Integrated Biobanking and Tumor Model Establishment of Human Colorectal Carcinoma Provides Excellent Tools for Preclinical Research
Abstract
1. Introduction
2. Results
2.1. Patient Cohort Characteristics
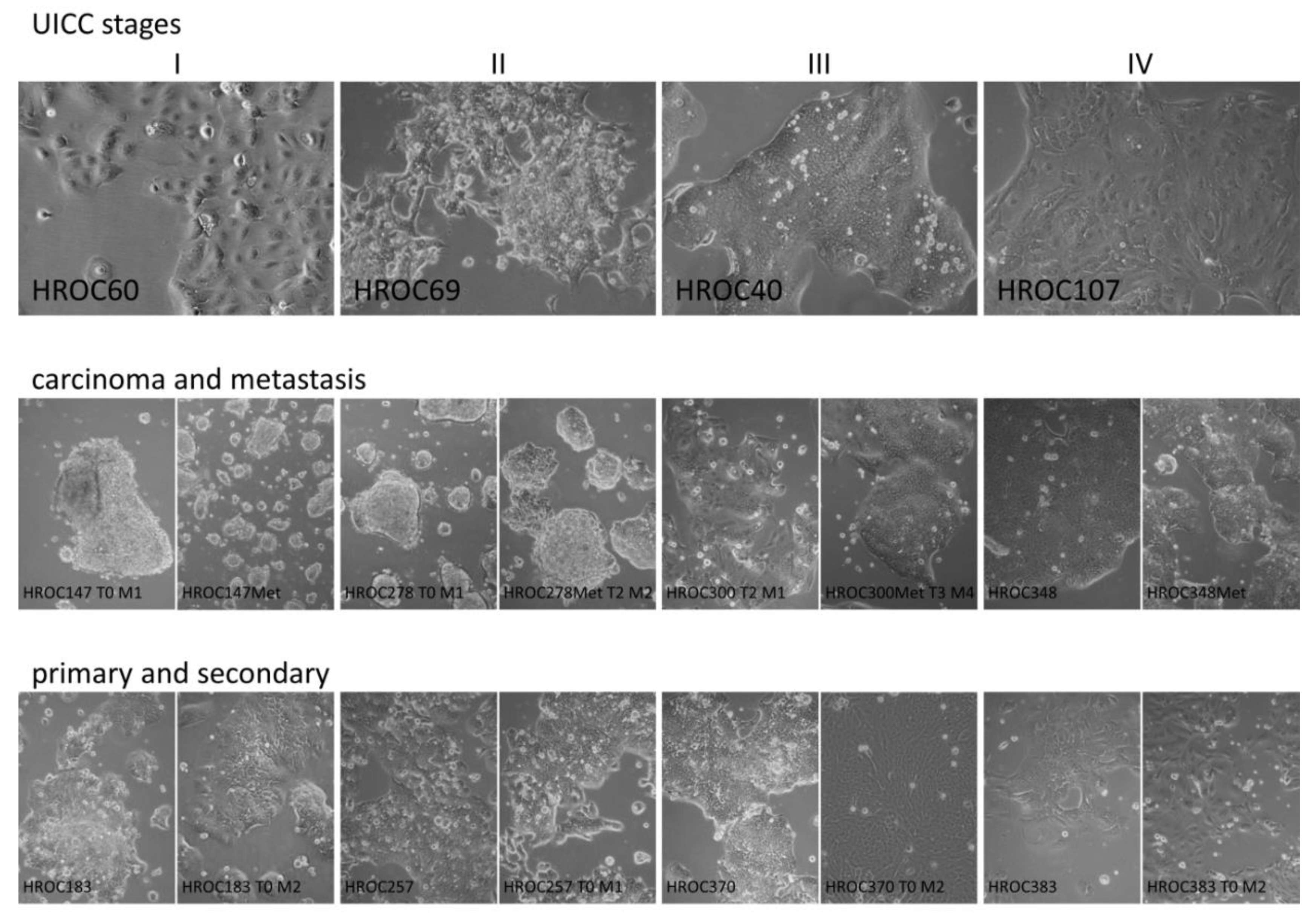
2.2. Establishment of Patient-Derived Cell Lines (Primary Cell Lines)
2.3. Establishment of Patient-Derived Xenografts (PDX)
2.4. Establishment of PDX-Derived Cell Lines (Secondary Cell Lines)
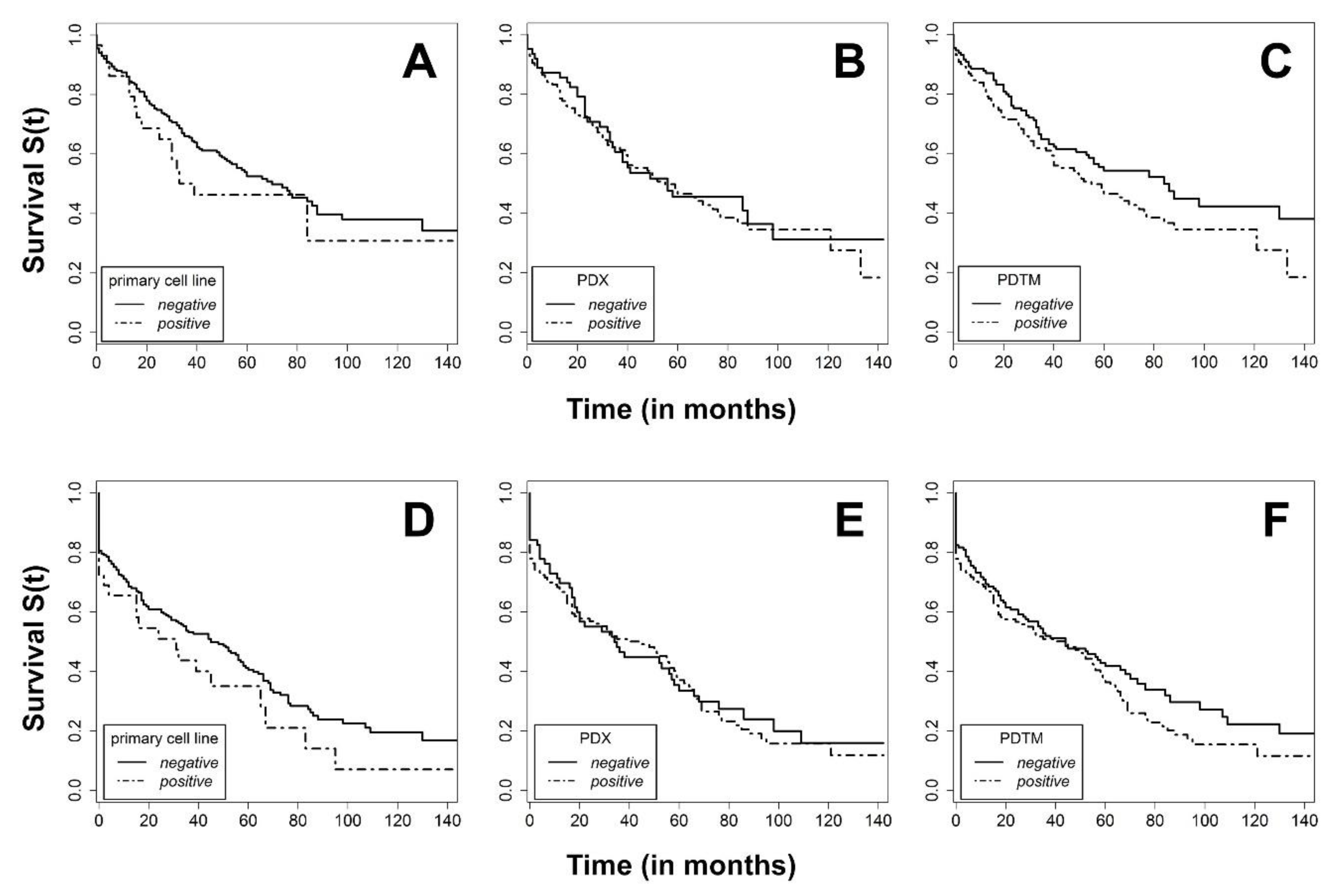
2.5. Global Patient and Model Analysis
3. Discussion
4. Materials and Methods
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- C, B.J. HeLa (for Henrietta Lacks). Science 1974, 184, 1268. [Google Scholar] [CrossRef] [PubMed]
- Pillai, S.P.S.; Uthamanthil, R.K. PDX Models: History and Development. In Patient Derived Tumor Xenograft Models, 1st ed.; Academic Press: Amsterdam, The Netherlands, 2017; pp. 1–12. ISBN 978-0-12-804010-2. [Google Scholar]
- Roschke, A.V.; Tonon, G.; Gehlhaus, K.S.; McTyre, N.; Bussey, K.J.; Lababidi, S.; Scudiero, D.A.; Weinstein, J.N.; Kirsch, I.R. Karyotypic complexity of the NCI-60 drug-screening panel. Cancer Res. 2003, 63, 8634–8647. [Google Scholar] [PubMed]
- Tan, D.S.; Thomas, G.V.; Garrett, M.D.; Banerji, U.; de Bono, J.S.; Kaye, S.B.; Workman, P. Biomarker-driven early clinical trials in oncology: A paradigm shift in drug development. Cancer J. 2009, 15, 406–420. [Google Scholar] [CrossRef]
- Horbach, S.P.J.M.; Halffman, W. The ghosts of HeLa: How cell line misidentification contaminates the scientific literature. PLoS ONE 2017, 12, e0186281. [Google Scholar] [CrossRef] [PubMed]
- Gao, H.; Korn, J.M.; Ferretti, S.; Monahan, J.E.; Wang, Y.; Singh, M.; Zhang, C.; Schnell, C.; Yang, G.; Zhang, Y.; et al. High-throughput screening using patient-derived tumor xenografts to predict clinical trial drug response. Nat. Med. 2015, 21, 1318–1325. [Google Scholar] [CrossRef] [PubMed]
- Bedra, M.; Vyskocil, T.; Emel, J.; Edwards, C.; Boutros, C. Synergetic role of integrating the departments of cancer registry and clinical research at an academic comprehensive cancer center. World J. Methodol. 2017, 7, 33–36. [Google Scholar] [CrossRef] [PubMed]
- Mullins, C.S.; Schneider, B.; Stockhammer, F.; Krohn, M.; Classen, C.F.; Linnebacher, M. Establishment and characterization of primary glioblastoma cell lines from fresh and frozen material: A detailed comparison. PLoS ONE 2013, 8, e71070. [Google Scholar] [CrossRef] [PubMed]
- Oberländer, M.; Linnebacher, M.; König, A.; Bogoevska, V.; Brodersen, C.; Kaatz, R.; Krohn, M.; Hackmann, M.; Ingenerf, J.; Christoph, J.; et al. ColoNet consortium. The “North German Tumor Bank of Colorectal Cancer”: Status report after the first 2 years of support by the German Cancer Aid Foundation. Langenbecks Arch. Surg. 2013, 398, 251–258. [Google Scholar] [CrossRef] [PubMed]
- Kolligs, F.T. Diagnostics and Epidemiology of Colorectal Cancer. Visc. Med. 2016, 32, 158–164. [Google Scholar] [CrossRef]
- Meza, R.; Jeon, J.; Renehan, A.G.; Luebeck, E.G. Colorectal cancer incidence trends in the United States and United kingdom: Evidence of right- to left-sided biological gradients with implications for screening. Cancer Res. 2010, 70, 5419–5429. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Fedewa, S.A.; Ahnen, D.J.; Meester, R.G.S.; Barzi, A.; Jemal, A. Colorectal cancer statistics, 2017. CA Cancer J. Clin. 2017, 67, 177–193. [Google Scholar] [CrossRef] [PubMed]
- Iversen, L.H.; Green, A.; Ingeholm, P.; Østerlind, K.; Gögenur, I. Improved survival of colorectal cancer in Denmark during 2001–2012—The efforts of several national initiatives. Acta Oncol. 2016, 55 (Suppl. 2), 10–23. [Google Scholar] [CrossRef] [PubMed]
- Gock, M.; Mullins, C.S.; Harnack, C.; Prall, F.; Ramer, R.; Göder, A.; Krämer, O.H.; Klar, E.; Linnebacher, M. Establishment, functional and genetic characterization of a colon derived large cell neuroendocrine carcinoma cell line. World J. Gastroenterol. 2018, 24, 3749–3759. [Google Scholar] [CrossRef] [PubMed]
- Dangles-Marie, V.; Pocard, M.; Richon Weiswald, L.B.; Assayag, F.; Saulnier, P.; Judde, J.G.; Janneau, J.L.; Auger, N.; Validire, P.; Dutrillaux, B.; et al. Establishment of human colon cancer cell lines from fresh tumors versus xenografts: Comparison of success rate and cell line features. Cancer Res. 2007, 67, 398–407. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Ory, V.; Chapman, S.; Yuan, H.; Albanese, C.; Kallakury, B.; Timofeeva, O.A.; Nealon, C.; Dakic, A.; Simic, V.; et al. ROCK inhibitor and feeder cells induce the conditional reprogramming of epithelial cells. Am. J. Pathol. 2012, 180, 599–607. [Google Scholar] [CrossRef] [PubMed]
- Bairoch, A. The Cellosaurus, a Cell-Line Knowledge Resource. J. Biomol. Tech. 2018, 29, 25–38. [Google Scholar] [CrossRef] [PubMed]
- Julien, S.; Merino-Trigo, A.; Lacroix, L.; Pocard, M.; Goéré, D.; Mariani, P.; Landron, S.; Bigot, L.; Nemati, F.; Dartigues, P.; et al. Characterization of a large panel of patient-derived tumor xenografts representing the clinical heterogeneity of human colorectal cancer. Clin. Cancer Res. 2012, 18, 5314–5328. [Google Scholar] [CrossRef] [PubMed]
- Oh, B.Y.; Lee, W.Y.; Jung, S.; Hong, H.K.; Nam, D.H.; Park, Y.A.; Huh, J.W.; Yun, S.H.; Kim, H.C.; Chun, H.K.; et al. Correlation between tumor engraftment in patient-derived xenograft models and clinical outcomes in colorectal cancer patients. Oncotarget 2015, 6, 16059–16068. [Google Scholar] [CrossRef]
- Katsiampoura, A.; Raghav, K.; Jiang, Z.Q.; Menter, D.G.; Varkaris, A.; Morelli, M.P.; Manuel, S.; Wu, J.; Sorokin, A.V.; Rizi, B.S.; et al. Modeling of Patient-Derived Xenografts in Colorectal Cancer. Mol. Cancer Ther. 2017, 16, 1435–1442. [Google Scholar] [CrossRef]
- Cybulska, M.; Olesinski, T.; Goryca, K.; Paczkowska, K.; Statkiewicz, M.; Kopczynski, M.; Grochowska, A.; Unrug-Bielawska, K.; Tyl-Bielicka, A.; Gajewska, M.; et al. Challenges in Stratifying the Molecular Variability of Patient-Derived Colon Tumor Xenografts. BioMed Res. Int. 2018, 2018, 2954208. [Google Scholar] [CrossRef]
- Bertotti, A.; Migliardi, G.; Galimi, F.; Sassi, F.; Torti, D.; Isella, C.; Corà, D.; Di Nicolantonio, F.; Buscarino, M.; Petti, C.; et al. A molecularly annotated platform of patient-derived xenografts (“xenopatients”) identifies HER2 as an effective therapeutic target in cetuximab-resistant colorectal cancer. Cancer Discov. 2011, 1, 508–523. [Google Scholar] [CrossRef] [PubMed]
- Pauli, C.; Hopkins, B.D.; Prandi, D.; Shaw, R.; Fedrizzi, T.; Sboner, A.; Sailer, V.; Augello, M.; Puca, L.; Rosati, R.; et al. Personalized In Vitro and In Vivo Cancer Models to Guide Precision Medicine. Cancer Discov. 2017, 7, 462–477. [Google Scholar] [CrossRef] [PubMed]
- Seol, H.S.; Kang, H.J.; Lee, S.I.; Kim, N.E.; Kim, T.I.; Chun, S.M.; Kim, T.W.; Yu, C.S.; Suh, Y.A.; Singh, S.R.; et al. Development and characterization of a colon PDX model that reproduces drug responsiveness and the mutation profiles of its original tumor. Cancer Lett. 2014, 345, 56–64. [Google Scholar] [CrossRef] [PubMed]
- Van de Wetering, M.; Francies, H.E.; Francis, J.M.; Bounova, G.; Iorio, F.; Pronk, A.; van Houdt, W.; van Gorp, J.; Taylor-Weiner, A.; Kester, L.; et al. Prospective derivation of a living organoid biobank of colorectal cancer patients. Cell 2015, 161, 933–945. [Google Scholar] [CrossRef] [PubMed]
- Gock, M.; Mullins, C.S.; Bergner, C.; Prall, F.; Ramer, R.; Göder, A.; Krämer, O.H.; Lange, F.; Krause, B.J.; Klar, E.; et al. Establishment, functional and genetic characterization of three novel patient-derived rectal cancer cell lines. World J. Gastroenterol. 2018, 24, 4880–4892. [Google Scholar] [CrossRef] [PubMed]
- Prall, F.; Maletzki, C.; Hühns, M.; Krohn, M.; Linnebacher, M. Colorectal carcinoma tumour budding and podia formation in the xenograft microenvironment. PLoS ONE 2017, 12, e0186271. [Google Scholar] [CrossRef] [PubMed]
- Kuehn, F.; Mullins, C.S.; Krohn, M.; Harnack, C.; Ramer, R.; Krämer, O.H.; Klar, E.; Huehns, M.; Linnebacher, M. Establishment and characterization of HROC69—A Crohn´s related colonic carcinoma cell line and its matched patient-derived xenograft. Sci. Rep. 2016, 6, 24671. [Google Scholar] [CrossRef]
- Maletzki, C.; Stier, S.; Gruenert, U.; Gock, M.; Ostwald, C.; Prall, F.; Linnebacher, M. Establishment, characterization and chemosensitivity of three mismatch repair deficient cell lines from sporadic and inherited colorectal carcinomas. PLoS ONE 2012, 7, e52485. [Google Scholar] [CrossRef] [PubMed]
- Lazzari, L.; Corti, G.; Picco, G.; Isella, C.; Montone, M.; Arcella, P.; Durinikova, E.; Zanella, E.R.; Novara, L.; Barbosa, F.; et al. Patient-Derived Xenografts and Matched Cell Lines Identify Pharmacogenomic Vulnerabilities in Colorectal Cancer. Clin. Cancer Res. 2019. [Google Scholar] [CrossRef]
- Russo, M.; Lamba, S.; Lorenzato, A.; Sogari, A.; Corti, G.; Rospo, G.; Mussolin, B.; Montone, M.; Lazzari, L.; Arena, S.; et al. Reliance upon ancestral mutations is maintained in colorectal cancers that heterogeneously evolve during targeted therapies. Nat. Commun. 2018, 9, 2287. [Google Scholar] [CrossRef]
- Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature 2012, 487, 330–337. [Google Scholar] [CrossRef] [PubMed]
- Puig, I.; Chicote, I.; Tenbaum, S.P.; Arqués, O.; Herance, J.R.; Gispert, J.D.; Jimenez, J.; Landolfi, S.; Caci, K.; Allende, H.; et al. A personalized preclinical model to evaluate the metastatic potential of patient-derived colon cancer initiating cells. Clin. Cancer Res. 2013, 19, 6787–6801. [Google Scholar] [CrossRef]
- Mullins, C.S.; Bock, S.; Krohn, M.; Linnebacher, M. Generation of Xenotransplants from Human Cancer Biopsies to Assess Anti-cancer Activities of HDACi. Methods Mol. Biol. 2017, 1510, 217–229. [Google Scholar] [PubMed]
- Hühns, M.; Krohn, S.; Murua Escobar, H.; Prall, F. Genomic heterogeneity in primary colorectal carcinomas and their metastases: Born bad or brought up a villain? Hum. Pathol. 2018, 74, 54–63. [Google Scholar] [CrossRef] [PubMed]
- Maletzki, C.; Huehns, M.; Bauer, I.; Ripperger, T.; Mork, M.M.; Vilar, E.; Klöcking, S.; Zettl, H.; Prall, F.; Linnebacher, M. Frameshift mutational target gene analysis identifies similarities and differences in constitutional mismatch repair-deficiency and Lynch syndrome. Mol. Carcinog. 2017, 56, 1753–1764. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018. [Google Scholar]
- Punt, C.J.; Buyse, M.; Köhne, C.H.; Hohenberger, P.; Labianca, R.; Schmoll, H.J.; Påhlman, L.; Sobrero, A.; Douillard, J.Y. Endpoints in adjuvant treatment trials: A systematic review of the literature in colon cancer and proposed definitions for future trials. J. Natl. Cancer Inst. 2007, 99, 998–1003. [Google Scholar] [CrossRef] [PubMed]
| Patients Characteristics (n = 382) | ||||||
|---|---|---|---|---|---|---|
| Male | Female | Mean Age | 5-year Follow-up Rate | 5-year Relapse-free Survival | 5-year Survival Rate | |
| Adenoma (n = 32) | 23 (71.9%) | 9 (28.1%) | 69.0 (28–85) | n.a. | n.a. | n.a. |
| CRC (n = 350 ◆) | 207 (59.1%) | 143 (40.9%) | 72.0 (21–98) | 236 (67.4%) | 43.2% | 51.3% |
| UICC I (n = 74; 21.1%) | 42 (56.8%) | 32 (43.2%) | 73.0 (28–98) | 47 (63.5%) | 61.7% | 72.3% |
| UICCII (n = 112; 32.0%) | 63 (56.2%) | 49 (43.8%) | 73.5 (21–92) | 71 (63.4%) | 62% | 66.2% |
| UICC III (n = 87; 24.9%) | 47 (54.0%) | 40 (46.0%) | 70.0 (40–88) | 65 (74.7%) | 43% | 53.8% |
| UICC IV (n = 76; 21.7%) | 54 (71.0%) | 22 (29.0%) | 70.0 (30–85) | 52 (68.4%) | 0% | 9.6% |
| Sample Properties (n = 315) | ||||||
| primary resected tumors | metastases | |||||
| Adenoma (n = 9) | 9 | / | ||||
| Adenocarcinoma (n = 306) | 262 | 44 | ||||
| Unsuccessful tissue acquisition (n = 107) | ||||||
| primary resected tumors | metastases | |||||
| Adenoma (n = 25) | 25 | / | ||||
| Adenocarcinoma (n = 82) | 72 | 10 | ||||
| CRC Metastases Characteristics (n = 44) | ||||||
| resection site | ||||||
| liver | lung | peritoneum | brain | lymph node | ||
| synchronous (n = 18) | 17 | 0 | 1 | 0 | 0 | |
| metachronous (n = 26) | 19 | 1 | 3 | 2 | 1 | |
| initial stage | UICC I | UICC II | UICC III | UICC IV | ||
| 5 | 4 | 10 | 7 | |||
| sets of primary tumor and one or more corresponding metastases | N = 20 | |||||
| A: Primary Cell Line | |||||
|---|---|---|---|---|---|
| patients | model success | p-value | |||
| n (%) | no | yes | univariate | multivariate | |
| age | 0.715 | ||||
| <61 | 54 (20.5) | 45 (83.3) | 9 (16.7) | ||
| 61–70 | 65 (24.7) | 58 (89.2) | 7 (10.8) | ||
| 71–80 | 105 (39.9) | 93 (88.6) | 12 (11.4) | ||
| >80 | 39 (14.8) | 33 (84.6) | 6 (15.4) | ||
| gender | 0.559 | ||||
| male | 159 (60.5) | 140 (88.1) | 19 (11.9) | ||
| female | 104 (39.5) | 89 (85.6) | 15 (14.4) | ||
| sample type | 0.197 | ||||
| primary | 221 (84.0) | 195 (88.2) | 26 (11.8) | ||
| metastasis | 42 (16.0) | 34 (81.0) | 8 (19.0) | ||
| localization | 0.156 | ||||
| left colon | 26 (9.9) | 23 (88.5) | 3 (11.5) | ||
| rectum | 29 (11.0) | 28 (96.6) | 1 (3.4) | ||
| right colon | 101 (38.4) | 84 (83.2) | 17 (16.8) | ||
| sigmoid | 54 (20.5) | 51 (94.4) | 3 (5.6) | ||
| transverse | 11 (4.2) | 9 (81.8) | 2 (18.2) | ||
| metastasis | 42 (16.0) | 34 (81.0) | 8 (19.0) | ||
| T | 0.778 | ||||
| T0-2 | 38 (17.4) | 34 (89.5) | 4 (10.5) | ||
| T3-4 | 181 (82.6) | 159 (87.8) | 22 (12.2) | ||
| N | 0.057 | 0.026 * | |||
| N0 | 113 (51.6) | 101 (89.4) | 12 (10.6) | ||
| N1 | 44 (20.1) | 42 (95.5) | 2 (4.5) | ||
| N2 | 62 (28.3) | 50 (80.6) | 12 (19.4) | ||
| M | 0.775 | ||||
| M0 | 165 (75.3) | 146 (88.5) | 19 (11.5) | ||
| M1 | 54 (24.7) | 47 (87.0) | 7 (13.0) | ||
| G | 0.007 ** | ||||
| G1-2 | 151 (68.9) | 139 (92.1) | 12 (7.9) | ||
| G3-4 | 68 (31.1) | 54 (79.4) | 14 (20.6) | ||
| R | 0.769 | ||||
| R0 | 172 (78.5) | 153 (89.0) | 19 (11.0) | ||
| R1 | 7 (3.2) | 6 (85.7) | 1 (14.3) | ||
| R2 | 40 (18.3) | 34 (85.0) | 6 (15.0) | ||
| L | 0.657 | ||||
| L0 | 155 (71.8) | 139 (88.0) | 17 (11.0) | ||
| L1 | 61 (28.2) | 53 (86.9) | 8 (13.1) | ||
| V | 0.769 | ||||
| V0 | 128 (59.3) | 112 (87.5) | 16 (12.5) | ||
| V1 | 82 (38.0) | 74 (90.2) | 8 (9.8) | ||
| V2 | 6 (2.7) | 5 (83.3) | 1 (16.7) | ||
| molecular type | 0.707 | ||||
| spStd | 84 (56.4) | 74 (88.1) | 10 (11.9) | ||
| spMSI-H | 25 (16.8) | 20 (80.0) | 5 (20.0) | ||
| CIMP-H | 30 (20.1) | 25 (83.3) | 5 (16.7) | ||
| Lynch | 10 (6.7) | 8 (80.0) | 2 (20.0) | ||
| UICC | 0.896 | ||||
| I | 35 (14.7) | 31 (88.6) | 4 (11.4) | ||
| II | 79 (33.2) | 71 (89.9) | 8 (10.1) | ||
| III | 65 (27.3) | 56 (86.2) | 9 (13.8) | ||
| IV | 59 (24.8) | 51 (86.4) | 8 (13.6) | ||
| MSI status | 0.297 | ||||
| MSS + MSI-L | 109 (75.7) | 95 (87.2) | 14 (12.8) | ||
| MSI-H | 35 (24.3) | 28 (80.0) | 7 (20.0) | ||
| TP53 | 0.121 | 0.105 | |||
| wt | 15 (40.5) | 13 (86.7) | 2 (13.3) | ||
| mut | 22 (59.5) | 14 (63.6) | 8 (36.4) | ||
| K-Ras | 0.248 | ||||
| wt | 106 (65.4) | 94 (88.7) | 12 (11.3) | ||
| mut | 56 (34.6) | 46 (82.1) | 10 (17.9) | ||
| B-Raf | 0.255 | ||||
| wt | 139 (85.3) | 122 (87.8) | 17 (12.2) | ||
| mut | 24 (14.7) | 19 (79.2) | 5 (20.8) | ||
| PIK3CA | |||||
| wt | 27 (100.0) | 23 (85.2) | 4 (14.8) | ||
| mut | 0 (0.0) | 0 (0.0) | 0 (0.0) | ||
| B: PDX | |||||
| patients | model success | p-value | |||
| n (%) | no | yes | univariate | multivariate | |
| age | 0.048 * | ||||
| <61 | 49 (22.4) | 22 (44.9) | 27 (55.1) | ||
| 61–70 | 51 (23.3) | 16 (31.4) | 35 (68.6) | ||
| 71–80 | 78 (35.6) | 17 (21.8) | 61 (78.2) | ||
| >80 | 41 (18.7) | 15 (36.6) | 26 (63.4) | ||
| gender | 0.196 | ||||
| male | 127 (58.0) | 45 (35.4) | 82 (64.6) | ||
| female | 92 (42.0) | 25 (27.2) | 67 (72.8) | ||
| sample type | 0.014 * | 0.092 | |||
| primary | 188 (85.8) | 66 (35.1) | 122 (64.9) | ||
| metastasis | 31 (14.2) | 4 (12.9) | 27 (87.1) | ||
| localization | 0.000 *** | ||||
| left colon | 19 (8.7) | 4 (21.1) | 15 (78.9) | ||
| rectum | 32 (14.6) | 17 (53.1) | 15 (46.9) | ||
| right colon | 82 (37.4) | 20 (24.4) | 62 (75.6) | ||
| sigmoid | 43 (19.6) | 24 (55.8) | 19 (44.2) | ||
| transverse | 12 (5.5) | 1 (8.3) | 11 (91.7) | ||
| metastasis | 31 (14.2) | 4 (12.9) | 27 (87.1) | ||
| T | 0.293 | ||||
| T0-2 | 27 (14.5) | 12 (44.4) | 15 (55.6) | ||
| T3-4 | 159 (85.5) | 54 (34.0) | 105 (66.0) | ||
| N | 0.062 | ||||
| N0 | 94 (50.5) | 41 (43.6) | 53 (56.4) | ||
| N1 | 39 (21.0) | 10 (25.6) | 29 (74.4) | 0.024 * | |
| N2 | 53 (28.5) | 15 (28.3) | 38 (71.7) | 0.200 | |
| M | 0.047 * | 0.035 * | |||
| M0 | 136 (73.1) | 54 (39.7) | 82 (60.3) | ||
| M1 | 50 (26.9) | 12 (24.0) | 38 (76.0) | ||
| G | 0.083 | 0.110 | |||
| G1-2 | 123 (66.1) | 49 (39.8) | 74 (60.2) | ||
| G3-4 | 63 (33.9) | 17 (27.0) | 46 (73.0) | ||
| R | 0.074 | ||||
| R0 | 140 (75.3) | 56 (40.0) | 84 (60.0) | ||
| R1 | 7 (3.8) | 2 (28.6) | 5 (71.4) | ||
| R2 | 39 (21.0) | 8 (20.5) | 31 (79.5) | ||
| L | 0.996 | ||||
| L0 | 133 (72.3) | 47 (35.3) | 86 (64.7) | ||
| L1 | 51 (27.7) | 18 (35.3) | 33 (64.7) | ||
| V | 0.506 | ||||
| V0 | 108 (58.7) | 35 (32.4) | 73 (67.6) | ||
| V1 | 72 (39.1) | 29 (40.3) | 43 (59.7) | ||
| V2 | 4 (2.2) | 1 (25.0) | 3 (75.0) | ||
| molecular type | 0.003 ** | ||||
| spStd | 82 (53.9) | 34 (41.5) | 48 (58.5) | ||
| spMSI-H | 27 (17.8) | 3 (11.1) | 24 (88.9) | ||
| CIMP-H | 34 (22.4) | 9 (26.5) | 25 (73.5) | ||
| Lynch | 9 (5.9) | 0 (0.0) | 9 (100.0) | ||
| UICC | 0.083 | ||||
| I | 22 (11.3) | 7 (31.8) | 15 (68.2) | ||
| II | 67 (34.5) | 31 (46.3) | 36 (53.7) | ||
| III | 53 (27.3) | 16 (30.2) | 37 (69.8) | ||
| IV | 52 (26.8) | 13 (25.0) | 39 (75.0) | ||
| MSI status | 0.001 *** | 0.070 | |||
| MSS + MSI-L | 112 (76.2) | 43 (38.4) | 69 (61.6) | ||
| MSI-H | 35 (23.8) | 3 (8.6) | 32 (91.4) | ||
| TP53 | 0.887 | ||||
| wt | 21 (47.7) | 4 (19.0) | 17 (81.0) | ||
| mut | 23 (52.3) | 4 (17.4) | 19 (82.6) | ||
| K-Ras | 0.019 * | 0.005 ** | |||
| wt | 105 (64.8) | 39 (37.1) | 66 (62.9) | ||
| mut | 57 (35.2) | 11 (19.3) | 46 (80.7) | ||
| B-Raf | 0.002 ** | 0.004 ** | |||
| wt | 139 (85.3) | 49 (35.3) | 90 (64.7) | ||
| mut | 24 (14.7) | 1 (4.2) | 23 (95.8) | ||
| PIK3CA | |||||
| wt | 16 (100.0) | 4 (25.0) | 12 (75.0) | ||
| mut | 0 (0.0) | 0 (0.0) | 0 (0.0) | ||
| C: PDTM | |||||
| patients | model success | p-value | |||
| n (%) | no | yes | univariate | Multivariate | |
| age | 0.730 | ||||
| <61 | 59 (19.8) | 32 (54.2) | 27 (45.8) | ||
| 61–70 | 73 (24.5) | 36 (49.3) | 37 (50.7) | ||
| 71–80 | 115 (38.6) | 52 (45.2) | 63 (54.8) | ||
| >80 | 51 (17.1) | 25 (49.0) | 26 (51.0) | ||
| gender | 0.131 | ||||
| male | 178 (59.7) | 93 (52.2) | 85 (47.8) | ||
| female | 120 (40.3) | 52 (43.3) | 68 (56.7) | ||
| sample type | 0.150 | ||||
| primary | 254 (85.2) | 128 (50.4) | 126 (49.6) | ||
| metastasis | 44 (14.8) | 17 (38.6) | 27 (61.4) | ||
| localization | 0.011 * | ||||
| left colon | 29 (9.7) | 14 (48.3) | 15 (51.7) | ||
| rectum | 35 (11.7) | 20 (57.1) | 15 (42.9) | ||
| right colon | 117 (39.3) | 52 (44.4) | 65 (55.6) | ||
| sigmoid | 59 (19.8) | 39 (66.1) | 20 (33.9) | ||
| transverse | 14 (4.7) | 3 (21.4) | 11 (78.6) | ||
| metastasis | 44 (14.8) | 17 (38.6) | 27 (61.4) | ||
| T | 0.061 | ||||
| T0-2 | 44 (17.5) | 28 (63.6) | 16 (36.4) | ||
| T3-4 | 208 (82.5) | 100 (48.1) | 108 (51.9) | ||
| N | 0.011 * | ||||
| N0 | 133 (52.8) | 79 (59.4) | 54 (40.6) | ||
| N1 | 53 (21.0) | 24 (45.3) | 29 (54.7) | 0.046 * | |
| N2 | 66 (26.2) | 25 (37.9) | 41 (62.1) | 0.110 | |
| M | 0.008 ** | 0.070 | |||
| M0 | 191 (75.8) | 106 (55.5) | 85 (44.5) | ||
| M1 | 61 (24.2) | 22 (36.1) | 39 (63.9) | ||
| G | 0.006 ** | ||||
| G1-2 | 173 (68.7) | 98 (56.6) | 75 (43.4) | ||
| G3-4 | 79 (31.3) | 30 (38.0) | 49 (62.0) | ||
| R | 0.026 * | ||||
| R0 | 195 (77.4) | 108 (55.4) | 87 (44.6) | ||
| R1 | 8 (3.2) | 3 (37.5) | 5 (62.5) | ||
| R2 | 49 (19.4) | 17 (34.7) | 32 (65.3) | ||
| L | 0.688 | ||||
| L0 | 183 (73.5) | 94 (51.4) | 89 (48.6) | ||
| L1 | 66 (26.5) | 32 (48.5) | 34 (51.5) | ||
| V | 0.601 | ||||
| V0 | 147 (59.0) | 71 (48.3) | 76 (51.7) | ||
| V1 | 94 (37.8) | 50 (53.2) | 44 (46.8) | ||
| V2 | 8 (3.2) | 5 (62.5) | 3 (37.5) | ||
| molecular type | 0.024 * | ||||
| spStd | 93 (53.8) | 44 (47.3) | 49 (52.7) | ||
| spMSI-H | 33 (19.1) | 9 (27.3) | 24 (72.7) | ||
| CIMP-H | 37 (21.4) | 11 (29.7) | 26 (70.3) | ||
| Lynch | 10 (5.8) | 1 (10.0) | 9 (90.0) | ||
| UICC | 0.023* | ||||
| I | 40 (14.8) | 24 (60.0) | 16 (40.0) | ||
| II | 93 (34.3) | 57 (61.3) | 36 (38.7) | ||
| III | 72 (26.6) | 33 (45.8) | 39 (54.2) | ||
| IV | 66 (24.4) | 26 (39.4) | 40 (60.6) | ||
| MSI status | 0.019 * | 0.070 | |||
| MSS + MSI-L | 127 (75.1) | 56 (44.1) | 71 (55.9) | ||
| MSI-H | 42 (24.9) | 10 (23.8) | 32 (76.2) | ||
| TP53 | 0.786 | ||||
| wt | 21 (45.7) | 4 (19.0) | 17 (81.0) | ||
| mut | 25 (54.9) | 4 (16.0) | 21 (84.0) | ||
| K-Ras | 0.014 * | 0.002 ** | |||
| wt | 123 (65.1) | 56 (45.5) | 67 (54.5) | ||
| mut | 66 (34.9) | 18 (27.3) | 48 (72.7) | ||
| B-Raf | 0.022 * | 0.026 * | |||
| wt | 162 (84.8) | 70 (43.2) | 92 (56.8) | ||
| mut | 29 (15.2) | 6 (20.7) | 23 (79.3) | ||
| PIK3CA | |||||
| wt | 28 (100.0) | 16 (57.1) | 12 (42.9) | ||
| mut | 0 (0.0) | 0 (0.0) | 0 (0.0) | ||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mullins, C.S.; Micheel, B.; Matschos, S.; Leuchter, M.; Bürtin, F.; Krohn, M.; Hühns, M.; Klar, E.; Prall, F.; Linnebacher, M. Integrated Biobanking and Tumor Model Establishment of Human Colorectal Carcinoma Provides Excellent Tools for Preclinical Research. Cancers 2019, 11, 1520. https://doi.org/10.3390/cancers11101520
Mullins CS, Micheel B, Matschos S, Leuchter M, Bürtin F, Krohn M, Hühns M, Klar E, Prall F, Linnebacher M. Integrated Biobanking and Tumor Model Establishment of Human Colorectal Carcinoma Provides Excellent Tools for Preclinical Research. Cancers. 2019; 11(10):1520. https://doi.org/10.3390/cancers11101520
Chicago/Turabian StyleMullins, Christina S., Bianca Micheel, Stephanie Matschos, Matthias Leuchter, Florian Bürtin, Mathias Krohn, Maja Hühns, Ernst Klar, Friedrich Prall, and Michael Linnebacher. 2019. "Integrated Biobanking and Tumor Model Establishment of Human Colorectal Carcinoma Provides Excellent Tools for Preclinical Research" Cancers 11, no. 10: 1520. https://doi.org/10.3390/cancers11101520
APA StyleMullins, C. S., Micheel, B., Matschos, S., Leuchter, M., Bürtin, F., Krohn, M., Hühns, M., Klar, E., Prall, F., & Linnebacher, M. (2019). Integrated Biobanking and Tumor Model Establishment of Human Colorectal Carcinoma Provides Excellent Tools for Preclinical Research. Cancers, 11(10), 1520. https://doi.org/10.3390/cancers11101520





