1. Introduction
Gammaherpesviruses (GHVs) are double-stranded DNA viruses. They belong to the Herpesviridae family, which includes three subfamilies:
Alpha-,
Beta- and
Gammaherpesvirinae [
1,
2]. GHVs can infect humans, establishing a lifelong persistent infection mostly without evident clinical signs [
3]. Host immunity is supposed to eliminate the infection, but frequently the virus undergoes latency with potential reactivation. GHV reactivation is suspected during co-infections or when cell-mediated immunity is compromised. In the latter cases, the virus can cause severe diseases that can be potentially fatal [
4,
5]. Two human GHVs are known to promote tumorigenesis in humans: Epstein–Barr virus and Kaposi’s sarcoma-associated herpesvirus [
6,
7,
8,
9]. Epstein–Barr virus is commonly identified in adult human beings worldwide. The virus is normally innocuous; however, in a few cases, it can cause lymphomas, carcinomas or other types of cancer. Loss of T-cell immunity and genetic predisposition are thought to be critical risk factors for the oncogenic potential of the Epstein–Barr virus [
4,
5].
GHVs are known to infect different mammalian species, and they are reported worldwide [
10,
11,
12,
13,
14,
15,
16,
17,
18]. Novel GHVs were identified among Primates, Artiodactyla, Perissodactyla, Carnivora, Scandentia, and Eulipotyphla using PCR with panherpesvirus DNA polymerase gene primers or genus-specific glycoprotein B (gB) gene primers [
19]. In 2014, the first gammaherpesvirus (named
Felis catus gammaherpesvirus 1, FcaGHV1 [
20]) was identified in domestic cats, followed by the identification of novel GHVs in other felids (bobcats, pumas, ocelots, leopard cats) in the USA and Japan [
20,
21,
22]. Since that first identification, different epidemiological studies have shown that FcaGHV1 infection is widely endemic in domestic cats. Most epidemiological studies are based on the detection of the genus-specific glycoprotein B gene by PCR [
19]. The reported worldwide prevalence of FcaGHV1 ranges from 1.3% to 23.6% in domestic cats [
20,
22,
23,
24,
25,
26,
27]. Direct comparison between prevalence studies is difficult. Differences in prevalence may reflect variations in the studied cat population, the study inclusion criteria (such as feral free-roaming cats captured for neutering programs, feral cats housed in animal shelters or privately-owned cats) and the health status of the sampled cats. In addition, it is important to remember that the identification of FcaGHV1 DNA material does not provide information on the infection status of the animal and cannot differentiate between virus-infected cells, virion particles or free DNA [
28]. More recently, a serological assay was developed to assess the exposure rate to FcaGHV1 within a cat population [
29]. The seroprevalence of FcaGHV1 was found to be higher than the molecular prevalence detected by PCR [
29], with approximately 50% of the FcaGHV1-seropositive cats being PCR-positive [
20,
27,
29].
The pathogenic potential of FcaGHV1 in cats remains unclear. It has been shown that FcaGHV1 is more frequently identified in sick cats than in healthy cats [
30]. In addition, age, sex, and concomitant co-infections have been identified as risk factors for FcaGHV1, with some regional variations. A significant association between FcaGHV1 and feline leukemia virus (FeLV) antigenemia was identified only in one study in Singapore [
27], but this association was not confirmed recently [
31]. The prevalence of FcaGHV1 is, however, increased in cats co-infected with feline immunodeficiency virus (FIV) and FeLV [
27,
31]. In addition, an association between FIV alone and FcaGHV1 has been reported in independent studies [
23,
25,
26]. FIV is a widespread feline retrovirus sharing similarity with human immunodeficiency virus (HIV) that may cause an acquired immunodeficiency syndrome (AIDS)-like syndrome in infected cats. Additionally, FIV and HIV are two viral infections that can increase the risk of high-grade B cell lymphomas [
30,
32,
33,
34,
35]. Interestingly, 15–20% of HIV-associated lymphomas are correlated with human gammaherpesviruses (Epstein–Barr virus and Kaposi’s sarcoma-associated herpesvirus) [
32,
36]. Thus, similar to human GHVs, FcaGHV1 in cats may be a cofactor for malignant transformation in lentivirus-infected individuals.
In the present study, we (i) assessed for the first time the molecular prevalence of FcaGHV1 in cats in Switzerland; (ii) evaluated potential risk factors associated with FcaGHV1 detection, such as geographic origin (canton), the age, sex and breed of the cat and co-infections with FeLV, FIV, and the three feline hemoplasma species; (iii) evaluated the presence of FcaGHV1 in neoplastic tissues of cats with lymphoma; and (iiii) sequenced a fragment (316 nucleotides) of the glycoprotein B gene of FcaGHV1 to further elucidate the phylogenetic relationship with other FcaGHV1 isolates and assess possible genetic variation in Switzerland.
Our study assessed the molecular prevalence of FcaGHV1 and risk factors associated with this infection in Switzerland for the first time. Remarkably, FcaGHV1 prevalence was similar in cats presented to veterinarians and in stray cats. In accordance with previous studies, factors significantly associated with FcaGHV1 detection included male sex, age >3 years, nonpedigree status and co-infections with FIV and hemoplasmas. Swiss FcaGHV1 isolates revealed homogeneity and >99.7% identity to previously published FcaGHV1 sequences.
3. Results
3.1. Efficiency, Sensitivity and Linear Range of Amplification of the FcaGHV1 Real-Time qPCR Assay
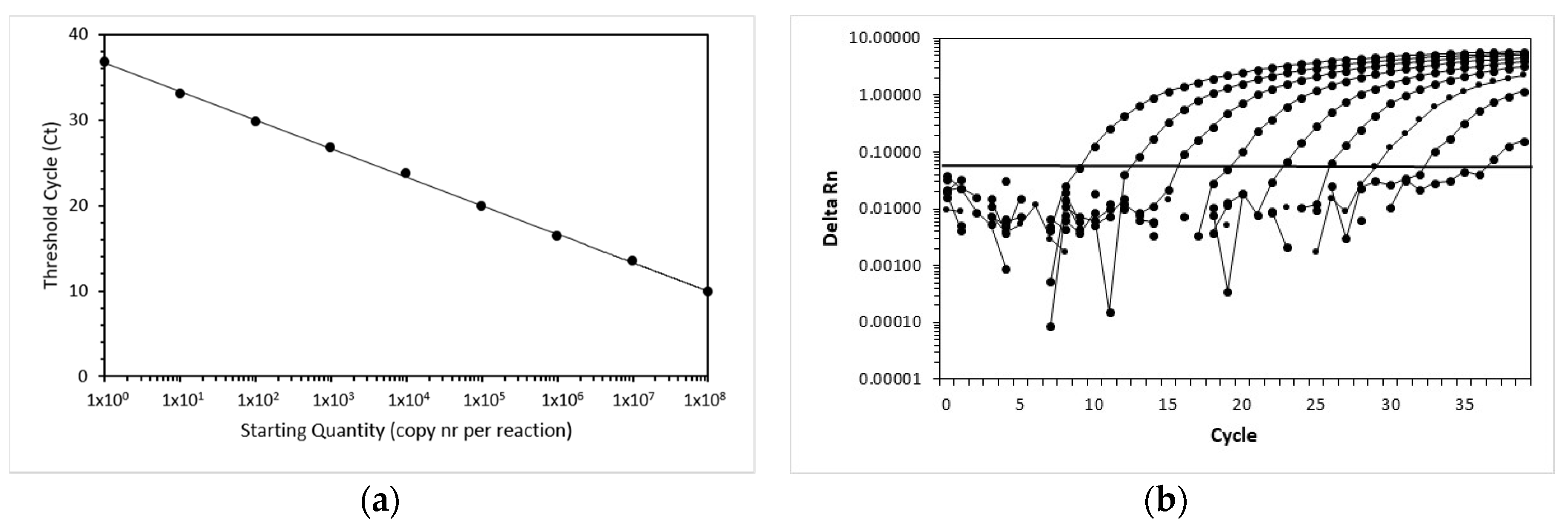
The efficiency and linear range of amplification of the FcaGHV1 real-time qPCR assay were determined using 10-fold serial dilutions of the standard. Amplification of FcaGHV1 showed linearity over the whole range of dilution (
Figure 1). The highest dilution still yielded a positive signal containing one copy per reaction. In the endpoint dilution experiment, the dilution with 1 copy yielded 8 positive reactions of 10. For the next lower dilution, none of the 8 reactions was positive. Thus, the lower limit of detection was 1 copy of template per 5 µL reaction, corresponding to 200 copies per milliliter of blood.
3.2. Sample Characteristics
In the first set of 881 samples from cats presented to veterinarians throughout all cantons of Switzerland (
Table 2), the samples originated from 160 pedigree and 479 nonpedigree cats, and no information was available for 242 samples (
Table 3). The sex and reproductive status data revealed 308 castrated male and 207 spayed female cats and 56 male and 70 female intact animals (the sex and reproductive status of 240 cats were unknown). The cats’ ages varied from 2 months to 21.5 years; the median age was 6.6 years (the age of 244 cats was unknown).
For the second set of samples collected from 91 stray cats in the canton of Jura, limited information was available: 48 cats were intact females and 37 cats were intact males; the information was unavailable for six cats. The ages of the 11 cats with available information were <1 year for five cats, 1–3 years for four cats and >3 years for four cats.
3.3. Sample Prevalence of FcaGHV1 in Swiss Domestic Cats
Of the 881 cats from all cantons of Switzerland, 53 tested as FcaGHV1-positive (6.0%; 95% CI, 4.5–7.8%). FcaGHV1 PCR-positive samples originated from 19 of the 26 Swiss cantons (
Table 2 and
Figure 2). Overall, although the molecular prevalence ranged from 0% to 16.7% in each canton, there was no significant difference in FcaGHV1 molecular prevalence among the different cantons of Switzerland (pChi2 = 0.1576;
Table 2;
Figure 2). From
Figure 2, however, it is evident that fewer FcaGHV1-positive cats were found in the Alps than in the other two major Swiss natural environments—the Swiss Plateau and the Jura region. The highest molecular prevalence of FcaGHV1 was found in the canton of Jura (16.7%; 95% CI, 4.7–37.4%), followed by the canton of Appenzell Innerrhoden (12.5%; 95% CI, 2.7–32.4%) and the canton of Ticino (12.0%; 95% CI, 4.5–24.3%;
Table 2).
We aimed to assess whether the density of the cat population in the different Swiss cantons plays a role in FcaGHV1 molecular prevalence. Since no data are available on the cat density in each Swiss canton, we used the human density as an approximation [
65]. The human population density in the three cantons showing the highest FcaGHV1 molecular prevalence (Jura, Appenzell Innerrhoden and Ticino) ranges from 87 to 129 people/km
2 [
65]. FcaGHV1 was not detected in any of the tested cats from seven cantons: Glarus, Solothurn, Neuchâtel, Sant Gallen, Thurgau, Vaud and Valais. The human population density in these cantons ranges from 59 to 341 people/km
2 [
65]. There was no statistical association between the human population density and the FcaGHV1 molecular prevalence in cats (
n = 26; pS = 0.4649;
r = 0.01818; 95% CI, −0.3822–0.4128).
Among the stray cats originating from the canton of Jura, five of 91 were FcaGHV1-positive (5.5%; 95% CI, 1.8–12.4%). There was no significant difference in the molecular prevalence of FcaGHV1 between stray cats (5.5%; 95% CI, 1.8–12.4%) and cats presented to veterinarians in the canton of Jura (4 of 24; 16.7%; 95% CI, 4.7–37.4%; pF = 0.4664).
3.4. Association of FcaGHV1 Detection with Nonpedigree Status, Male Sex and Age >3 Years
The association of FcaGHV1 detection with breed, sex and age was assessed in the 881 cats presented to veterinarians. Pedigree cats were significantly less frequently FcaGHV1-positive (2/160; 1.3%; 95% CI, 0.2–4.4%) than nonpedigree cats (39/479; 8.1%; 95% CI, 6.0–10.9%; pF = 0.0012; odds ratio (OR) 7.0; 95% CI, 1.9–29.8;
Table 3). The two pedigree cats that were FcaGHV1-positive were one Sphinx and one Maine Coon cat. There was a significant difference in sex and reproductive status between FcaGHV1-positive and negative cats (pChi2 = 0.0223;
Table 3). Male cats (intact and castrated) were more frequently FcaGHV1-infected (33/364; 9.1%; 95% CI, 6.5–12.5%) than female cats (intact and spayed; 10/277; 3.6%; 95% CI, 2.0–6.5%; pF = 0.0064; OR 2.7; 95% CI, 1.3–5.4). There was a significant difference in the age distribution between FcaGHV1-positive and negative cats (pChi2 = 0.0422;
Table 3). When the molecular prevalence in the different age groups was examined, it became evident and was confirmed by statistical analysis that cats older than 3 years were significantly more frequently FcaGHV1-positive (39/440; 8.9%; 95% CI, 6.6–11.9%) than cats less than 3 years of age (3/196; 1.5%; 95% CI, 0.4–4.4%); pF = 0.0002; OR 6.3; 95% CI, 2.0–19.5).
3.5. Association of FcaGHV1 with Other Infectious Agents
In the first set of feline samples (
n = 881), 47 samples were FeLV provirus-positive (5.3%; 95% CI, 3.9–7.0%), of which four were also FcaGHV1-positive (
Table 4); in addition, 18 samples were viremic (2.0%; 95% CI, 1.2–3.2%), of which three were also FcaGHV1-positive (
Table 4). FeLV-viremic (p27 antigen-positive) cats tended to be more frequently FcaGHV1-positive (3 of 18; 16.7%) than FeLV-aviremic cats (50 of 863; 5.8%; pF = 0.0884; OR 3.3; 95% CI, 0.97–11.1;
Table 4). There was no statistical difference between the molecular prevalence of FcaGHV1 in FeLV provirus-positive cats and that in FeLV provirus-negative cats (pF = 0.5202;
Table 4).
In addition, 104 of the 881 samples were tested for FIV, including 52 FcaGHV1-positive samples and 52 of the FcaGHV1-negative matched samples. Among the 104 tested samples, 23 were positive for FIV by Western blot analysis (22.1%; 95% CI, 14.6–31.3%). FIV-positive cats were more frequently FcaGHV1-positive (20 of 23; 87.0%) than FIV-negative cats (32 of 81; 39.5%; pF <0.0001; OR 10.2; 95% CI, 2.8–37.2;
Table 5). Overall, 106 of the 881 samples were also tested for feline hemoplasma infections, including the 53 FcaGHV1-positive and 53 matched FcaGHV1-negative samples. Among these 106 samples, 27 were positive for at least one hemoplasma species (25.5%; 95% CI, 17.5–34.9%). Cats positive for any feline hemoplasma species were significantly more frequently FcaGHV1-positive (19 of 28; 67.9%) than hemoplasma-negative cats (34 of 78; 43.6%; pF = 0.0463; OR 3.1; 95% CI, 1.2–8.0;
Table 5). The same pattern held for CMhm-positive cats, since all hemoplasma-positive cats were also positive for CMhm: cats positive for CMhm were significantly more frequently FCaGHV1-positive than CMhm-negative cats (pF = 0.0463; OR 3.1; 95% CI, 1.2–8.0). None of the cats had a single infection with Mhf or CMt. Of the 19 cats positive for both hemoplasma and FcaGHV1, four were concurrently PCR-positive for more than one hemoplasma species, including three cats co-infected with CMt and CMhm and one cat co-infected with CMhm and Mhf. All the FcaGHV1-negative cats were PCR-positive only for CMhm. None of the FcaGHV1-negative cats were PCR-positive for more than one feline hemoplasma. No blood samples of the cats presented to veterinarians tested positive for all three hemoplasma species.
In the second set of feline samples (
n = 91 stray cats from the canton of Jura), 7 samples were FeLV provirus-positive (7.7%; 95% CI, 3.1–15.2%), but none of the FeLV provirus-positive cats were also FcaGHV1 PCR-positive. No serum samples were available to test for FIV. Seventeen of the 91 whole blood samples were positive for at least one hemoplasma species (18.7%; 95% CI, 11.3-28.2%). Notably, all five FcaGHV1-positive cats were also hemoplasma-positive (
Table 6), and hemoplasma PCR-positive cats were more frequently FcaGHV1-positive (5 of 17; 29.4%) than FcaGHV1-negative (0 of 74; 0%; pF = 0.0001; OR 65.6; 95% CI 3.4–1261). Among the 91 tested samples, seven tested PCR-positive for Mhf (7.7%; 95% CI, 3.1-15.2%), 12 for CMhm (13.1%; 95% CI, 7.0-22%), and two for CMt (2.2%; CI, 0.3-7.7%). Among those samples, two were concurrently PCR-positive for all three hemoplasma species (Mhf, CMhm, and CMt). One of these two samples was also FcaGHV1 PCR-positive.
3.6. FcaGHV1 Blood Loads
The FcaGHV1 blood loads in this study ranged from 200 (the detection limit) to 3.9 × 105 copies/mL of blood. There was no significant difference in the blood loads between the samples from the 53 FcaGHV1-positive cats presented to veterinarians and the five FcaGHV1-positive stray cats (pMW = 0.4956). When the FcaGHV1 blood loads in the 881 cats presented to veterinarians were grouped into “low” (<800 copies/mL; n = 31) and “high” loads (>1000 copies/mL; n = 22), FeLV provirus-positive cats significantly more frequently exhibited high FcaGHV1 loads (4/4; 100%; 95% CI, 40–100%) than FeLV provirus-negative cats (18/49; 37%; 95% CI, 23–52%; pF = 0.0250; OR 15.3; 95% CI, 0.8–301). There was also a tendency for FeLV-viremic cats to more frequently exhibit high FcaGHV1 loads (3/3; 100%; 95% CI, 29–100%) than FeLV-aviremic cats (19/50; 38%; 95% CI, 25–53%; pF = 0.0657; OR 11.3; 95% CI 0.6–231). In contrast, hemoplasma-positive cats significantly less frequently exhibited high FcaGHV1 loads (4/19; 21%; 95% CI, 6.1–46%) than hemoplasma-negative cats (18/34; 52%; 95% CI, 35–70%; pF = 0.0408; OR 0.23; 95% CI, 0.07–0.86). There was no statistical difference in the FcaGHV1 loads between FIV-positive and FIV-negative cats (pMW = 0.1295). In addition, there was no statistical association between the FcaGHV1 blood loads and the age of the positive cats (n = 42; pS = 0.5495; r = 0.095; 95% CI, -0.224–0.396).
3.7. Clinical Presentation of Some FcaGHV1-Positive Cats
Clinical information (presenting clinical signs, pretreatment, and diagnosis) was available for selected FcaGHV1 PCR-positive cats presented to the Clinic for Small Animal Internal Medicine (n = 7). This information was compared to that of age-, sex- and breed-matched FcaGHV1 PCR-negative cats presented at the same clinic in this study (n = 12).
Four of the seven FcaGHV1-positive cats had a history of pretreatment with glucocorticoids for one or several days, and only two were current on their vaccinations against feline parvovirus, feline herpesvirus-1 and feline calicivirus. Five of the seven FcaGHV1-positive cats had cardiorespiratory signs, with final diagnoses of hypertrophic cardiomyopathy (HCM) (n = 3) and/or bronchopneumonia (n = 2), pyothorax (n = 1), and feline asthma (n = 1). In the latter cat, squamous cell carcinoma of the cheek with metastasis to the regional lymph node was confirmed after surgical excision. In one of the two cats without cardiorespiratory signs (cat 2223639), immune-mediated hemolytic anemia was diagnosed, which, after 4 weeks of glucocorticoid treatment, was determined to be acute leukemia. Further characterization of the acute leukemia was not possible. The other positive cat (cat 2236869) was 20 years old with chronic kidney disease (CKD) and secondary chronic vomiting and weight loss.
In comparison, only 2 of the 12 FcaGHV1-negative cats with clinical information had a history of pretreatment with glucocorticoids (one day), and all except one were current on their vaccinations. None of those cats showed cardiorespiratory signs; 7/12 presented because of gastrointestinal signs (vomiting, diarrhea and/or anorexia). The final diagnoses in these cats were pancreatitis (
n = 2), food-responsive inflammatory bowel disease (
n = 1), cholangitis (
n = 1), GI bleeding (
n = 1) and lymphoma (
n = 2). Three of the other 5 cats showed neurological signs (permethrin intoxication,
n = 1; otitis media,
n = 1; meningioma,
n = 1). The final diagnoses in 2 of the 5 cats were diabetes mellitus and fibrosarcoma (showing a large mass on the lateral abdomen). There was no difference in outdoor access between the FcaGHV1 PCR-positive and -negative cats. The clinical diagnoses of the seven selected FcaGHV1-positive cats are shown in
Table 7.
3.8. Sequence Analysis
The results of the FcaGHV1 qPCR assay were confirmed by the sequencing of a 316 bp fragment of the glycoprotein B gene of FcaGHV1 from three random samples. Two samples (BE26 and AR3) showed identical sequences, and the third sample (LZ11) differed from the others by one nucleotide (a synonymous nucleotide polymorphism). The sequences of the former samples were 100% identical to the FcaGHV1 sequence from Australian infected cats (GenBank KJ561572 [
27]) and sequences from Germany, Austria, the USA and Japan (KP862648, KP862649, KF840717, and LC198234, respectively [
20,
23,
25]) and 99.7% identical to that from a Singaporean cat (KJ561573 [
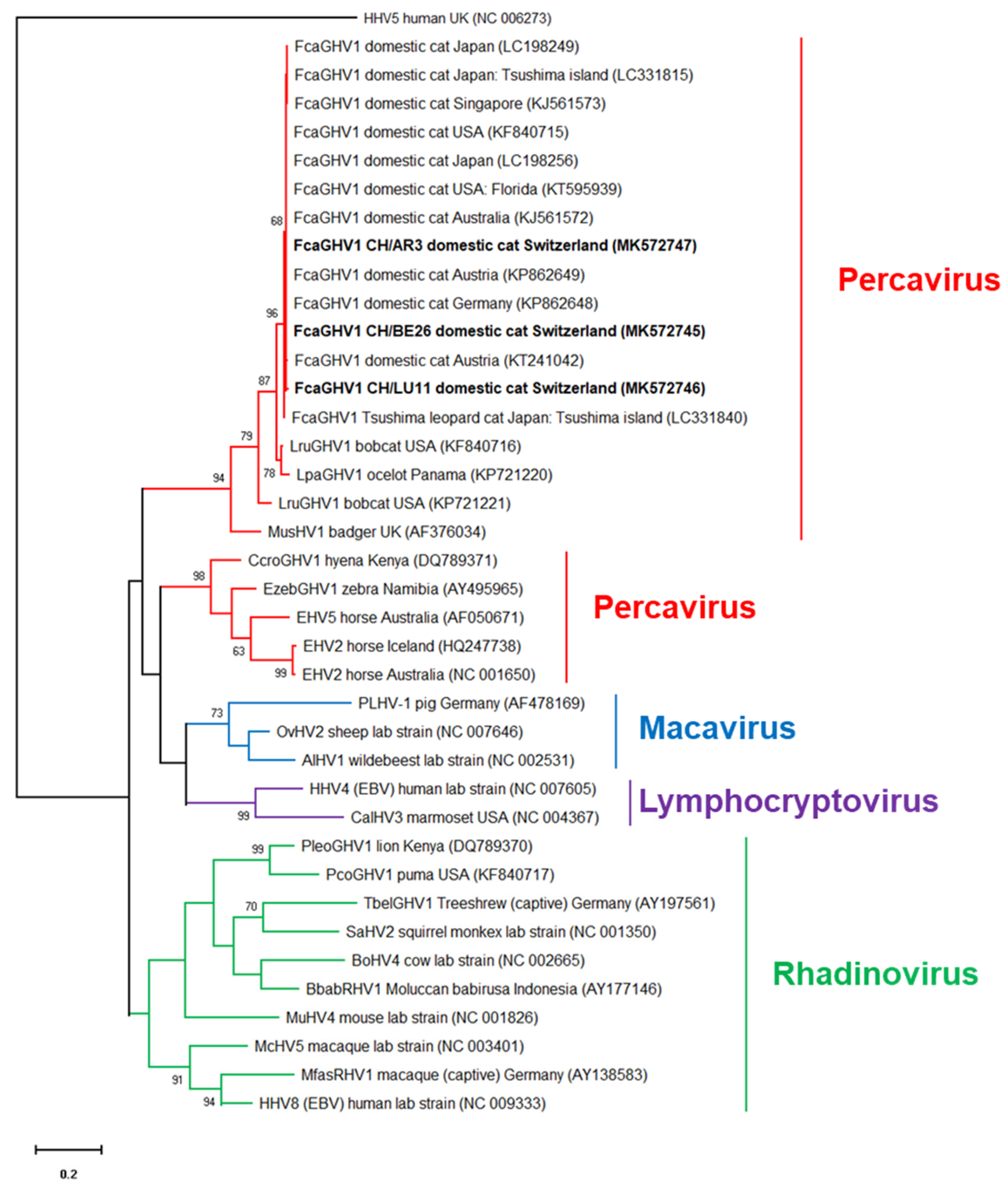
27]). Phylogenetic analysis showed that the Swiss FcaGHV1 sequences (MK572745-MK572747) clustered together with FcaGHV1 sequences from domestic cats, bobcats and ocelots, in the genus Percavirus (
Figure 3).
3.9. Absence of FcaGHV1 in Lymphomas
None of the analyzed tumor or bone marrow samples from the 17 cats with lymphoma were positive for FcaGHV1. This sample set included samples from nine SPF cats experimentally infected with FeLV, two cats coinfected with FeLV and FIV, and six samples collected from clinical cases in field cats. The latter six samples collected from the clinical cases tested PCR-negative for FeLV and FIV. The lymphoma cases included both B- and T-cell lymphoma.
4. Discussion
This study is the first to investigate the presence of FcaGHV1 in Switzerland. A molecular prevalence of 6.0% (95% CI, 4.5–7.8%) was found for FcaGHV1 in domestic cats presented to veterinarians throughout Switzerland. A similar molecular prevalence was noted in stray cats in Switzerland (5.5%; 95% CI, 1.8–12.4%). FcaGHV1 PCR-positive cats originated from 19 of the 26 Swiss cantons, and no significant difference in the prevalence of FcaGHV1 infection was observed among the different cantons. However, there was an obvious geographic clustering of FcaGHV1 detection in the Swiss Plateau and Jura region of Switzerland compared to the Alps. This pattern could indicate that the climate or certain vectors could play a role in FcaGHV1 transmission. We also investigated whether there was an association with population density; as no data are available on the cat density in each Swiss canton, we used the human density as an approximation [
65], assuming that a higher number of people indicates a higher number of cats. There was no association between the density of the human population and the molecular prevalence of FcaGHV1 infection in domestic cats. Thus, either our assumption was wrong and many of the cats in more highly populated regions do not have outdoor access providing contact to other cats, or cat density is not associated with the FcaGHV1 detection.
In contrast, we did find an association between FcaGHV1 detection and the sex of the cats. Male cats were more likely to be infected with FcaGHV1 than female cats. A higher FcaGHV1 infection risk for male cats was reported in prevalence studies from the USA, Europe, Brazil, Tsushima Island (Japan) and Australia [
20,
22,
23,
24,
26,
27]. Male cats were also more likely to be seropositive than female cats, using the newly developed serological assay for FcaGHV1 [
29]. It has been speculated that male cats may be at higher risk of viral exposure due to their strong territorial behavior, which often leads to aggressive interactions among cats [
20,
23,
27,
28]. Biological factors such as hormonal differences between male and female cats could also potentially influence host immunity or viral replication [
20,
66]. Cross-sectional epidemiological data using structural equation models further showed that host phenotypic traits such as aggressive male phenotypes are real FcaGHV1 disease risk factors and not simply correlated events [
67]. This important epidemiological finding suggests horizontal transmission by direct animal contact (e.g., during territorial aggression or through contaminated objects) as one possible route of FcaGHV1 infection. This hypothesis is supported by the recent identification of FcaGHV1 in oronasal swabs and tissues collected from infected cats [
68]. Oronasal secretions, therefore, might play an important role in the transmission of FcaGHV1 among cats.
We found that cats greater than three years of age were at increased risk of being FcaGHV1 PCR-positive. The age of the cats also seems to play a role in the FcaGHV1 prevalence in other studies, where adult cats were more frequently infected than young cats [
20,
23,
25,
26,
27,
29]. The observation that adult cats are more frequently infected than young animals supports the assumption that FcaGHV1 infection cannot be cleared by the cat’s immune system, leading to lifelong infection and increasing infection rates with increasing age of the cats. Interestingly, it was recently reported that cats can be infected with FcaGHV1 from two months of age [
68]. None of the younger cats (< 2 months of age) were FcaGHV1 PCR-positive [
68].
FcaGHV1 infections have been associated with several other infectious agents [
23,
24,
25,
26,
27]. In the present study, FIV was identified as a risk factor for FcaGHV1 detection. FIV was previously recognized as a risk factor, and higher FcaGHV1 blood loads were found in FIV-infected cats than in FIV-negative cats in Europe, Australia [
23,
27], Brazil [
26] and Japan [
25]. The association between FcaGHV1 and FIV may be because FIV may lead to immunosuppression in infected cats, and this, in turn, may facilitate FcaGHV1 replication or escape from latency [
27]. FeLV viremia also tended to be associated with FcaGHV1 detection. We found a tendency for FeLV viremic cats to be more frequently FcaGHV1-infected than FeLV aviremic cats. Progressive FeLV infection with persistent FeLV viremia was previously reported to induce loss of cell (neutrophil and lymphocyte) function and changes in cytokine patterns, leading to the clear dysfunction of the immune system and a decrease in tumor surveillance mechanisms, causing an increased risk of tumor development in infected cats [
69]. These alterations of the immune system in viremic FeLV-infected cats may explain the tendency toward the increased frequency of FcaGHV1 detection in FeLV-viremic cats compared to that in FeLV-aviremic cats. Interestingly, we also found that FeLV-viremic and FeLV provirus-positive cats more frequently exhibited high FcaGHV1 blood loads than uninfected cats. FeLV was not consistently identified as a risk factor for FcaGHV1 alone [
27,
31], but cats co-infected with FIV and FeLV were identified as having an increased risk [
31]. It is known that FeLV–FIV co-infections have a more severe influence on a cat’s immune system than FIV or FeLV alone [
43,
69,
70]. The present study showed that both single FIV and single FeLV infection may have an effect on FcaGHV1 detection. It cannot be determined whether this effect is actually due to an immunosuppressive effect of feline retroviruses, since immunological parameters such as CD4+ counts have not been investigated in FcaGHV1-infected cats, or whether the association could also be based on similar transmission routes. However, in stray cats, no association was found between FeLV infection and FcaGHV1 detection; this lack of association could imply that not a similar method of transmission but rather actual immunosuppression, which is more detrimental and probably more quickly fatal in stray cats than in privately-owned cats presented to veterinarians, is the basis of this association. Alternatively, the small number of stray cats may have limited the statistical analysis; however, this possibility seems less likely since the percentage of FeLV provirus-positive stray cats (7.7%) was rather high and was higher than that of cats presented to veterinarians (5.3%).
We also identified hemoplasma infections as risk factors for FcaGHV1 in cats presented to veterinarians as well as in stray cats. Hemoplasmas were also previously recognized as risk factors for FcaGHV1 [
24,
27]. Again, the real meaning of the association between hemoplasma and FcaGHV1 infections has not yet been identified; however, these two infections may indeed share similar transmission routes, such as aggressive contact between cats in an outdoor environment or vector-borne transmission. In addition, we found two cats concurrently infected with all three feline hemoplasma species: Mhf, CMhm and CMt. Both cats were stray cats, and one of the two cats was also FcaGHV1 PCR-positive. Concurrent infections with all three feline hemoplasmas have rarely been reported previously.
The pathogenic potential of FcaGHV1 in domestic cats is not completely understood. It was shown that FcaGHV1 PCR-positive cats are 2.8 times more likely to be ill than healthy [
27]. However, this association (FcaGHV1 and illness) could not be shown in other studies [
23]. In the present study, clinical information was available only for a limited number of cats; none of the FcaGHV1-positive cats had lymphoma, but two of the matched FcaGHV1-negative cats did. This finding was consistent with the negative results of all our tested lymphoma tissue samples. Interestingly, the majority of the FcaGHV1-positive cats with available clinical data showed cardiorespiratory signs, with a confirmed diagnosis of hypertrophic cardiomyopathy in three of these cats. Although one of these cats was a Maine Coon cat, a breed in which the incidence of hypertrophic cardiomyopathy is known to be increased, none of the three FcaGHV1-negative Maine Coon cats exhibited hypertrophic cardiomyopathy. An association between FcaGHV1 and cardiomyopathy has not been reported previously. However, Epstein–Barr virus, another GHV, was detected in humans with cardiomyopathy, suggesting a possible association between the virus and the otherwise unexplained cardiomyopathy [
71]. FcaGHV1 DNA was detected in multiple tissues from infected cats, including heart tissue [
27]. More studies are needed to further evaluate the possible association of FcaGHV1 and cardiomyopathy in a larger number of cats. Whether glucocorticoid or immunosuppressive treatment and vaccination status have any influence on FcaGHV1 infection and pathogenesis can only be speculated, as our number of cases was small.
In humans, two GHVs (Epstein–Barr virus and Kaposi’s sarcoma-associated herpesvirus) are associated with the development of cancer [
72]. FcaGHV1 was isolated in T and B lymphocytes of cats but not in feline myeloid cells [
73]. This pattern supports the hypothesis that FcaGHV1 is lymphotropic, as reported for the other GHVs. In a recent study analyzing 122 feline lymphoma cases (including histological and/or cytological preparations), no association was found between FcaGHV1 and the development of lymphoma in cats [
74]. The absence of an association, however, does not rule out a pathogenic role of the virus [
74]. To further evaluate the role of FcaGHV1 in lymphomagenesis, transcriptome analysis and in situ hybridization were performed on tissue samples of cats with lymphoma [
75]. In that study, FcaGHV1 was detected in some FIV-associated lymphomas, and a positive FcaGHV1 intranuclear signal was visible by in situ hybridization in >90% of the cells in a case of feline large granular lymphocyte lymphoma. These results demonstrated that FcaGHV1 was present in feline lymphoma but only at low copy numbers, impairing the possibility of evaluating the role of the virus in lymphomagenesis [
75]. High FcaGHV1 tissue loads (48 FcaGHV1 genomes per cell) were previously detected in a lymph node from a cat with high-grade T-cell intestinal lymphoma [
76], suggesting a certain variability of the viral loads within feline lymphoma subtypes. Further studies investigating the cellular location and expression of FcaGHV1 antigens within neoplasias are needed to clarify this possibility. In the present study, all tissue samples from cats with lymphoma—six clinical cases testing negative for FeLV and FIV and 11 cats experimentally infected with FeLV and/or FIV—were FcaGHV1-negative. One of the cats also had large granular lymphocyte lymphoma. It is important to mention that the lack of association between FcaGHV1 and the development of lymphoma seen in the present study does not rule out a role of FcaGHV1 in lymphomagenesis, especially considering the inclusion of the tissue samples from the 11 SPF-cats that had been kept in a research facility under barrier conditions. However, when the cats had been purchased from an accredited SPF breeding facility some years ago, FcaGHV1 had not yet been discovered and no screening assays were available to detect the virus. Thus, FcaGHV1 was not on the list of specified pathogens yet, and the cats had not been tested for FcaGHV1 prior to the FeLV and/or FIV infection or throughout their lifespan. Thus, we were pleased to recognize that the cats were free of FcaGHV1, and therefore FcaGHV1 was not the underlying reason for the rather high frequency of lymphoma observed in our FeLV-infected SPF cats. However, we recognize that the samples from the SPF cats may not be representative to assess the contribution of FcaGHV1 to the development of lymphoma in field cats.