Serological Screening for Coronavirus Infections in Cats
Abstract
:1. Introduction
2. Materials and Methods
2.1. Serum Samples
2.2. Cells and Viruses
2.3. Plasmids Constructs and Recombinant Protein Expression
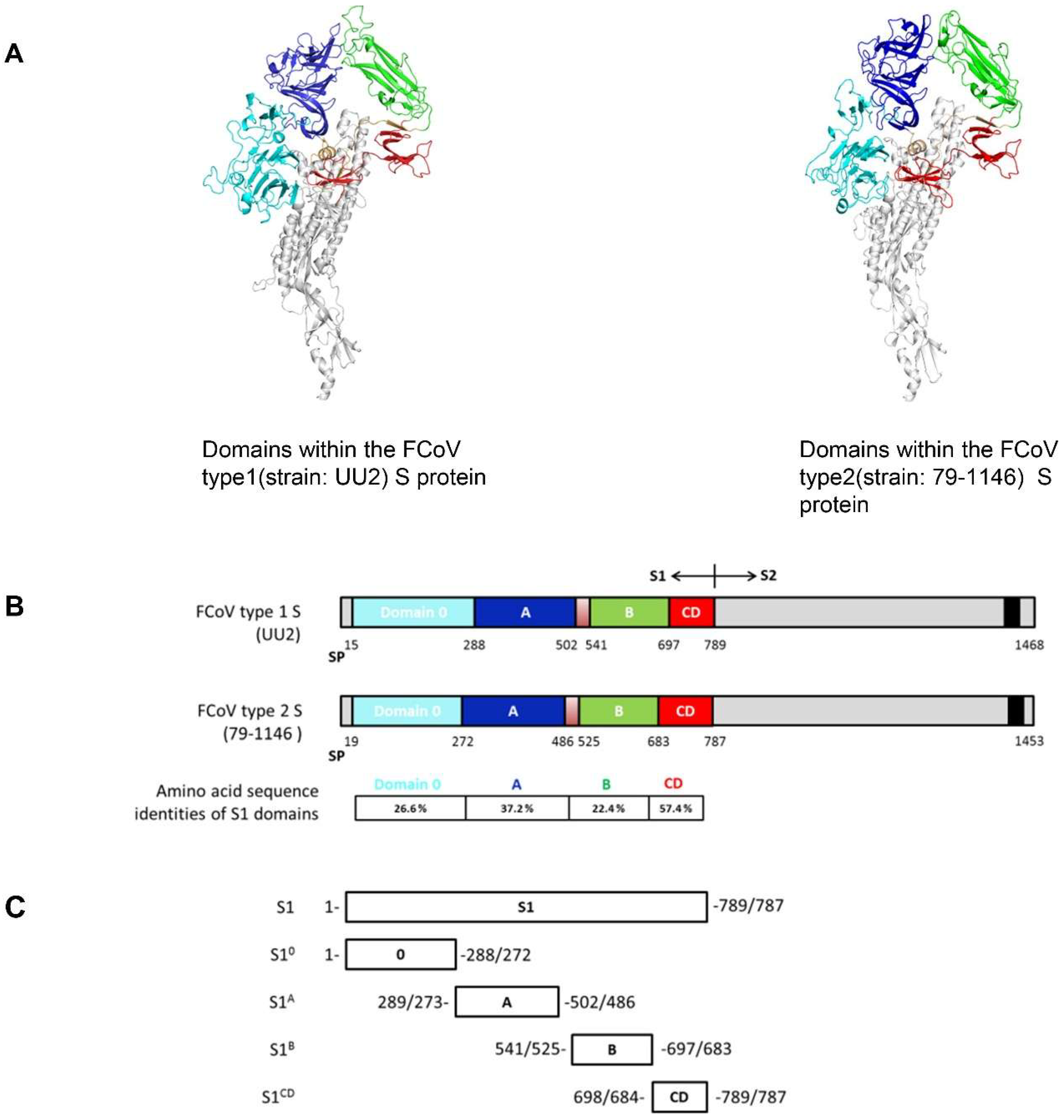
2.4. FCoV S Structure Modelling and S1 Domain Expression
2.5. Enzyme-Linked Immunosorbent Assay (ELISA)
2.6. Virus Neutralization Assay
3. Results
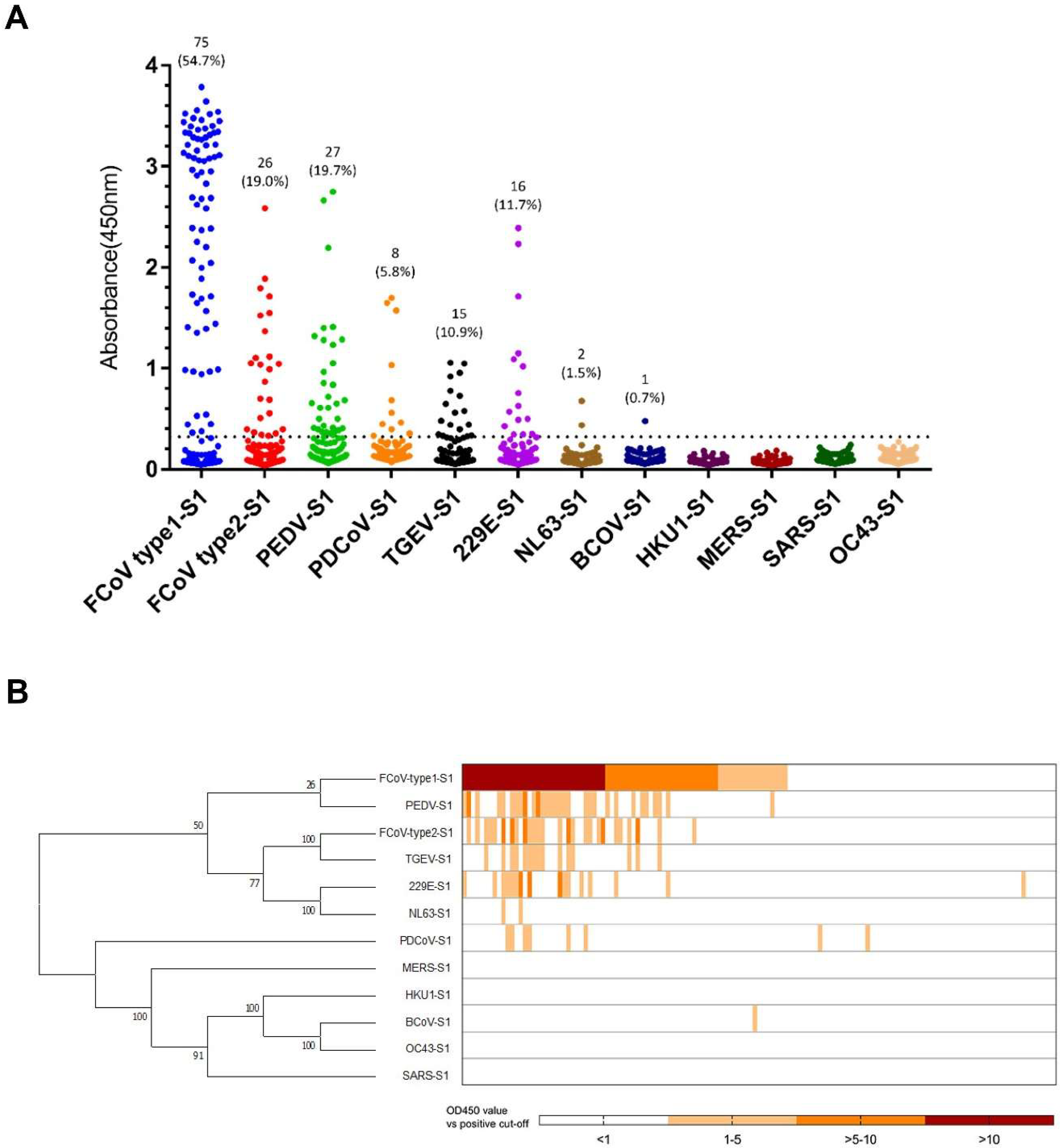
3.1. CoVs Seroprevalence in Cats
3.2. Assessment of Cross-Reactivity for CoV S1s
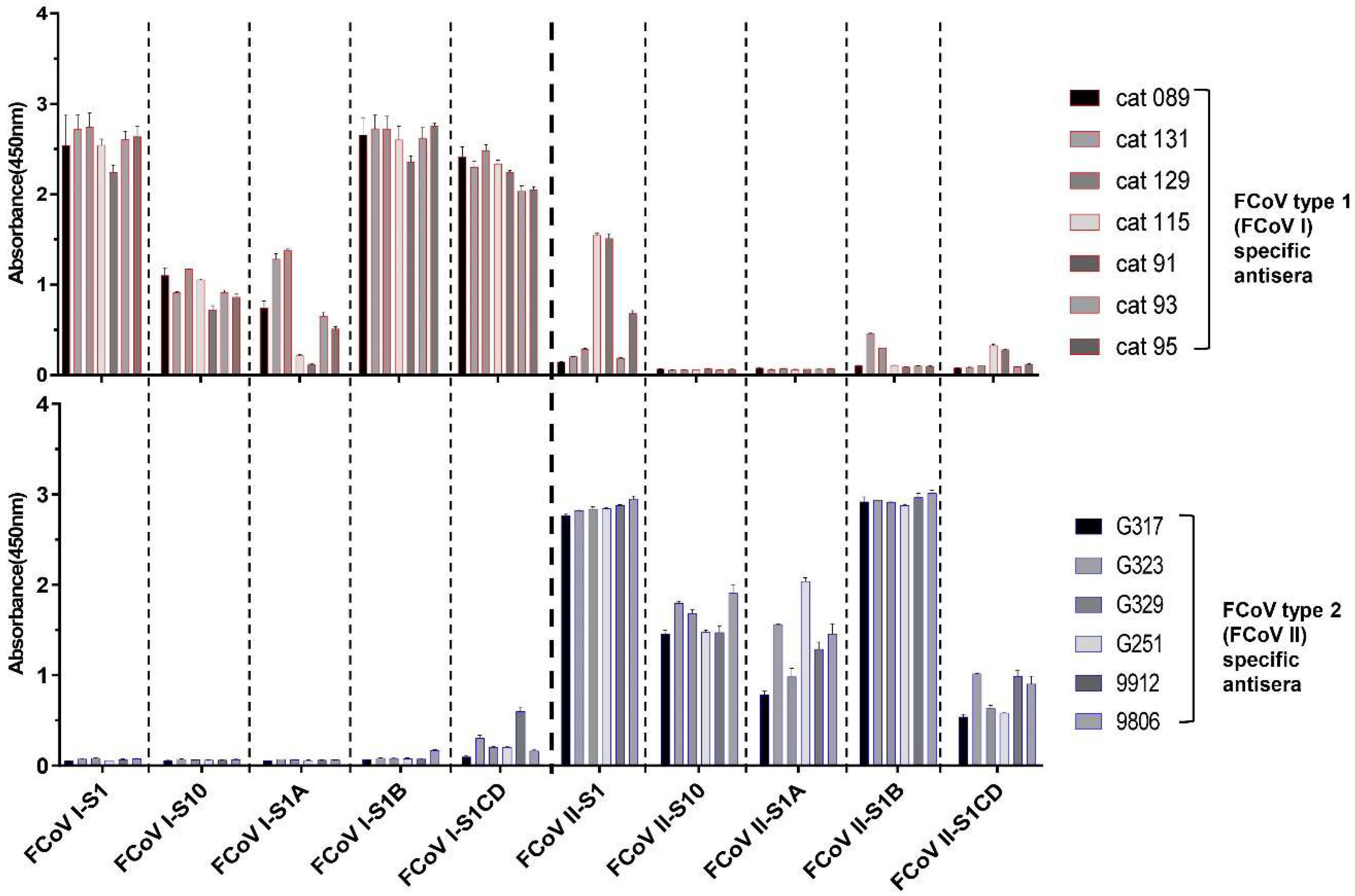
3.3. S1 Domain Mapping of Conserved Epitopes Shared between Type 1 and Type 2 Feline Coronaviruses
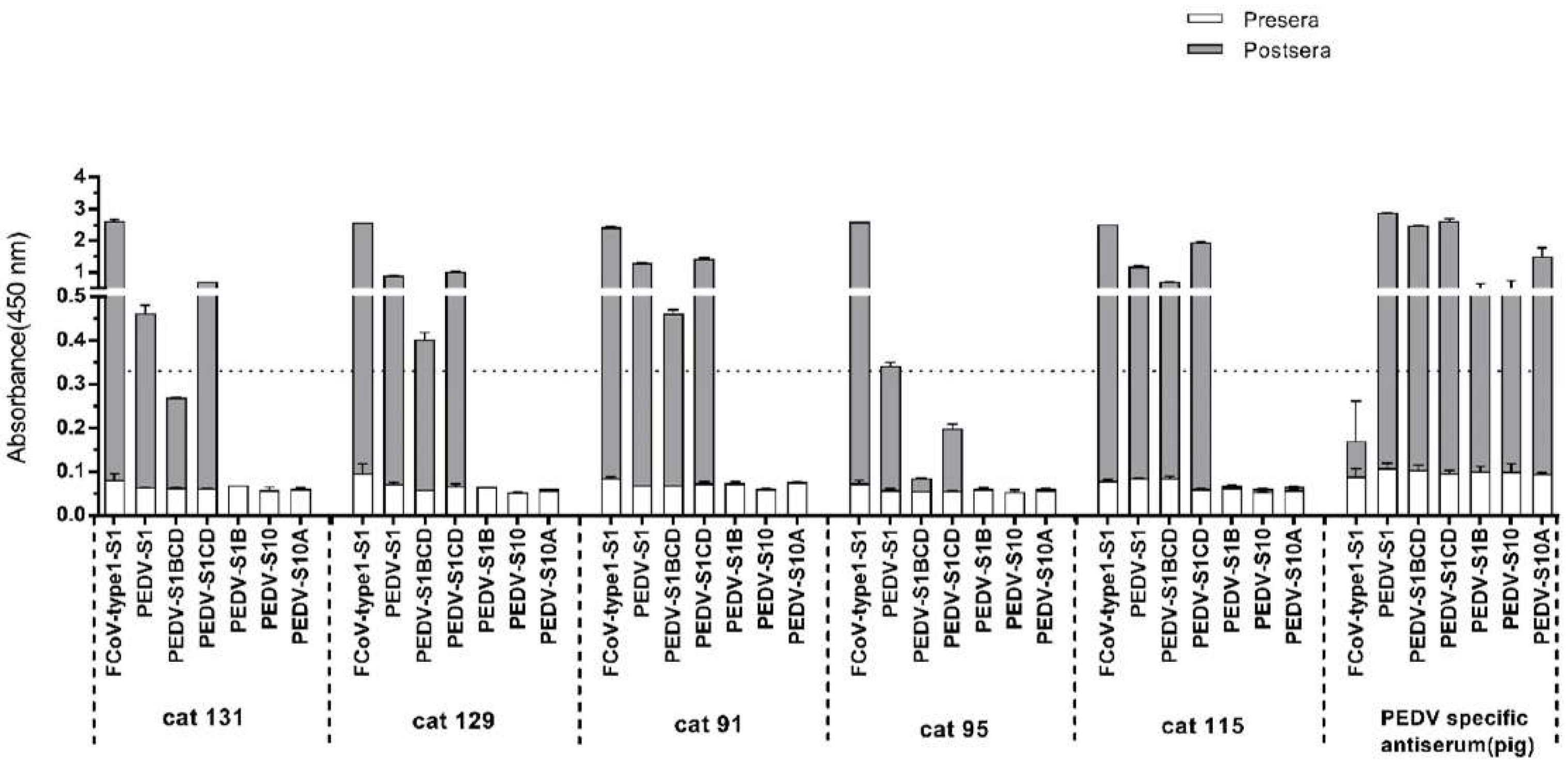
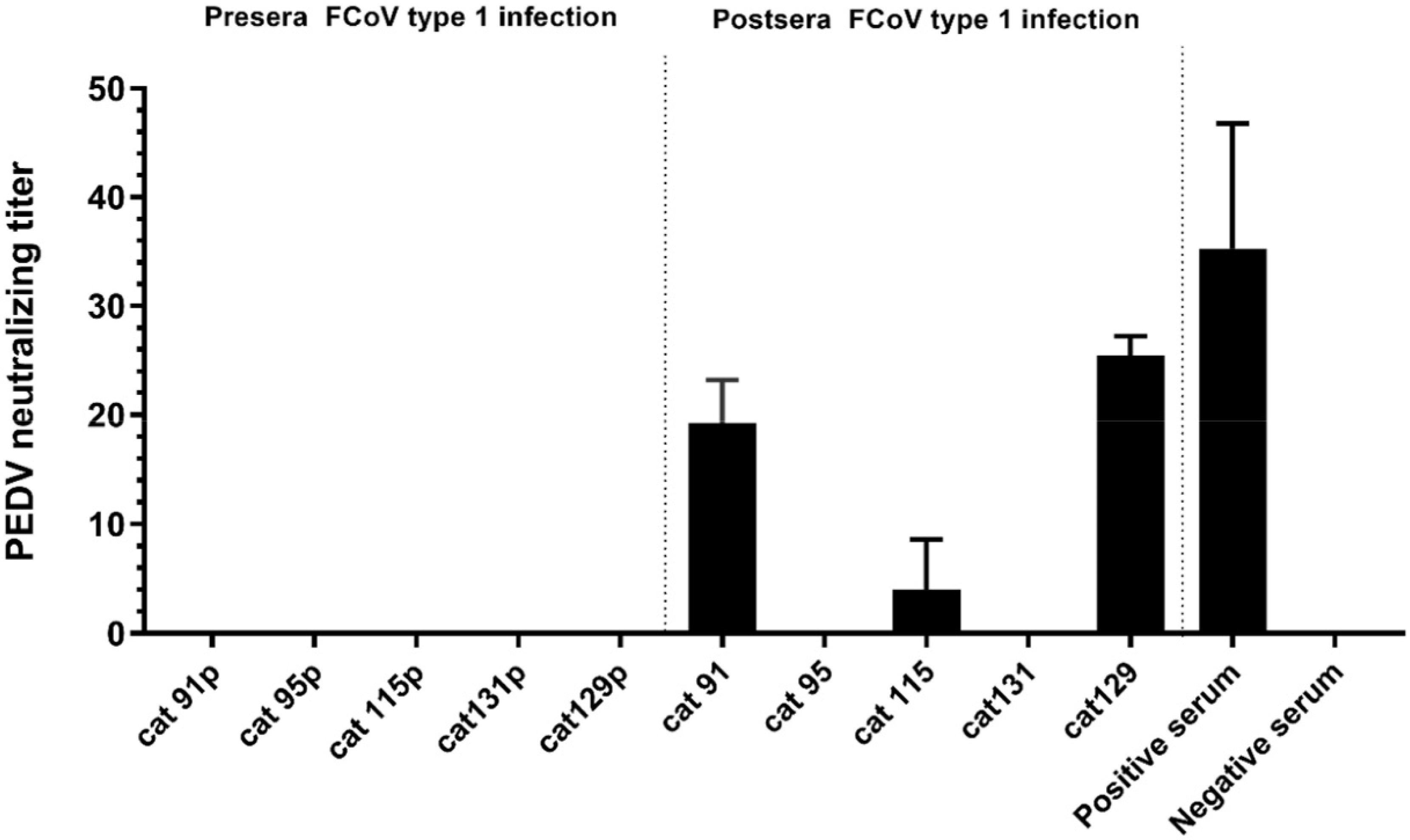
3.4. Assessment of Cross-Reactivity between FCoV Type 1 and PEDV
3.5. Evaluation of Virus Neutralization Capacity of HCoV-229E and PDCoV Positive Serum Samples
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Kirchdoerfer, R.N.; Cottrell, C.A.; Wang, N.; Pallesen, J.; Yassine, H.M.; Turner, H.L.; Corbett, K.S.; Graham, B.S.; McLellan, J.S.; Ward, A.B. Pre-fusion structure of a human coronavirus spike protein. Nature 2016, 531, 118–121. [Google Scholar] [CrossRef] [Green Version]
- Woo, P.C.; Lau, S.K.; Lam, C.S.; Lau, C.C.; Tsang, A.K.; Lau, J.H.; Bai, R.; Teng, J.L.; Tsang, C.C.; Wang, M.; et al. Discovery of seven novel Mammalian and avian coronaviruses in the genus deltacoronavirus supports bat coronaviruses as the gene source of alphacoronavirus and betacoronavirus and avian coronaviruses as the gene source of gammacoronavirus and deltacoronavirus. J. Virol. 2012, 86, 3995–4008. [Google Scholar] [CrossRef]
- Walls, A.C.; Tortorici, M.A.; Bosch, B.J.; Frenz, B.; Rottier, P.J.M.; DiMaio, F.; Rey, F.A.; Veesler, D. Cryo-electron microscopy structure of a coronavirus spike glycoprotein trimer. Nature 2016, 531, 114–117. [Google Scholar] [CrossRef] [Green Version]
- De Wit, E.; van Doremalen, N.; Falzarano, D.; Munster, V.J. SARS and MERS: recent insights into emerging coronaviruses. Nat. Rev. Microbiol. 2016, 14, 523. [Google Scholar] [CrossRef]
- Bolles, M.; Donaldson, E.; Baric, R. SARS-CoV and emergent coronaviruses: viral determinants of interspecies transmission. Curr. Opin. Virol. 2011, 1, 624–634. [Google Scholar] [CrossRef]
- Zumla, A.; Hui, D.S.; Perlman, S. Middle East respiratory syndrome. Lancet 2015, 386, 995–1007. [Google Scholar] [CrossRef] [Green Version]
- Al-Tawfiq, J.A.; Memish, Z.A. Middle East respiratory syndrome coronavirus: transmission and phylogenetic evolution. Trends Microbiol. 2014, 22, 573–579. [Google Scholar] [CrossRef] [Green Version]
- Corman, V.M.; Eckerle, I.; Memish, Z.A.; Liljander, A.M.; Dijkman, R.; Jonsdottir, H.; Juma Ngeiywa, K.J.; Kamau, E.; Younan, M.; Al Masri, M.; et al. Link of a ubiquitous human coronavirus to dromedary camels. Proc. Natl. Acad. Sci. USA 2016, 113, 9864–9869. [Google Scholar] [CrossRef] [Green Version]
- Corman, V.M.; Baldwin, H.J.; Tateno, A.F.; Zerbinati, R.M.; Annan, A.; Owusu, M.; Nkrumah, E.E.; Maganga, G.D.; Oppong, S.; Adu-Sarkodie, Y.; et al. Evidence for an Ancestral Association of Human Coronavirus 229E with Bats. J. Virol. 2015, 89, 11858–11870. [Google Scholar] [CrossRef] [Green Version]
- Vijgen, L.; Keyaerts, E.; Lemey, P.; Maes, P.; Van Reeth, K.; Nauwynck, H.; Pensaert, M.; Van Ranst, M. Evolutionary history of the closely related group 2 coronaviruses: porcine hemagglutinating encephalomyelitis virus, bovine coronavirus, and human coronavirus OC43. J. Virol. 2006, 80, 7270–7274. [Google Scholar] [CrossRef]
- Tao, Y.; Shi, M.; Chommanard, C.; Queen, K.; Zhang, J.; Markotter, W.; Kuzmin, I.V.; Holmes, E.C.; Tong, S. Surveillance of Bat Coronaviruses in Kenya Identifies Relatives of Human Coronaviruses NL63 and 229E and Their Recombination History. J. Virol. 2017, 91, e01953-16. [Google Scholar] [CrossRef] [Green Version]
- Perlman, S.; Netland, J. Coronaviruses post-SARS: Update on replication and pathogenesis. Nat. Rev. Microbiol. 2009, 7, 439–450. [Google Scholar] [CrossRef]
- Le Poder, S. Feline and canine coronaviruses: Common genetic and pathobiological features. Adv. Virol. 2011, 2011, 609465. [Google Scholar] [CrossRef]
- Katz, J.M.; Hancock, K.; Xu, X. Serologic assays for influenza surveillance, diagnosis and vaccine evaluation. Expert Rev. Anti. Infect. 2011, 9, 669–683. [Google Scholar] [CrossRef] [Green Version]
- Hohdatsu, T.; Okada, S.; Ishizuka, Y.; Yamada, H.; Koyama, H. The prevalence of types I and II feline coronavirus infections in cats. J. Vet. Med. Sci. 1992, 54, 557–562. [Google Scholar] [CrossRef]
- Addie, D.; Belak, S.; Boucraut-Baralon, C.; Egberink, H.; Frymus, T.; Gruffydd-Jones, T.; Hartmann, K.; Hosie, M.J.; Lloret, A.; Lutz, H.; et al. Feline infectious peritonitis. ABCD guidelines on prevention and management. J. Feline Med. Surg. 2009, 11, 594–604. [Google Scholar] [CrossRef]
- Shiba, N.; Maeda, K.; Kato, H.; Mochizuki, M.; Iwata, H. Differentiation of feline coronavirus type I and II infections by virus neutralization test. Vet. Microbiol 2007, 124, 348–352. [Google Scholar] [CrossRef]
- Herrewegh, A.A.P.M.; Smeenk, I.; Horzinek, M.C.; Rottier, P.J.M.; de Groot, R.J. Feline coronavirus type II strains 79-1683 and 79-1146 originate from a double recombination between feline coronavirus type I and canine coronavirus. J. Virol. 1998, 72, 4508–4514. [Google Scholar]
- Terada, Y.; Matsui, N.; Noguchi, K.; Kuwata, R.; Shimoda, H.; Soma, T.; Mochizuki, M.; Maeda, K. Emergence of Pathogenic Coronaviruses in Cats by Homologous Recombination between Feline and Canine Coronaviruses. PLoS ONE 2014, 9, e106534. [Google Scholar] [CrossRef]
- Pedersen, N.C. An update on feline infectious peritonitis: Virology and immunopathogenesis. Vet. J. 2014, 201, 123–132. [Google Scholar] [CrossRef] [Green Version]
- Tresnan, D.B.; Levis, R.; Holmes, K.V. Feline aminopeptidase N serves as a receptor for feline, canine, porcine, and human coronaviruses in serogroup I. J. Virol. 1996, 70, 8669–8674. [Google Scholar] [Green Version]
- Dye, C.; Temperton, N.; Siddell, S.G. Type I feline coronavirus spike glycoprotein fails to recognize aminopeptidase N as a functional receptor on feline cell lines. J. Gen. Virol. 2007, 88, 1753. [Google Scholar] [CrossRef]
- Barlough, J.E.; Stoddart, C.A.; Sorresso, G.P.; Jacobson, R.H.; Scott, F.W. Experimental inoculation of cats with canine coronavirus and subsequent challenge with feline infectious peritonitis virus. Lab. Anim. Sci. 1984, 34, 592–597. [Google Scholar]
- Barlough, J.E.; Johnson-Lussenburg, C.M.; Stoddart, C.A.; Jacobson, R.H.; Scott, F.W. Experimental inoculation of cats with human coronavirus 229E and subsequent challenge with feline infectious peritonitis virus. Can. J. Comp. Med. 1985, 49, 303–307. [Google Scholar]
- Tresnan, D.B.; Holmes, K.V. Feline aminopeptidase N is a receptor for all group I coronaviruses. In Coronaviruses and Arteriviruses; Springer: Boston, MA, USA, 1998; Volume 440, pp. 69–75. [Google Scholar]
- Wang, Y.T.; Chueh, L.L.; Wan, C.H. An eight-year epidemiologic study based on baculovirus-expressed type-specific spike proteins for the differentiation of type I and II feline coronavirus infections. BMC Vet. Res. 2014, 10, 186. [Google Scholar] [CrossRef]
- Tekelioglu, B.K.; Berriatua, E.; Turan, N.; Helps, C.R.; Kocak, M.; Yilmaz, H. A retrospective clinical and epidemiological study on feline coronavirus (FCoV) in cats in Istanbul, Turkey. Prev. Vet. Med. 2015, 119, 41–47. [Google Scholar] [CrossRef]
- Kummrow, M.; Meli, M.L.; Haessig, M.; Goenczi, E.; Poland, A.; Pedersen, N.C.; Hofmann-Lehmann, R.; Lutz, H. Feline coronavirus serotypes 1 and 2: Seroprevalence and association with disease in Switzerland. Clin. Diagn. Lab. Immunol. 2005, 12, 1209–1215. [Google Scholar] [CrossRef]
- Hulswit, R.; de Haan, C.; Bosch, B.-J. Coronavirus spike protein and tropism changes. In Advances in Virus Research; Elsevier: Amsterdam, The Netherlands, 2016; Volume 96, pp. 29–57. [Google Scholar]
- Belouzard, S.; Millet, J.K.; Licitra, B.N.; Whittaker, G.R. Mechanisms of coronavirus cell entry mediated by the viral spike protein. Viruses 2012, 4, 1011–1033. [Google Scholar] [CrossRef]
- Reusken, C.; Mou, H.; Godeke, G.J.; van der Hoek, L.; Meyer, B.; Muller, M.A.; Haagmans, B.; de Sousa, R.; Schuurman, N.; Dittmer, U.; et al. Specific serology for emerging human coronaviruses by protein microarray. Eurosurveillance 2013, 18, 20441. [Google Scholar] [CrossRef] [Green Version]
- Leung, G.M.; Chung, P.-H.; Tsang, T.; Lim, W.; Chan, S.K.; Chau, P.; Donnelly, C.A.; Ghani, A.C.; Fraser, C.; Riley, S. SARS-CoV antibody prevalence in all Hong Kong patient contacts. Emerg. Infect. Dis. 2004, 10, 1653. [Google Scholar] [CrossRef]
- Reusken, C.B.; Haagmans, B.L.; Muller, M.A.; Gutierrez, C.; Godeke, G.J.; Meyer, B.; Muth, D.; Raj, V.S.; Smits-De Vries, L.; Corman, V.M.; et al. Middle East respiratory syndrome coronavirus neutralising serum antibodies in dromedary camels: A comparative serological study. Lancet Infect. Dis. 2013, 13, 859–866. [Google Scholar] [CrossRef]
- Dortmans, J.C.F.M.; Li, W.; van der Wolf, P.J.; Buter, G.J.; Franssen, P.J.M.; van Schaik, G.; Houben, M.; Bosch, B.J. Porcine epidemic diarrhea virus (PEDV) introduction into a naive Dutch pig population in 2014. Vet. Microbiol. 2018, 221, 13–18. [Google Scholar] [CrossRef] [Green Version]
- Haijema, B.J.; Volders, H.; Rottier, P.J.M. Live, attenuated coronavirus vaccines through the directed deletion of group-specific genes provide protection against feline infectious peritonitis. J. Virol. 2004, 78, 3863–3871. [Google Scholar] [CrossRef]
- Vogel, L.; Van der Lubben, M.; te Lintelo, E.G.; Bekker, C.P.; Geerts, T.; Schuijff, L.S.; Grinwis, G.C.; Egberink, H.F.; Rottier, P.J. Pathogenic characteristics of persistent feline enteric coronavirus infection in cats. Vet. Res. 2010, 41, 71. [Google Scholar] [CrossRef]
- Li, W.; Hulswit, R.J.G.; Kenney, S.P.; Widjaja, I.; Jung, K.; Alhamo, M.A.; van Dieren, B.; van Kuppeveld, F.J.M.; Saif, L.J.; Bosch, B.J. Broad receptor engagement of an emerging global coronavirus may potentiate its diverse cross-species transmissibility. Proc. Natl Acad. Sci. USA 2018, 115, E5135–E5143. [Google Scholar] [CrossRef] [Green Version]
- Li, C.H.; Li, Z.; Zou, Y.; Wicht, O.; van Kuppeveld, F.J.M.; Rottier, P.J.M.; Bosch, B.J. Manipulation of the Porcine Epidemic Diarrhea Virus Genome Using Targeted RNA Recombination. PLoS ONE 2013, 8, e69997. [Google Scholar] [CrossRef]
- Wicht, O.; Li, W.; Willems, L.; Meuleman, T.J.; Wubbolts, R.W.; van Kuppeveld, F.J.; Rottier, P.J.; Bosch, B.J. Proteolytic activation of the porcine epidemic diarrhea coronavirus spike fusion protein by trypsin in cell culture. J. Virol. 2014, 88, 7952–7961. [Google Scholar] [CrossRef]
- Li, C.; Li, W.; Lucio de Esesarte, E.; Guo, H.; van den Elzen, P.; Aarts, E.; van den Born, E.; Rottier, P.J.M.; Bosch, B.J. Cell Attachment Domains of the Porcine Epidemic Diarrhea Virus Spike Protein Are Key Targets of Neutralizing Antibodies. J. Virol. 2017, 91, e00273-17. [Google Scholar] [CrossRef] [Green Version]
- Li, W.; Hulswit, R.J.G.; Widjaja, I.; Raj, V.S.; McBride, R.; Peng, W.; Widagdo, W.; Tortorici, M.A.; van Dieren, B.; Lang, Y.; et al. Identification of sialic acid-binding function for the Middle East respiratory syndrome coronavirus spike glycoprotein. Proc. Natl Acad. Sci. USA 2017, 114, E8508–E8517. [Google Scholar] [CrossRef] [Green Version]
- Biasini, M.; Bienert, S.; Waterhouse, A.; Arnold, K.; Studer, G.; Schmidt, T.; Kiefer, F.; Cassarino, T.G.; Bertoni, M.; Bordoli, L.; et al. SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. 2014, 42, W252–W258. [Google Scholar] [CrossRef]
- Hohdatsu, T.; Okada, S.; Koyama, H. Characterization of Monoclonal-Antibodies against Feline Infectious Peritonitis Virus Type-II and Antigenic Relationship between Feline, Porcine, and Canine Coronaviruses. Arch. Virol. 1991, 117, 85–95. [Google Scholar] [CrossRef]
- Pedersen, N.C. A review of feline infectious peritonitis virus infection: 1963–2008. J. Feline Med. Surg. 2009, 11, 225–258. [Google Scholar] [CrossRef]
- Bell, E.T.; Toribio, J.A.; White, J.D.; Malik, R.; Norris, J.M. Seroprevalence study of feline coronavirus in owned and feral cats in Sydney, Australia. Aust. Vet. J. 2006, 84, 74–81. [Google Scholar] [CrossRef]
- An, D.J.; Jeoung, H.Y.; Jeong, W.; Park, J.Y.; Lee, M.H.; Park, B.K. Prevalence of Korean cats with natural feline coronavirus infections. Virol. J. 2011, 8, 455. [Google Scholar] [CrossRef]
- Addie, D.D.; Schaap, I.A.; Nicolson, L.; Jarrett, O. Persistence and transmission of natural type I feline coronavirus infection. J. Gen. Virol. 2003, 84, 2735–2744. [Google Scholar] [CrossRef]
- Chang, H.W.; Egberink, H.F.; Halpin, R.; Spiro, D.J.; Rottier, P.J. Spike protein fusion peptide and feline coronavirus virulence. Emerg. Infect. Dis. 2012, 18, 1089–1095. [Google Scholar] [CrossRef]
- Okda, F.A.; Lawson, S.; Singrey, A.; Nelson, J.; Hain, K.S.; Joshi, L.R.; Christopher-Hennings, J.; Nelson, E.A.; Diel, D.G. The S2 glycoprotein subunit of porcine epidemic diarrhea virus contains immunodominant neutralizing epitopes. Virology 2017, 509, 185–194. [Google Scholar] [CrossRef]
- Decaro, N.; Mari, V.; Campolo, M.; Lorusso, A.; Camero, M.; Elia, G.; Martella, V.; Cordioli, P.; Enjuanes, L.; Buonavoglia, C. Recombinant canine coronaviruses related to transmissible gastroenteritis virus of Swine are circulating in dogs. J. Virol. 2009, 83, 1532–1537. [Google Scholar] [CrossRef]
- Truong, Q.L.; Seo, T.W.; Yoon, B.I.; Kim, H.C.; Han, J.H.; Hahn, T.W. Prevalence of swine viral and bacterial pathogens in rodents and stray cats captured around pig farms in Korea. J. Vet. Med. Sci. 2013, 75, 1647–1650. [Google Scholar] [CrossRef]
- Kandeil, A.; Gomaa, M.; Shehata, M.; El-Taweel, A.; Kayed, A.E.; Abiadh, A.; Jrijer, J.; Moatasim, Y.; Kutkat, O.; Bagato, O.; et al. Middle East respiratory syndrome coronavirus infection in non-camelid domestic mammals. Emerg. Microbes. Infect. 2019, 8, 103–108. [Google Scholar] [CrossRef] [Green Version]
| S1 Antigens of Different Coronaviruses | Number of Cats (Total = 137) | |||||||
|---|---|---|---|---|---|---|---|---|
| FCoV Type1-S1 | FCoV Type2-S1 | PEDV-S1 | PDCoV-S1 | TGEV-S1 | 229E-S1 | NL63-S1 | BCoV-S1 | |
| 59 | ||||||||
| 32 | ||||||||
| 6 | ||||||||
| 10 | ||||||||
| 1 | ||||||||
| 1 | ||||||||
| 2 | ||||||||
| 4 | ||||||||
| 2 | ||||||||
| 2 | ||||||||
| 1 | ||||||||
| 1 | ||||||||
| 1 | ||||||||
| 1 | ||||||||
| 1 | ||||||||
| 1 | ||||||||
| 1 | ||||||||
| 1 | ||||||||
| 1 | ||||||||
| 1 | ||||||||
| 4 | ||||||||
| 4 | ||||||||
| 75 | 26 | 27 | 8 | 15 | 16 | 2 | 1 | |
| Sequence No. | Protein (Accession No.) | % Amino Acid Sequence Similarity with Protein of Sequence No.: | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | ||
| 1 | FCoV type1-S1 (FJ938060.1) | |||||||||||
| 2 | FCoV type2-S1 (AY994055.1) | 28.5 | ||||||||||
| 3 | PEDV-S1(AOG30832.1) | 32.8 | 30.2 | |||||||||
| 4 | PDCoV-S1 (AML40825.1) | 23.1 | 26.7 | 26.5 | ||||||||
| 5 | TGEV-S1 (ABG89325.1) | 28.5 | 70.4 | 31.8 | 27.4 | |||||||
| 6 | HCoV-229E-S1 (NP_073551.1) | 30.4 | 38.6 | 34.5 | 27.7 | 37.2 | ||||||
| 7 | HCoV-NL63-S1 (YP_003767.1) | 28.4 | 30.4 | 29.6 | 26.7 | 31.7 | 51.9 | |||||
| 8 | BCoV-S1 (P15777.1) | 9.6 | 11.3 | 8.5 | 10.8 | 11.2 | 14 | 10.6 | ||||
| 9 | HCoV-HKU1-S1 (YP_173238.1) | 11 | 12.1 | 9.5 | 12.4 | 11.4 | 13.5 | 11.8 | 61.2 | |||
| 10 | MERS-S1 (YP_009047204.1) | 9.2 | 9.2 | 7 | 6.6 | 9.4 | 7.9 | 8.1 | 15.6 | 17 | ||
| 11 | SARS-S1 (AAX16192.1) | 6.5 | 8.9 | 9.1 | 8.6 | 8.9 | 11.4 | 8.5 | 17 | 17.9 | 15.5 | |
| 12 | HCoV-OC43-S1 (AAR01015.1) | 8.9 | 11.3 | 7.7 | 10.2 | 10.7 | 13.1 | 9.6 | 90.9 | 59.7 | 15.7 | 16.4 |
| Amino Acid % Identity to FCoV Type 1(UU2) | |||||
|---|---|---|---|---|---|
| S1 | S10 | S1A | S1B | S1CD | |
| FCoV type 1 (RM) | 89.3 | 88.3 | 85 | 94.9 | 91.2 |
| FCoV type 2 | 28.5 | 26.6 | 37.2 | 22.4 | 57.4 |
| TGEV | 28.5 | 24 | 38.2 | 23.1 | 57.3 |
| PEDV | 32.8 | 20.4 | 36.5 | 24.8 | 51.2 |
| HCoV-229E | 30.4 | NA | 36.5 | 21.4 | 35.6 |
| HCoV-NL63 | 28.4 | 19.3 | 37.7 | 20.6 | 44.9 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, S.; Li, W.; Schuurman, N.; van Kuppeveld, F.; Bosch, B.-J.; Egberink, H. Serological Screening for Coronavirus Infections in Cats. Viruses 2019, 11, 743. https://doi.org/10.3390/v11080743
Zhao S, Li W, Schuurman N, van Kuppeveld F, Bosch B-J, Egberink H. Serological Screening for Coronavirus Infections in Cats. Viruses. 2019; 11(8):743. https://doi.org/10.3390/v11080743
Chicago/Turabian StyleZhao, Shan, Wentao Li, Nancy Schuurman, Frank van Kuppeveld, Berend-Jan Bosch, and Herman Egberink. 2019. "Serological Screening for Coronavirus Infections in Cats" Viruses 11, no. 8: 743. https://doi.org/10.3390/v11080743
APA StyleZhao, S., Li, W., Schuurman, N., van Kuppeveld, F., Bosch, B.-J., & Egberink, H. (2019). Serological Screening for Coronavirus Infections in Cats. Viruses, 11(8), 743. https://doi.org/10.3390/v11080743






