Main Human Urinary Metabolites after Genipap (Genipa americana L.) Juice Intake
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethical Approval and Subject Recruitment
2.2. Juice Preparation
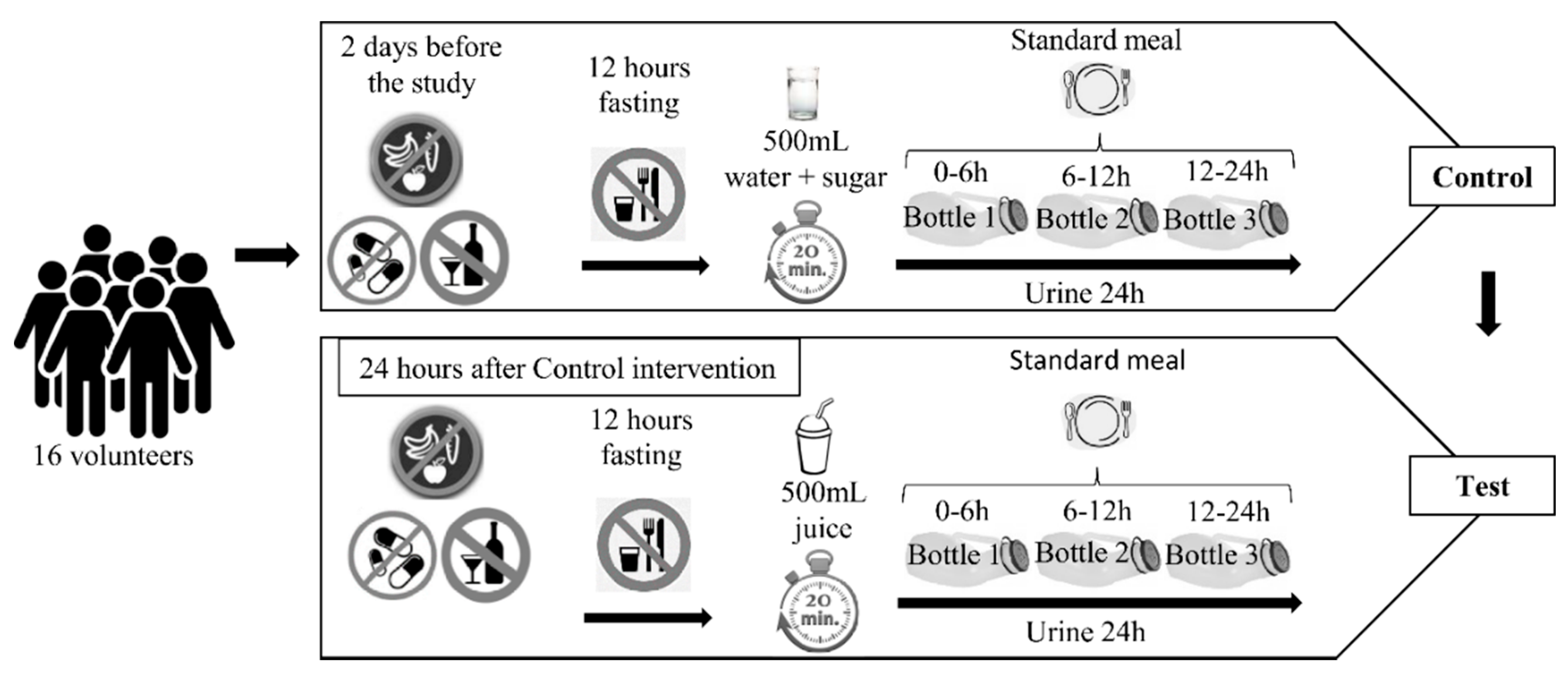
2.3. Study Design
2.4. Sample Preparation
2.5. UHPLC-HESI-Orbitrap-MS Analysis
2.6. Preprocessing and Pretreatment of Data
2.7. Data Analysis and Multi-Metabolite Biomarker Model
2.8. Annotation, Identification, and Interpretation
3. Results
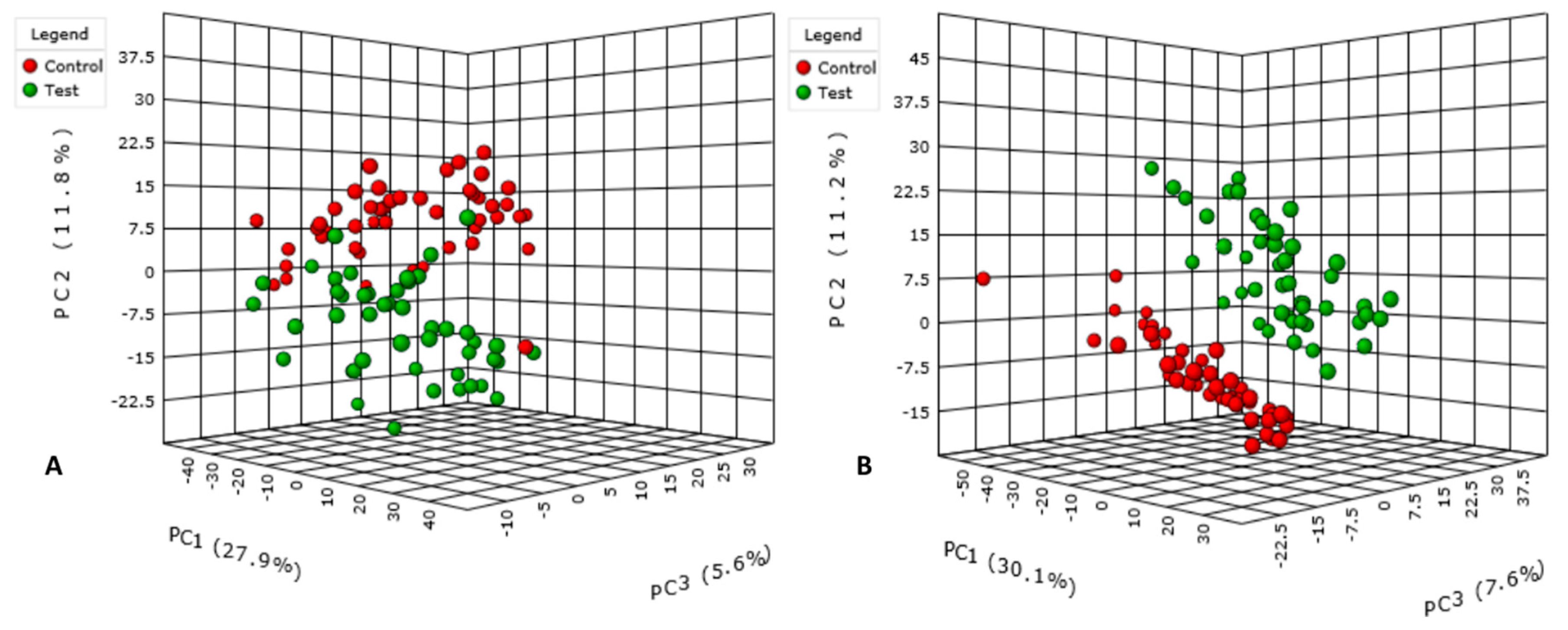
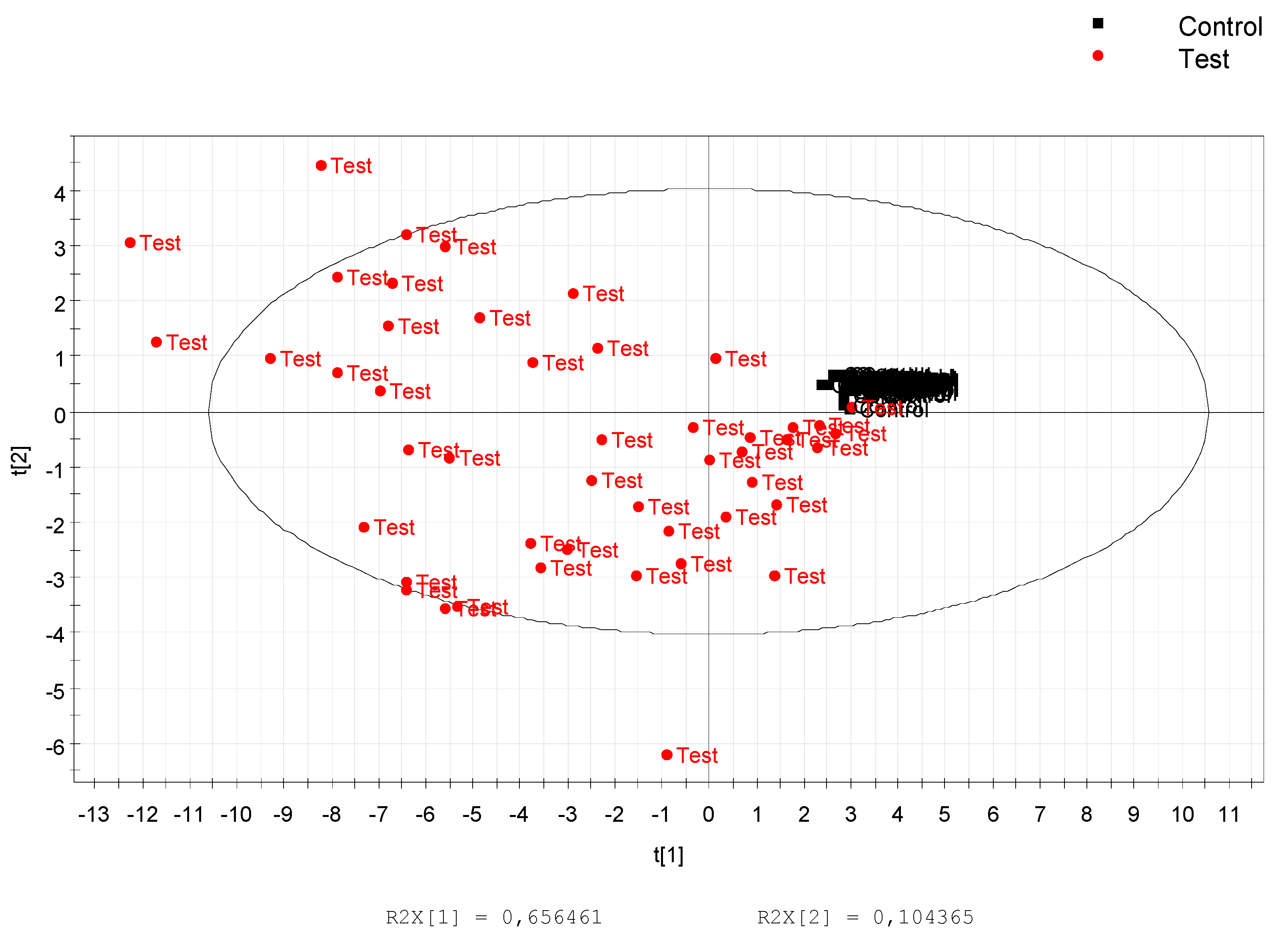
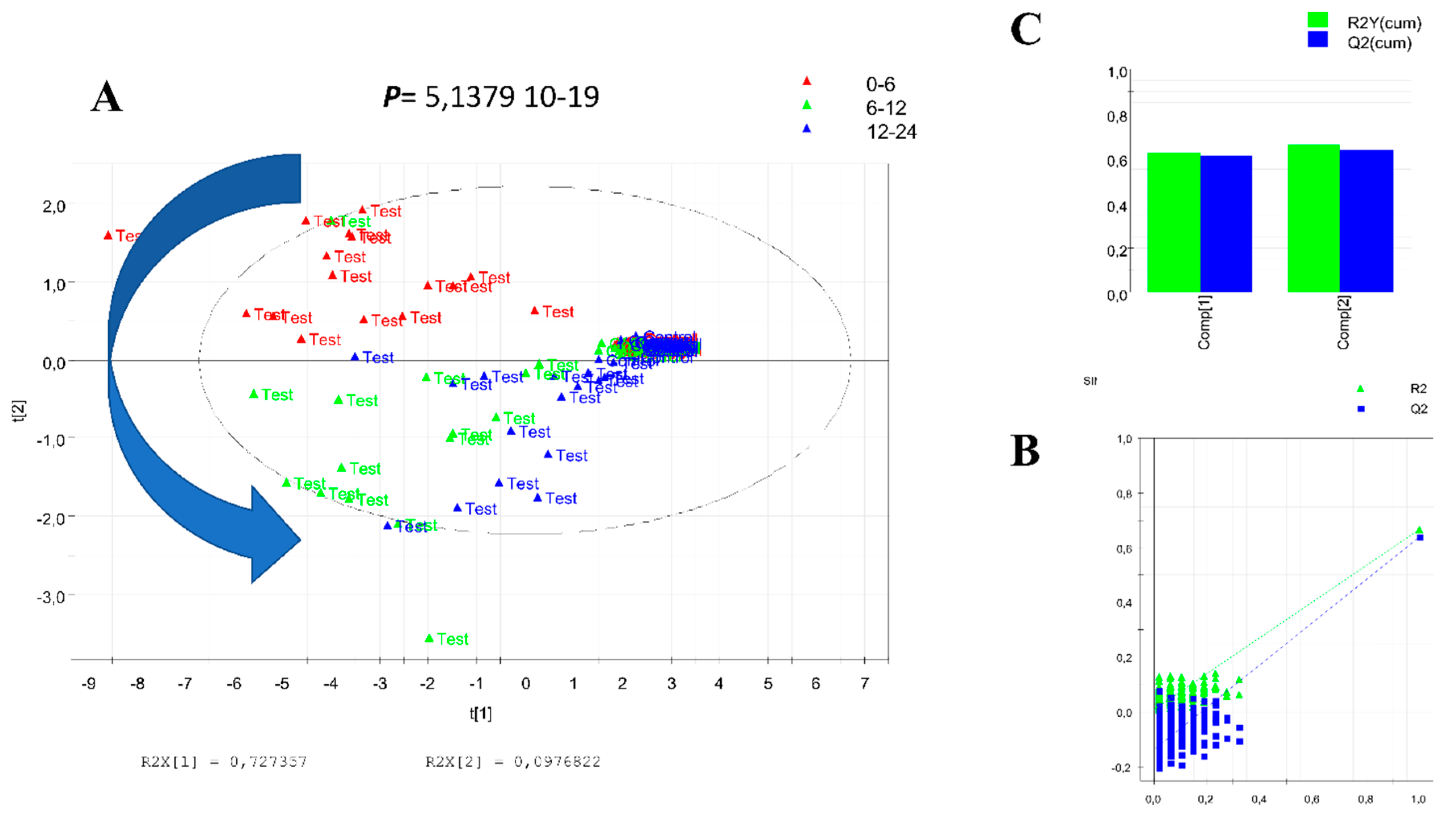
3.1. Data Treatment and Analysis
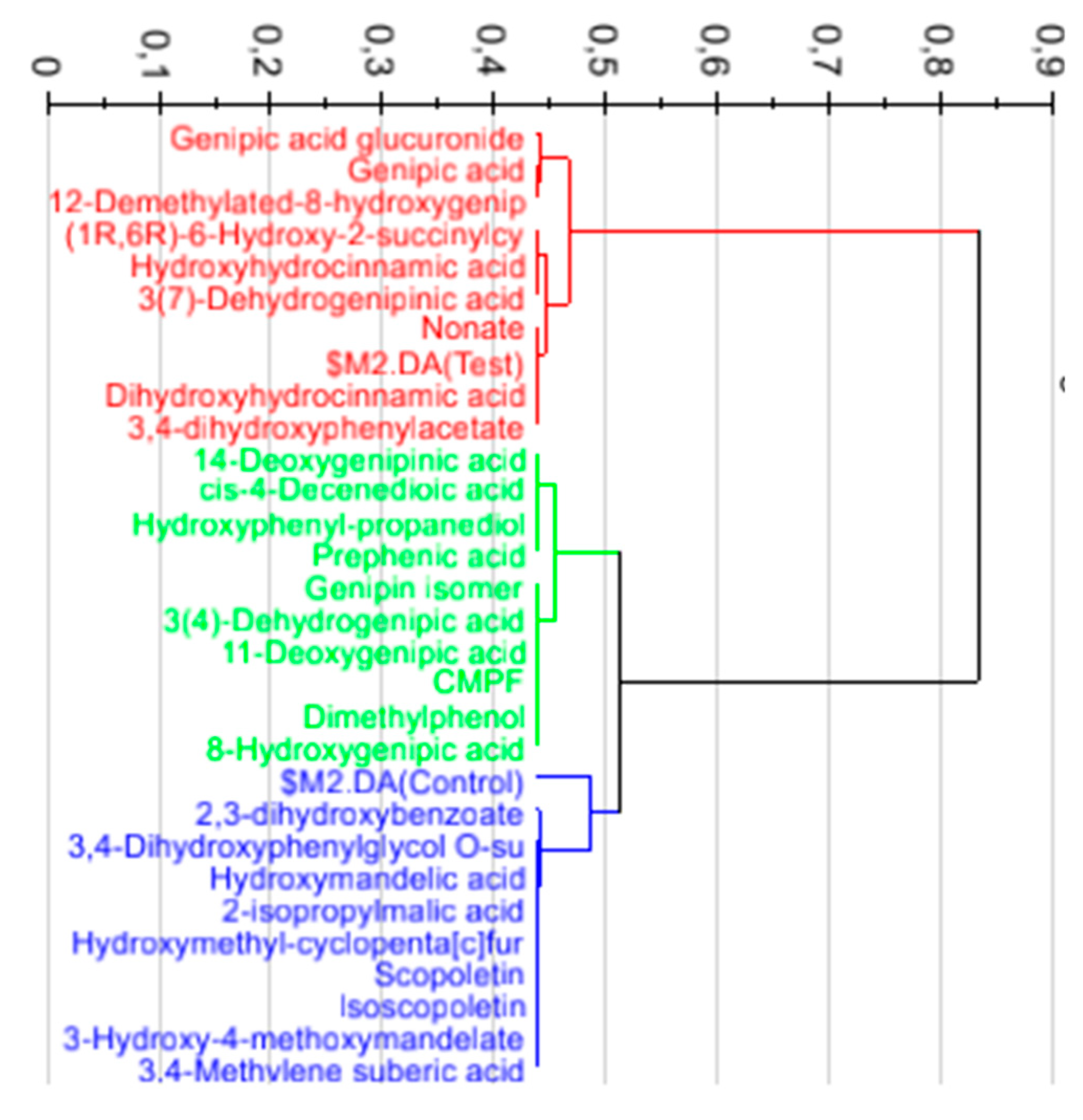
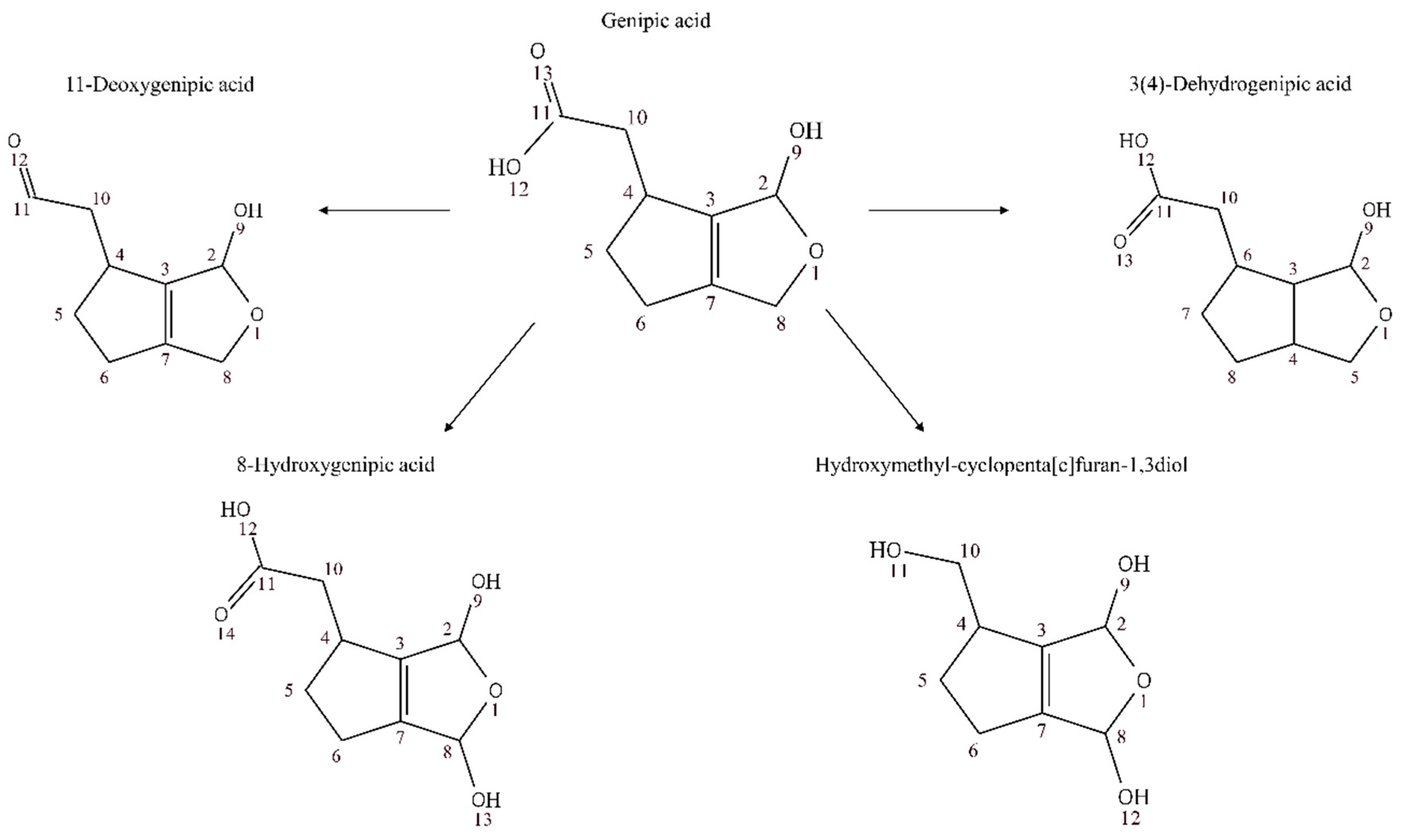
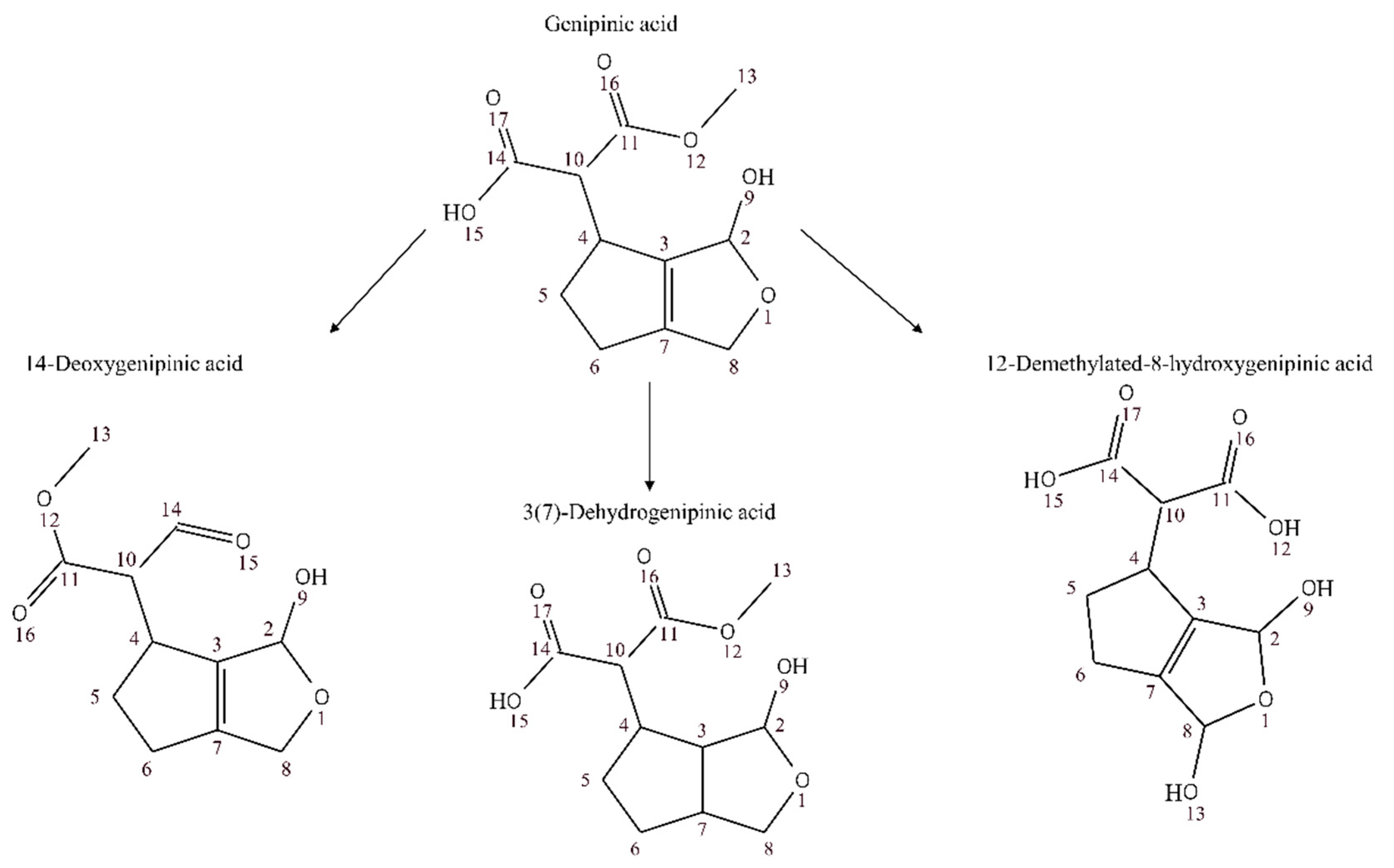
3.2. Annotation and Tentative Identification
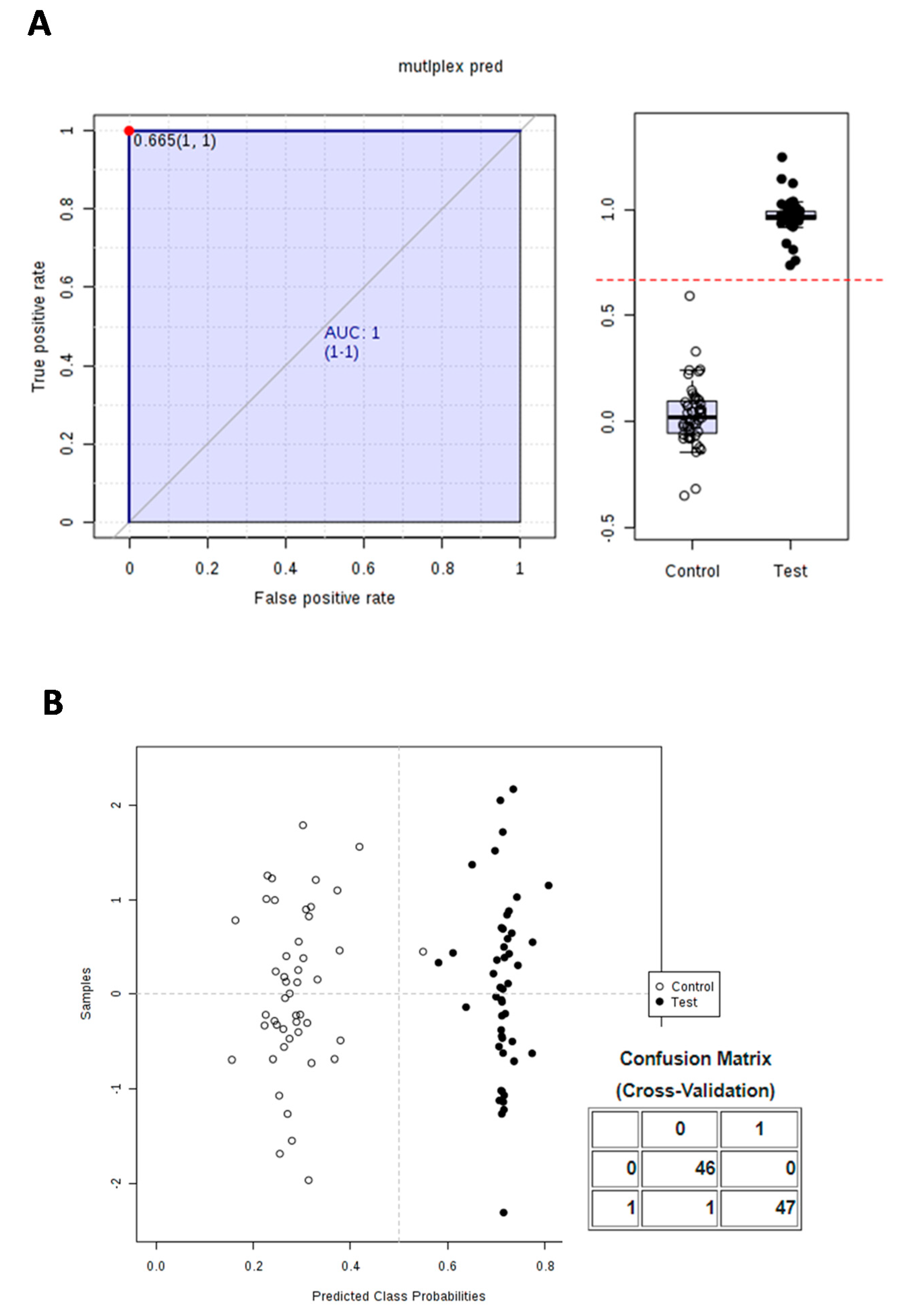
3.3. Calculation and Validation of a Multiplex Biomarker of Genipap Exposure
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Athersuch, T. Metabolome analyses in exposome studies: Profiling methods for a vast chemical space. Arch. Biochem. Biophys. 2016, 589, 177–186. [Google Scholar] [CrossRef] [PubMed]
- Wild, C.P. The exposome: From concept to utility. Int. J. Epidemiol. 2012, 41, 24–32. [Google Scholar] [CrossRef] [PubMed]
- Dragsted, L.O.; Gao, Q.; Praticò, G.; Manach, C.; Wishart, D.S.; Scalbert, A.; Feskens, E.J.M. Dietary and health biomarkers—Time for an update. Genes Nutr. 2017, 12, 24. [Google Scholar] [CrossRef] [PubMed]
- Rappaport, S.M. Biomarkers intersect with the exposome. Biomarkers 2012, 17, 483–489. [Google Scholar] [CrossRef] [PubMed]
- Bando, K.; Kawahara, R.; Kunimatsu, T.; Sakai, J.; Kimura, J.; Funabashi, H.; Seki, T.; Bamba, T.; Fukusaki, E. Influences of biofluid sample collection and handling procedures on gc–ms based metabolomic studies. J. Biosci. Bioeng. 2010, 110, 491–499. [Google Scholar] [CrossRef] [PubMed]
- Saude, E.J.; Sykes, B.D. Urine stability for metabolomic studies: Effects of preparation and storage. Metabolomics 2007, 3, 19–27. [Google Scholar] [CrossRef]
- Wishart, D.S. Metabolomics: Applications to food science and nutrition research. Trends Food Sci. Technol. 2008, 19, 482–493. [Google Scholar] [CrossRef]
- Martins, D.; Nunez, C. Secondary Metabolites from Rubiaceae Species. Molecules 2015, 20, 13422–13495. [Google Scholar] [CrossRef] [PubMed]
- De Figueiredo, R.W.; Maia, G.A.; de Holanda, L.F.F.; Monteiro, J.C.S. Características físicas e químicas do jenipapo. Pesquisa Agropecuária Brasileira 1986, 21, 421–428. [Google Scholar]
- Orwa, C.; Mutua, A.; Kindt, R.; Jamnadass, R.; Anthony, S. Genipa Americana. In Agroforestree Database: A Tree Reference and Selection Guide Version 4.0; World Agroforestry Centre: Nairobi, Kenya, 2009. [Google Scholar]
- Zappi, D. GENIPA L. Flora Fanerogâmica do Estado de São Paulo; Instituto de Botânica: São Paulo, Brazil, 2007; Volume 5, pp. 344–345. [Google Scholar]
- Da Conceição, A.O.; Rossi, M.H.; de Oliveira, F.F.; Takser, L.; Lafond, J. Genipa americana (Rubiaceae) fruit extract affects mitogen-activated protein kinase cell pathways in human trophoblast–derived BeWo cells: Implications for placental development. J. Med. Food 2011, 14, 483–494. [Google Scholar]
- Morton, J. Fruits of Warm Climates; Creative Resource Systems: Miami, FL, USA, 1987. [Google Scholar]
- De Sousa Bentes, A.; Mercadante, A.Z. Influence of the Stage of Ripeness on the Composition of Iridoids and Phenolic Compounds in Genipap (Genipa americana L.). J. Agric. Food Chem. 2014, 62, 10800–10808. [Google Scholar] [CrossRef] [PubMed]
- Alves, L.; Ming, L.C. Chemistry and pharmacology of some plants mentioned in the letter of Pero Vaz de Caminha. Etnobiol. Conserv. 2015, 4, 1–15. [Google Scholar] [CrossRef]
- Mitra, M. Gray Special Feature. Asia Pac. Biotech. News 2007, 11, 689–743. [Google Scholar] [CrossRef]
- Revilla, J. Apontamentos para a Cosmética Amazônica/Juan Revilla, 2nd ed.; INPA, Instituto Nacional de Pesquisas da Amazônia: Manaus, Brazil, 2002. [Google Scholar]
- Tundis, R.; Loizzo, M.R.; Menichini, F.; Statti, G.A.; Menichini, F. Biological and pharmacological activities of iridoids: Recent developments. Mini Rev. Med. Chem. 2008, 8, 399–420. [Google Scholar] [CrossRef] [PubMed]
- Ono, M.; Ishimatsu, N.; Masuoka, C.; Yoshimitsu, H.; Tsuchihashi, R.; Okawa, M.; Kinjo, J.; Ikeda, T.; Nohara, T. Three new monoterpenoids from the fruit of Genipa americana. Chem. Pharm. Bull. 2007, 55, 632–634. [Google Scholar] [CrossRef] [PubMed]
- Ono, M.; Ueno, M.; Masuoka, C.; Ikeda, T.; Nohara, T. Iridoid glucosides from the fruit of Genipa americana. Chem. Pharm. Bull. 2005, 53, 1342–1344. [Google Scholar] [CrossRef] [PubMed]
- Shanmugam, M.K.; Shen, H.; Tang, F.R.; Arfuso, F.; Rajesh, M.; Wang, L.; Kumar, A.P.; Bian, J.; Goh, B.C.; Bishayee, A.; et al. Potential role of genipin in cancer therapy. Pharmacol. Res. 2018, 133, 195–200. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Zhang, T.-B.; Jia, D.-H.; Sun, W.-Q.; Wang, C.-L.; Gu, A.-Z.; Yang, X.-M. Genipin inhibits the growth of human bladder cancer cells via inactivation of PI3K/Akt signaling. Oncol. Lett. 2018, 15, 2619–2624. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Yin, F.; Guo, L.; Deng, X.; Hu, Y. Neuroprotection of geniposide against hydrogen peroxide induced PC12 cells injury: Involvement of PI3 kinase signal pathway. Acta Pharmacol. Sin. 2009, 30, 159–165. [Google Scholar] [CrossRef] [PubMed]
- Son, M.; Lee, M.; Ryu, E.; Moon, A.; Jeong, C.-S.; Jung, Y.W.; Park, G.H.; Sung, G.-H.; Cho, H.; Kang, H. Genipin as a novel chemical activator of EBV lytic cycle. J. Microbiol. 2015, 53, 155–165. [Google Scholar] [CrossRef] [PubMed]
- Hwa, J.S.; Mun, L.; Kim, H.J.; Seo, H.G.; Lee, J.H.; Kwak, J.H.; Lee, D.-U.; Chang, K.C. Genipin selectively inhibits TNF-α-activated VCAM-1 but not ICAM-1 expression by upregulation of PPAR-γ in human endothelial cells. Korean J. Physiol. Pharmacol. 2011, 15, 157–162. [Google Scholar] [CrossRef] [PubMed]
- Viljoen, A.; Mncwangi, N.; Vermaak, I. Anti-Inflammatory Iridoids of Botanical Origin. Curr. Med. Chem. 2012, 19, 2104–2127. [Google Scholar] [CrossRef] [PubMed]
- Koo, H.-J.; Song, Y.S.; Kim, H.-J.; Lee, Y.-H.; Hong, S.-M.; Kim, S.-J.; Kim, B.-C.; Jin, C.; Lim, C.-J.; Park, E.-H. Antiinflammatory effects of genipin, an active principle of gardenia. Eur. J. Pharmacol. 2004, 495, 201–208. [Google Scholar] [CrossRef] [PubMed]
- Mikami Masaki; Takikawa Hajime Effect of genipin on the biliary excretion of cholephilic compounds in rats. Hepatol. Res. 2008, 38, 614–621. [CrossRef] [PubMed]
- Wu, S.; Wang, G.; Liu, Z.; Rao, J.; Lü, L.; Xu, W.; Wu, S.; Zhang, J. Effect of geniposide, a hypoglycemic glucoside, on hepatic regulating enzymes in diabetic mice induced by a high-fat diet and streptozotocin. Acta Pharmacol. Sin. 2009, 30, 202–208. [Google Scholar] [CrossRef] [PubMed]
- Team, R.C. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2013. [Google Scholar]
- Kessner, D.; Chambers, M.; Burke, R.; Agus, D.; Mallick, P. ProteoWizard: Open source software for rapid proteomics tools development. Bioinformatics 2008, 24, 2534–2536. [Google Scholar] [CrossRef] [PubMed]
- Smith, C.A.; Want, E.J.; O’Maille, G.; Abagyan, R.; Siuzdak, G. XCMS: processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal. Chem. 2006, 78, 779–787. [Google Scholar] [CrossRef] [PubMed]
- Xia, J.; Wishart, D.S. Using metaboanalyst 3.0 for comprehensive metabolomics data analysis. In Current Protocols in Bioinformatics; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2002. [Google Scholar]
- Aidoud, N.; Delplanque, B.; Baudry, C.; Garcia, C.; Moyon, A.; Balasse, L.; Guillet, B.; Antona, C.; Darmaun, D.; Fraser, K.; et al. A combination of lipidomics, MS imaging, and PET scan imaging reveals differences in cerebral activity in rat pups according to the lipid quality of infant formulas. FASEB J. 2018. [Google Scholar] [CrossRef] [PubMed]
- Sumner, L.W.; Amberg, A.; Barrett, D.; Beale, M.H.; Beger, R.; Daykin, C.A.; Fan, T.W.-M.; Fiehn, O.; Goodacre, R.; Griffin, J.L.; et al. Proposed minimum reporting standards for chemical analysis. Metabolomics 2007, 3, 211–221. [Google Scholar] [CrossRef] [PubMed]
- Afgan, E.; Baker, D.; van den Beek, M.; Blankenberg, D.; Bouvier, D.; Čech, M.; Chilton, J.; Clements, D.; Coraor, N.; Eberhard, C.; et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2016 update. Nucleic Acids Res. 2016, 44, W3–W10. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S.; Jewison, T.; Guo, A.C.; Wilson, M.; Knox, C.; Liu, Y.; Djoumbou, Y.; Mandal, R.; Aziat, F.; Dong, E.; et al. HMDB 3.0—The Human Metabolome Database in 2013. Nucleic Acids Res. 2013, 41, D801–D807. [Google Scholar] [CrossRef] [PubMed]
- Smith, C.A.; O’Maille, G.; Want, E.J.; Qin, C.; Trauger, S.A.; Brandon, T.R.; Custodio, D.E.; Abagyan, R.; Siuzdak, G. METLIN: A metabolite mass spectral database. Ther. Drug Monit. 2005, 27, 747–751. [Google Scholar] [CrossRef] [PubMed]
- Ruttkies, C.; Schymanski, E.L.; Wolf, S.; Hollender, J.; Neumann, S. MetFrag relaunched: Incorporating strategies beyond in silico fragmentation. J. Cheminformatics 2016, 8, 3. [Google Scholar] [CrossRef] [PubMed]
- Gerlich, M.; Neumann, S. MetFusion: Integration of compound identification strategies. J. Mass Spectrom 2013, 48. [Google Scholar] [CrossRef] [PubMed]
- Chagoyen, M.; Pazos, F. Tools for the functional interpretation of metabolomic experiments. Brief. Bioinform. 2013, 14, 737–744. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Furumichi, M.; Tanabe, M.; Sato, Y.; Morishima, K. KEGG: New perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2017, 45, D353–D361. [Google Scholar] [CrossRef] [PubMed]
- Caspi, R.; Billington, R.; Ferrer, L.; Foerster, H.; Fulcher, C.A.; Keseler, I.M.; Kothari, A.; Krummenacker, M.; Latendresse, M.; Mueller, L.A.; et al. The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res. 2016, 44, D471–D480. [Google Scholar] [CrossRef] [PubMed]
- Xia, J.; Broadhurst, D.I.; Wilson, M.; Wishart, D.S. Translational biomarker discovery in clinical metabolomics: An introductory tutorial. Metabolomics 2013, 9, 280–299. [Google Scholar] [CrossRef] [PubMed]
- Akao, T.; Kobashi, K.; Aburada, M. Enzymic studies on the animal and intestinal bacterial metabolism of geniposide. Biol. Pharm. Bull. 1994, 17, 1573–1576. [Google Scholar] [CrossRef] [PubMed]
- Cao, H.; Feng, Q.; Xu, W.; Li, X.; Kang, Z.; Ren, Y.; Du, L. Genipin induced apoptosis associated with activation of the c-Jun NH2-terminal kinase and p53 protein in hela cells. Biol. Pharm. Bull. 2010, 33, 1343–1348. [Google Scholar] [CrossRef] [PubMed]
- Nam, K.N.; Choi, Y.-S.; Jung, H.-J.; Park, G.H.; Park, J.-M.; Moon, S.-K.; Cho, K.-H.; Kang, C.; Kang, I.; Oh, M.S.; et al. Genipin inhibits the inflammatory response of rat brain microglial cells. Int. Immunopharmacol. 2010, 10, 493–499. [Google Scholar] [CrossRef] [PubMed]
- Miyagoshi, M.; Amagaya, S.; Ogihara, Y. Choleretic Actions of Iridoid Compounds. J. Pharmacobiodyn. 1988, 11, 186–190. [Google Scholar] [CrossRef] [PubMed]
- Tallent, W.H. Two new antibiotic cyclopentanoid monoterpenes of plant origin. Tetrahedron 1964, 20, 1781–1787. [Google Scholar] [CrossRef]
- Ueda, S.; Iwahashi, Y.; Tokuda, H. Production of Anti-Tumor-Promoting Iridoid Glucosides in Genipa americana and Its Cell Cultures. J. Nat. Prod. 1991, 54, 1677–1680. [Google Scholar] [CrossRef] [PubMed]
- Abadio Finco, F.D.B.; Böser, S.; Graeve, L. Antiproliferative activity of bacaba (Oenocarpus bacaba) and jenipapo (Genipa americana L.) phenolic extracts: A comparison of assays. Nutr. Food Sci. 2013, 43, 98–106. [Google Scholar] [CrossRef]
- Baeza, G.; Sarriá, B.; Mateos, R.; Bravo, L. Dihydrocaffeic acid, a major microbial metabolite of chlorogenic acids, shows similar protective effect than a yerba mate phenolic extract against oxidative stress in HepG2 cells. Food Res. Int. 2016, 87, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Moon, J.-H.; Terao, J. Antioxidant activity of caffeic acid and dihydrocaffeic acid in lard and human low-density lipoprotein. J. Agric. Food Chem. 1998, 46, 5062–5065. [Google Scholar] [CrossRef]
- Gröber, U.; Reichrath, J.; Holick, M.; Kisters, K. Vitamin K: An old vitamin in a new perspective. Dermatoendocrinol. 2014, 6, e968490. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Sinclair, A.J.; Faiza, M.; Li, D.; Han, X.; Yin, H.; Wang, Y. Furan fatty acids—Beneficial or harmful to health? Prog. Lipid Res. 2017, 68, 119–137. [Google Scholar] [CrossRef] [PubMed]
- Xu, T.; Fang, Y.; Rong, A.; Wang, J. Flexible combination of multiple diagnostic biomarkers to improve diagnostic accuracy. BMC Med. Res. Methodol. 2015, 15, 94. [Google Scholar] [CrossRef] [PubMed]

| Meal | Food |
|---|---|
| Breakfast | Bread with butter, or bread with cheese, or bread with cheese and ham, or toast with butter, or cream crackers + yogurt * |
| Snack | Cream crackers + yogurt * |
| Lunch | Rice or pasta + beans + grilled (or roasted) chicken or meat + mashed potatoes Dessert: gelatin |
| Snack | Sandwich without salad + yogurt or other beverage * |
| Dinner | Pasta |
| Time Range | Metabolites | AUC | p-Value (t-Test) | Cut-Off Value |
|---|---|---|---|---|
| All times | 1R,6R-6-Hydroxy-2-succinylcyclohexa-2,4-diene-1-carboxylate | 1 | 2.0454 × 10−21 | 5 |
| Hydroxyhydrocinnamic acid | 1 | 5.7497 × 10−28 | 2.64 | |
| multiplex pred | 1 | 6.8817 × 10−57 | 0.665 | |
| 3,4-dihydroxyphenylacetate | 0.99593 | 1.35 × 10−14 | 0.63 | |
| 3(7-dehydro)genipinic acid | 0.98008 | 7.1576 × 10−14 | 4.3 | |
| Nonate | 0.97737 | 3.9231 × 10−17 | 2.47 | |
| 12-demethylated-8-hydroxygenipinic acid | 0.95292 | 5.7703 × 10−7 | 0.00878 | |
| Dihydroxyhydrocinnamic acid | 0.93074 | 3.0562 × 10−18 | 37.8 | |
| Genipic acid | 0.85785 | 1.0072 × 10−9 | 0.322 | |
| Genipic acid glucuronide | 0.74423 | 0.55998 | 0.218 | |
| 0 to 6 h | Dihydroxyhydrocinnamic acid | 1 | 1.9091 × 10−14 | 48.9 |
| 1R,6R-6-Hydroxy-2-succinylcyclohexa-2,4-diene-1-carboxylate | 1 | 9.9186 × 10−10 | 3.95 | |
| Hydroxyhydrocinnamic acid | 1 | 7.7957 × 10−12 | 3.56 | |
| Nonate | 1 | 8.2316 × 10−9 | 5.85 | |
| 3,4-dihydroxyphenylacetate | 1 | 1.6602 × 10−7 | 0.244 | |
| multiplex pred | 1 | 1.3766 × 10−22 | 0.633 | |
| Genipic acid | 0.98333 | 7.4173 × 10−8 | 0.342 | |
| 3(7-dehydro)genipinic acid | 0.95 | 5.951 × 10−7 | 4.14 | |
| 12-demethylated-8-hydroxygenipinic acid | 0.9375 | 2.574 × 10−4 | 0.0159 | |
| Genipic acid glucuronide | 0.93333 | 2.0461 × 10−4 | 0.17 | |
| 6 to 12 h | 1R,6R-6-Hydroxy-2-succinylcyclohexa-2,4-diene-1-carboxylate | 1 | 6.1416 × 10−9 | 4.83 |
| Hydroxyhydrocinnamic acid | 1 | 1.7482 × 10−9 | 2.64 | |
| 12-Demethylated-8-hydroxygenipinic acid | 1 | 6.9629 × 10−5 | 0.00947 | |
| Nonate | 1 | 9.5177 × 10−6 | 3.09 | |
| 3,4-dihydroxyphenylacetate | 1 | 1.1069 × 10−4 | 0.892 | |
| multiplex pred | 1 | 2.2156 × 10−21 | 0.486 | |
| 3(7-dehydro)genipinic acid | 0.97917 | 1.3988 × 10−5 | 3.5 | |
| Dihydroxyhydrocinnamic acid | 0.93333 | 3.7546 × 10−6 | 36.8 | |
| Genipic acid | 0.77083 | 0.0073173 | 0.322 | |
| Genipic acid glucuronide | 0.7625 | 0.92915 | 0.374 | |
| 12 to 24 h | 1R,6R-6-Hydroxy-2-succinylcyclohexa-2,4-diene-1-carboxylate | 1 | 6.279 × 10−6 | 6.12 |
| Hydroxyhydrocinnamic acid | 1 | 2.1664 × 10−10 | 3.85 | |
| 3,4-dihydroxyphenylacetate | 1 | 9.8555 × 10−6 | 0.63 | |
| multiplex pred | 1 | 1.4956 × 10−16 | 0.665 | |
| 3(7-dehydro)genipinic acid | 0.99219 | 1.2466 × 10−5 | 1.74 | |
| 12-demethylated-8-hydroxygenipinic acid | 0.92929 | 0.002416 | 0.0106 | |
| Nonate | 0.92578 | 1.7992 × 10−5 | 7.13 | |
| Dihydroxyhydrocinnamic acid | 0.86528 | 1.8911 × 10−4 | 50 | |
| Genipic acid | 0.83984 | 8.338 × 10−4 | 0.318 | |
| Genipic acid glucuronide | 0.55469 | 0.22484 | 0.242 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dickson, L.; Tenon, M.; Svilar, L.; Fança-Berthon, P.; Lugan, R.; Martin, J.-C.; Vaillant, F.; Rogez, H. Main Human Urinary Metabolites after Genipap (Genipa americana L.) Juice Intake. Nutrients 2018, 10, 1155. https://doi.org/10.3390/nu10091155
Dickson L, Tenon M, Svilar L, Fança-Berthon P, Lugan R, Martin J-C, Vaillant F, Rogez H. Main Human Urinary Metabolites after Genipap (Genipa americana L.) Juice Intake. Nutrients. 2018; 10(9):1155. https://doi.org/10.3390/nu10091155
Chicago/Turabian StyleDickson, Livia, Mathieu Tenon, Ljubica Svilar, Pascale Fança-Berthon, Raphael Lugan, Jean-Charles Martin, Fabrice Vaillant, and Hervé Rogez. 2018. "Main Human Urinary Metabolites after Genipap (Genipa americana L.) Juice Intake" Nutrients 10, no. 9: 1155. https://doi.org/10.3390/nu10091155
APA StyleDickson, L., Tenon, M., Svilar, L., Fança-Berthon, P., Lugan, R., Martin, J.-C., Vaillant, F., & Rogez, H. (2018). Main Human Urinary Metabolites after Genipap (Genipa americana L.) Juice Intake. Nutrients, 10(9), 1155. https://doi.org/10.3390/nu10091155





