Comparing 30 Tree Biomass Models to Estimate Forest Biomass in the Amazon
Abstract
1. Introduction
2. Materials and Methods
2.1. Data and Study Area
2.2. Analytical Procedure
2.2.1. Succession Stages of the Amazon Forest
2.2.2. Plot Simulation and Modeling
- (1)
- From the Amazon dataset, select without replacement a random within the limits defined in methodology, calculate as in Equation (1), and average H as in Equation (2).
- (2)
- For the plot simulated in the previous step, consider approving that plot such that both and fall into the limits defined in methodology, and go to the next step. For not-approved plots, redo step 1. For the j-th plot approved in the previous step, obtain the observed and predicted through the biomass models shown in Table 2.
- (3)
- For every simulated plot and m-th biomass model, obtain the plot AGB error in Mg ha−1 (Equation (5)), plot AGB absolute error in Mg ha−1 (Equation (6)), and plot AGB relative error in % (Equation (7)).
- (4)
- Replicate steps 1–4 until a set with 100 simulated plots is obtained.
- (5)
- For the set of simulated plots and m-th biomass model (Table 2), estimate the mean error in Mg ha−1 (Equation (8)), mean absolute error in Mg ha−1 (Equation (9)), mean relative error in % (Equation (10)), and root mean square error in Mg ha−1 (Equation (11)).
- (6)
- Repeat steps 1–6 for all succession stages.
| M. | Biomass Models | Author |
|---|---|---|
| 1 | [10] | |
| 2 | [11] | |
| 3 | [11] | |
| 4 | [11] | |
| 5 | [11] | |
| 6 | [12] | |
| 7 | [13] | |
| 8 | [13] | |
| 9 | [14] | |
| 10 | [14] | |
| 11 | [15] | |
| 12 | [15] | |
| 13 | [15] | |
| 14 | [16] | |
| 15 | [16] | |
| 16 | [16] | |
| 17 | [16] | |
| 18 | [17] | |
| 19 | [17] | |
| 20 | [17] | |
| 21 | [18] | |
| 22 | [18] | |
| 23 | [19] | |
| 24 | [20] | |
| 25 | [20] | |
| 26 | [20] | |
| 27 | [20] | |
| 28 | [21] | |
| 29 | [21] | |
| 30 | [21] |
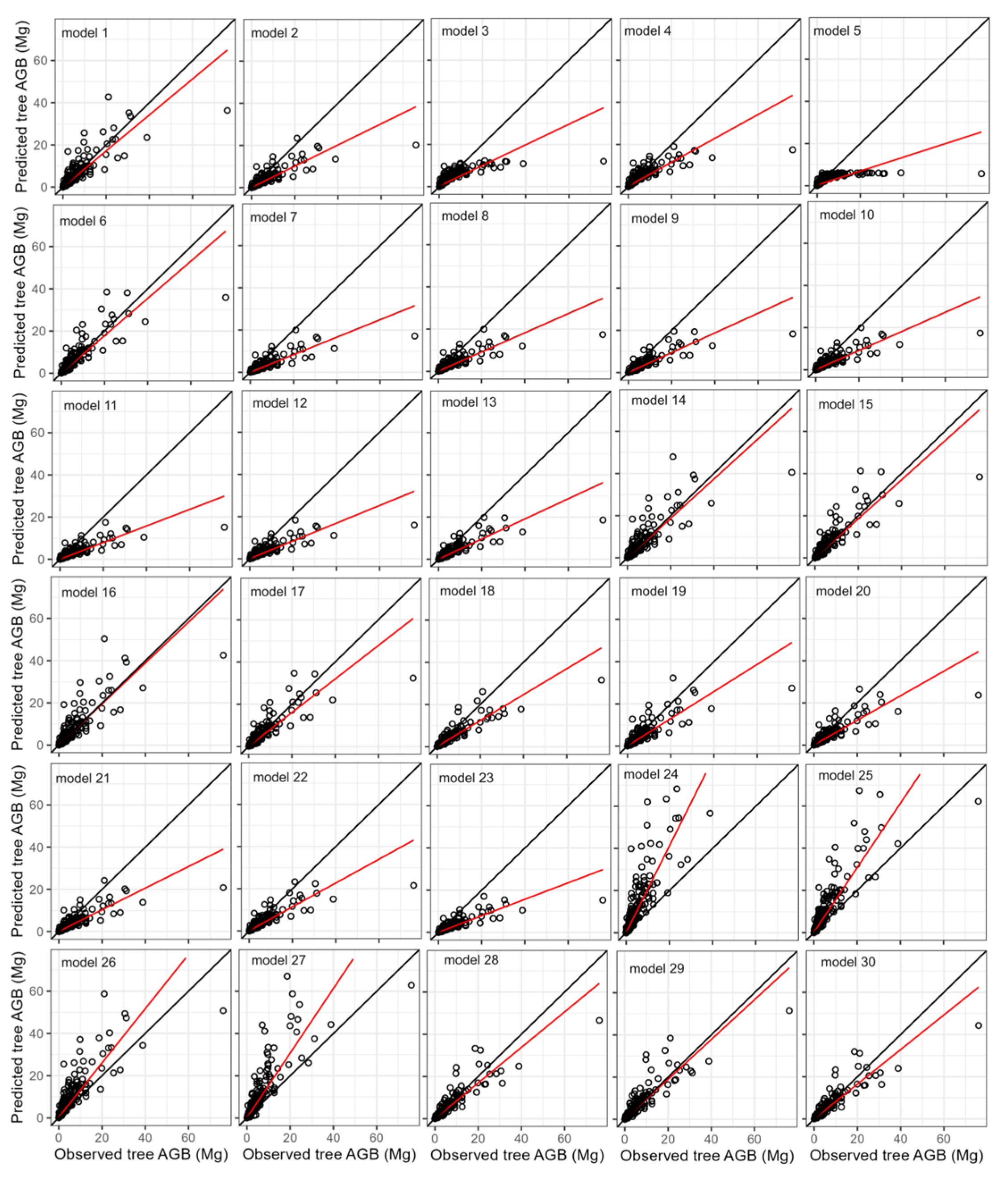
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Malhi, Y.; Wood, D.; Baker, T.R.; Wright, J.; Phillips, O.L.; Cochrane, T.; Meir, P.; Chave, J.; Almeida, S.; Arroyo, L.; et al. The regional variation of aboveground live biomass in old-growth Amazonian forests. Glob. Change Biol. 2006, 12, 1107–1138. [Google Scholar] [CrossRef]
- Bastin, J.F.; Rutishauser, E.; Kellner, J.R.; Saatchi, S.; Pélissier, R.; Hérault, B.; Réjou-Méchain, M. Pan-tropical prediction of forest structure from the largest trees. Glob. Ecol. Biogeogr. 2018, 27, 1366–1383. [Google Scholar] [CrossRef]
- Slik, J.W.F.; Aiba, S.I.; Brearley, F.Q.; Cannon, C.H.; Forshed, O.; Kitayama, K.; Nagamasu, H.; Nilo, R.; Payne, J.; Paoli, G.; et al. Environmental correlates of tree biomass, basal area, wood specific gravity, and stem density gradients in Borneo tropical forests. Glob. Ecol. Biogeogr. 2010, 19, 50–60. [Google Scholar] [CrossRef]
- Chave, J.; Réjou-Méchain, M.; Búrquez, A.; Chidumayo, E.; Colgan, M.S.; Delitti, W.B.C.; Vieilledent, G.; Eid, T.; Fearnside, P.M.; Goodman, R.C.; et al. Improved allometric models to estimate the aboveground biomass of tropical trees. Glob. Change Biol. 2014, 20, 3177–3190. [Google Scholar] [CrossRef]
- Avitabile, V.; Herold, M.; Heuvelink, G.B.M.; Lewis, S.L.; Phillips, O.L.; Asner, G.P.; Willcock, S. An integrated pan-tropical biomass map using multiple reference datasets. Glob. Change Biol. 2016, 22, 1406–1420. [Google Scholar] [CrossRef] [PubMed]
- Moran, E.F.; Brondizio, E.S.; Tucker, J.; Silva-Forsberg, M.D.; Falesi, I.; McCracken, S. Strategies for Amazonian Forest Restoration: Evidence for Afforestation in Five Regions of the Brazilian Amazon. In Amazônia at the Crossroads: The Challenge of Sustainable Development; Institute for Latin American Studies/University of London: London, UK, 2000; pp. 129–149. [Google Scholar]
- David, H.C.; Barbosa, R.I.; Vibrans, A.C.; Watzlawick, L.F.; Trautenmuller, J.W.; Balbinot, R.; Ribeiro, S.C.; Jacovine, L.A.G.; Corte, A.P.D.; Sanquetta, C.R.; et al. The tropical biomass & carbon project—An application for forest biomass and carbon estimates. Ecol. Model. 2022, 472, 110067. [Google Scholar] [CrossRef]
- David, H.C.; Carvalho, J.O.P.; Pires, I.P.; Santos, L.S.; Barbosa, E.S.; Braga, N.S. A 20-year tree liberation experiment in the Amazon: Highlights for diameter growth rates and species-specific management. For. Ecol. Manag. 2019, 453, 117584. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2024. [Google Scholar]
- Nogueira, E.M.; Fearnside, P.M.; Nelson, B.W.; Barbosa, R.I.; Keiser, E.W.H. Estimates of forest biomass in the Brazilian Amazon: New allometric equations and adjustments to biomass from wood-volume inventories. For. Ecol. Manag. 2008, 256, 1853–1867. [Google Scholar] [CrossRef]
- Chambers, J.Q.; Dos Santos, J.; Ribeiro, R.J.; Higuchi, N. Tree damage, allometric relationships, and above-ground net primary production in central Amazon forest. For. Ecol. Manag. 2001, 152, 73–84. [Google Scholar] [CrossRef]
- Brown, I.F.; Martinelli, L.A.; Thomas, W.W.; Moreira, M.Z.; Ferreira, C.A.C.; Victoria, R.A. Uncertainty in the biomass of Amazonian forests: An example from Rondônia, Brazil. For. Ecol. Manag. 1995, 75, 175–189. [Google Scholar] [CrossRef]
- Brown, S. Estimating Biomass and Biomass Change of Tropical Forests: A Primer. In FAO Forestry Paper; Food and Agriculture Organization: Rome, Italy, 1997; Volume 134. Available online: www.fao.org/docrep/w4095e/w4095e00.HTM (accessed on 7 November 2025).
- Brown, S.; Iverson, L.R. Biomass estimates for tropical forests. World Resour. Rev. 1992, 4, 366–384. [Google Scholar]
- Brown, S.; Gillespie, A.; Lugo, A.E. Biomass Estimation Methods for Tropical Forests with Applications to Forest Inventory Data. For. Sci. 1989, 35, 881–902. [Google Scholar] [CrossRef]
- Overman, J.P.M.; Witte, H.J.L.; Saldarriaga, J.G. Evaluation of regression models for above-ground biomass determination in Amazon rainforest. J. Trop. Ecol. 1994, 10, 207–218. [Google Scholar] [CrossRef]
- Nelson, B.W.; Mesquita, R.; Pereira, J.L.G.; Souza, S.G.A.; Batista, G.T.; Couto, L.B. Allometric regressions for improved estimate of secondary forest biomass in the central Amazon. For. Ecol. Manag. 1999, 117, 149–167. [Google Scholar] [CrossRef]
- Ducey, M.J.; Zarin, D.J.; Vasconcelos, S.S.; Araújo, M.M. Biomass equations for forest regrowth in the eastern Amazon using randomized branch sampling. Acta Amaz. 2009, 39, 349–360. [Google Scholar] [CrossRef][Green Version]
- Uhl, C.; Buschbacher, R.; Serrao, E.A.S. Abandoned Pastures in Eastern Amazonia. I. Patterns of Plant Succession. J. Ecol. 1988, 76, 663. [Google Scholar] [CrossRef]
- Higuchi, N.; Santos, J.; Ribeiro, R.J.; Minette, L.; Biot, Y. Biomassa da parte aérea da vegetação da Floresta Tropical úmida de terra firme da Amazônia Brasileira. Acta Amaz. 1998, 28, 153. [Google Scholar] [CrossRef]
- Chave, J.; Andalo, C.; Brown, S.; Cairns, M.A.; Chambers, J.Q.; Eamus, D.; Folster, H.; Fromard, F.; Higuchi, N.; Kira, T.; et al. Tree allometry and improved estimation of carbon stocks and balance in tropical forests. Oecologia 2005, 145, 87–99. [Google Scholar] [CrossRef]
- Leão, F.M.; Nascimento, R.G.M.; Emmert, F.; Santos, G.G.A.; Caldeira, N.A.M.; Miranda, I.S. How many trees are necessary to fit an accurate volume model for the Amazon forest? A site-dependent analysis. For. Ecol. Manag. 2021, 480, 118652. [Google Scholar] [CrossRef]
- Ketterings, Q.M.; Coe, R.; Noordwijk, M.V.; Ambagau’, Y.; Palm, C.A. Reducing uncertainty in the use of allometric biomass equations for predicting above-ground tree biomass in mixed secondary forests. For. Ecol. Manag. 2001, 146, 199–209. [Google Scholar] [CrossRef]
- Marra, D.M.; Higuchi, N.; Trumbore, S.E.; Ribeiro, G.H.P.M.; Santos, J.; Carneiro, V.M.C.; Lima, A.J.N.; Chambers, J.Q.; Negrón-Juárez, R.I.; Holzwarth, F. Predicting biomass of hyperdiverse and structurally complex central Amazonian forests—A virtual approach using extensive field data. Biogeosciences 2016, 13, 1553–1570. [Google Scholar] [CrossRef]
- David, H.C. Are tropical forests an extremistan environment? CERNE 2024, 30, 10338. [Google Scholar] [CrossRef]



| Variables | Minimum | Mean | Maximum | CV (%) |
|---|---|---|---|---|
| DBH (cm) | 5.0 | 19.0 | 138.0 | 84.6 |
| H (m) | 6.0 | 17.4 | 54.1 | 42.5 |
| Biomass (kg) | 5.1 | 558.0 | 20,416.0 | 306 |
| Density (g/cm3) | 0.22 | 0.61 | 1.08 | 22.8 |
| Model | ||||||||
|---|---|---|---|---|---|---|---|---|
| Advanced secondary forest | Mature forest | |||||||
| M1 | 2.1 | 3.7 | 4.2% | 4.6 | 8.8 | 15.8 | 9.7% | 20.2 |
| M2 | −12.6 | 12.6 | −23.8% | 13.4 | −35.9 | 36 | −29.0% | 40.8 |
| M3 | 11.7 | 11.7 | 22.8% | 12.8 | −12.4 | 17.4 | −8.1% | 24.2 |
| M4 | 10.5 | 10.6 | 20.6% | 11.6 | −7.5 | 14.8 | −3.9% | 20.7 |
| M5 | 7.8 | 8.1 | 15.5% | 9.1 | −31.4 | 31.5 | −24.5% | 38.8 |
| M6 | 5.1 | 5.4 | 10.0% | 6.4 | 13.4 | 17.7 | 13.6% | 20.8 |
| M7 | −23.0 | 23.0 | −43.9% | 23.9 | −53.5 | 53.5 | −44.4% | 57.5 |
| M8 | −15.7 | 15.7 | −29.9% | 16.5 | −42.8 | 42.8 | −35.0% | 47.4 |
| M9 | −17.6 | 17.6 | −33.5% | 18.4 | −44.3 | 44.3 | −36.3% | 48.3 |
| M10 | −15.7 | 15.7 | −29.8% | 16.5 | −42.8 | 42.8 | −35.0% | 47.3 |
| M11 | −22.2 | 22.2 | −42.2% | 23 | −54.2 | 54.2 | −44.9% | 58.2 |
| M12 | −16.5 | 16.5 | −31.4% | 17.4 | −46.8 | 46.8 | −38.4% | 51.1 |
| M13 | −16.5 | 16.5 | −31.5% | 17.3 | −42.4 | 42.4 | −34.8% | 46.6 |
| M14 | 2.7 | 4.0 | 5.3% | 5.0 | 16.3 | 20.3 | 16.1% | 24.9 |
| M15 | 2.5 | 3.6 | 5.0% | 4.5 | 14.4 | 18.3 | 14.3% | 21.5 |
| M16 | −30.9 | 30.9 | −58.9% | 31.9 | −72.7 | 72.7 | −61.0% | 76.3 |
| M17 | −0.9 | 2.9 | −1.5% | 3.6 | 0.1 | 12.1 | 2.1% | 16.0 |
| M18 | −7.1 | 7.2 | −13.3% | 8.2 | −14.9 | 18.0 | −10.9% | 23.5 |
| M19 | −11.3 | 11.3 | −21.4% | 12.1 | −22.2 | 23.4 | −17.2% | 28.9 |
| M20 | −11.5 | 11.5 | −21.8% | 12.3 | −28.2 | 28.5 | −22.4% | 33.6 |
| M21 | −13.6 | 13.6 | −25.7% | 14.4 | −3.06 | 36.1 | −29.1% | 40.9 |
| M22 | −5.5 | 5.7 | −10.1% | 6.6 | −24.4 | 24.7 | −19.0% | 30.3 |
| M23 | −20.7 | 20.7 | −39.4% | 21.5 | −53.7 | 53.7 | −44.5% | 57.6 |
| M24 | 59.0 | 59.0 | 113.1% | 61.2 | 165.1 | 165.1 | 145.2% | 170.2 |
| M25 | 46.8 | 46.8 | 89.9% | 48.5 | 108.8 | 108.8 | 96.3% | 111.6 |
| M26 | 55.2 | 55.2 | 106.5% | 57.1 | 96.6 | 96.6 | 86.3% | 99.6 |
| M27 | 30.8 | 30.8 | 58.9% | 32.3 | 98.6 | 98.6 | 86.7% | 102.0 |
| M28 | −1.8 | 2.7 | −3.1% | 3.4 | −0.9 | 10.8 | 0.7% | 14.1 |
| M29 | 4.0 | 4.4 | 7.9% | 5.2 | 15.3 | 18.0 | 14.8% | 22.3 |
| M30 | 0.1 | 2.4 | 0.6% | 3.0 | −0.9 | 10.7 | 0.8% | 14.1 |
| Model Characteristics | Advanced Secondary Forest | Mature Forest | N | ||
|---|---|---|---|---|---|
| Correlation (r) | p-Value | Correlation (r) | p-Value | ||
| DBH Min | 0.19 | 0.32 | 0.16 | 0.40 | 30 |
| DBH Max | 0.38 | 0.04 | 0.42 | 0.02 | 30 |
| Range of DBH | 0.33 | 0.08 | 0.37 | 0.05 | 30 |
| R2 | 0.33 | 0.07 | 0.34 | 0.07 | 30 |
| RMSE (unit: kg) | 0.60 (a) | 0.40 (a) | 0.40 (a) | 0.60 (a) | 4 |
| RMSE (unit: log) | 0.57 (b) | 0.18 (b) | 0.21 (b) | 0.64 (b) | 26 |
| Number of predictors | 0.10 | 0.60 | –0.03 | 0.99 | 30 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Garcia, R.A.; Galvão, L.M.R.; Chivale, X.S.; Almeida, T.C.; Pereira, F.R.; Martins-Neto, R.P.; Sanquetta, C.R.; David, H.C. Comparing 30 Tree Biomass Models to Estimate Forest Biomass in the Amazon. Forests 2026, 17, 213. https://doi.org/10.3390/f17020213
Garcia RA, Galvão LMR, Chivale XS, Almeida TC, Pereira FR, Martins-Neto RP, Sanquetta CR, David HC. Comparing 30 Tree Biomass Models to Estimate Forest Biomass in the Amazon. Forests. 2026; 17(2):213. https://doi.org/10.3390/f17020213
Chicago/Turabian StyleGarcia, Rebecca A., Lina M. R. Galvão, Xavier S. Chivale, Thaís C. Almeida, Fabiano R. Pereira, Rorai Pereira Martins-Neto, Carlos R. Sanquetta, and Hassan C. David. 2026. "Comparing 30 Tree Biomass Models to Estimate Forest Biomass in the Amazon" Forests 17, no. 2: 213. https://doi.org/10.3390/f17020213
APA StyleGarcia, R. A., Galvão, L. M. R., Chivale, X. S., Almeida, T. C., Pereira, F. R., Martins-Neto, R. P., Sanquetta, C. R., & David, H. C. (2026). Comparing 30 Tree Biomass Models to Estimate Forest Biomass in the Amazon. Forests, 17(2), 213. https://doi.org/10.3390/f17020213







