Mycosporine-Like Amino Acids: Making the Foundation for Organic Personalised Sunscreens
Abstract
1. Introduction
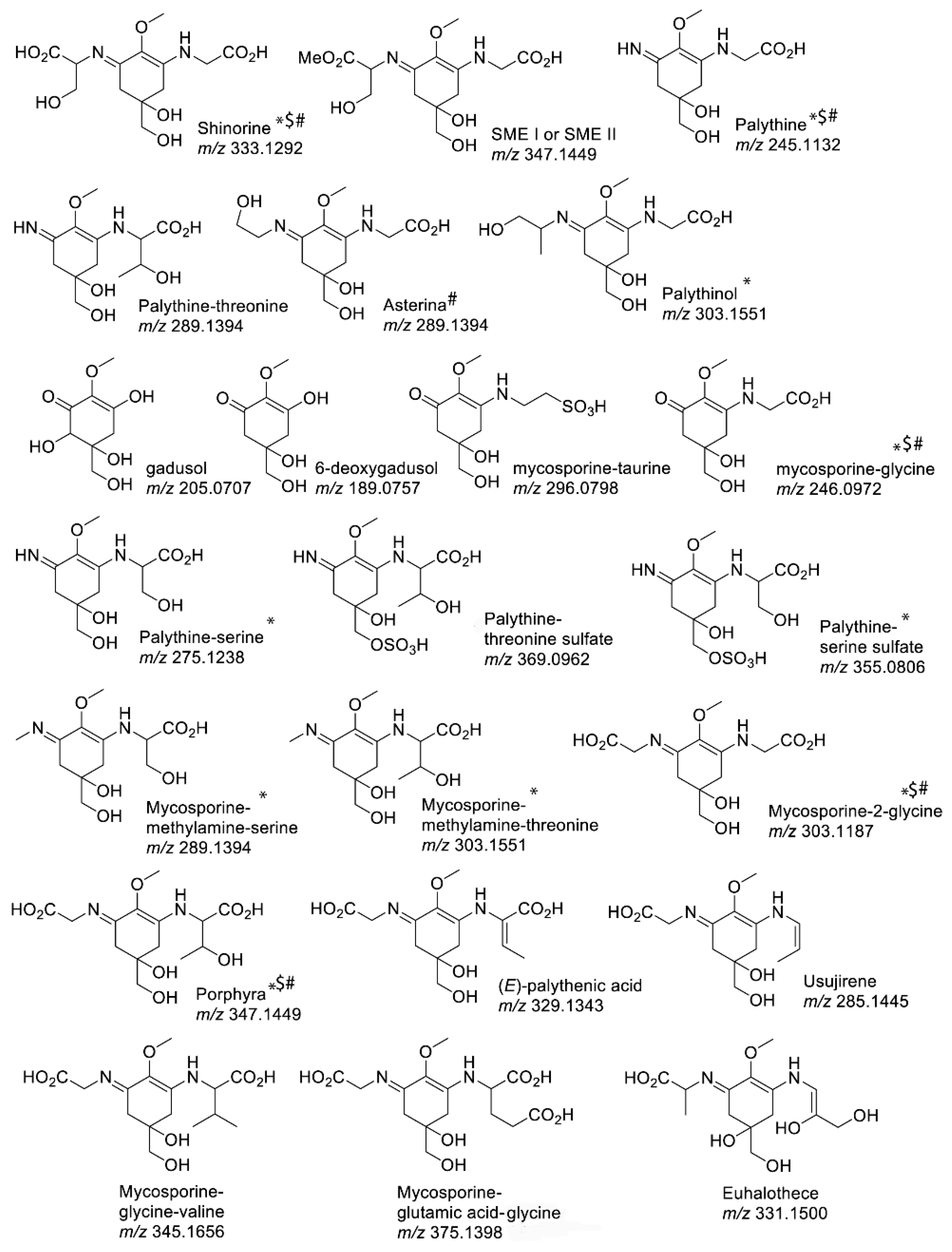
2. MAA Diversity
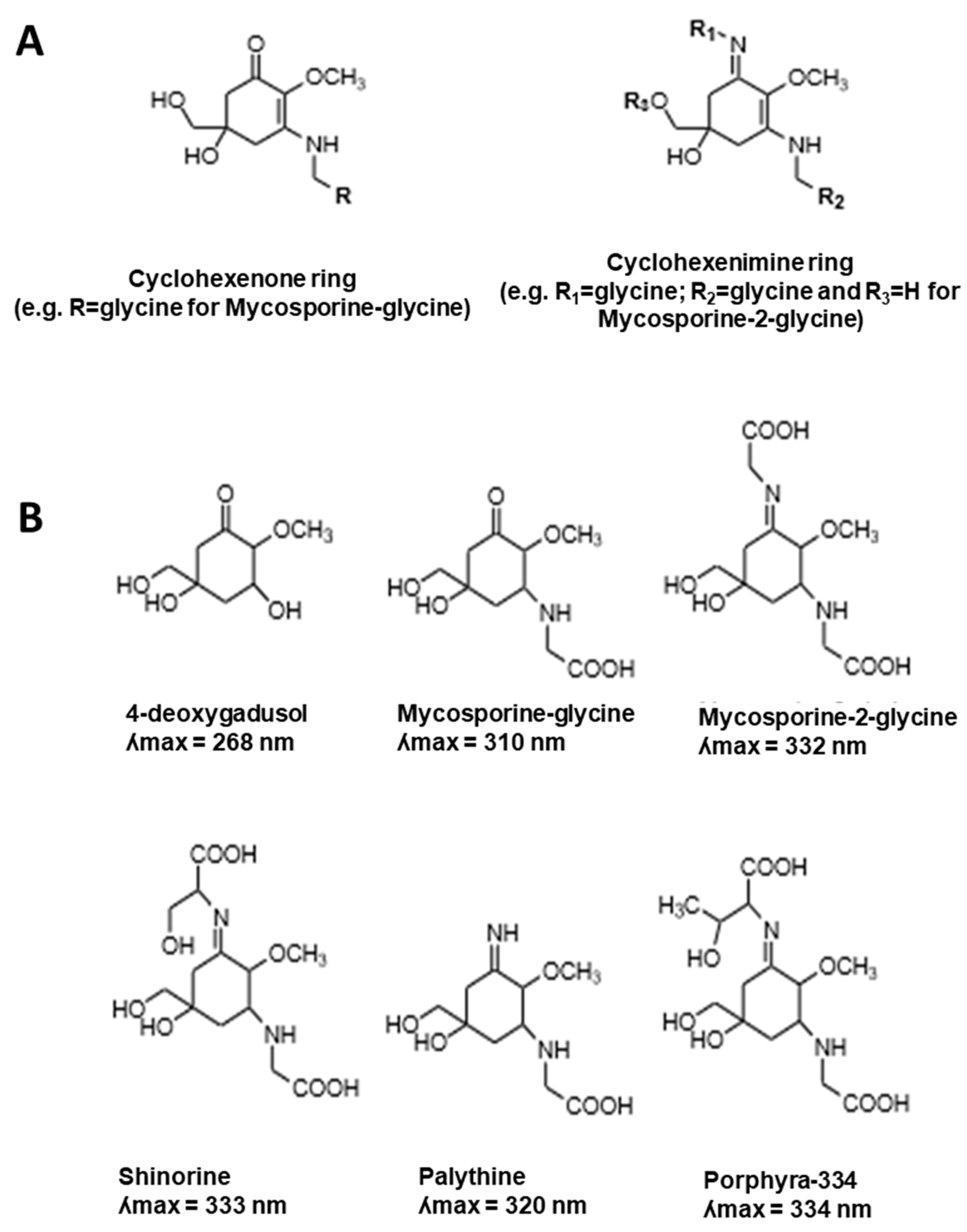
2.1. MAA UV-Absorbing Features
2.2. MAA Antioxidative Properties
2.3. MAA Anti-Inflammatory and Antiaging Properties
3. Fluctuation of the MAA Content in Aquatic Species
4. Genes from the MAA Biosynthetic Pathway and Their Regulation
5. Potential Use of MAAs in Human Skin Protection
6. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- Watanabe, S.; Sudo, K.; Nagashima, T.; Takemura, T.; Kawase, H.; Nozawa, T. Future projections of surface UV-B in a changing climate. J. Geophys. Res. Atmos. 2011, 116. [Google Scholar] [CrossRef]
- Reef, R.; Dunn, S.; Levy, O.; Dove, S.; Shemesh, E.; Bhckner, I.; Leggat, W.; Hoegh-Guldberg, O. Photoreactivation is the main repair pathway for UV-induced DNA damage in coral planulae. J. Exp. Biol. 2009, 212, 2760–2766. [Google Scholar] [CrossRef] [PubMed]
- Sinha, R.P.; Häder, D.P. UV-induced DNA damage and repair: A review. Photochem. Photobiol. Sci. 2002, 1, 225–236. [Google Scholar] [CrossRef] [PubMed]
- Korbee, N.; Teresa Mata, M.; Figueroa, F.L. Photoprotection mechanisms against ultraviolet radiation in Heterocapsa sp. (Dinophyceae) are influenced by nitrogen availability: Mycosporine-like amino acids vs. xanthophyll cycle. Limnol. Oceanogr. 2010, 55, 899–908. [Google Scholar] [CrossRef]
- Singh, S.P.; Klisch, M.; Sinha, R.P.; Häder, D.P. Genome mining of mycosporine-like amino acid (MAA) synthesizing and non-synthesizing cyanobacteria: A bioinformatics study. Genomics 2010, 95, 120–128. [Google Scholar] [CrossRef] [PubMed]
- Shick, J.M.; Dunlap, W.C. Mycosporine-like amino acids and related gadusols: Biosynthesis, accumulation; UV-protective functions in aquatic organisms. Annu. Rev. Physiol. 2002, 64, 223–262. [Google Scholar] [CrossRef] [PubMed]
- Sinha, R.P.; Singh, S.P.; Häder, D.P. Database on mycosporines and mycosporine-like amino acids (MAAs) in fungi, cyanobacteria, macroalgae, phytoplankton and animals. J. Photochem. Photobiol. B Biol. 2007, 89, 29–35. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.P.; Kumari, S.; Rastogi, R.P.; Singh, K.L.; Sinha, R.P. Mycosporine-like amino acids (MAAs): Chemical structure, biosynthesis and significance as UV-absorbing/screening compounds. Indian J. Exp. Biol. 2008, 46, 7–17. [Google Scholar] [PubMed]
- Rosic, N.N. Phylogenetic analysis of genes involved in mycosporine-like amino acid biosynthesis in symbiotic dinoflagellates. Appl. Microbiol. Biotechnol. 2012, 94, 29–37. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Pichel, F.; Castenholz, R.W. Occurrence of UV-absorbing, mycosporine-like compounds among cyanobacterial isolates and an estimate of their screening capacity. Appl. Environ. Microbiol. 1993, 59, 163–169. [Google Scholar] [PubMed]
- Garcia-Pichel, F.; Castenholz, R.W. Characterization and biological implications of scytonemin, a cyanobacterial sheath pigment. J. Phycol. 1991, 27, 395–409. [Google Scholar] [CrossRef]
- Klassen, J.L. Phylogenetic and evolutionary patterns in microbial carotenoid biosynthesis are revealed by comparative genomics. PLoS ONE 2010, 5, e11257. [Google Scholar] [CrossRef] [PubMed]
- Britton, G. Structure and properties of carotenoids in relation to function. FASEB J. 1995, 9, 1551–1558. [Google Scholar] [CrossRef] [PubMed]
- Dunlap, W.C.; Chalker, B.E. Identification and quantitation of near-UV absorbing compounds (S-320) in a hermatypic scleractinian. Coral Reefs 1986, 5, 155–159. [Google Scholar] [CrossRef]
- Nakamura, H.; Kobayashi, J.; Hirata, Y. Separation of mycosporine-like amino acids in marine organisms using reversed-phase high-performance liquid chromatography. J. Chromatogr. A 1982, 250, 113–118. [Google Scholar] [CrossRef]
- Carreto, J.I.; Carignan, M.O. Mycosporine-like amino acids: Relevant secondary metabolites. chemical and ecological aspects. Mar. Drugs 2011, 9, 387–446. [Google Scholar] [CrossRef] [PubMed]
- Rastogi, R.P.; Sinha, R.P.; Singh, S.P.; Häder, D.P. Photoprotective compounds from marine organisms. J. Ind. Microbiol. Biotechnol. 2010, 37, 537–558. [Google Scholar] [CrossRef] [PubMed]
- Bandaranayake, W.M. Mycosporines: Are they nature’s sunscreens? Nat. Prod. Rep. 1998, 15, 159–172. [Google Scholar] [CrossRef] [PubMed]
- Llewellyn, C.A.; Airs, R.L. Distribution and abundance of MAAs in 33 species of microalgae across 13 classes. Mar. Drugs 2010, 8, 1273–1291. [Google Scholar] [CrossRef] [PubMed]
- Sinha, R.P.; Häder, D.P. UV-protectants in cyanobacteria. Plant Sci. 2008, 174, 278–289. [Google Scholar] [CrossRef]
- Shibata, K. Pigments and a UV-absorbing substance in corals and a blue-green alga living in the great barrier reef. Plant Cell Physiol. 1969, 10, 325–335. [Google Scholar]
- Jain, S.; Prajapat, G.; Agrawal, A. A taxonomical study of mycosporine-like amino acids producing cyanobacteria. In Sunscreens: Source, Formulations, Efficacy and Recommendations; Rastogi, R.P., Ed.; Nova Science Publishers, Inc.: Hauppauge, NY, USA, 2018; pp. 131–151. [Google Scholar]
- Karsten, U.; Sawall, T.; Hanelt, D.; Bischof, K.; Figueroa, F.L.; Flores-Moya, A.; Wiencke, C. An inventory of UV-absorbing mycosporine-like amino acids in macroalgae from polar to warm-temperate regions. Bot. Mar. 1998, 41, 443–453. [Google Scholar] [CrossRef]
- Dunlap, W.C.; Yamamoto, Y. Small-molecule antioxidants in marine organisms: Antioxidant activity of mycosporine-glycine. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 1995, 112, 105–114. [Google Scholar] [CrossRef]
- Figueroa, F.L.; Bueno, A.; Korbee, N.; Santos, R.; Mata, L.; Schuenhoff, A. Accumulation of mycosporine-like amino acids in asparagopsis armata grown in tanks with fishpond effluents of gilthead sea bream, Sparus aurata. J. World Aquac. Soc. 2008, 39, 692–699. [Google Scholar] [CrossRef]
- Figueroa, F.L.; Escassi, L.; Pérez-Rodríguez, E.; Korbee, N.; Giles, A.D.; Johnsen, G. Effects of short-term irradiation on photoinhibition and accumulation of mycosporine-like amino acids in sun and shade species of the red algal genus Porphyra. J. Photochem. Photobiol B Biol. 2003, 69, 21–30. [Google Scholar] [CrossRef]
- Lalegerie, F.; Lajili, S.; Bedoux, G.; Taupin, L.; Stiger-Pouvreau, V.; Connan, S. Photoprotective compounds in red macroalgae from Brittany: Considerable diversity in mycosporine-like amino acids (MAAs). Mar. Environ. Res. 2019, 147, 37–48. [Google Scholar] [CrossRef] [PubMed]
- Orfanoudaki, M.; Hartmann, A.; Karsten, U.; Ganzera, M. Chemical profiling of mycosporine-like amino acids in twenty-three red algal species. J. Phycol. 2019, 55, 393–403. [Google Scholar] [CrossRef] [PubMed]
- Favre-Bonvin, J.; Bernillon, J.; Salin, N.; Arpin, N. Biosynthesis of mycosporines: Mycosporine glutaminol in Trichothecium roseum. Phytochemistry 1987, 26, 2509–2514. [Google Scholar] [CrossRef]
- Figueroa, F.L.; Bonomi Barufi, J.; Malta, E.J.; Conde-Álvarez, R.; Nitschke, U.; Arenas, F.; Mata, M.; Connan, S.; Abreu, M.H.; Marquardt, R.; et al. Short-term effects of increasing CO2, nitrate and temperature on three mediterranean macroalgae: Biochemical composition. Aquat. Biol. 2014, 22, 177–193. [Google Scholar] [CrossRef]
- Suh, S.S.; Hwang, J.; Park, M.; Seo, H.H.; Kim, H.S.; Lee, J.H.; Moh, S.H.; Lee, T.K. Anti-inflammation activities of mycosporine-like amino acids (MAAs) in response to UV radiation suggest potential anti-skin aging activity. Mar. Drugs 2014, 12, 5174–5187. [Google Scholar] [CrossRef] [PubMed]
- Banaszak, A.T.; Barba Santos, M.G.; LaJeunesse, T.C.; Lesser, M.P. The distribution of mycosporine-like amino acids (MAAs) and the phylogenetic identity of symbiotic dinoflagellates in cnidarian hosts from the Mexican Caribbean. J. Exp. Mar. Biol. Ecol. 2006, 337, 131–146. [Google Scholar] [CrossRef]
- Vale, P. Effects of Light and Salinity Stresses in Production of Mycosporine-Like Amino Acids by Gymnodinium catenatum (Dinophyceae). Photochem. Photobiol. 2015, 91, 1112–1122. [Google Scholar] [CrossRef] [PubMed]
- Vale, P. Can Mycosporine-Like Amino Acids Act as Multifunctional Compounds in Gymnodinium catenatum (Dinophyceae)? Photochem. Photobiol. 2016, 92, 264–275. [Google Scholar] [CrossRef] [PubMed]
- Bandaranayake, W.M.; Bemis, J.E.; Bourne, D.J. Ultraviolet absorbing pigments from the marine sponge Dysidea herbacea: Isolation and structure of a new mycosporine. Comp. Biochem. Physiol. Part C Pharmacol. Toxicol. Endocrinol. 1996, 115, 281–286. [Google Scholar] [CrossRef]
- Oda, Y.; Zhang, Q.; Matsunaga, S.; Fujita, M.J.; Sakai, R. Two new mycosporine-like amino acids LC-343 and mycosporine-ethanolamine from the micronesian marine sponge Lendenfeldia chondrodes. Chem. Lett. 2017, 46, 1272–1274. [Google Scholar] [CrossRef]
- Ferrier-Pagès, C.; Richard, C.; Forcioli, D.; Allemand, D.; Pichon, M.; Shick, J.M. Effects of temperature and UV radiation increases on the photosynthetic efficiency in four scleractinian coral species. Biol. Bull. 2007, 213, 76–87. [Google Scholar] [CrossRef] [PubMed]
- Yakovleva, I.M.; Baird, A.H. Ontogenetic change in the abundance of mycosporine-like amino acids in non-zooxanthellate coral larvae. Coral Reefs 2005, 24, 443–452. [Google Scholar] [CrossRef]
- Adams, N.L.; Shick, J.M. Mycosporine-like amino acids prevent UVB-induced abnormalities during early development of the green sea urchin Strongylocentrotus droebachiensis. Mar. Biol. 2001, 138, 267–280. [Google Scholar] [CrossRef]
- Carroll, A.K.; Shick, J.M. Dietary accumulation of UV-absorbing mycosporine-like amino acids (MAAs) by the green sea urchin (Strongylocentrotus droebachiensis). Mar. Biol. 1996, 124, 561–569. [Google Scholar] [CrossRef]
- Eckes, M.; Dove, S.; Siebeck, U.E.; Grutter, A.S. Fish mucus versus parasitic gnathiid isopods as sources of energy and sunscreens for a cleaner fish. Coral Reefs 2015, 34, 823–833. [Google Scholar] [CrossRef]
- Newman, S.J.; Dunlap, W.C.; Nicol, S.; Ritz, D. Antarctic krill (Euphausia superba) acquire a UV-absorbing mycosporine-like amino acid from dietary algae. J. Exp. Mar. Biol. Ecol. 2000, 255, 93–110. [Google Scholar] [CrossRef]
- Osborn, A.R.; Almabruk, K.H.; Holzwarth, G.; Asamizu, S.; LaDu, J.; Kean, K.M.; Karplus, P.A.; Tanguay, R.L.; Bakalinsky, A.T.; Mahmud, T. De novo synthesis of a sunscreen compound in vertebrates. Elife 2015, 4, e05919. [Google Scholar] [CrossRef] [PubMed]
- Brotherton, C.A.; Balskus, E.P. Shedding light on sunscreen biosynthesis in zebrafish. eLife 2015, 4, e07961. [Google Scholar] [CrossRef] [PubMed]
- Rosic, N.N.; Braun, C.; Kvaskoff, D. Extraction and Analysis of Mycosporine-Like Amino Acids in Marine Algae. Methods Mol. Biol. 2015, 1308, 119–129. [Google Scholar] [PubMed]
- Orfanoudaki, M.; Hartmann, A.; Miladinovic, H.; Ngoc, H.N.; Karsten, U.; Ganzera, M. Bostrychines A–F, Six Novel Mycosporine-Like Amino-Acids and a Novel Betaine from the Red Alga Bostrychia scorpioides. Mar. Drugs 2019, 17, 356. [Google Scholar] [CrossRef] [PubMed]
- Rastogi, R.P.; Incharoensakdi, A. UV radiation-induced biosynthesis, stability and antioxidant activity of mycosporine-like amino acids (MAAs) in a unicellular cyanobacterium Gloeocapsa sp. CU2556. J. Photochem. Photobiol. B Biol. 2014, 130, 287–292. [Google Scholar] [CrossRef] [PubMed]
- Shinzato, C.; Shoguchi, E.; Kawashima, T.; Hamada, M.; Hisata, K.; Tanaka, M.; Fujie, M.; Fujiwara, M.; Koyanagi, R.; Ikuta, T.; et al. Using the Acropora digitifera genome to understand coral responses to environmental change. Nature 2011, 476, 320–323. [Google Scholar] [CrossRef] [PubMed]
- Carignan, M.O.; Cardozo, K.H.M.; Oliveira-Silva, D.; Colepicolo, P.; Carreto, J.I. Palythine-threonine, a major novel mycosporine-like amino acid (MAA) isolated from the hermatypic coral Pocillopora capitata. J. Photochem. Photobiol. B Biol. 2009, 94, 191–200. [Google Scholar] [CrossRef] [PubMed]
- D’Agostino, P.M.; Javalkote, V.S.; Mazmouz, R.; Pickford, R.; Puranik, P.R.; Neilan, B.A. Comparative profiling and discovery of novel glycosylated mycosporine-like amino acids in two strains of the cyanobacterium Scytonema cf. crispum. Appl. Environ. Microbiol. 2016, 82, 5951–5959. [Google Scholar] [CrossRef] [PubMed]
- Ngoennet, S.; Nishikawa, Y.; Hibino, T.; Waditee-Sirisattha, R.; Kageyama, H. A method for the isolation and characterization of mycosporine-like amino acids from cyanobacteria. Methods Protoc. 2018, 1, 46. [Google Scholar] [CrossRef] [PubMed]
- Kageyama, H.; Waditee-Sirisattha, R. Antioxidative, anti-inflammatory; anti-aging properties of mycosporine-like amino acids: Molecular and cellular mechanisms in the protection of skin-aging. Mar. Drugs 2019, 17, 222. [Google Scholar] [CrossRef] [PubMed]
- Kaidbey, K.H.; Agin, P.P.; Sayre, R.M.; Kligman, A.M. Photoprotection by melanin—A comparison of black and Caucasian skin. J. Am. Acad. Dermatol. 1979, 1, 249–260. [Google Scholar] [CrossRef]
- Brenner, M.; Hearing, V.J. The protective role of melanin against UV damage in human skin. Photochem. Photobiol. 2008, 84, 539–549. [Google Scholar] [CrossRef] [PubMed]
- Jain, S.; Prajapat, G.; Abrar, M.; Ledwani, L.; Singh, A.; Agrawal, A. Cyanobacteria as efficient producers of mycosporine-like amino acids. J. Basic Microbiol. 2017, 57, 715–727. [Google Scholar] [CrossRef] [PubMed]
- Rastogi, R.P.; Sinha, R.P.; Incharoensakdi, A. Partial characterization, UV-induction and photoprotective function of sunscreen pigment, scytonemin from Rivularia sp. HKAR-4. Chemosphere 2013, 93, 1874–1878. [Google Scholar] [CrossRef] [PubMed]
- Rastogi, R.P.; Sonani, R.R.; Madamwar, D. Cyanobacterial Sunscreen Scytonemin: Role in Photoprotection and Biomedical Research. Appl. Biochem. Biotechnol. 2015, 176, 1551–1563. [Google Scholar] [CrossRef] [PubMed]
- Pathak, J.; Pandey, A.; Maurya, P.K.; Rajneesh, R.; Sinha, R.P.; Singh, S.P. Cyanobacterial Secondary Metabolite Scytonemin: A Potential Photoprotective and Pharmaceutical Compound. Proc. Natl. Acad. Sci. India Sect. B Biol. Sci. 2019, 89, 1–15. [Google Scholar] [CrossRef]
- Garcia-Pichel, F.; Bebout, B.M. Penetration of ultraviolet radiation into shallow water sediments: High exposure for photosynthetic communities. Mar. Ecol. Prog. Ser. 1996, 131, 257–262. [Google Scholar] [CrossRef]
- Wada, N.; Sakamoto, T.; Matsugo, S. Mycosporine-like amino acids and their derivatives as natural antioxidants. Antioxidants 2015, 4, 603–646. [Google Scholar] [CrossRef] [PubMed]
- De La Coba, F.; Aguilera, J.; Figueroa, F.L.; De Gálvez, M.V.; Herrera, E. Antioxidant activity of mycosporine-like amino acids isolated from three red macroalgae and one marine lichen. J. Appl. Phycol. 2009, 21, 161–169. [Google Scholar] [CrossRef]
- Rastogi, R.P.; Richa; Singh, S.P.; Häder, D.P.; Sinha, R.P. Mycosporine-like amino acids profile and their activity under PAR and UVR in a hot-spring cyanobacterium Scytonema sp. HKAR-3. Aust. J. Bot. 2010, 58, 286–293. [Google Scholar] [CrossRef]
- Suh, H.J.; Lee, H.W.; Jung, J. Mycosporine Glycine Protects Biological Systems Against Photodynamic Damage by Quenching Singlet Oxygen with a High Efficiency. Photochem. Photobiol. 2003, 78, 109–113. [Google Scholar] [CrossRef]
- Gacesa, R.; Lawrence, K.P.; Georgakopoulos, N.D.; Yabe, K.; Dunlap, W.C.; Barlow, D.J.; Wells, G.; Young, A.R.; Long, P.F. The mycosporine-like amino acids porphyra-334 and shinorine are antioxidants and direct antagonists of Keap1-Nrf2 binding. Biochimie 2018, 154, 35–44. [Google Scholar] [CrossRef] [PubMed]
- Becker, K.; Hartmann, A.; Ganzera, M.; Fuchs, D.; Gostner, J.M. Immunomodulatory Effects of the Mycosporine-Like Amino Acids Shinorine and Porphyra-334. Mar. Drugs 2016, 14, 119. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, A.; Gostner, J.; Fuchs, J.E.; Chaita, E.; Aligiannis, N.; Skaltsounis, L.; Ganzera, M. Inhibition of Collagenase by Mycosporine-like Amino Acids from Marine Sources. Planta Med. 2015, 81, 813–820. [Google Scholar] [CrossRef] [PubMed]
- Yoshiki, M.; Tsuge, K.; Tsuruta, Y.; Yoshimura, T.; Koganemaru, K.; Sumi, T.; Matsui, T.; Matsumoto, K. Production of new antioxidant compound from mycosporine-like amino acid, porphyra-334 by heat treatment. Food Chem. 2009, 113, 1127–1132. [Google Scholar] [CrossRef]
- Ryu, J.; Park, S.J.; Kim, I.H.; Choi, Y.H.; Nam, T.J. Protective effect of porphyra-334 on UVA-induced photoaging in human skin fibroblasts. Int. J. Mol. Med. 2014, 34, 796–803. [Google Scholar] [CrossRef] [PubMed]
- Cheewinthamrongrod, V.; Kageyama, H.; Palaga, T.; Takabe, T.; Waditee-Sirisattha, R. DNA damage protecting and free radical scavenging properties of mycosporine-2-glycine from the Dead Sea cyanobacterium in A375 human melanoma cell lines. J. Photochem. Photobiol. B Biol. 2016, 164, 289–295. [Google Scholar] [CrossRef] [PubMed]
- Tarasuntisuk, S.; Palaga, T.; Kageyama, H.; Waditee-Sirisattha, R. Mycosporine-2-glycine exerts anti-inflammatory and antioxidant effects in lipopolysaccharide (LPS)-stimulated RAW 264.7 macrophages. Arch. Biochem. Biophys. 2019, 662, 33–39. [Google Scholar] [CrossRef] [PubMed]
- Lawrence, K.P.; Gacesa, R.; Long, P.F.; Young, A.R. Molecular photoprotection of human keratinocytes in vitro by the naturally occurring mycosporine-like amino acid palythine. Br. J. Dermatol. 2018, 178, 1353–1363. [Google Scholar] [CrossRef] [PubMed]
- Dunlap, W.C.; Shick, J.M. Ultraviolet radiation-absorbing mycosporine-like amino acids in coral reef organisms: A biochemical and environmental perspective. J. Phycol. 1998, 34, 418–430. [Google Scholar] [CrossRef]
- Suzuki, T.; Yamamoto, M. Molecular basis of the Keap1–Nrf2 system. Free Radic. Biol. Med. 2015, 88, 93–100. [Google Scholar] [CrossRef] [PubMed]
- Panich, U.; Sittithumcharee, G.; Rathviboon, N.; Jirawatnotai, S. Ultraviolet radiation-induced skin aging: The role of DNA damage and oxidative stress in epidermal stem cell damage mediated skin aging. Stem Cells Int. 2016, 2016. [Google Scholar] [CrossRef] [PubMed]
- Yakovleva, I.; Bhagooli, R.; Takemura, A.; Hidaka, M. Differential susceptibility to oxidative stress of two scleractinian corals: Antioxidant functioning of mycosporine-glycine. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 2004, 139, 721–730. [Google Scholar] [CrossRef] [PubMed]
- Ma, Q. Role of Nrf2 in oxidative stress and toxicity. Annu. Rev. Pharmacol. Toxicol. 2013, 53, 401–426. [Google Scholar] [CrossRef] [PubMed]
- Siomek, A. NF-κB signaling pathway and free radical impact. Acta Biochim. Pol. 2012, 59, 323–331. [Google Scholar] [CrossRef] [PubMed]
- Álvarez-Gómez, F.; Korbee, N.; Casas-Arrojo, V.; Abdala-Díaz, R.T.; Figueroa, F.L. UV photoprotection, cytotoxicity and immunology capacity of red algae extracts. Molecules 2019, 24, 341. [Google Scholar] [CrossRef] [PubMed]
- Grimbaldeston, M.A.; Simpson, A.; Finlay-Jones, J.J.; Hart, P.H. The effect of ultraviolet radiation exposure on the prevalence of mast cells in human skin. Br. J. Dermatol. 2003, 148, 300–306. [Google Scholar] [CrossRef] [PubMed]
- Bhandari, R.; Sharma, P.K. Effect of UV-B and high visual radiation on photosynthesis in freshwater (Nostoc spongiaeforme) and marine (Phormidium corium) cyanobacteria. Indian J. Biochem. Biophys. 2007, 44, 231–239. [Google Scholar] [PubMed]
- Gao, K.; Xu, J. Effects of solar UV radiation on diurnal photosynthetic performance and growth of Gracilaria lemaneiformis (Rhodophyta). Eur. J. Phycol. 2008, 43, 297–307. [Google Scholar] [CrossRef]
- Rui, Y.; Zhaohui, Z.; Wenshan, S.; Bafang, L.; Hu, H. Protective effect of MAAs extracted from Porphyra tenera against UV irradiation-induced photoaging in mouse skin. J. Photochem. Photobiol. B Biol. 2019, 192, 26–33. [Google Scholar] [CrossRef] [PubMed]
- Oren, A.; Gunde-Cimerman, N. Mycosporines and mycosporine-like amino acids: UV protectants or multipurpose secondary metabolites? FEMS Microbiol. Lett. 2007, 269, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Michalek-Wagner, K.; Willis, B.L. Impacts of bleaching on the soft coral Lobophytum compactum. Ii. Biochemical changes in adults and their eggs. Coral Reefs 2001, 19, 240–246. [Google Scholar] [CrossRef]
- Al-Utaibi, A.A.; Niaz, G.R.; Al-Lihaibi, S.S. Mycosporine-like amino acids in six scleractinian coral species. Oceanologia 2009, 51, 93–104. [Google Scholar] [CrossRef]
- Bohm, G.A.; Pfleiderer, W.; Boger, P.; Scherer, S. Structure of a novel oligosaccharide-mycosporine-amino acid ultraviolet A/B sunscreen pigment from the terrestrial cyanobacterium Nostoc commune. J. Biol. Chem. 1995, 270, 8536–8539. [Google Scholar] [CrossRef] [PubMed]
- Hernando, M.; Carreto, J.I.; Carignan, M.O.; Ferreyra, G.A.; Gross, C. Effects of solar radiation on growth and mycosporine-like amino acids content in Thalassiosira sp, an Antarctic diatom. Polar Biol. 2002, 25, 12–20. [Google Scholar]
- Laurion, I.; Roy, S. Growth and photoprotection in three dinoflagellates (including two strains of Alexandrium tamarense) and one diatom exposed to four weeks of natural and enhanced ultraviolet-b radiation. J. Phycol. 2009, 45, 16–33. [Google Scholar] [CrossRef] [PubMed]
- Riegger, L.; Robinson, D. Photoinduction of UV-absorbing compounds in Antarctic diatoms and Phaeocystis antarctica. Mar. Ecol. Prog. Ser. 1997, 160, 13–25. [Google Scholar] [CrossRef]
- Taira, H.; Aoki, S.; Yamanoha, B.; Taguchi, S. Daily variation in cellular content of UV-absorbing compounds mycosporine-like amino acids in the marine dinoflagellate Scrippsiella sweeneyae. J. Photochem. Photobiol. B Biol. 2004, 75, 145–155. [Google Scholar] [CrossRef]
- Franklin, L.A.; Kräbs, G.; Kuhlenkamp, R. Blue light and UV-A radiation control the synthesis of mycosporine-like amino acids in Chondrus crispus (Florideophyceae). J. Phycol. 2001, 37, 257–270. [Google Scholar] [CrossRef]
- Sinha, R.P.; Ambasht, N.K.; Sinha, J.P.; Klisch, M.; Häder, D.P. UV-B-induced synthesis of mycosporine-like amino acids in three strains of Nodularia (cyanobacteria). J. Photochem. Photobiol. B Biol. 2003, 71, 51–58. [Google Scholar] [CrossRef] [PubMed]
- Lesser, M.P. Depth-dependent photoacclimatization to solar ultraviolet radiation in the Caribbean coral Montastraea faveolata. Mar. Ecol. Prog. Ser. 2000, 192, 137–151. [Google Scholar] [CrossRef]
- Shick, J.M.; Romaine-Lioud, S.; Ferrier-Pagès, C.; Gattuso, J.P. Ultraviolet-B radiation stimulates shikimate pathway-dependent accumulation of mycosporine-like amino acids in the coral Stylophora pistillata despite decreases in its population of symbiotic dinoflagellates. Limnol. Oceanogr. 1999, 44, 1667–1682. [Google Scholar] [CrossRef]
- Banaszak, A.T.; Lesser, M.P. Effects of solar ultraviolet radiation on coral reef organisms. Photochem. Photobiol. Sci. 2009, 8, 1276–1294. [Google Scholar] [CrossRef] [PubMed]
- Portwich, A.; Garcia-Pichel, F. Biosynthetic pathway of mycosporines (mycosporine-like amino acids) in the cyanobacterium Chlorogloeopsis sp. strain PCC 6912. Phycologia 2003, 42, 384–392. [Google Scholar] [CrossRef]
- Rath, J.; Mandal, S.; Adhikary, S.P. Ecophysiology of the estuarine cyanobacterium Lyngbya aestuarii to varying salinity in vitro. Acta Physiol. Plant. 2014, 36, 409–419. [Google Scholar] [CrossRef]
- Singh, S.P.; Klisch, M.; Sinha, R.P.; Häder, D.P. Effects of abiotic stressors on synthesis of the mycosporine-like amino acid shinorine in the cyanobacterium Anabaena variabilis PCC 7937. Photochem. Photobiol. 2008, 84, 1500–1505. [Google Scholar] [CrossRef] [PubMed]
- Whitehead, K.; Hedges, J.I. Electrospray ionization tandem mass spectrometric and electron impact mass spectrometric characterization of mycosporine-like amino acids. Rapid Commun. Mass Spectrom. 2003, 17, 2133–2138. [Google Scholar] [CrossRef] [PubMed]
- Waditee-Sirisattha, R.; Kageyama, H.; Sopun, W.; Tanaka, Y.; Takabe, T. Identification and upregulation of biosynthetic genes required for accumulation of mycosporine-2-glycine under salt stress conditions in the halotolerant cyanobacterium Aphanothece halophytica. Appl. Environ. Microbiol. 2014, 80, 1763–1769. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.N.; Zhang, Z.C.; Feng, J.L.; Qiu, B.S. Effects of UV-B radiation and periodic desiccation on the morphogenesis of the edible terrestrial cyanobacterium Nostoc flagelliforme. Appl. Environ. Microbiol. 2012, 78, 7075–7081. [Google Scholar] [CrossRef] [PubMed]
- Tartarotti, B.; Sommaruga, R. Seasonal and ontogenetic changes of mycosporine-like amino acids in planktonic organisms from an alpine lake. Limnol. Oceanogr. 2006, 51, 1530–1541. [Google Scholar] [CrossRef] [PubMed]
- Rosic, N.N.; Dove, S. Mycosporine-like amino acids from coral dinoflagellates. Appl. Environ. Microbiol. 2011, 77, 8478–8486. [Google Scholar] [CrossRef] [PubMed]
- Balskus, E.P.; Walsh, C.T. The genetic and molecular basis for sunscreen biosynthesis in cyanobacteria. Science 2010, 329, 1653–1656. [Google Scholar] [CrossRef] [PubMed]
- Miyamoto, K.T.; Komatsu, M.; Ikeda, H. Discovery of gene cluster for mycosporine-like amino acid biosynthesis from Actinomycetales microorganisms and production of a novel mycosporine-like amino acid by heterologous expression. Appl. Environ. Microbiol. 2014, 80, 5028–5036. [Google Scholar] [CrossRef] [PubMed]
- D’Agostino, P.M.; Woodhouse, J.N.; Liew, H.T.; Sehnal, L.; Pickford, R.; Wong, H.L.; Burns, B.P.; Neilan, B.A. Bioinformatic, phylogenetic and chemical analysis of the UV-absorbing compounds scytonemin and mycosporine-like amino acids from the microbial mat communities of Shark Bay, Australia. Environ. Microbiol. 2019, 21, 702–715. [Google Scholar] [CrossRef] [PubMed]
- Gao, Q.; Garcia-Pichel, F. An ATP-Grasp ligase involved in the last biosynthetic step of the iminomycosporine shinorine in Nostoc punctiforme ATCC 29133. J. Bacteriol. 2011, 193, 5923–5928. [Google Scholar] [CrossRef] [PubMed]
- Miller, A.J.; Mihm, M.C., Jr. Melanoma. N. Engl. J. Med. 2006, 355, 51–65. [Google Scholar] [CrossRef] [PubMed]
- Savoye, I.; Olsen, C.M.; Whiteman, D.C.; Bijon, A.; Wald, L.; Dartois, L.; Clavel-Chapelon, F.; Boutron-Ruault, M.C.; Kvaskoff, M. Patterns of Ultraviolet Radiation Exposure and Skin Cancer Risk: The E3N-SunExp Study. J. Epidemiol. 2018, 28, 27–33. [Google Scholar] [CrossRef] [PubMed]
- Ichihashi, M.; Ueda, M.; Budiyanto, A.; Bito, T.; Oka, M.; Fukunaga, M.; Tsuru, K.; Horikawa, T. UV-induced skin damage. Toxicology 2003, 189, 21–39. [Google Scholar] [CrossRef]
- Ikehata, H.; Ono, T. The mechanisms of UV mutagenesis. J. Radiat. Res. 2011, 52, 115–125. [Google Scholar] [CrossRef] [PubMed]
- Napolitano, A.; Panzella, L.; Monfrecola, G.; d’Ischia, M. Pheomelanin-induced oxidative stress: Bright and dark chemistry bridging red hair phenotype and melanoma. Pigment. Cell Melanoma Res. 2014, 27, 721–733. [Google Scholar] [CrossRef] [PubMed]
- Premi, S.; Wallisch, S.; Mano, C.M.; Weiner, A.B.; Bacchiocchi, A.; Wakamatsu, K.; Bechara, E.J.; Halaban, R.; Douki, T.; Brash, D.E. Photochemistry. Chemiexcitation of melanin derivatives induces DNA photoproducts long after UV exposure. Science 2015, 347, 842–847. [Google Scholar] [CrossRef] [PubMed]
- Rykowska, I.; Wasiak, W. Research trends on emerging environment pollutants—A review. Open Chem. 2015, 13, 1353–1370. [Google Scholar] [CrossRef]
- Downs, C.A.; Kramarsky-Winter, E.; Segal, R.; Fauth, J.; Knutson, S.; Bronstein, O.; Ciner, F.R.; Jeger, R.; Lichtenfeld, Y.; Woodley, C.M.; et al. Toxicopathological Effects of the Sunscreen UV Filter, Oxybenzone (Benzophenone-3), on Coral Planulae and Cultured Primary Cells and Its Environmental Contamination in Hawaii and the U.S. Virgin Islands. Arch. Environ. Contam. Toxicol. 2016, 70, 265–288. [Google Scholar] [CrossRef] [PubMed]
- Chrapusta, E.; Kaminski, A.; Duchnik, K.; Bober, B.; Adamski, M.; Bialczyk, J. Mycosporine-Like Amino Acids: Potential health and beauty ingredients. Mar. Drugs 2017, 15, 326. [Google Scholar] [CrossRef] [PubMed]
- Lawrence, K.P.; Long, P.F.; Young, A.R. Mycosporine-like amino acids for skin photoprotection. Curr. Med. Chem. 2018, 25, 5512–5527. [Google Scholar] [CrossRef] [PubMed]
- Gröniger, A.; Häder, D.P. Stability of mycosporine-like amino acids. Recent Res. Dev. Photochem. Photobiol. 2000, 4, 247–252. [Google Scholar]
- Schmidt, E.W. An Enzymatic Route to Sunscreens. ChemBioChem 2011, 12, 363–365. [Google Scholar] [CrossRef] [PubMed]

| MAAs | Activity: UV-Absorbing | Activity: Antioxidative | Activity: Anti- Inflammatory | Activity: Antiaging | Sources of MAAs and References |
|---|---|---|---|---|---|
| Mycosporine-glycine | Yes ʎmax = 310 nm | Yes [24,31,51,61,63] | Yes [31] | Yes [31] | Red alga Porphyra tenera [24] Green alga Chlamydomonas hedleyi [31] Ascidian Lissoclinum patella [63] Marine lichen Lichina pygmaea [61] |
| Shinorine | Yes ʎmax = 333 nm | Yes [24,51,64] No [31] | Yes [31] No [65] | Yes [31,66] | Red alga Chondrus yendoi [64] Red alga Porphyra sp. [65] Green alga Chlamydomonas hedleyi [31] Red algae Porphyra sp. & Palmaria palmate [66] |
| Porphyra-334 | Yes ʎmax = 334 nm | Yes [24,51,64,67,68] No [31] | Yes [65,68] No [31] | Yes [31,66] No [31] | Green alga Chlamydomonas hedleyi [31] Red algae Porphyra sp. & Palmaria palmate [66] Red alga Porphyra yezoensis [68] |
| Mycosporine-2-glycine | Yes ʎmax = 332 nm | Yes [51,69,70] | Yes [70] | Yes [70] | Cyanobacterium Aphanothece halophytica [51,69,70] |
| Palythine | Yes ʎmax = 320 nm | Yes [71] No [69] | - | Yes [66] | Red alga Chrondus yendoi [71] Cyanobacterium Aphanothece halophytica [69] Red algae Porphyra sp. & Palmaria palmate [66] Cyanobacterium Aphanothece halophytica [69] |
© 2019 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rosic, N.N. Mycosporine-Like Amino Acids: Making the Foundation for Organic Personalised Sunscreens. Mar. Drugs 2019, 17, 638. https://doi.org/10.3390/md17110638
Rosic NN. Mycosporine-Like Amino Acids: Making the Foundation for Organic Personalised Sunscreens. Marine Drugs. 2019; 17(11):638. https://doi.org/10.3390/md17110638
Chicago/Turabian StyleRosic, Nedeljka N. 2019. "Mycosporine-Like Amino Acids: Making the Foundation for Organic Personalised Sunscreens" Marine Drugs 17, no. 11: 638. https://doi.org/10.3390/md17110638
APA StyleRosic, N. N. (2019). Mycosporine-Like Amino Acids: Making the Foundation for Organic Personalised Sunscreens. Marine Drugs, 17(11), 638. https://doi.org/10.3390/md17110638





