Long Non-Coding RNAs: Significant Drivers of Carcinogenesis Mechanisms in Head and Neck Squamous Cell Carcinoma
Abstract
1. Introduction
2. Mechanisms of lncRNA-Mediated Carcinogenesis in HNSCC
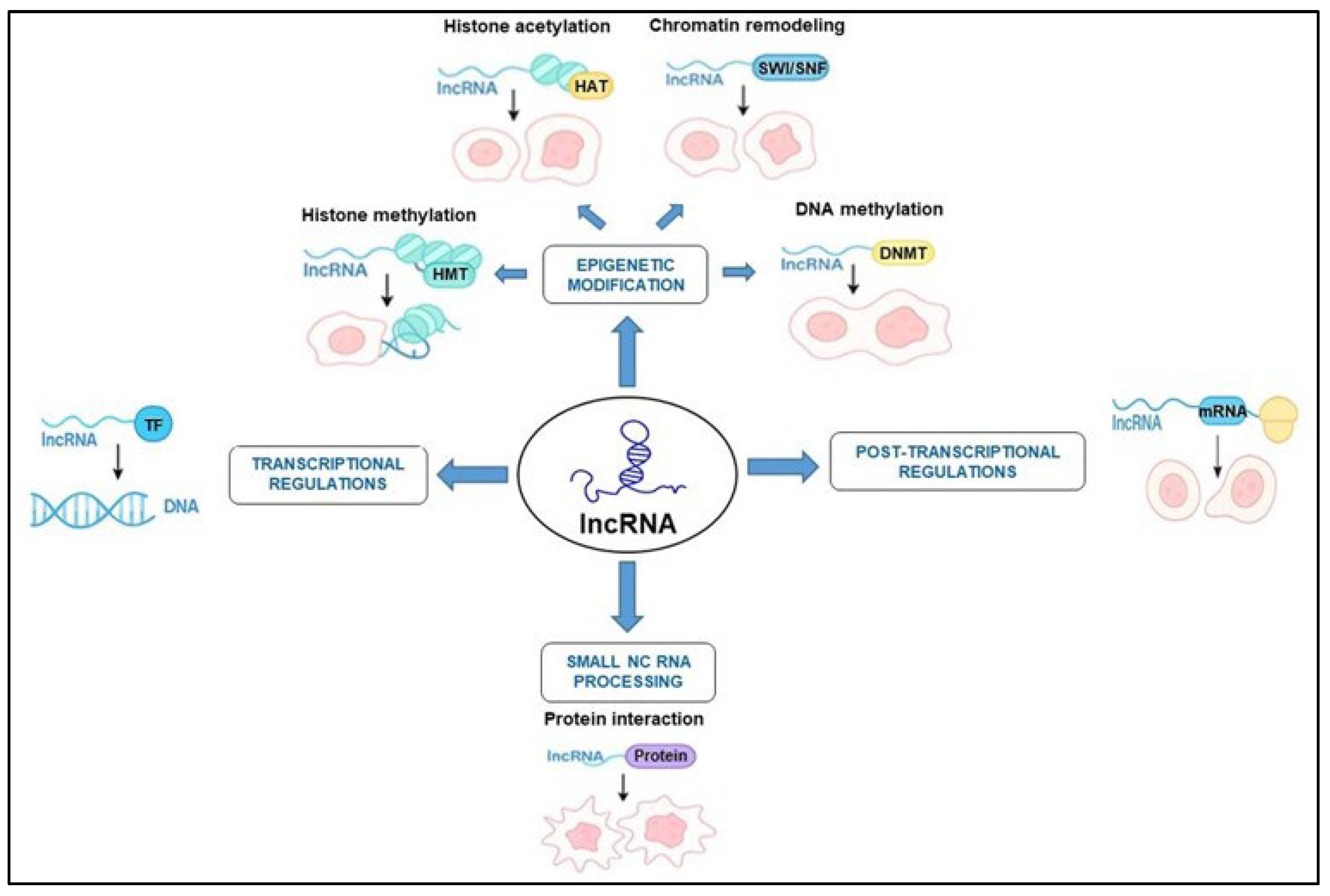
2.1. Epigenetic Regulation of HNSCC by lncRNAs
2.2. Role of lncRNAs in Transcriptional and Post-Transcriptional Regulation in HNSCC
2.2.1. Transcriptional Regulation
2.2.2. Post-Transcriptional Control
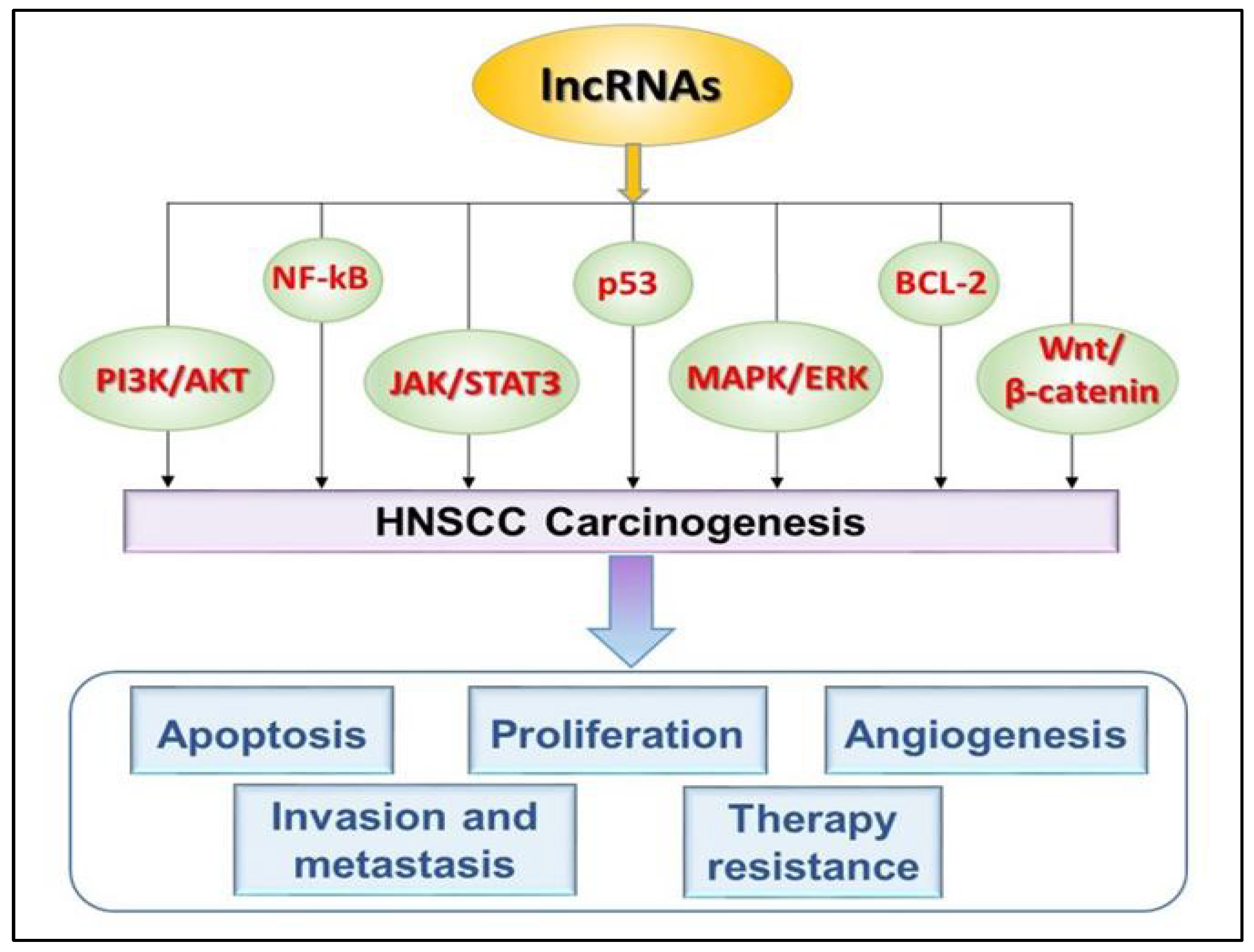
3. Impact of lncRNAs on the Modulation of Signaling Pathways That Govern Cellular Functions in HNSCC
3.1. lncRNAs’ Modulation of Signaling Pathways Involved in HNSCC Apoptosis
3.2. lncRNAs’ Modulation of Signaling Pathways Involved in HNSCC Proliferation
3.3. Impact of lncRNAs on the Signaling Pathways Involved in HNSCC Angiogenesis
3.4. lncRNAs’ Modulation of Signaling Pathways Involved in HNSCC Invasion and Metastasis
4. Impact of lncRNAs on Therapy Resistance in HNSCC
5. Challenges and Future Perspectives
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| HNSCC | Head and neck squamous cell carcinoma |
| lncRNAs | Long non-coding RNAs |
| HPV | Human papillomavirus |
| HOTAIR | HOX transcript antisense RNA |
| DNMT1 | DNA methyltransferase 1 |
| PVT1 | Plasmacytoma variant translocation 1 |
| PRC2 | Polycomb repressive complex 2 |
| EZH2 | Enhancer of zeste homolog 2 |
| PTEN | Phosphatase and TENsin homolog |
| OSCC | Oral squamous cell carcinoma |
| MALAT | Metastasis-associated lung adenocarcinoma transcript 1 |
| EMT | Epithelial–mesenchymal transition |
| CCAT1 | Colon cancer associated transcript 1 |
| RBPs | RNA-binding proteins |
| hnRNPs | Heterogeneous nuclear ribonucleoproteins |
| SRSFs | Serine/arginine-rich splicing factors |
| TINCR | TINCR Ubiquitin Domain Containing |
| ceRNA | Competing endogenous RNA |
| MEG3 | Maternally expressed 3 |
| BCL-2 | B-cell lymphoma 2 |
| BCL-XL | B-cell lymphoma-extra large |
| MYC | Myelocytoma oncogene |
| BAX | Bcl-2-associated X protein |
| PUMA | p53 upregulated modulator of apoptosis |
| NOXA | PMAIP1 (phorbol-12-myristate-13-acetate-induced protein 1) |
| BAK | Bcl-2 homologous antagonist/killer |
| XIAP | X-linked inhibitor of apoptosis protein |
| EGFR | Epidermal growth factor receptor |
| UCA1 | Urothelial cancer-associated 1 |
| PI3K | Phosphatidylinositol 3-kinase |
| AKT | Serine/threonine kinase 1 |
| GAS5 | Growth arrest-specific 5 lncRNA |
| mTOR | Mammalian target of rapamycin |
| ASOs | Antisense oligonucleotides |
| siRNAs | Small interfering RNAs |
| CRISPR | Clustered regularly interspaced short palindromic repeats |
| HNF1A-AS1 | HNF1A antisense RNA 1 lncRNA |
| MAPK/ERK | Mitogen-Activated Protein Kinase/Extracellular Signal-Regulated Kinase |
| IκB | Inhibitor of κB |
| NF-κB | Nuclear factor kappa-light-chain-enhancer of activated B cells |
| STAT3 | Signal transducer and activator of transcription 3 |
| JAK | Janus kinase |
| LINC00152 | Long intergenic non-protein coding RNA 152 |
| BAD | BCL2-associated agonist of cell death |
| BIM | Bcl-2 Interacting Mediator of cell death |
| AFAP1-AS1 | Actin filament associated protein 1 antisense RNA 1 |
| FAS | Fas cell surface death receptor |
| TNF | Tumor necrosis factor |
| TRAIL | TNF-related apoptosis-inducing ligand |
| c-FLIP | Cellular FLICE-inhibitory protein |
| CDKs | Cyclin-dependent kinases |
| Wnt | Wingless-related integration site |
| VEGF | Vascular endothelial growth factor |
| Ang2 | Angiopoietin 2 |
| MMP | Matrix metalloproteinases |
| ECM | Extracellular matrix |
| NPC | Nasopharyngeal cancer |
| FOXCUT | FOXC1 upstream transcript |
| LINC00319 | Long intergenic non-protein coding RNA 319 |
| CCL18 | Chemokine ligand 18 |
| CAFs | Carcinoma-associated fibroblasts |
| FENDRR | FOXF1 adjacent non-coding developmental regulatory RNA |
| TGF-β | Transforming growth factor-beta |
| MYO6 | Myosin VI |
| VHL | von Hippel–Lindau protein |
| ZEB | zinc-finger E-box-binding |
| Snail | Zinc finger protein SNAI1 |
| TWIST1/2 | Twist-related protein 1 and 2 |
| TSCC | Tongue squamous cell carcinomas |
| LSCC | Laryngeal squamous cell carcinoma |
| LINC00473 | Long intergenic non-protein coding RNA 473 |
| ZFAS1 | ZNFX1 antisense RNA 1 |
| NORAD | Non-coding RNA activated by DNA damage |
| ESCC | Esophageal squamous cell carcinoma |
| NKILA | NF-κB interacting lncRNA |
| TUG1 | Taurine upregulated gene 1 |
| 5-FU | 5-fluorouracil |
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef]
- Johnson, D.E.; Burtness, B.; Leemans, C.R.; Lui, V.W.Y.; Bauman, J.E.; Grandis, J.R. Head and neck squamous cell carcinoma. Nat. Rev. Dis. Primers 2020, 6, 92. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Han, J.; Zhu, Y.; Huang, N.; Qu, N. New advances in the therapeutic strategy of head and neck squamous cell carcinoma: A review of latest therapies and cutting-edge research. Biochim. Biophys. Acta (BBA)—Rev. Cancer 2025, 1880, 189230. [Google Scholar] [CrossRef]
- Goel, B.; Tiwari, A.K.; Pandey, R.K.; Singh, A.P.; Kumar, S.; Sinha, A.; Jain, S.K.; Khattri, A. Therapeutic approaches for the treatment of head and neck squamous cell carcinoma-An update on clinical trials. Transl. Oncol. 2022, 21, 101426. [Google Scholar] [CrossRef]
- Bhan, A.; Soleimani, M.; Mandal, S.S. Long noncoding RNA and cancer: A new paradigm. Cancer Res. 2017, 77, 3965–3981. [Google Scholar] [CrossRef]
- Zhong, Z.; Hong, M.; Chen, X.; Xi, Y.; Xu, Y.; Kong, D.; Deng, J.; Li, Y.; Hu, R.; Sun, C.; et al. Transcriptome analysis reveals the link between lncRNA-mRNA co-expression network and tumor immune microenvironment and overall survival in head and neck squamous cell carcinoma. BMC Med. Genom. 2020, 13, 57. [Google Scholar] [CrossRef] [PubMed]
- Lin, W.; Zhou, Q.; Wang, C.Q.; Zhu, L.; Bi, C.; Zhang, S.; Wang, X.; Jin, H. LncRNAs regulate metabolism in cancer. Int. J. Biol. Sci. 2020, 16, 1194–1206. [Google Scholar] [CrossRef] [PubMed]
- Lei, C.S.; Kung, H.J.; Shih, J.W. Long non-coding RNAs as functional codes for oral cancer: Translational potential, progress and promises. Int. J. Mol. Sci. 2021, 22, 4903. [Google Scholar] [CrossRef]
- Mattick, J.S.; Amaral, P.P.; Carninci, P.; Carpenter, S.; Chang, H.Y.; Chen, L.L.; Chen, R.; Dean, C.; Dinger, M.E.; Fitzgerald, K.A.; et al. Long non-coding RNAs: Definitions, functions, challenges and recommendations. Nat. Rev. Mol. Cell Biol. 2023, 24, 430–447. [Google Scholar] [CrossRef]
- Peng, W.X.; Koirala, P.; Mo, Y.Y. LncRNA-mediated regulation of cell signaling in cancer. Oncogene 2017, 36, 5661–5667. [Google Scholar] [CrossRef]
- Bhan, A.; Mandal, S.S. LncRNA HOTAIR: A master regulator of chromatin dynamics and cancer. Biochim. Biophys. Acta 2015, 1856, 151–164. [Google Scholar] [CrossRef]
- Chen, X.; Song, J.; Wang, X.; Sun, D.; Liu, Y.; Jiang, Y. LncRNA LINC00460: Function and mechanism in human cancer. Thorac. Cancer 2022, 13, 3–14. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Cao, W.; Wu, K.; Qin, X.; Wang, X.; Li, Y.; Yu, B.; Zhang, Z.; Wang, X.; Yan, M.; et al. LncRNA LINC00460 promotes EMT in head and neck squamous cell carcinoma by facilitating peroxiredoxin-1 into the nucleus. J. Exp. Clin. Cancer Res. CR 2019, 38, 365. [Google Scholar] [CrossRef] [PubMed]
- Ye, D.; Deng, Y.; Shen, Z. The role and mechanism of MALAT1 long non-coding RNA in the diagnosis and treatment of head and neck squamous cell carcinoma. Onco Targets Ther. 2021, 14, 4127–4136. [Google Scholar] [CrossRef]
- Wu, F.; Zhu, Y.; Zhou, C.; Gui, W.; Li, H.; Lin, H. Regulation mechanism and pathogenic role of lncRNA plasmacytoma variant translocation 1 (PVT1) in human diseases. Genes Dis. 2023, 10, 901–914. [Google Scholar] [CrossRef]
- Patty, B.J.; Hainer, S.J. Non-Coding RNAs and Nucleosome Remodeling Complexes: An Intricate Regulatory Relationship. Biology 2020, 9, 213. [Google Scholar] [CrossRef]
- Prozzillo, Y.; Santopietro, M.V.; Messina, G.; Patrizio, D. Unconventional roles of chromatin remodelers and long non-coding RNAs in cell division. Cell. Mol. Life Sci. 2023, 80, 365. [Google Scholar] [CrossRef]
- Oo, J.A.; Warwick, T.; Pálfi, K.; Lam, F.; McNicoll, F.; Prieto-Garcia, C.; Gunther, S.; Cao, C.; Zhou, Y.; Gavrilov, A.A.; et al. Long non-coding RNAs direct the SWI/SNF complex to cell type-specific enhancers. Nat. Commun. 2025, 16, 131. [Google Scholar] [CrossRef]
- Ahadi, A. Functional roles of lncRNAs in the pathogenesis and progression of cancer. Genes Dis. 2021, 8, 424–437. [Google Scholar] [CrossRef] [PubMed]
- Liz, J.; Esteller, M. lncRNAs and microRNAs with a role in cancer development. Biochim. Biophys. Acta 2016, 1859, 169–176. [Google Scholar]
- Zhou, M.; Mao, Y.; Yu, S.; Li, Y.; Yin, R.; Zhang, Q.; Lu, T.; Sun, R.; Lin, S.; Qian, Y.; et al. LINC00673 Represses CDKN2C and Promotes the Proliferation of Esophageal Squamous Cell Carcinoma Cells by EZH2-Mediated H3K27 Trimethylation. Front. Oncol. 2020, 10, 1546. [Google Scholar] [CrossRef] [PubMed]
- Bohrer, C.; Varon, E.; Peretz, E.; Gita Reinitz, G.; Kinor, N.; Halle, D.; Nissan, A.; Shav-Tal, Y. CCAT1 lncRNA is chromatin-retained and post-transcriptionally spliced. Histochem. Cell Biol. 2024, 162, 91–107. [Google Scholar] [CrossRef]
- Segal, D.; Dostie, J. The talented LncRNAs: Meshing into transcriptional regulatory networks in cancer. Cancers 2023, 15, 3433. [Google Scholar] [CrossRef]
- Deshpande, A.; Mahale, S.; Kanduri, C. Beyond the Transcript: Translating Non-Coding RNAs and Their Impact on Cellular Regulation. Cancers 2025, 17, 1555. [Google Scholar] [CrossRef]
- He, R.Z.; Luo, D.X.; Mo, Y.Y. Emerging Roles of lncRNAs in Post-Transcriptional Regulation in Cancer. Genes Dis. 2019, 6, 6–15. [Google Scholar] [CrossRef]
- Xu, D.; Wang, W.; Wang, D.; Ding, J.; Zhou, Y.; Zhang, W. Long noncoding RNA MALAT-1: A versatile regulator in cancer progression, metastasis, immunity, and therapeutic resistance. Non-Coding RNA Res. 2024, 9, 388–406. [Google Scholar] [CrossRef] [PubMed]
- Cui, S.; Peng, Q.; Ma, Q.; Xu, X.; Zhang, W.; Jiang, X.; Tan, S.; Yang, W.; Han, Y.; Oyang, L.; et al. Crosstalk between RNA-binding proteins and non-coding RNAs in tumors: Molecular mechanisms, and clinical significance. Int. J. Biol. Sci. 2025, 21, 2991–3010. [Google Scholar] [CrossRef] [PubMed]
- Akbari Dilmaghani, N.; Khoshsirat, S.; Shanaki-Bavarsad, M.; Pourbagheri-Sigaroodi, A.; Bashash, D. The contributory role of long non-coding RNAs (lncRNAs) in head and neck cancers: Possible biomarkers and therapeutic targets? Eur. J. Pharmacol. 2021, 900, 174053. [Google Scholar] [CrossRef]
- Braga, E.A.; Fridman, M.V.; Burdennyy, A.M.; Loginov, V.I.; Dmitriev, A.A.; Pronina, I.V.; Morozov, S.G. Various LncRNA Mechanisms in Gene Regulation Involving miRNAs or RNA-Binding Proteins in Non-Small-Cell Lung Cancer: Main Signaling Pathways and Networks. Int. J. Mol. Sci. 2023, 24, 13617. [Google Scholar] [CrossRef]
- Fernandes, J.C.R.; Acuna, S.M.; Aoki, J.I.; Floeter-Winter, L.M.; Muxel, S.M. Long non-coding RNAs in the regulation of gene expression: Physiology and disease. Noncoding RNA 2019, 5, 17. [Google Scholar] [CrossRef]
- Lopez-Urrutia, E.; Bustamante Montes, L.P.; Ladron de Guevara Cervantes, D.; Perez-Plasencia, C.; Campos-Parra, A.D. Crosstalk between long non-coding RNAs, micro-RNAs and mRNAs: Deciphering molecular mechanisms of master regulators in cancer. Front. Oncol. 2019, 9, 669. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.; Kansara, S.; Mahajan, M.; Yadav, B.; Garg, M.; Pandey, A.K. Long non-coding RNAs orchestrate various molecular and cellular processes by modulating epithelial-mesenchymal transition in head and neck squamous cell carcinoma. Biochim. Biophys. Acta (BBA)—Mol. Basis Dis. 2021, 1867, 166240. [Google Scholar] [CrossRef] [PubMed]
- Rossi, M.N.; Antonangeli, F. LncRNAs: New Players in Apoptosis Control. Int. J. Cell Biol. 2014, 2014, 473857. [Google Scholar] [CrossRef]
- Kong, L.; Zhou, X.; Wu, Y.; Wang, Y.; Chen, L.; Li, P.; Liu, S.; Sun, S.; Ren, Y.; Mei, M.; et al. Targeting HOTAIR Induces Mitochondria Related Apoptosis and Inhibits Tumor Growth in Head and Neck Squamous Cell Carcinoma in vitro and in vivo. Curr. Mol. Med. 2015, 15, 952–960. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Qin, Y.; Zhen, Z.; Shen, H.; Cong, T.; Schiferle, E.; Xiao, S. Long non-coding RNA HOTAIR/microRNA-206 sponge regulates STC2 and further influences cell biological functions in head and neck squamous cell carcinoma. Cell Prolif. 2019, 52, e12651, Correction in 2022, 55, e13200. [Google Scholar] [CrossRef]
- Ow, T.J.; Thomas, C.; Fulcher, C.D.; Chen, J.; López, A.; Reyna, D.E.; Prystowsky, M.B.; Smith, R.V.; Schiff, B.A.; Rosenblatt, G.; et al. Apoptosis signaling molecules as treatment targets in head and neck squamous cell carcinoma. Laryngoscope 2020, 130, 2643–2649. [Google Scholar] [CrossRef]
- Aubrey, B.J.; Kelly, G.L.; Janic, A.; Herold, M.J.; Strasser, A. How does p53 induce apoptosis and how does this relate to p53-mediated tumour suppression? Cell Death Differ. 2018, 25, 104–113. [Google Scholar] [CrossRef]
- Heravi, M.; Bala, S. Emerging role of non-coding RNA in oral cancer. Cell. Signal. 2018, 42, 134–143. [Google Scholar] [CrossRef]
- Qian, S.; Wei, Z.; Yang, W.; Huang, J.; Yang, Y.; Wang, J. The role of BCL-2 family proteins in regulating apoptosis and cancer therapy. Front. Oncol. 2022, 12, 985363. [Google Scholar] [CrossRef]
- Zhang, L.; Zhao, F.; Li, W.; Song, G.; Kasim, V.; Wu, S. The Biological Roles and Molecular Mechanisms of Long Non-Coding RNA MEG3 in the Hallmarks of Cancer. Cancers 2022, 14, 6032. [Google Scholar] [CrossRef]
- Nguyen, L.N.T.; Pyburn, J.S.; Nguyen, N.L.; Schank, M.B.; Zhao, J.; Wang, L.; Leshaodo, T.O.; El Gazzar, M.; Moorman, J.P.; Yao, Z.Q. Epigenetic Regulation by lncRNA GAS5/miRNA/mRNA Network in Human Diseases. Int. J. Mol. Sci. 2025, 26, 1377. [Google Scholar] [CrossRef] [PubMed]
- Ghorbani, V.; Ghorbian, S. Gene expression profiling of lncRNA-HOTAIR and lncRNA-MALAT1 in esophageal cancer: Uncovering links to lifestyle factors and diagnostic significance. Discov. Onc. 2025, 16, 630. [Google Scholar] [CrossRef]
- Saddam, M.; Paul, S.K.; Habib, M.A.; Fahim, M.A.; Mimi, A.; Islam, S.; Paul, B.; Helal, M.U. Emerging biomarkers and potential therapeutics of the BCL-2 protein family: The apoptotic and anti-apoptotic context. Egypt. J. Med. Hum. Genet. 2024, 25, 12. [Google Scholar] [CrossRef]
- Kaloni, D.; Diepstraten, S.T.; Strasser, A.; Kelly, L.G. BCL-2 protein family: Attractive targets for cancer therapy. Apoptosis 2023, 28, 20–38. [Google Scholar] [CrossRef]
- Guo, Q.; Jin, Y.; Chen, X.; Ye, X.; Shen, X.; Lin, M.; Zeng, C.; Zhou, T.; Zhang, J. NF-κB in biology and targeted therapy: New insights and translational implications. Signal Transduct. Target. Ther. 2024, 9, 53. [Google Scholar] [CrossRef]
- Fu, S.; Wang, Y.; Li, H.; Chen, L.; Liu, Q. Regulatory Networks of LncRNA MALAT-1 in Cancer. Cancer Manag. Res. 2020, 12, 10181–10198. [Google Scholar] [CrossRef] [PubMed]
- Zhan, Y.; Li, Y.; Guan, B.; Wang, Z.; Peng, D.; Chen, Z.; He, A.; He, S.; Gong, Y.; Li, X.; et al. Long non-coding RNA HNF1A-AS1 promotes proliferation and suppresses apoptosis of bladder cancer cells through upregulating Bcl-2. Oncotarget 2017, 8, 76656–76665. [Google Scholar] [CrossRef]
- Price, R.L.; Bhan, A.; Mandal, S.S. HOTAIR beyond repression: In protein degradation, inflammation, DNA damage response, and cell signaling. DNA Repair 2021, 105, 103141. [Google Scholar] [CrossRef]
- Masrour, M.; Khanmohammadi, S.; Fallahtafti, P.; Rezaei, N. Long non-coding RNA as a potential diagnostic biomarker in head and neck squamous cell carcinoma: A systematic review and meta-analysis. PLoS ONE 2023, 18, e0291921. [Google Scholar] [CrossRef]
- Sur, S.; Davray, D.; Basu, S.; Kheur, S.; Pal, J.K.; Nagar, S.; Sanap, A.; Rudagi, B.M.; Gupta, S. Novel insights on oral squamous cell carcinoma management using long non-coding RNAs. Oncol. Res. 2024, 32, 1589–1612. [Google Scholar] [CrossRef] [PubMed]
- Assal, R.A.; Rashwan, H.H.; Zakaria, Z.I.; Sweillam, J.H.; Fouda, Y.M.; Abdelhamid, A.M.; Youness, R.A. Deciphering the mysteries of MEG3 LncRNA and its implications in genitourinary cancers. Front. Oncol. 2025, 15, 1519103. [Google Scholar] [CrossRef]
- Tang, R.; Mei, X.; Wang, Y.C.; Cui, X.B.; Zhang, G.; Li, W.; Chen, S.Y. LncRNA GAS5 regulates vascular smooth muscle cell cycle arrest and apoptosis via p53 pathway. Biochim. Biophys. Acta Mol. Basis Dis. 2019, 1865, 2516–2525. [Google Scholar] [CrossRef]
- Bahar, M.E.; Kim, H.J.; Kim, D.R. Targeting the RAS/RAF/MAPK pathway for cancer therapy: From mechanism to clinical studies. Signal Transduct. Target. Ther. 2023, 8, 455. [Google Scholar] [CrossRef]
- Constantin, M.; Chifiriuc, M.C.; Bleotu, C.; Vrancianu, C.O.; Cristian, R.E.; Bertesteanu, S.V.; Grigore, R.; Bertesteanu, G. Molecular pathways and targeted therapies in head and neck cancers pathogenesis. Front. Oncol. 2024, 14, 1373821. [Google Scholar] [CrossRef]
- Rong, C.; Muller, M.F.; Xiang, F.; Jensen, A.; Weichert, W.; Major, G.; Plinkert, P.K.; Hess, J.; Affolter, A. Adaptive ERK signalling activation in response to therapy and in silico prognostic evaluation of EGFR-MAPK in HNSCC. Br. J. Cancer 2020, 123, 288–297. [Google Scholar] [CrossRef]
- Song, H.; Liu, D.; Dong, S.; Zeng, L.; Wu, Z.; Zhao, P.; Zhang, L.; Chen, Z.S.; Zou, C. Epitranscriptomics and epiproteomics in cancer drug resistance: Therapeutic implications. Signal Transduct. Target. Ther. 2020, 5, 193. [Google Scholar] [CrossRef] [PubMed]
- Zhu, C.; Wang, X.; Wang, Y.; Wang, K. Functions and underlying mechanisms of lncRNA HOTAIR in cancer chemotherapy resistance. Cell Death Discov. 2022, 8, 383. [Google Scholar] [CrossRef] [PubMed]
- Amodio, N.; Raimondi, L.; Juli, G.; Stamato, M.A.; Caracciolo, D.; Tagliaferri, P.; Tassone, P. MALAT1: A druggable long non-coding RNA for targeted anti-cancer approaches. J. Hematol. Oncol. 2018, 11, 63. [Google Scholar] [CrossRef]
- Kaur, J.; Salehen, N.; Norazit, A.; Rahman, A.A.; Murad, N.A.A.; Rahman, N.M.A.N.A.; Ibrahim, K. Tumor Suppressive Effects of GAS5 in Cancer Cells. Non-Coding RNA 2022, 8, 39. [Google Scholar] [CrossRef]
- Matthews, H.K.; Bertoli, C.; de Bruin, R.A.M. Cell cycle control in cancer. Nat. Rev. Mol. Cell Biol. 2022, 23, 74–88. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Li, S.; Oyang, L.; Cui, S.; Zhang, W.; Yang, W.; Peng, M.; Tan, S.; Xia, L.; Lin, J.; et al. Novel insights into lncRNAs as key regulators of post-translational modifications in cancer: Mechanisms and therapeutic potential. Cell. Oncol. 2025. [Google Scholar] [CrossRef]
- Karimian, A.; Ahmadi, Y.; Yousefi, B. Multiple functions of p21 in cell cycle, apoptosis and transcriptional regulation after DNA damage. DNA Repair 2016, 42, 63–71. [Google Scholar] [CrossRef]
- Hakami, M.A.; Hazazi, A.; Abdulaziz, O.; Almasoudi, H.H.; Alhazmi, A.Y.M.; Alkhalil, S.S.; Alharthi, N.S.; Alhuthali, H.M.; Almalki, W.H.; Gupta, G.; et al. HOTAIR: A key regulator of the Wnt/β-catenin signaling cascade in cancer progression and treatment. Pathol. Res. Pract. 2024, 253, 154957. [Google Scholar] [CrossRef]
- Hashemi, M.; Moosavi, M.S.; Abed, H.M.; Dehghani, M.; Aalipour, M.; Heydari, E.L.; Behroozaghdam, M.; Entezari, M.; Salimimoghadam, S.; Gunduz, E.S.; et al. Long non-coding RNA (lncRNA) H19 in human cancer: From proliferation and metastasis to therapy. Pharmacol. Res. 2022, 184, 106418. [Google Scholar] [CrossRef] [PubMed]
- Xia, Y.; Pei, T.; Zhao, J.; Wang, Z.; Shen, Y.; Yang, Y.; Liang, J. Long noncoding RNA H19: Functions and mechanisms in regulating programmed cell death in cancer. Cell Death Discov. 2024, 10, 76. [Google Scholar] [CrossRef]
- Xiang, Y.; Hua, Q. The Role and Mechanism of Long Non-Coding RNA HOTAIR in the Oncogenesis, Diagnosis, and Treatment of Head and Neck Squamous Cell Carcinoma. Clin. Med. Insights Oncol. 2023, 17, 11795549231169099. [Google Scholar] [CrossRef]
- Kaur, G.; Roy, B. Decoding Tumor Angiogenesis for Therapeutic Advancements: Mechanistic Insights. Biomedicines 2024, 12, 827. [Google Scholar] [CrossRef]
- Liu, X.; Zhang, J.; Yi, T.; Li, H.; Tang, X.; Liu, D.; Wu, D.; Li, Y. Decoding tumor angiogenesis: Pathways, mechanisms, and future directions in anti-cancer strategies. Biomark Res. 2025, 13, 62. [Google Scholar] [CrossRef] [PubMed]
- Fu, W.M.; Lu, Y.F.; Hu, B.G.; Liang, W.C.; Zhu, X.; Yang, H.D.; Li, G.; Zhang, J.F. Long noncoding RNA Hotair mediated angiogenesis in nasopharyngeal carcinoma by direct and indirect signaling pathways. Oncotarget 2016, 7, 4712–4723. [Google Scholar] [CrossRef] [PubMed]
- Kong, X.P.; Yao, J.; Luo, W.; Feng, F.K.; Ma, J.T.; Ren, Y.P.; Wang, D.L.; Bu, R.F. The expression and functional role of a FOXC1 related mRNA-lncRNA pair in oral squamous cell carcinoma. Mol. Cell. Biochem. 2014, 394, 177–186. [Google Scholar] [CrossRef] [PubMed]
- Jiang, X.; Liu, J.; Li, S.; Jia, B.; Huang, Z.; Shen, J.; Luo, H.; Zhao, J. CCL18-induced LINC00319 promotes proliferation and metastasis in oral squamous cell carcinoma via the miR-199a-5p/FZD4 axis. Cell Death Dis. 2020, 11, 777. [Google Scholar] [CrossRef]
- Yang, C.; Shu, J.; Li, Y.; Zhao, N.; Liu, X.; Tian, X.; Sun, Z.; Tabish, M.S.; Hong, Y.; Chen, K.; et al. Long non-coding RNAs are involved in the crosstalk between cancer associated fibroblasts and tumor cells. Front. Immunol. 2024, 15, 1469918. [Google Scholar] [CrossRef]
- Yu, Z.; Zhao, H.; Feng, X.; Li, H.; Qiu, C.; Yi, X.; Tang, H.; Zhang, J. Long Non-Coding RNA FENDRR Acts as a miR-423-5p Sponge to Suppress the Treg-Mediated Immune Escape of Hepatocellular Carcinoma Cells. Mol. Ther. Nucleic Acids 2019, 17, 516–529. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Jiang, E.; Shao, Z.; Shang, Z. LncRNA FENDRR in Carcinoma-Associated Fibroblasts Regulates the Angiogenesis of Oral Squamous Cell Carcinoma Through the PI3K/AKT Pathway. Front. Oncol. 2021, 11, 616576. [Google Scholar] [CrossRef]
- Feng, C.; Liu, S.; Shang, Z. Identification and Validation of an EMT Related LncRNA Signature for HNSCC to Predict Survival and Immune Landscapes. Front. Cell Dev. Biol. 2022, 9, 798898. [Google Scholar] [CrossRef]
- Luo, X.; Qiu, Y.; Jiang, Y.; Chen, F.; Jiang, L.; Dan, H.; Zeng, X.; Lei, Y.L.; Chen, Q. Long non-coding RNA implicated in the invasion and metastasis of head and neck cancer: Possible function and mechanisms. Mol. Cancer 2018, 17, 14. [Google Scholar] [CrossRef]
- Loh, C.Y.; Chai, J.Y.; Tang, T.F.; Wong, W.F.; Sethi, G.; Shanmugam, M.K.; Chong, P.P.; Looi, C.Y. The E-Cadherin and N-Cadherin switch in epithelial-to-mesenchymal transition: Signaling, therapeutic implications, and challenges. Cells 2019, 8, 1118. [Google Scholar] [CrossRef]
- Hou, J.; Yan, D.; Liu, Y.; Huang, P.; Cui, H. The roles of integrin α5β1 in human cancer. Onco Targets Ther. 2020, 13, 13329–13344. [Google Scholar] [CrossRef] [PubMed]
- Elango, J.; Hou, C.; Bao, B.; Wang, S.; de Val Maté Sánchez, J.E.; Wenhui, W. The molecular interaction of collagen with cell receptors for biological function. Polymers 2022, 14, 876. [Google Scholar] [CrossRef]
- Imodoye, S.O.; Adedokun, K.A.; Muhammed, A.O.; Bello, I.O.; Muhibi, M.A.; Oduola, T.; Oyenike, M.A. Understanding the complex milieu of epithelial-mesenchymal transition in cancer metastasis: New insight into the roles of transcription factors. Front. Oncol. 2021, 11, 762817. [Google Scholar] [CrossRef]
- Ahmad, M.; Weiswald, L.B.; Poulain, L.; Denoyelle, C.; Meryet-Figuiere, M. Involvement of lncRNAs in cancer cells migration, invasion and metastasis: Cytoskeleton and ECM crosstalk. J. Exp. Clin. Cancer Res. 2023, 42, 173. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.T.; Wu, K.J. Epigenetic regulation of epithelial-mesenchymaltransition: Focusing on hypoxia and TGF-β signaling. J. Biomed. Sci. 2020, 27, 39. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wang, S.; Ren, Y.; Zhou, X. The Role of lncRNA Crosstalk in Leading Cancer Metastasis of Head and Neck Squamous Cell Carcinoma. Front. Oncol. 2020, 10, 561833. [Google Scholar] [CrossRef]
- Lambert, A.W.; Weinberg, R.A. Linking EMT programmes to normal and neoplastic epithelial stem cells. Nat. Rev. Cancer 2021, 21, 325–338. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wu, C.; Zhang, C.; Li, Z.; Zhu, T.; Chen, J.; Ren, Y.; Wang, X.; Zhang, L.; Zhou, X. TGF-beta-induced STAT3 overexpression promotes human head and neck squamous cell carcinoma invasion and metastasis through malat1/miR-30a interactions. Cancer Lett. 2018, 436, 52–62. [Google Scholar] [CrossRef]
- Duan, Y.; Yue, K.; Ye, B.; Chen, P.; Zhang, J.; He, Q.; Wu, Y.; Lai, Q.; Li, H.; Wu, Y.; et al. LncRNA MALAT1 promotes growth and metastasis of head and neck squamous cell carcinoma by repressing VHL through a non-canonical function of EZH2. Cell Death Dis. 2023, 14, 522. [Google Scholar] [CrossRef]
- Zhou, X.; Liu, S.; Cai, G.; Kong, L.; Zhang, T.; Ren, Y. Long non coding RNA MALAT1 promotes tumor growth and metastasis by inducing epithelial- mesenchymal transition in oral squamous cell carcinoma. Sci. Rep. 2015, 5, 15972. [Google Scholar] [CrossRef]
- Wu, Y.S.; Zhang, L.; Zhang, L.; Wang, Y.; Li, H.; Ren, X.; Wei, F.; Yu, W.; Liu, T.; Wang, X.; et al. Long non-coding RNA HOTAIR promotes tumor cell invasion and metastasis by recruiting EZH2 and repressing E-cadherin in oral squamous cell carcinoma. Int. J. Oncol. 2015, 46, 2586–2594. [Google Scholar] [CrossRef]
- Lu, M.Y.; Liao, Y.W.; Chen, P.Y.; Hsieh, P.L.; Fang, C.Y.; Wu, C.Y.; Yen, M.L.; Peng, B.Y.; Wang, D.P.; Cheng, H.C.; et al. Targeting LncRNA HOTAIR suppresses cancer stemness and metastasis in oral carcinomas stem cells through modulation of EMT. Oncotarget 2017, 8, 98542–98552. [Google Scholar] [CrossRef]
- Yao, F.; Wang, Q.; Wu, Q. The prognostic value and mechanisms of lncRNA UCA1 in human cancer. Cancer Manag. Res. 2019, 11, 7685–7696. [Google Scholar] [CrossRef]
- Zhang, T.H.; Liang, L.Z.; Liu, X.L.; Wu, J.N.; Su, K.; Chen, J.Y.; Zheng, Q.Y. LncRNA UCA1/miR-124 axis modulates TGF1-induced epithelial-mesenchymal transition and invasion of tongue cancer cells through JAG1/Notch signaling. J. Cell. Biochem. 2019, 120, 10495–10504. [Google Scholar] [CrossRef] [PubMed]
- Duan, Q.; Xu, M.; Wu, M.; Zhang, X.; Gan, M.; Jiang, H. Long noncoding RNA UCA1 promotes cell growth, migration, and invasion by targeting miR-143-3p in oral squamous cell carcinoma. Cancer Med. 2020, 9, 3115–3129. [Google Scholar] [CrossRef] [PubMed]
- Sun, S.; Gong, C.; Yuan, K. LncRNA UCA1 promotes cell proliferation, invasion and migration of laryngeal squamous cell carcinoma cells by activating Wnt/β-catenin signaling pathway. Exp. Ther. Med. 2019, 17, 1182–1189. [Google Scholar] [CrossRef]
- Han, P.B.; Ji, X.J.; Zhang, M.; Gao, L.Y. Upregulation of lncRNA LINC00473 promotes radioresistance of HNSCC cells through activating Wnt/b-catenin signaling pathway. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 7305–7313. [Google Scholar] [PubMed]
- Kolenda, T.; Guglas, K.; Kopczyńska, M.; Teresiak, A.; Bliźniak, R.; Mackiewicz, A.; Lamperska, K.; Mackiewicz, J. Oncogenic role of ZFAS1 lncRNA in head and neck squamous cell carcinomas. Cells 2019, 8, 366. [Google Scholar] [CrossRef]
- Hu, W.; Zhao, Y.; Su, L.; Wu, Z.; Jiang, W.; Jiang, X.; Liu, M. Silencing the lncRNA NORAD inhibits EMT of head and neck squamous cell carcinoma stem cells via miR-26a-5p. Mol. Med. Rep. 2021, 24, 743. [Google Scholar] [CrossRef]
- Yang, J.; Qi, M.; Fei, X.; Wang, X.; Wang, K. LncRNA H19: A novel oncogene in multiple cancers. Int. J. Biol. Sci. 2021, 17, 3188–3208. [Google Scholar] [CrossRef]
- Ji, Y.; Feng, G.; Hou, Y.; Yu, Y.; Wang, R.; Yuan, H. Long noncoding RNA MEG3 decreases the growth of head and neck squamous cell carcinoma by regulating the expression of miR-421 and E-cadherin. Cancer Med. 2020, 9, 3954–3963. [Google Scholar] [CrossRef]
- Hussen, B.M.; Azimi, T.; Hidayat, H.J.; Taheri, M.; Ghafouri-Fard, S. NF-KappaB interacting LncRNA: Review of its roles in neoplastic and non-neoplastic conditions. Biomed. Pharmacother. 2021, 139, 111604. [Google Scholar] [CrossRef]
- Wang, W.; Liu, Y.; Wu, J. The roles of lncRNAs in the development of drug resistance of oral cancers. Biomed. Pharmacother. 2024, 180, 117458. [Google Scholar] [CrossRef] [PubMed]
- Ramasubbu, S.K.; Pasricha, R.K.; Nath, U.K.; Rawat, V.S.; Das, B. Quality of life and factors affecting it in adult cancer patients undergoing cancer chemotherapy in a tertiary care hospital. Cancer Rep. 2021, 4, e1312. [Google Scholar] [CrossRef]
- Jeyachandran, S. Interlink between pharyngeal radiation dose and long-term quality of life in HNSCC patients. Oral Oncol. Rep. 2024, 10, 100514. [Google Scholar] [CrossRef]
- Gauss, C.; Stone, L.D.; Ghafouri, M.; Quan, D.; Johnson, J.; Fribley, A.M.; Amm, H.M. Overcoming Resistance to Standard-of-Care Therapies for Head and Neck Squamous Cell Carcinomas. Cells 2024, 13, 1018. [Google Scholar] [CrossRef]
- Qu, Y.; Tan, H.Y.; Chan, Y.T.; Jiang, H.; Wang, N.; Wang, D. The functional role of long noncoding RNA in resistance to anticancer treatment. Ther. Adv. Med. Oncol. 2020, 12, 1758835920927850. [Google Scholar] [CrossRef]
- Da, M.; Zhuang, J.; Zhou, Y.; Qi, Q.; Han, S. Role of long noncoding RNA taurine-upregulated gene 1 in cancers. Mol. Med. 2021, 27, 51. [Google Scholar] [CrossRef]
- Mi, J.; Wang, Y.; He, S.; Qin, X.; Li, Z.; Zhang, T.; Huang, W.; Wang, R. LncRNA HOTAIRM1 promotes radioresistance in nasopharyngeal carcinoma by modulating FTO acetylation-dependent alternative splicing of CD44. Neoplasia 2024, 56, 101034. [Google Scholar] [CrossRef]
- Li, G.; Jiang, Y.; Li, G.; Qiao, Q. Comprehensive analysis of radiosensitivity in head and neck squamous cell carcinoma. Radiother. Oncol. J. Eur. Soc. Ther. Radiol. Oncol. 2021, 159, 126–135. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Zhu, C.; Jin, Y. The oncogenic and tumor suppressive functions of the long noncoding RNA MALAT1: An emerging controversy. Front Genet. 2020, 11, 93. [Google Scholar] [CrossRef]
- Jiang, M.C.; Ni, J.J.; Cui, W.Y.; Wang, B.Y.; Zhuo, W. Emerging roles of lncRNA in cancer and therapeutic opportunities. Am. J. Cancer Res. 2019, 9, 1354–1366. [Google Scholar] [PubMed]
| Pathway | lncRNAs | Mechanisms | Effect on HNSCC Apoptosis |
|---|---|---|---|
| PI3K/AKT/mTOR | HOTAIR, UCA1, MALAT1 | miRNA sponging, PTEN silencing | Inhibit apoptosis |
| p53 | MEG3, HOTAIR, MALAT | Epigenetic silencing and miRNA sponging | Enhance or suppress apoptosis |
| BCL-2 family | MALAT1, HOTAIR | Epigenetic and ceRNA mechanisms | Block mitochondrial apoptosis |
| MEG3, GAS5 | Pro-apoptotic signaling | Enhance apoptosis | |
| NF-κB | MALAT1, HNF1A-AS1, HOTAIR | IκB degradation → NF-κB activation | Promote anti-apoptotic genes |
| JAK/STAT3 | LINC00152, MALAT1 | ↑ STAT3 phosphorylation | Upregulate anti-apoptotic genes |
| MAPK/ERK | HOTAIR, MALAT, UCA1, AFAP1-AS1 | Increase ERK phosphorylation | Anti-apoptotic signaling |
| MEG3, GAS5 | Inhibit ERK activity | Restore apoptotic sensitivity | |
| Death receptor pathway | HOTAIR, MALAT1 | Epigenetic silencing, miRNA sponging | Reduces extrinsic apoptosis |
| lncRNA | Signaling Pathway | Effect on HNSCC Proliferation |
|---|---|---|
| MALAT1 | PI3K/AKT, MAPK, NF-κB | ↑ cyclin D1 and CDK4; promotes cell cycle progression and survival |
| HOTAIR | Wnt/β-catenin | ↑ cyclin D1; epigenetically silences p21; enhances proliferation |
| UCA1 | PI3K/AKT | ↑ cyclin D1; suppresses p21/p27; increases proliferation |
| H19 | Wnt/β-catenin | ↑ cyclin D1; sponges miRNAs (e.g., miR-675); promotes stemness and proliferation |
| lncRNA | Mechanisms | Effect on HNSCC Angiogenesis |
|---|---|---|
| HOTAIR | Increases VEGF-A and Ang2 | Pro-angiogenic, correlated with advanced stage and poor survival |
| LINC00319 | Increases CCL18, VEGFA, and MMP-9 | Enhances proliferation, metastasis, EMT, and angiogenesis |
| FOXCUT | Increases MMP2, MMP7, MMP9, and VEGF-A | Enhances angiogenesis |
| FENDRR | Inhibits PI3K/AKT pathway, decreases MMP-9 | Anti-angiogenic, inhibits tumor progression |
| lncRNA | Mechanism of Invasion/Metastasis | Effect in HNSCC |
|---|---|---|
| MALAT1 | TGF-β and STAT3 overexpression, activation of Wnt/β-catenin and NF-κB pathways, vimentin and N-cadherin overexpression | Promotes EMT and enhances cell proliferation, invasion, and metastasis |
| HOTAIR | Downregulates E-cadherin and activates vimentin and Snail expression | Induces EMT, predicts metastasis advanced stage marker |
| H19 | Inhibits E-cadherin, modulates the STAT3/EZH2/β-catenin pathway | Promotes EMT, increases invasion |
| UCA1 | miRNA sponging, Wnt/β-catenin pathway activation | Promotes EMT, migration, and metastasis |
| LINC00473 | Activates the Wnt/b-catenin pathway | Induces invasion and resistance to radiotherapy |
| ZFAS1 | Activates the Wnt/b-catenin, TGF-β, and JAK/STAT pathways | Promotes cell proliferation and invasion |
| NORAD | Downregulates E-cadherin and upregulates MMP-2, MMP-9, N-cadherin, and vimentin | Promotes EMT, migration, and metastasis |
| MEG3 | Upregulates E-cadherin levels | Inhibits EMT, cell proliferation, and invasion |
| NKILA | Upregulates E-cadherin and downregulates N-cadherin and vimentin | Inhibits EMT, suppresses migration and invasion |
| lncRNA | Mechanisms | Therapy Resistance in HNSCC |
|---|---|---|
| PVT1 | MYC, PI3K/AKT activation, apoptosis inhibition | Poor chemotherapy response |
| TUG1 | BCL-2 overexpression | Associated with cisplatin resistance |
| HOTAIR | Epigenetic silencing of tumor suppressors | Poor radiotherapy response |
| H19 | miRNA sponging, Wnt/β-catenin activation | Induces cisplatin resistance |
| MALAT1 | PI3K/AKT activation, apoptosis inhibition | Poor prognosis, cisplatin resistance, radioresistance |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hotnog, C.M.; Bostan, M.; Anghelescu, M.; Roman, V.; Bleotu, C.; Hainarosie, R.; Voiosu, C.; Marineata, S.; Bostan, I.-S.; Diaconu, C.C.; et al. Long Non-Coding RNAs: Significant Drivers of Carcinogenesis Mechanisms in Head and Neck Squamous Cell Carcinoma. Curr. Issues Mol. Biol. 2025, 47, 698. https://doi.org/10.3390/cimb47090698
Hotnog CM, Bostan M, Anghelescu M, Roman V, Bleotu C, Hainarosie R, Voiosu C, Marineata S, Bostan I-S, Diaconu CC, et al. Long Non-Coding RNAs: Significant Drivers of Carcinogenesis Mechanisms in Head and Neck Squamous Cell Carcinoma. Current Issues in Molecular Biology. 2025; 47(9):698. https://doi.org/10.3390/cimb47090698
Chicago/Turabian StyleHotnog, Camelia Mia, Marinela Bostan, Matei Anghelescu, Viviana Roman, Coralia Bleotu, Razvan Hainarosie, Catalina Voiosu, Stefania Marineata, Ioana-Stefania Bostan, Carmen Cristina Diaconu, and et al. 2025. "Long Non-Coding RNAs: Significant Drivers of Carcinogenesis Mechanisms in Head and Neck Squamous Cell Carcinoma" Current Issues in Molecular Biology 47, no. 9: 698. https://doi.org/10.3390/cimb47090698
APA StyleHotnog, C. M., Bostan, M., Anghelescu, M., Roman, V., Bleotu, C., Hainarosie, R., Voiosu, C., Marineata, S., Bostan, I.-S., Diaconu, C. C., & Mihaila, M. (2025). Long Non-Coding RNAs: Significant Drivers of Carcinogenesis Mechanisms in Head and Neck Squamous Cell Carcinoma. Current Issues in Molecular Biology, 47(9), 698. https://doi.org/10.3390/cimb47090698









