Group-1 Grass Pollen Allergens with Near-Identical Sequences Identified in Species of Subtropical Grasses Commonly Found in Southeast Asia
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. RNA Extraction and cDNA Synthesis
2.3. RACE PCR Cloning
2.4. Cloning and Sequencing
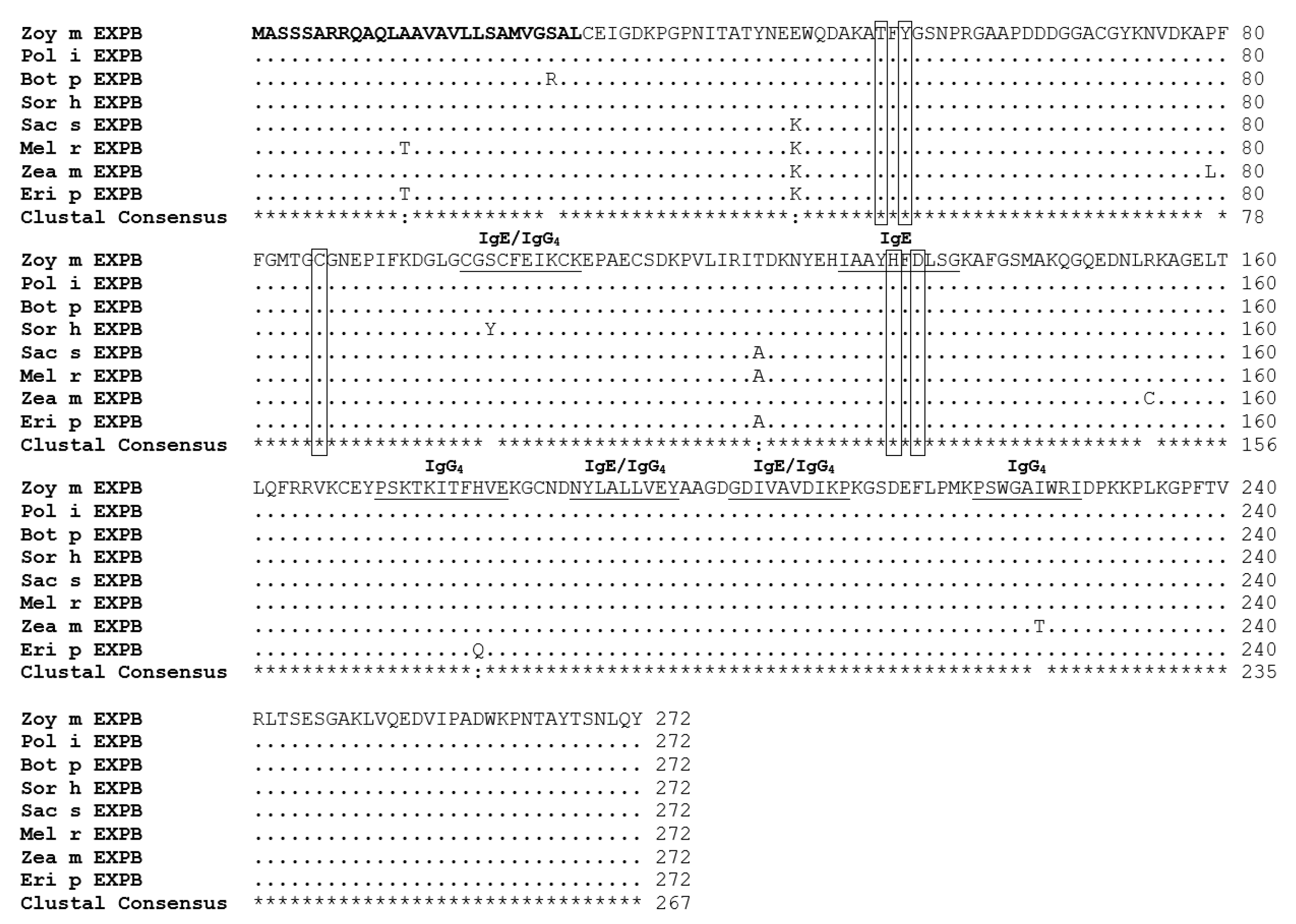
2.5. Sequence Analysis
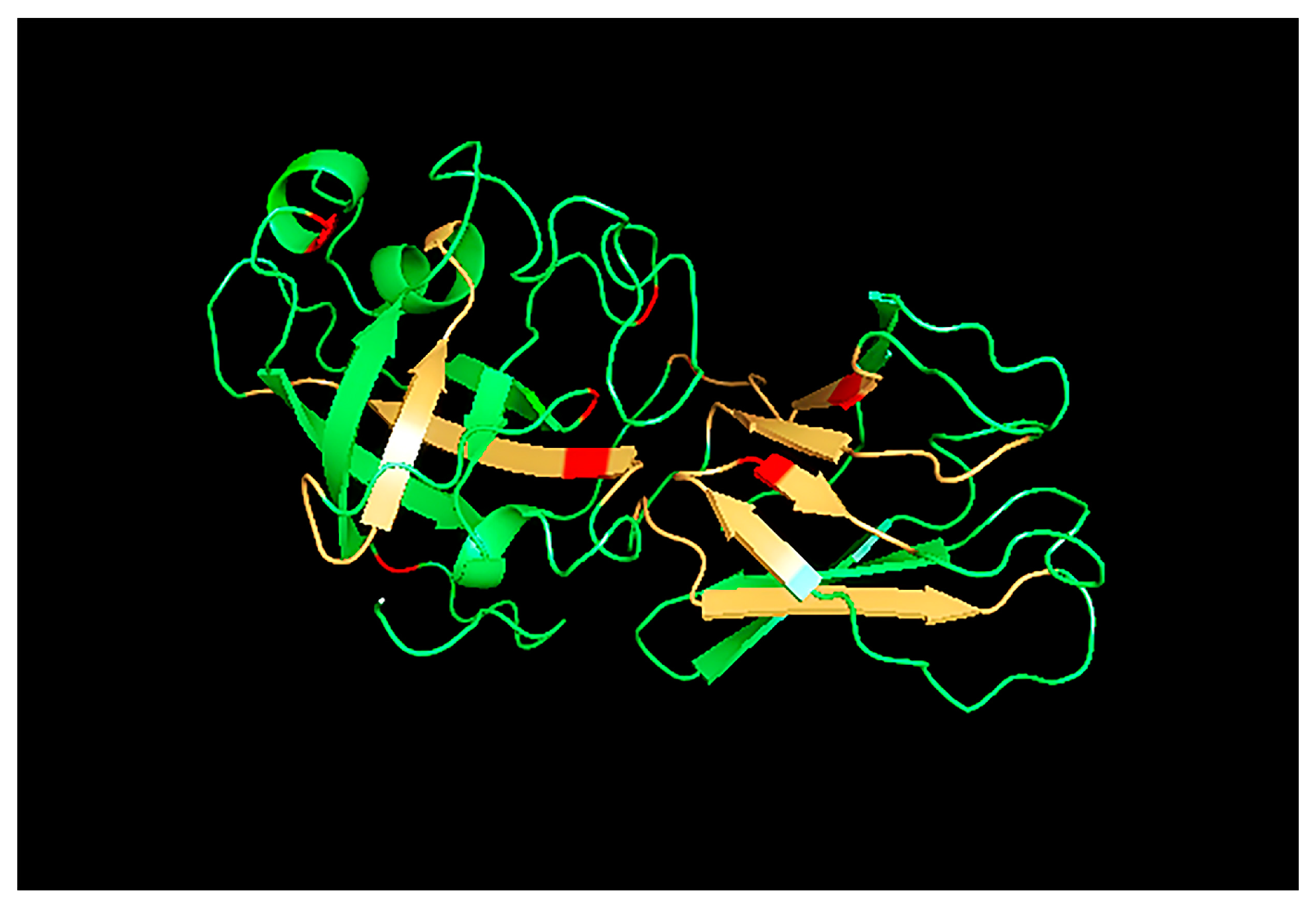
2.6. Protein Structure Prediction
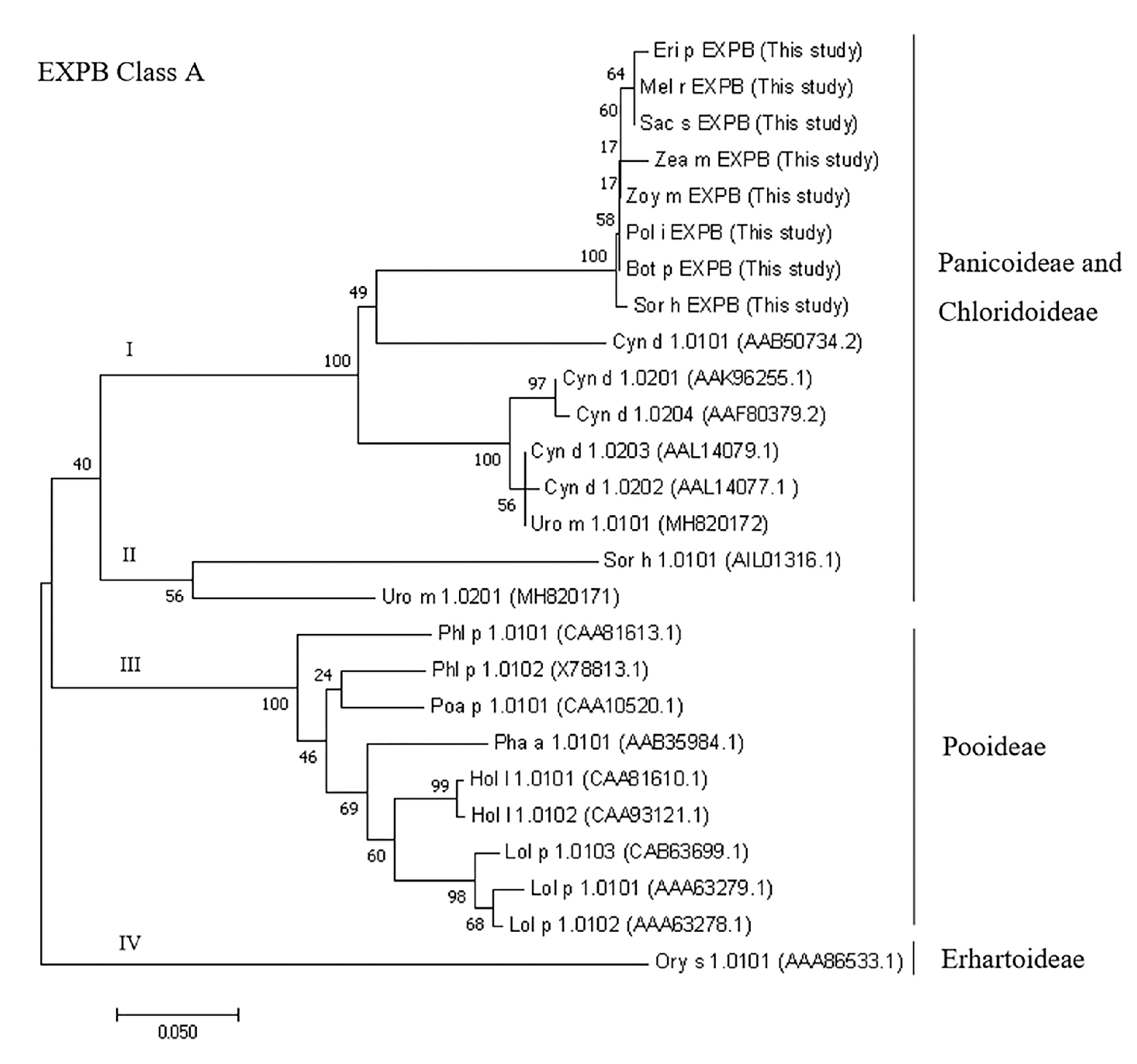
2.7. Phylogenetic Analysis
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hrabina, M.; Peltre, G.; Van Ree, R.; Moingeon, P. Grass pollen allergens. Clin. Exp. Allergy Rev. 2008, 8, 7–11. [Google Scholar]
- Valenta, R.; Vrtala, S.; Ebner, C.; Kraft, D.; Scheiner, O. Diagnosis of grass pollen allergy with recombinant timothy grass (Phleum pratense) pollen allergens. Int. Arch. Allergy Immunol. 1992, 97, 287–294. [Google Scholar]
- Sastre, J. Molecular diagnosis in allergy. Clin. Exp. Allergy J. Br. Soc. Allergy Clin. Immunol. 2010, 40, 1442–1460. [Google Scholar]
- Kende, H.; Bradford, K.; Brummell, D.; Cho, H.-T.; Cosgrove, D.; Fleming, A.; Gehring, C.; Lee, Y.; McQueen-Mason, S.; Rose, J.; et al. Nomenclature for Members of the Expansin Superfamily of Genes and Proteins; Kluwer Academic Publishers: Alphen aan den Rijn, The Netherlands, 2004; Volume 55, pp. 311–314. [Google Scholar]
- Cosgrove, D.J. Loosening of plant cell walls by expansins. Nature 2000, 407, 321–326. [Google Scholar]
- Yennawar, N.H.; Li, L.-C.; Dudzinski, D.M.; Tabuchi, A.; Cosgrove, D.J. Crystal structure and activities of EXPB1 (Zea m 1), a β-expansin and group-1 pollen allergen from maize. Proc. Natl. Acad. Sci. USA 2006, 103, 14664–14671. [Google Scholar]
- Cosgrove, D.J.; Bedinger, P.; Durachko, D.M. Group I allergens of grass pollen as cell wall-loosening agents. Proc. Natl. Acad. Sci. USA 1997, 94, 6559–6564. [Google Scholar]
- Gangl, K.; Niederberger, V.; Valenta, R.; Nandy, A. Marker Allergens and Panallergens in Tree and Grass Pollen Allergy; Springer International Publishing: Cham, Switzerland, 2015; Volume 24, pp. 158–169. [Google Scholar]
- Morrison, I.C. The changing face of pollen allergy in Queensland. Med. J. Aust. 1984, 141 (Suppl. 5), S12–S13. [Google Scholar]
- Pumhirun, P.; Towiwat, P.; Mahakit, P. Aeroallergen sensitivity of Thai patients with allergic rhinitis. Asian Pac. J. Allergy Immunol. 1997, 15, 183–185. [Google Scholar]
- Liang, K.L.; Su, M.C.; Shiao, J.Y.; Wu, S.H.; Li, Y.H.; Jiang, R.S. Role of pollen allergy in Taiwanese patients with allergic rhinitis. J. Formos. Med Assoc. 2010, 109, 879–885. [Google Scholar]
- Prasad, R.; Verma, S.K.; Dua, R.; Kant, S.; Kushwaha, R.A.; Agarwal, S.P. A study of skin sensitivity to various allergens by skin prick test in patients of nasobronchial allergy. Lung India Off. Organ Indian Chest Soc. 2009, 26, 70–73. [Google Scholar]
- Bunnag, C. Airborne pollens in Bangkok, Thailand: An update. Asian Rhinol. J. 2016, 3, 47–51. [Google Scholar]
- Sorensen, H.; Ashoor, A.A.; Maglad, S. Perennial rhinitis in Saudi Arabia. A prospective study. Ann Allergy 1986, 56, 76–80. [Google Scholar]
- Sam, C.K.; Kesavan, P.; Liam, C.K.; Soon, S.C.; Lim, A.L.; Ong, E.K. A study of pollen prevalence in relation to pollen allergy in Malaysian asthmatics. Asian Pac. J. Allergy Immunol. 1998, 16, 1–4. [Google Scholar]
- Singh, A.B.; Shahi, S. Aeroallergens in clinical practice of allergy in India—ARIA Asia Pacific Workshop report. Asian Pac. J. Allergy Immunol. 2008, 26, 245–256. [Google Scholar]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- McWilliam, H.; Li, W.; Uludag, M.; Squizzato, S.; Park, Y.M.; Buso, N.; Cowley, A.P.; Lopez, R. Analysis Tool Web Services from the EMBL-EBI. Nucleic Acids Res. 2013, 41, W597–W600. [Google Scholar]
- Smith, P.M.; Suphioglu, C.; Griffith, I.J.; Theriault, K.; Knox, R.B.; Singh, M.B. Cloning and expression in yeast Pichia pastoris of a biologically active form of Cyn d 1, the major allergen of Bermuda grass pollen. J. Allergy Clin. Immunol. 1996, 98, 331–343. [Google Scholar]
- Chang, Z.N.; Peng, H.J.; Lee, W.C.; Chen, T.S.; Chua, K.Y.; Tsai, L.C.; Chi, C.W.; Han, S.H. Sequence polymorphism of the group 1 allergen of Bermuda grass pollen. Clin. Exp. Allergy J. Br. Soc. Allergy Clin. Immunol. 1999, 29, 488–496. [Google Scholar]
- Au, L.C.; Lin, S.T.; Peng, H.J.; Liang, C.C.; Lee, S.S.; Liao, C.D.; Chang, Z.N. Molecular cloning and sequence analysis of full-length cDNAs encoding new group of Cyn d 1 isoallergens. Allergy 2002, 57, 215–220. [Google Scholar]
- Yuan, H.C.; Wu, K.G.; Chen, C.J.; Su, S.N.; Shen, H.D.; Chen, Y.J.; Peng, H.J. Mapping of IgE and IgG4 antibody-binding epitopes in Cyn d 1, the major allergen of Bermuda grass pollen. Int. Arch. Allergy Immunol. 2012, 157, 125–135. [Google Scholar]
- Schwede, T.; Kopp, J.; Guex, N.; Peitsch, M.C. SWISS-MODEL: an automated protein homology-modeling server. Nucleic Acids Res. 2003, 31, 3381–3385. [Google Scholar]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar]
- Saitou, N.; Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar]
- Zuckerkandl, E.; Pauling, L. Molecules as documents of evolutionary history. J. Theor. Biol. 1965, 8, 357–366. [Google Scholar]
- Yampolsky, L.Y.; Stoltzfus, A. The Exchangeability of Amino Acids in Proteins. Genetics 2005, 170, 1459–1472. [Google Scholar]
- Davies, J.M.; Dang, T.D.; Voskamp, A.; Drew, A.C.; Biondo, M.; Phung, M.; Upham, J.W.; Rolland, J.M.; O’Hehir, R.E. Functional immunoglobulin E cross-reactivity between Pas n 1 of Bahia grass pollen and other group 1 grass pollen allergens. Clin. Exp. Allergy 2011, 41, 281–291. [Google Scholar]
- Campbell, B.C.; Gilding, E.K.; Timbrell, V.; Guru, P.; Loo, D.; Zennaro, D.; Mari, A.; Solley, G.; Hill, M.M.; Godwin, I.D.; et al. Total transcriptome, proteome, and allergome of Johnson grass pollen, which is important for allergic rhinitis in subtropical regions. J. Allergy Clin. Immunol. 2015, 135, 133–142. [Google Scholar]
- Perez, M.; Ishioka, G.Y.; Walker, L.E.; Chesnut, R.W. cDNA cloning and immunological characterization of the rye grass allergen Lol p I. J. Biol. Chem. 1990, 265, 16210–16215. [Google Scholar]
- Griffith, I.J.; Smith, P.M.; Pollock, J.; Theerakulpisut, P.; Avjioglu, A.; Davies, S.; Hough, T.; Singh, M.B.; Simpson, R.J.; Ward, L.D.; et al. Cloning sequencing of Lol pI, the major allergenic protein of rye-grass pollen. Febs Lett. 1991, 279, 210–215. [Google Scholar]
- Jones, D.H.A.; Christopher, F.; Franklin, H.; Thomas, C.M. Molecular analysis of the operon which encodes the RNA polymerase sigma factor O54 of Escherichia coli. Microbiology 1994, 140, 1035–1043. [Google Scholar]
- Schramm, G.; Petersen, A.; Bufe, A.; Schlaak, M.; Becker, W.M. Identification and characterization of the major allergens of velvet grass (Holcus lanatus), Hol l 1 and Hol l 5. Int. Arch. Allergy Immunol. 1996, 110, 354–363. [Google Scholar]
- Petersen, A.; Becker, W.M.; Moll, H.; Blumke, M.; Schlaak, M. Studies on the carbohydrate moieties of the timothy grass pollen allergen Phl p I. Electrophoresis 1995, 16, 869–875. [Google Scholar]
- Suphioglu, C.; Singh, M.B.; Simpson, R.J.; Ward, L.D.; Knox, R.B. Identification of canary grass (Phalaris aquatica) pollen allergens by immunoblotting: IgE and IgG antibody-binding studies. Allergy 1993, 48, 273–281. [Google Scholar]
- Johansen, N.; Weber, R.W.; Ipsen, H.; Barber, D.; Broge, L.; Hejl, C. Extensive IgE cross-reactivity towards the Pooideae grasses substantiated for a large number of grass-pollen-sensitized subjects. Int. Arch. Allergy Immunol. 2009, 150, 325–334. [Google Scholar]
- Davies, J.M. Grass pollen allergens globally: The contribution of subtropical grasses to burden of allergic respiratory diseases. Clin. Exp. Allergy 2014, 44, 790–801. [Google Scholar]
- Radauer, C.; Nandy, A.; Ferreira, F.; Goodman, R.E.; Larsen, J.N.; Lidholm, J.; Pomes, A.; Raulf-Heimsoth, M.; Rozynek, P.; Thomas, W.R.; et al. Update of the WHO/IUIS Allergen Nomenclature Database based on analysis of allergen sequences. Allergy 2014, 69, 413–419. [Google Scholar]
| Scientific Name | Common Name | Reported Allergenicity (Patient Group) | Near Identical EXPB (This Study) | GPS Coordinates |
|---|---|---|---|---|
| Zoysia matrella (L.) Merr. | Manila grass | No report | Zoy m EXPB | 14.079971, 99.702437 |
| Polytrias indica (Houtt.) Veldkamp | Batiki bluegrass | No report | Pol i EXPB | 13.850821, 100.527323 |
| Bothriochloa pertusa (L.) A.Camus | Hurricane grass | No report | Bot p EXPB | 14.113808, 99.150451 |
| Sorghum halepense (L.) Pers. | Johnson grass | 77% (48 GP- allergic), 21% (100 AR with + SPT to common inhalant allergens), 10% (419 AR) [9,10,11] | Sor h EXPB | 14.047468, 99.780618 |
| Saccharum spontaneum L. | Canne sauvage | No report | Sac s EXPB | 13.921569, 100.490671 |
| Zea mays L. | Maize, corn | 6.25% (48 patients suspected to have nasobronchial allergy) [12] | Zea m EXPB | 14.071024, 99.729950 |
| Melinis repens (Willd.) Zizka | Natal grass | No report | Mel r EXPB | 14.129449, 99.161192 |
| Eriochloa procera (Retz.) C.E.Hubb. | Cup grass | No report | Eri p EXPB | 13.850821, 100.527323 |
| Urochloa mutica (Forssk.) T.Q.Nguyen | Para grass | 53.2% (2,383 AR with + SPT to pollen) [13] | - | 13.740637, 99.920501 |
| Cynodon dactylon (L.) Pers. | Bermuda grass | 84% (48 GP-allergic and AR), 2.1% (419 AR), 12.5% (48 GP- allergic and AR), 54.9% (2,383 AR), 61% (54 Perennial AR), 20.5% (200 As), 52% (133 GP-allergic with + IDST) [9,11,12,13,14,15,16] | - | 14.075435, 99.698798 |
| Amino Acid | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
 | ||||||||||
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | |||
Nucleotide | 1 | 100.00 | 100.00 | 99.63 | 99.63 | 99.26 | 98.90 | 98.53 | 98.53 | 1. Zoy m EXPB |
| 2 | 99.76 | 100.00 | 99.63 | 99.63 | 99.26 | 98.90 | 98.53 | 98.53 | 2. Pol i EXPB | |
| 3 | 99.39 | 99.63 | 100.00 | 99.26 | 98.90 | 98.53 | 98.16 | 98.16 | 3. Bot p EXPB | |
| 4 | 99.63 | 99.88 | 99.51 | 100.00 | 98.90 | 98.53 | 98.16 | 98.16 | 4. Sor h EXPB | |
| 5 | 99.63 | 99.39 | 99.02 | 99.27 | 100.00 | 99.63 | 98.53 | 99.26 | 5. Sac s EXPB | |
| 6 | 99.63 | 99.39 | 99.02 | 99.27 | 99.76 | 100.00 | 98.16 | 99.63 | 6. Mel r EXPB | |
| 7 | 99.51 | 99.27 | 98.90 | 99.15 | 99.39 | 99.39 | 100.00 | 97.79 | 7. Zea m EXPB | |
| 8 | 99.39 | 99.15 | 98.78 | 99.02 | 99.51 | 99.76 | 99.15 | 100.00 | 8. Eri p EXPB | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aud-in, S.; Somkid, K.; Songnuan, W. Group-1 Grass Pollen Allergens with Near-Identical Sequences Identified in Species of Subtropical Grasses Commonly Found in Southeast Asia. Medicina 2019, 55, 193. https://doi.org/10.3390/medicina55050193
Aud-in S, Somkid K, Songnuan W. Group-1 Grass Pollen Allergens with Near-Identical Sequences Identified in Species of Subtropical Grasses Commonly Found in Southeast Asia. Medicina. 2019; 55(5):193. https://doi.org/10.3390/medicina55050193
Chicago/Turabian StyleAud-in, Sirirat, Koravit Somkid, and Wisuwat Songnuan. 2019. "Group-1 Grass Pollen Allergens with Near-Identical Sequences Identified in Species of Subtropical Grasses Commonly Found in Southeast Asia" Medicina 55, no. 5: 193. https://doi.org/10.3390/medicina55050193
APA StyleAud-in, S., Somkid, K., & Songnuan, W. (2019). Group-1 Grass Pollen Allergens with Near-Identical Sequences Identified in Species of Subtropical Grasses Commonly Found in Southeast Asia. Medicina, 55(5), 193. https://doi.org/10.3390/medicina55050193




