. Biol. Cell 2005, 16, 5013–5025. [Google Scholar] [CrossRef]
- Hallek, M.; Cheson, B.D.; Catovsky, D.; Caligaris-Cappio, F.; Dighiero, G.; Döhner, H.; Hillmen, P.; Keating, M.J.; Montserrat, E.; Rai, K.R.; et al. Guidelines for the diagnosis and treatment of chronic lymphocytic leukemia: A report from the International Workshop on Chronic Lymphocytic Leukemia updating the National Cancer Institute-Working Group 1996 guidelines. Blood 2008, 111, 5446–5456. [Google Scholar] [CrossRef]
- Gehrke, I.; Bouchard, E.D.J.; Beiggi, S.; Poeppl, A.G.; Johnston, J.B.; Gibson, S.B.; Banerji, V. On-Target Effect of FK866, a Nicotinamide Phosphoribosyl Transferase Inhibitor, by Apoptosis-Mediated Death in Chronic Lymphocytic Leukemia Cells. Clin. Cancer Res. 2014, 20, 4861–4872. [Google Scholar] [CrossRef]
- Yoon, J.Y.; Ishdorj, G.; Graham, B.A.; Johnston, J.B.; Gibson, S.B. Valproic acid enhances fludarabine-induced apoptosis mediated by ROS and involving decreased AKT and ATM activation in B-cell-lymphoid neoplastic cells. Apoptosis 2014, 19, 191–200. [Google Scholar] [CrossRef]
- Johnston, J.B.; Kabore, A.F.; Strutinsky, J.; Hu, X.; Paul, J.T.; Kropp, D.M.; Kuschak, B.; Begleiter, A.; Gibson, S.B. Role of the TRAIL/APO2-L death receptors in chlorambucil- and fludarabine-induced apoptosis in chronic lymphocytic leukemia. Oncogene 2003, 22, 8356–8369. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Okkenhaug, K.; Bilancio, A.; Priddle, H.; Sancho, S.; Peskett, E.; Pearce, W.; Meek, S.E.; Salpekar, A.; Waterfield, M.D. Impaired B and T Cell Antigen Receptor Signaling in p110delta PI 3-Kinase Mutant Mice.pdf. Science (80-. ) 2002, 297, 1031–1034. [Google Scholar]
- Katyal, S.; Lee, Y.; Nitiss, K.C.; Downing, S.M.; Li, Y.; Shimada, M.; Zhao, J.; Russell, H.R.; Petrini, J.H.J.; Nitiss, J.L.; et al. Aberrant topoisomerase-1 DNA lesions are pathogenic in neurodegenerative genome instability syndromes. Nat. Neurosci. 2014, 17, 813–821. [Google Scholar] [CrossRef] [PubMed]
- Amrein, L.; Hernandez, T.A.; Ferrario, C.; Johnston, J.; Gibson, S.B.; Panasci, L.; Aloyz, R. Dasatinib sensitizes primary chronic lymphocytic leukaemia lymphocytes to chlorambucil and fludarabine in vitro. Br. J. Haematol. 2008, 143, 698–706. [Google Scholar] [CrossRef]
- Chou, T. Theoretical basis, experimental design, and computerized simulation of synergism and antagonism in drug combination studies. Pharmacol. Rev. 2006, 58, 621–681. [Google Scholar] [CrossRef]
- Chou, T.-C. Drug Combination Studies and Their Synergy Quantification Using the Chou-Talalay Method. Cancer Res. 2010, 70, 440–446. [Google Scholar] [CrossRef]

| Sample | IC50 (µM) | BEN/IDE CI Value | Patient Characteristics | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| IDE | IBR | BEN | CLB | FLU | Age (years) | Gender | Rai Stage | WBC (× 103/µL) | LDT (Months) | IgVh Status | Zap-70 Status | CD38 Status | Previous Treatment | FISH Status (%) | ||
| 1 | 1.6 | 0.6 | 24.2 | 14.4 | 4.9 | 61 | M | II | 54 | >12 | U | 15% | 16% | None | Del 13q14 (81) | |
| 2 | 3.3 | 1.6 | 69.1 | 30.4 | >10 | 1.2 | 77 | F | 0 | 141 | >12 | M | 2.2% | 0% | Yes 1 | Del 17p (43), 13q14 (24), ×2 (56) |
| 3 | 3.7 | 2.1 | 9.9 | 3.4 | 1.2 | 0.5 | 70 | M | I | 50 | >12 | M | 0% | 100% | None | Tri 12 (90) |
| 4 | 4.1 | 3.8 | 23.3 | 7.6 | 0.7 | 0.5 | 60 | M | II | 89 | >12 | M | 10% | 0% | None | Del 13q14 (94) |
| 5 | 4.9 | 2.0 | 38.8 | 13.5 | 1.7 | 0.7 | 77 | M | IV | 53 | >12 | M | 37% | 45% | None | Del 13q14 (93) |
| 6 | 5.3 | 2.4 | 15.4 | 10.0 | 1.2 | 64 | M | II | 61 | >12 | M | 3% | 45% | None | Normal | |
| 7 | 6.1 | 3.5 | 23.8 | 16.1 | 3.1 | 0.4 | 65 | M | II | 228 | <12 | U | 40% | 100% | Yes 2 | Del 11q (93), Tri 12 (94) |
| 8 | 6.6 | 3.8 | 15.4 | 8.8 | 45 | M | I | 31 | >12 | M | 1% | 0% | None | Normal | ||
| 9 | 7.0 | 3.3 | 139.4 | 23.4 | 4.1 | 0.5 | 67 | F | III | 75 | <12 | U | 50% | 100% | Yes 3 | Del 17p (14), 11q (13) |
| 10 | 7.6 | 6.1 | 6.2 | 3.8 | 0.4 | 0.6 | 65 | M | 0 | 37 | >12 | M | 2.80% | 0% | None | Del 13q14 (31) |
| 11 | 7.8 | 1.4 | 12.4 | 7.6 | 0.5 | 0.7 | 54 | M | II | 73 | >12 | M | 5% | 2% | None | Del 13q14 (25) |
| 12 | 8.0 | 1.8 | 78.9 | 54.7 | >10 | 60 | M | 0 | 75 | >12 | U | N/A | N/A | None | Del 11q (88), 13q14 (92) | |
| 13 | 8.1 | 3.8 | 14.0 | 10.1 | 7.4 | 0.6 | 74 | M | I | 103 | >12 | M | N/A | 0% | None | Del 13q14 (79) |
| 14 | 9.0 | 4.7 | 14.1 | 9.1 | 0.6 | 0.6 | 80 | F | 0 | 54 | >12 | M | 0% | 0% | None | Del 13q14 (15) |
| 15 | 9.3 | 2.8 | 31.9 | 14.4 | 4.1 | 0.5 | 69 | M | II | 124 | >12 | M | 3% | 3% | None | Del 13q14 (31), ×2 (8) |
| 16 | 13.2 | 4.5 | 23.9 | 19.3 | 2.0 | 0.6 | 86 | M | I | 48 | >12 | M | 0% | 0% | None | Del 13q14 × 2 (77) |
| 17 | 13.3 | 5.5 | 31.0 | 26.8 | >10 | 0.7 | 68 | M | II | 109 | <12 | M | 6% | 1% | None | Del 13q14 × 2 (92) |
| 18 | 13.5 | 5.7 | 51.5 | 35.5 | 6.3 | 0.5 | 57 | F | II | 391 | <12 | U | N/A | 0% | None | Del 11q (96), 13q14 (98) |
| 19 | 14.1 | 3.6 | 71.5 | 37.6 | >10 | 0.5 | 53 | M | IV | 94 | <12 | U | 36% | 31% | Yes 4 | Normal |
| 20 | 14.8 | 5.0 | 16.5 | 8.1 | 0.6 | 0.7 | 64 | M | 0 | 52 | >12 | M | 15% | 0% | None | Normal |
| 21 | 15.7 | 5.2 | 19.9 | 13.1 | 0.7 | 65 | M | II | 45 | >12 | M | 21% | 42% | None | Normal | |
| 22 | 17.9 | 4.6 | 29.8 | 13.1 | 1.0 | 63 | M | II | 21 | >12 | M | 0% | 0% | None | Del 13q14 (90) | |
| 23 | 19.8 | 3.0 | 15.4 | 8.1 | 0.6 | 0.4 | 82 | F | 0 | 129 | >12 | M | 3% | 7% | None | Normal |
| 24 | 19.9 | 5.1 | 32.4 | 15.7 | 1.2 | 0.6 | 59 | M | I | 145 | <12 | U | 0% | 0% | None | Del 13q14 (92) |
| 25 | 22.7 | 6.5 | 19.8 | 17.3 | >10 | 0.7 | 65 | M | III | 82 | >12 | U | 0% | 0% | None | Del 17p (9), 11q (48), 13q14 (39), ×2 (20) |
| 26 | 25.4 | 6.3 | 87.6 | 64.7 | >10 | 0.4 | 73 | F | II | 437 | <12 | U | 31% | 27% | None | Del 13q14 (86) |
| 27 | 26.5 | 7.5 | 134.3 | >80 | >10 | 0.8 | 71 | M | II | 135 | >12 | U | 9% | 0% | None | Del 17p (84), 13q14 × 2 (97) |
| 28 | 27.3 | 7.0 | 102.4 | 65.6 | >10 | 0.6 | 79 | M | III | 80 | >12 | M | 7.7% | 2% | None | Del 11q (21), 13q14 (93) |
| 29 | 30.3 | 5.5 | 10.3 | 4.6 | 1.2 | 0.7 | 78 | M | I | 49 | >12 | M | 4% | 0% | None | Del 13q14 (76.19), × 2 (6.19) |
| 30 | 32.6 | 3.0 | 49.2 | 38.3 | >10 | 0.8 | 67 | M | I | 408 | <12 | M | 90% | 0% | Yes 5 | Normal |
| 31 | 45.1 | 4.4 | 48.7 | 24.3 | 4.1 | 0.7 | 91 | F | 0 | 30 | >12 | M | 0% | 98% | None | Tri 12 (87) |
| 32 | 51.6 | 9.9 | 22.7 | 16.7 | 1.9 | 0.6 | 61 | M | I | 119 | >12 | U | N/A | N/A | None | Normal |
| Med | 13.3 | 4.1 | 24.0 | 14.4 | 1.2 | 0.59 | ||||||||||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kost, S.E.F.; Saleh, A.; Mejia, E.M.; Mostafizar, M.; Bouchard, E.D.J.; Banerji, V.; Marshall, A.J.; Gibson, S.B.; Johnston, J.B.; Katyal, S. Transcriptional Modulation by Idelalisib Synergizes with Bendamustine in Chronic Lymphocytic Leukemia. Cancers 2019, 11, 1519. https://doi.org/10.3390/cancers11101519
Kost SEF, Saleh A, Mejia EM, Mostafizar M, Bouchard EDJ, Banerji V, Marshall AJ, Gibson SB, Johnston JB, Katyal S. Transcriptional Modulation by Idelalisib Synergizes with Bendamustine in Chronic Lymphocytic Leukemia. Cancers. 2019; 11(10):1519. https://doi.org/10.3390/cancers11101519
Chicago/Turabian StyleKost, Sara E. F., Ali Saleh, Edgard M. Mejia, Marina Mostafizar, Eric D. J. Bouchard, Versha Banerji, Aaron J. Marshall, Spencer B. Gibson, James B. Johnston, and Sachin Katyal. 2019. "Transcriptional Modulation by Idelalisib Synergizes with Bendamustine in Chronic Lymphocytic Leukemia" Cancers 11, no. 10: 1519. https://doi.org/10.3390/cancers11101519
APA StyleKost, S. E. F., Saleh, A., Mejia, E. M., Mostafizar, M., Bouchard, E. D. J., Banerji, V., Marshall, A. J., Gibson, S. B., Johnston, J. B., & Katyal, S. (2019). Transcriptional Modulation by Idelalisib Synergizes with Bendamustine in Chronic Lymphocytic Leukemia. Cancers, 11(10), 1519. https://doi.org/10.3390/cancers11101519





