Synthesis, Molecular Docking, and Biological Evaluation of Novel Anthranilic Acid Hybrid and Its Diamides as Antispasmodics
Abstract
:1. Introduction
2. Results and Discussion
2.1. In Silico Predictions and Synthesis
2.2. In Silico Analysis
2.3. In Vitro Biological Activity
2.3.1. Antimicrobial Activity
2.3.2. Cytotoxicity
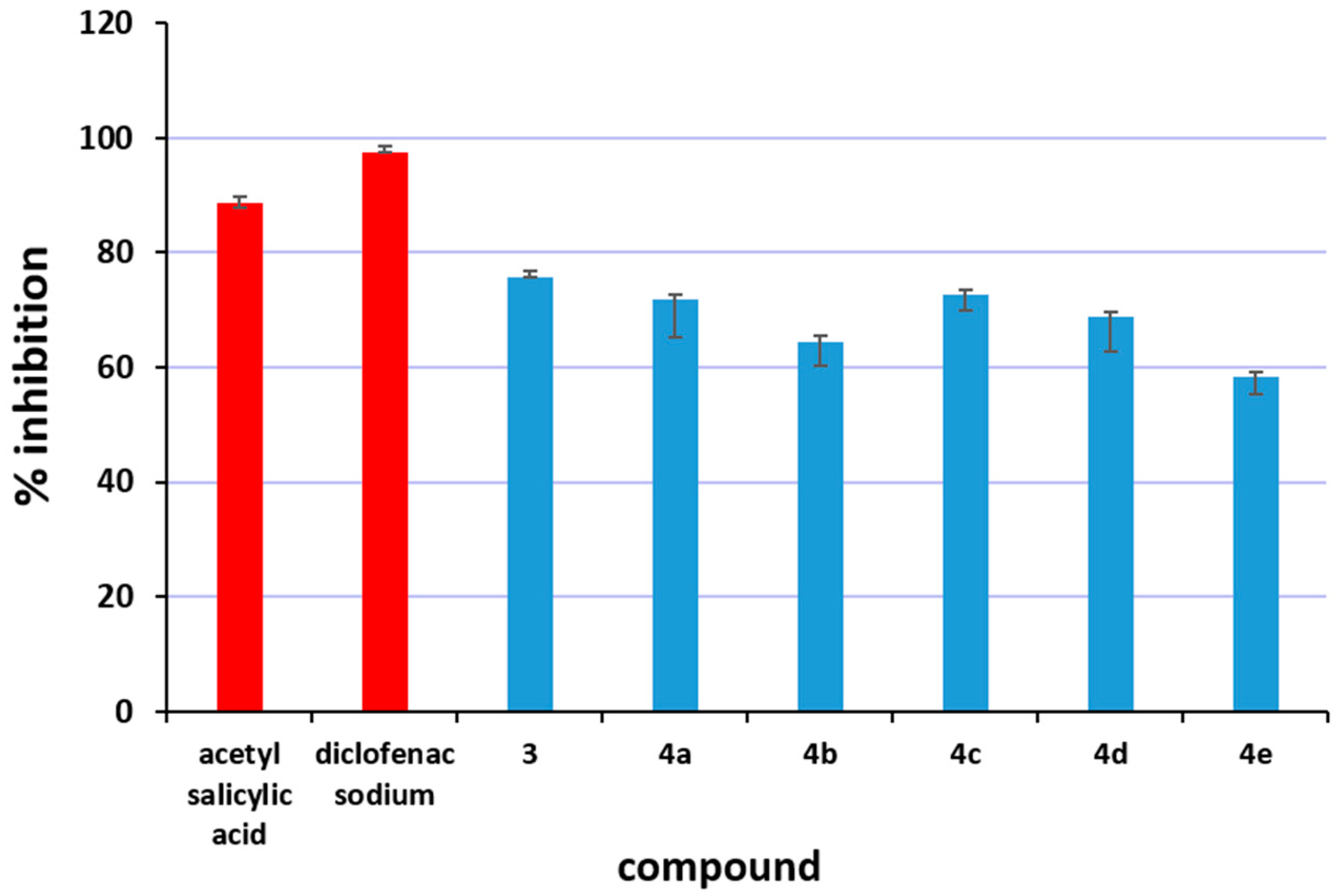
2.3.3. Inhibition of Albumin Denaturation
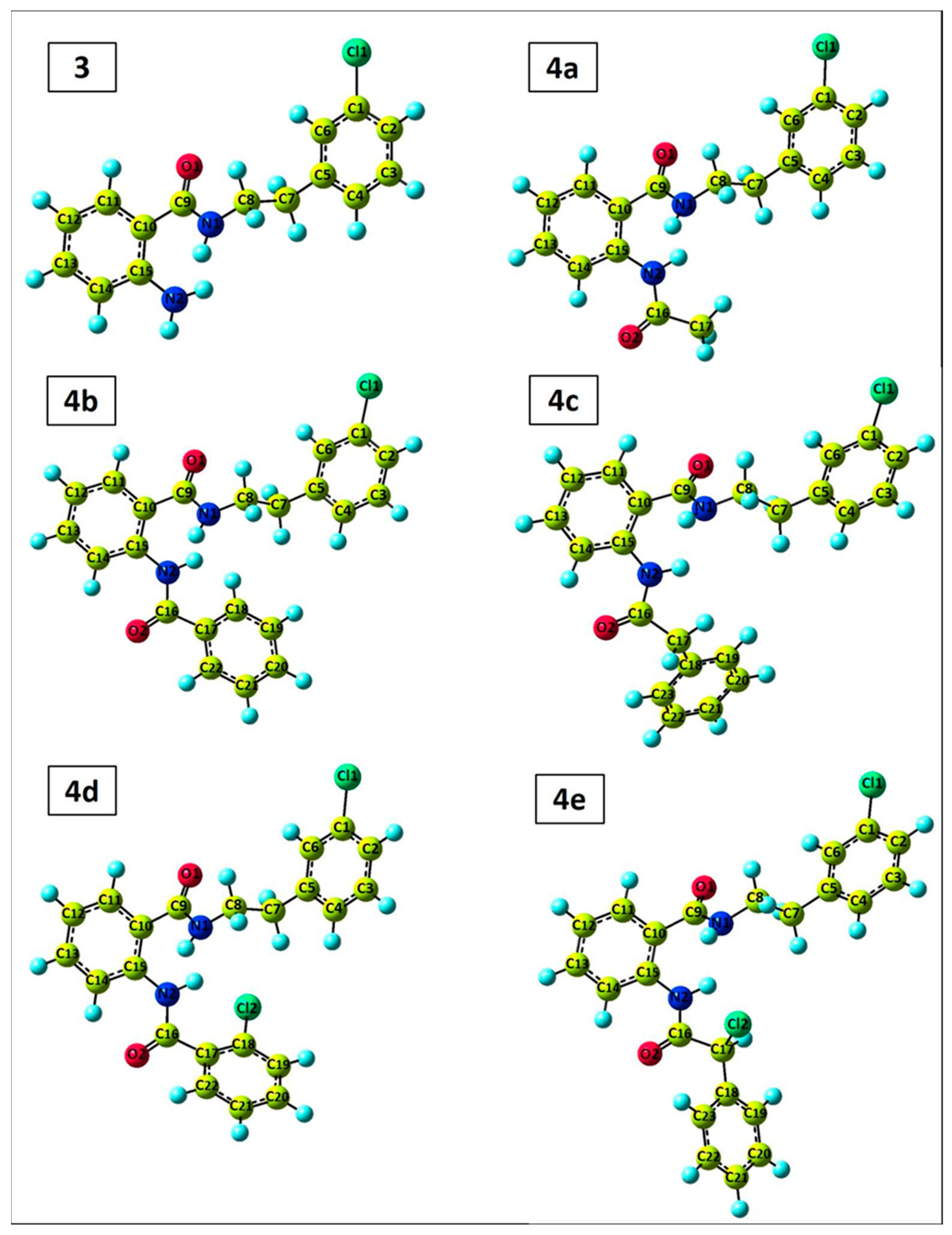
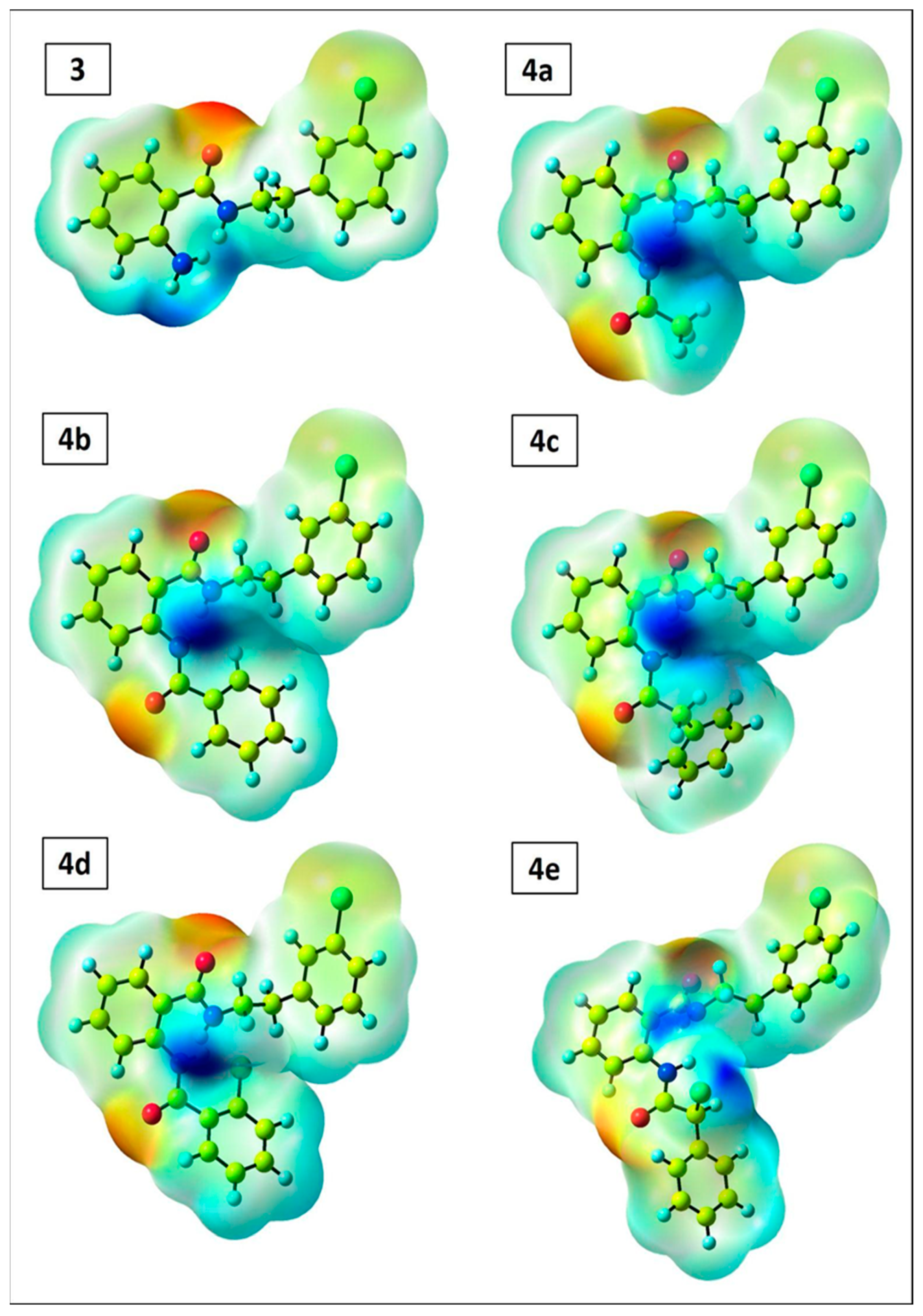
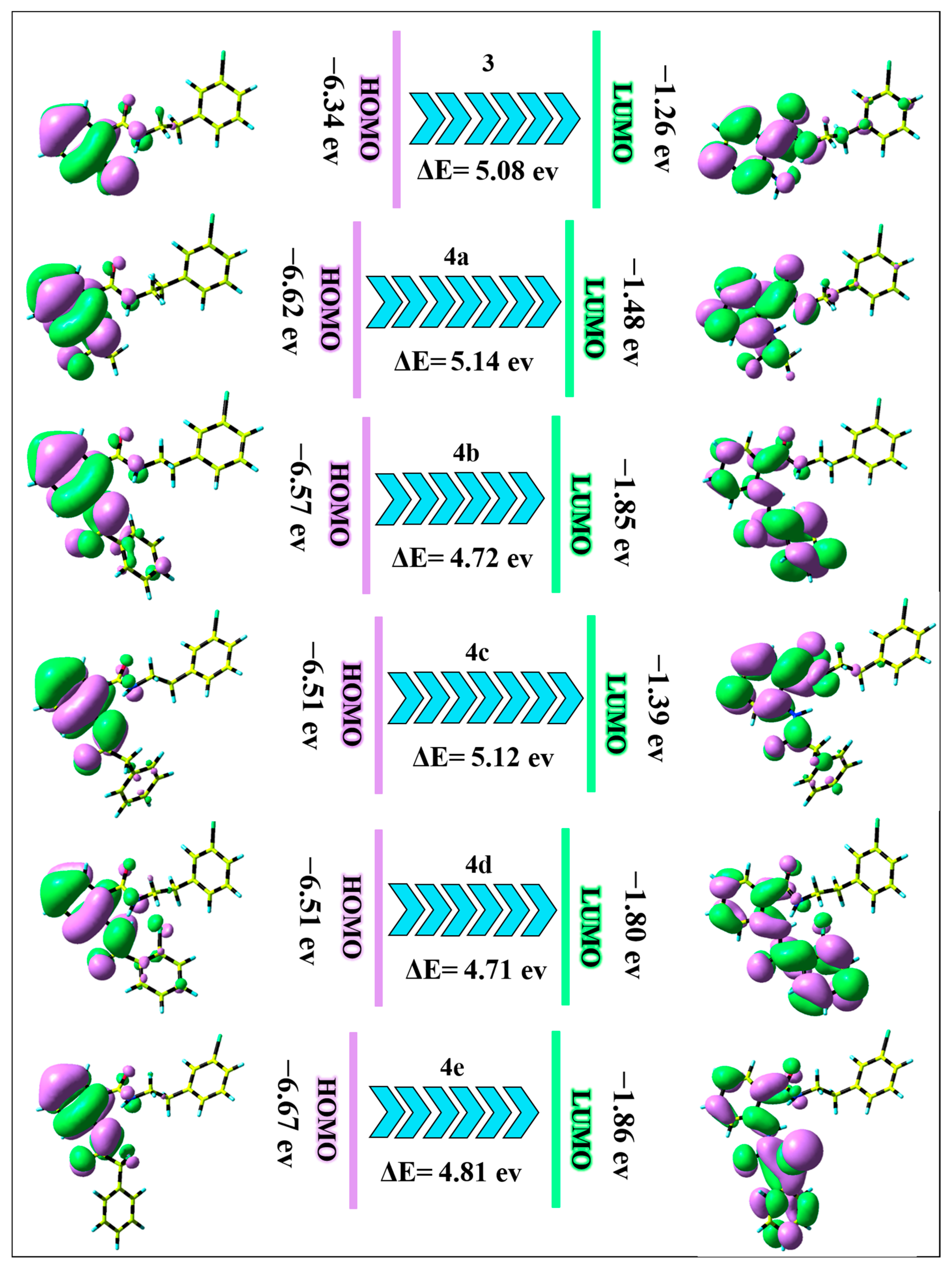
2.4. DFT Perspective
2.5. Molecular Docking Simulation
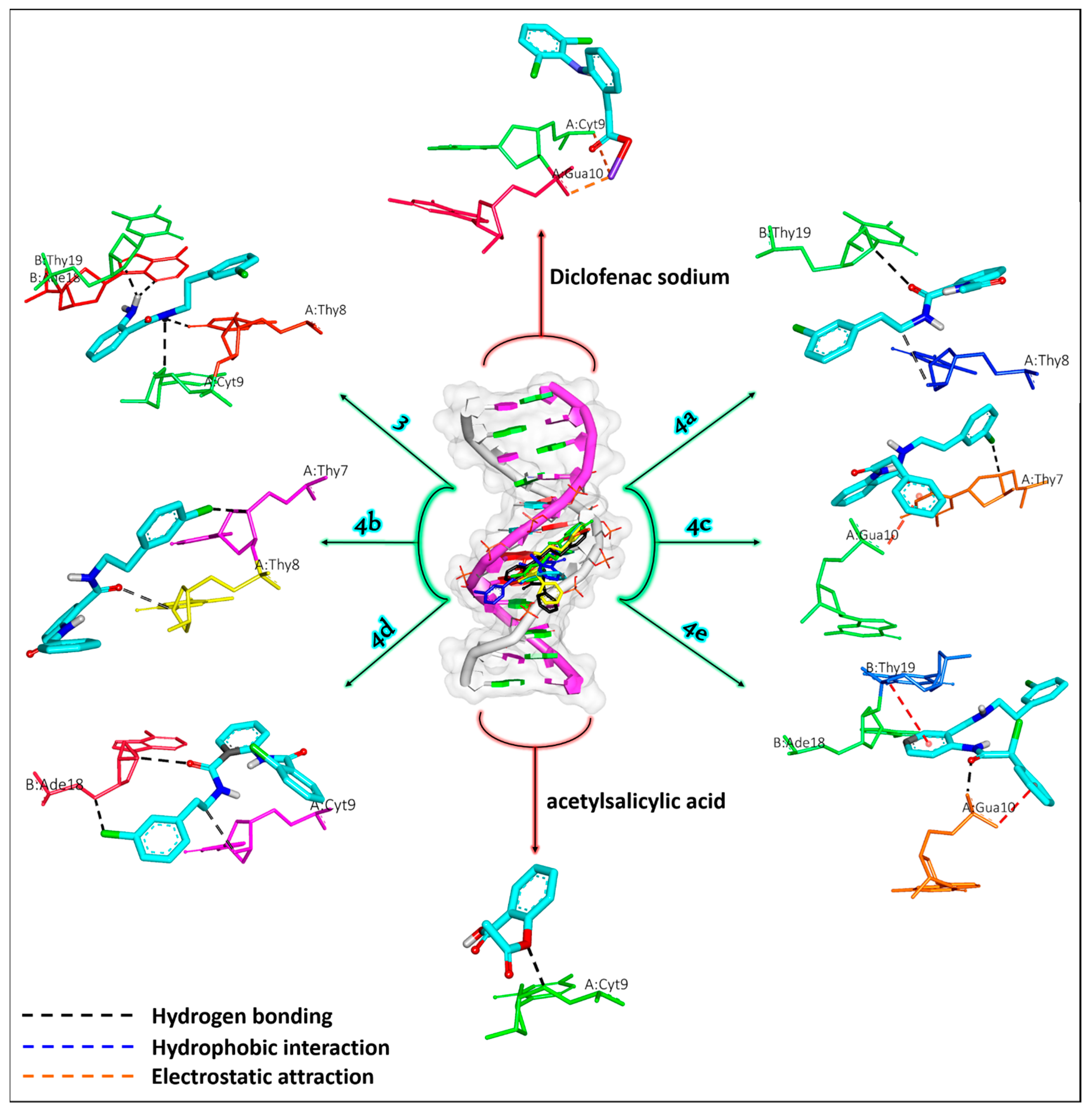
2.5.1. DNA Simulation
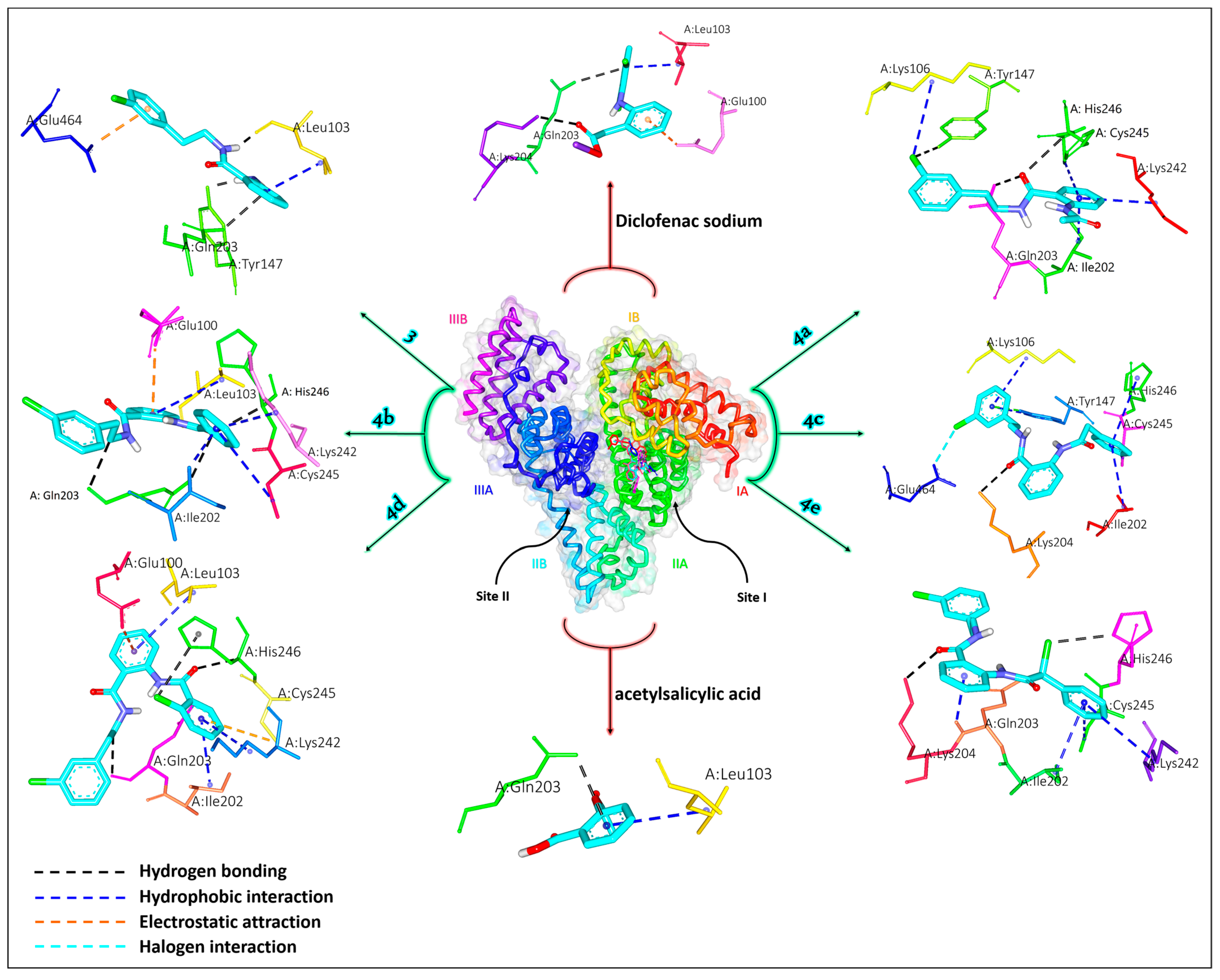
2.5.2. Albumin Simulation
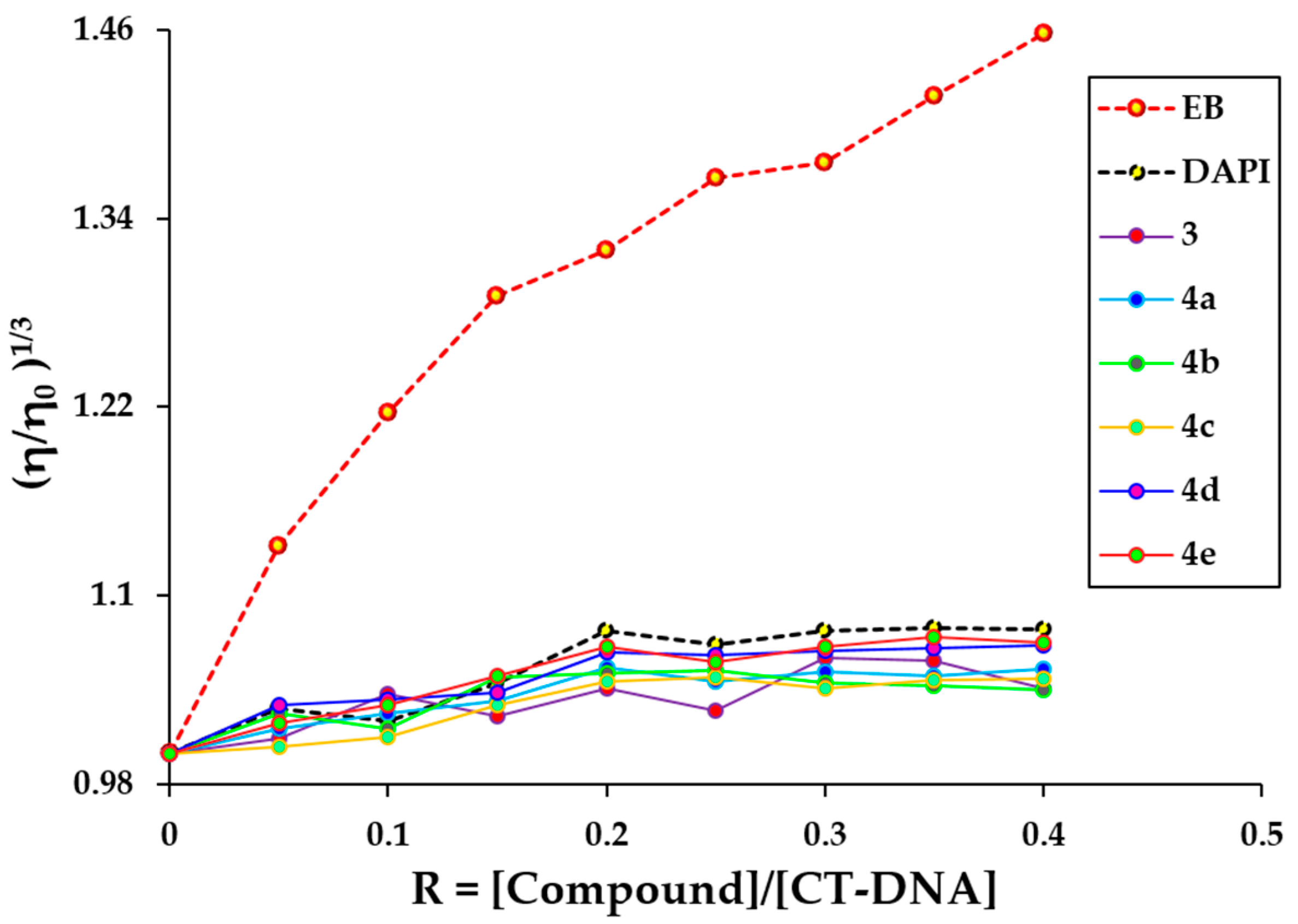
2.6. Viscosity Measurement
2.7. Ex Vivo Smooth Muscle Relaxant Activity
3. Materials and Methods
3.1. Chemicals
3.2. Synthetic Methods Experimental Protocols and Spectral Data
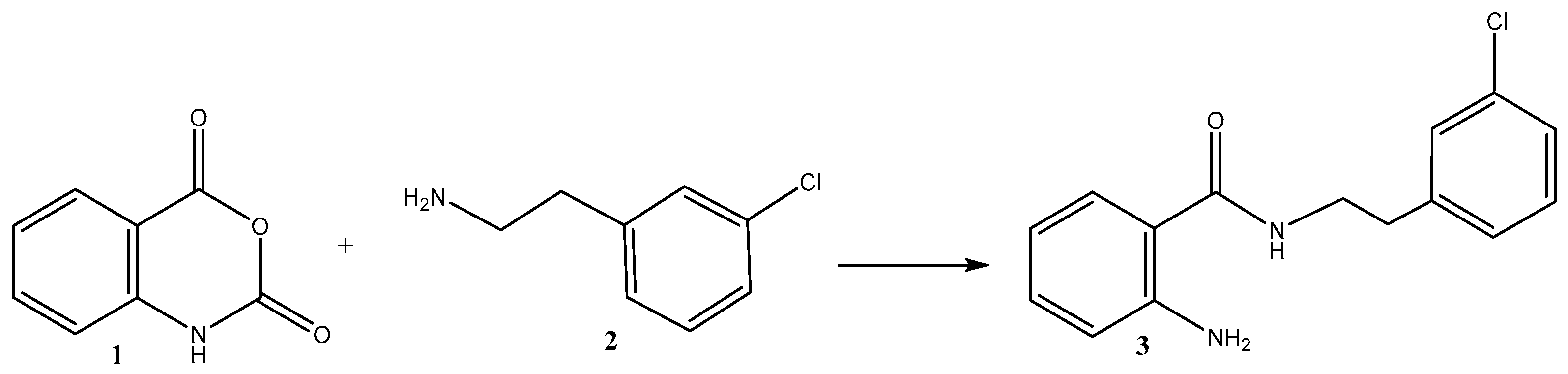
3.2.1. Synthesis of the Hybrid Molecule 2-Amino-N-(3-chlorophenethyl)benzamide 3
3.2.2. Preparation of Diamides of the Hybrid Molecule 4a–e; Typical Procedure
3.3. In Silico Pharmacokinetic Profiling and Toxicity Analysis
3.3.1. Theoretical Prediction of Pharmacokinetic Parameters (ADME)
3.3.2. PASS Online Predictions
3.3.3. OSIRIS
3.3.4. Theoretical Prediction of Toxicity
3.4. DFT Perspective
3.5. Molecular Docking Simulation
3.6. Viscosity Measurement
3.7. Microbiological Tests
3.7.1. Tested Microorganisms
3.7.2. Culture Media
Luria-Bertani Agar Medium Supplemented with Glucose (LBG Agar)
Malt Extract Agar (MEA)
3.7.3. Antimicrobial Activity Assay
3.8. Cytotoxic Activity
3.8.1. Cytotoxicity Assay
3.8.2. MTT Cell Viability Assay
3.9. Determination of the Anti-Inflammatory Activity: Inhibition of Albumin Denaturation
3.10. Smooth Muscle Activity
3.10.1. Ex Vivo Experiments on Gastric Smooth Muscle Preparations (SMPs) from Wistar Rats
3.10.2. Method of Studying a Mechanical Activity of Isolated SMPs
3.11. Ethics Statement
3.12. Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Elshaarawy, R.F.M.; Janiak, C. Antibacterial Susceptibility of New Copper(II) N-Pyruvoyl Anthranilate Complexes against Marine Bacterial Strains—In Search of New Antibiofouling Candidate. Arab. J. Chem. 2016, 9, 825–834. [Google Scholar] [CrossRef]
- Jayanthi, M.; Rajakumar, P. Synthesis and Antimicrobial Activity of Unsymmetrical Dendrimers with Indazole, Salicylates and Anthranilates as Surface Units. J. Het. Chem. 2017, 54, 1963–1973. [Google Scholar] [CrossRef]
- Merk, D.; Lamers, C.; Weber, J.; Flesch, D.; Gabler, M.; Proschak, E.; Schubert-Zsilavecz, M. Anthranilic Acid Derivatives as Nuclear Receptor Modulators—Development of Novel PPAR Selective and Dual PPAR/FXR Ligands. Bioorg. Med. Chem. 2015, 23, 499–514. [Google Scholar] [CrossRef] [PubMed]
- Patrone, J.D.; Pelz, N.F.; Bates, B.S.; Souza-Fagundes, E.M.; Vangamudi, B.; Camper, D.V.; Kuznetsov, A.G.; Browning, C.F.; Feldkamp, M.D.; Frank, A.O.; et al. Identification and Optimization of Anthranilic Acid-Based Inhibitors of Replication Protein A. ChemMedChem 2016, 11, 893–899. [Google Scholar] [CrossRef]
- Kwon, I.-S.; Jong Hwan, K.; Pyo, S.; Lee, H.-W.; Kim, A.; Schmitz, F.J. Oscarellin, an Anthranilic Acid Derivative from a Philippine Sponge, Oscarella Stillans, as an Inhibitor of Inflammatory Cytokines in Macrophages. J. Nat. Prod. 2017, 80, 149–155. [Google Scholar] [CrossRef]
- Teponno, R.B.; Noumeur, S.R.; Helaly, S.E.; Hüttel, S.; Harzallah, D.; Stadler, M. Furanones and Anthranilic Acid Derivatives from the Endophytic Fungus Dendrothyrium Variisporum. Molecules 2017, 22, 1674. [Google Scholar] [CrossRef]
- Schrey, H.; Freya Janina, M.; Harz, P.; Zeljka, R.; Stadler, M.; Spiteller, P. Nematicidal Anthranilic Acid Derivatives from Laccaria Species. Phytochemistry. 2019, 160, 85–91. [Google Scholar] [CrossRef]
- Oxenkrug, G.; van der Hart, M.; Roeser, J.; Summergrad, P. Anthranilic Acid: A Potential Biomarker and Treatment Target for Schizophrenia. Ann. Psychiatry Ment. Health 2016, 4, 1059. [Google Scholar]
- Prasher, P.; Sharma, M. “Azole” as Privileged Heterocycle for Targeting the Inducible Cyclooxygenase Enzyme. Drug Dev. Res. 2020, 82, 167–197. [Google Scholar] [CrossRef]
- Prasher, P.; Sharma, M.; Zacconi, F.; Gupta, G.; Aljabali, A.A.A.; Mishra, V.; Tambuwala, M.M.; Kapoor, D.N.; Negi, P.; de Pinto, T.J.A.; et al. Synthesis and Anticancer Properties of “Azole” Based Chemotherapeutics as Emerging Chemical Moieties: A Comprehensive Review. Curr. Org. Chem. 2021, 25, 654–668. [Google Scholar]
- Varnavas, A.; Lassiani, L.; Valenta, V.; Mennuni, L.; Makovec, F.; Hadjipavlou-Litina, D. Anthranilic Acid Based CCK1 Receptor Antagonists: Preliminary Investigation on Their Second “Touch Point”. Eur. J. Med. Chem. 2005, 40, 563–581. [Google Scholar] [CrossRef] [PubMed]
- Crawley, J.N.; Corwin, R.L. Biological Actions of Cholecystokinin. Peptides 1994, 15, 731–755. [Google Scholar] [CrossRef] [PubMed]
- Wank, S.A.I. CCK Receptors: An Exemplary Family. Am. J. Physiol. Gastrointest. Liver Physiol. 1998, 274, G607–G613. [Google Scholar] [CrossRef] [PubMed]
- Nichols, D.E. Hallucinogens. Pharmacol. Ther. 2004, 101, 131–181. [Google Scholar] [CrossRef]
- Berger, M.; Gray, J.A.; Roth, B.L. The Expanded Biology of Serotonin. Annu. Rev. Med. 2009, 60, 355–366. [Google Scholar] [CrossRef]
- Ramage, A.G.; Villalón, C.M. 5-Hydroxytryptamine and Cardiovascular Regulation. Trends Pharmacol. Sci. 2008, 29, 472–481. [Google Scholar] [CrossRef]
- Monti, J.M.; Jantos, H. The Roles of Dopamine and Serotonin, and of Their Receptors, in Regulating Sleep and Waking. Prog. Brain Res. 2008, 172, 625–646. [Google Scholar] [CrossRef]
- Guzel, T.; Mirowska-Guzel, D. The Role of Serotonin Neurotransmission in Gastrointestinal Tract and Pharmacotherapy. Molecules 2022, 27, 1680. [Google Scholar] [CrossRef]
- Szymaszkiewicz, A.; Zieli’nska, M. Irritable bowel syndrome: Current therapies and future perspectives. In A Comprehensive Overview of Irritable Bowel Syndrome; Fichna, J., Ed.; Elsevier: London, UK, 2020; pp. 129–144. ISBN 978-0-12-821324-7. [Google Scholar] [CrossRef]
- Milusheva, M.; Gledacheva, V.; Batmazyan, M.; Nikolova, S.; Stefanova, I.; Dimitrova, D.; Saracheva, K.; Tomov, D.; Chaova-Gizdakova, V. Ex Vivo and In Vivo Study of Some Isoquinoline Precursors. Sci. Pharm. 2022, 90, 37. [Google Scholar] [CrossRef]
- Milusheva, M.; Gledacheva, V.; Stefanova, I.; Pencheva, M.; Mihaylova, R.; Tumbarski, Y.; Nedialkov, P.; Cherneva, E.; Todorova, M.; Nikolova, S. In Silico, In Vitro, and Ex Vivo Biological Activity of Some Novel Mebeverine Precursors. Biomedicines 2023, 11, 605. [Google Scholar] [CrossRef]
- Xiang, J.; Wang, H.; Ma, C.; Zhou, M.; Wu, Y.; Wang, L.; Guo, S.; Chen, T.; Shaw, C. Ex Vivo Smooth Muscle Pharmacological Effects of a Novel Bradykinin-Related Peptide, and Its Analogue, from Chinese Large Odorous Frog, Odorrana Livida Skin Secretions. Toxins 2016, 8, 283. [Google Scholar] [CrossRef] [PubMed]
- Wright, D.; Sharma, P.; Ryu, M.-H.; Rissé, P.-A.; Ngo, M.; Maarsingh, H.; Koziol-White, C.; Jha, A.; Halayko, A.J.; West, A.R. Models to Study Airway Smooth Muscle Contraction In Vivo, Ex Vivo and In Vitro: Implications in Understanding Asthma. Pulm. Pharmacol. Ther. 2013, 26, 24–36. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Lin, Z. Vascular Smooth Muscle Cells Mechanosensitive Regulators and Vascular Remodeling. J. Vasc. Res. 2021, 59, 90–113. [Google Scholar] [CrossRef] [PubMed]
- Hashitani, H.; Brading, A.F.; Suzuki, H. Correlation between Spontaneous Electrical, Calcium and Mechanical Activity in Detrusor Smooth Muscle of the Guinea-Pig Bladder. Br. J. Pharmacol. 2004, 141, 183–193. [Google Scholar] [CrossRef] [PubMed]
- Ivanov, I.; Nikolova, S.; Aladjov, D.; Stefanova, I.; Zagorchev, P. Synthesis and Contractile Activity of Substituted 1,2,3,4-Tetrahydroisoquinolines. Molecules 2011, 16, 7019–7042. [Google Scholar] [CrossRef]
- Sagorchev, P.; Lukanov, J. Effects of 1.8-Cineole (Eucalyptol) on the Spontaneous Contractile Activity of Smooth Muscles Fibre. J. Med. Plants Res. 2015, 9, 486–493. [Google Scholar] [CrossRef]
- Gledacheva, V.; Pencheva, M.; Nikolova, S.; Stefanova, I. Ability of 2-Chloro-N-(1-(3,4-dimethoxyphenyl)propan-2-yl)-2-phenylacetamide to Stimulate Endogenous Nitric Oxide Synthesis. Appl. Sci. 2022, 12, 4473. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A Free Web Tool to Evaluate Pharmacokinetics, Drug-Likeness and Medicinal Chemistry Friendliness of Small Molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef]
- Anzali, S.; Barnickel, G.; Cezanne, B.; Krug, M.; Filimonov, D.; Poroikov, V. Discriminating between Drugs and Nondrugs by Prediction of Activity Spectra for Substances (PASS). J. Med. Chem. 2001, 44, 2432–2437. [Google Scholar] [CrossRef]
- Mathew, B.; Suresh, J.; Anbazhagan, S. Synthesis and PASS-Assisted in Silico Approach of Some Novel 2-Substituted Benzimidazole Bearing a Pyrimidine-2, 4, 6(Trione) System as Mucomembranous Protector. J. Pharm. Bioallied Sci. 2013, 5, 39–43. [Google Scholar] [CrossRef]
- Ekins, S.; Olechno, J.; Williams, A.J. Dispensing Processes Impact Apparent Biological Activity as Determined by Computational and Statistical Analyses. PLoS ONE 2013, 8, e62325. [Google Scholar] [CrossRef]
- Roughley, S.D.; Jordan, A.M. The Medicinal Chemist’s Toolbox: An Analysis of Reactions Used in the Pursuit of Drug Candidates. J. Med. Chem. 2011, 54, 3451–3479. [Google Scholar] [CrossRef]
- Seavill, P.W.; Wilden, J.D. The Preparation and Applications of Amides Using Electrosynthesis. Green Chem. 2020, 22, 7737–7759. [Google Scholar] [CrossRef]
- Wang, X. Challenges and Outlook for Catalytic Direct Amidation Reactions. Nat. Catal. 2019, 2, 98–102. [Google Scholar] [CrossRef]
- Bray, B.L. Large-Scale Manufacture of Peptide Therapeutics by Chemical Synthesis. Nat. Rev. Drug Discov. 2003, 2, 587–593. [Google Scholar] [CrossRef] [PubMed]
- Sangamwar, A.; Deshpande, U.; Pekamwar, S.; Vadvalkar, S. Improving Decision Making for Drug Candidates: A Computational Approach for Benzthiazoles as Antifungal. Indian J. Biotechnol. 2007, 6, 389–396. [Google Scholar]
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 2001, 46, 3–26. [Google Scholar] [CrossRef]
- Prasanna, S.; Doerksen, R. Topological Polar Surface Area: A Useful Descriptor in 2D-QSAR. Curr. Med. Chem. 2009, 16, 21–41. [Google Scholar] [CrossRef]
- Maximo da Silva, M.; Comin, M.; Santos Duarte, T.; Foglio, M.; de Carvalho, J.; do Carmo Vieira, M.; Nazari Formagio, A. Synthesis, Antiproliferative Activity and Molecular Properties Predictions of Galloyl Derivatives. Molecules 2015, 20, 5360–5373. [Google Scholar] [CrossRef]
- Ottaviani, G.; Gosling, D.J.; Patissier, C.; Rodde, S.; Zhou, L.; Faller, B. What Is Modulating Solubility in Simulated Intestinal Fluids? Eur. J. Pharm. Sci. 2010, 41, 452–457. [Google Scholar] [CrossRef]
- Hollenberg, P.F. Characteristics and Common Properties of Inhibitors, Inducers, and Activators of CYP Enzymes. Drug Metab. Rev. 2002, 34, 17–35. [Google Scholar] [CrossRef] [PubMed]
- Zafirah Ismail, N.; Annamalai, N.; Mohamad Zain, N.N.; Arsad, H. Molecular Docking of Selected Compounds from Clinacanthus Nutans with Bcl-2, P53, Caspase-3 and Caspase-8 Proteins in the Apoptosis Pathway. J. Biol. Sci. Opin. 2020, 8, 4–11. [Google Scholar] [CrossRef]
- Banerjee, P.; Eckert, A.O.; Schrey, A.K.; Preissner, R. ProTox-II: A Webserver for the Prediction of Toxicity of Chemicals. Nucleic Acids Res. 2018, 46, W257–W263. [Google Scholar] [CrossRef] [PubMed]
- Mazumder, K.; Emran Hossain, M.; Aktar, A.; Mohiuddin, M.W.; Kishore Kumar, S.; Biswas, B.; Abdullah Nur, A.; Ahsan Abid, M.; Fukase, K. In Silico Analysis and Experimental Evaluation of Ester Prodrugs of Ketoprofen for Oral Delivery: With a View to Reduce Toxicity. Processes 2021, 9, 2221. [Google Scholar] [CrossRef]
- Trognon, J.; Vera, G.; Rima, M.; Stigliani, J.-L.; Amielet, L.; El Hage, S.; Lajoie, B.; Roques, C.; El Garah, F. Investigation of Direct and Retro Chromone-2-Carboxamides Based Analogs of Pseudomonas Aeruginosa Quorum Sensing Signal as New Anti-Biofilm Agents. Pharmaceuticals 2022, 15, 417. [Google Scholar] [CrossRef]
- De Oliveira, D.M.P.; Forde, B.M.; Kidd, T.J.; Harris, P.N.A.; Schembri, M.A.; Beatson, S.A.; Paterson, D.L.; Walker, M.J. Antimicrobial Resistance in ESKAPE Pathogens. Clin. Microbiol. Rev. 2020, 33, e00181-19. [Google Scholar] [CrossRef]
- Chen, Y.; Tang, J.; Tang, X.; Wang, C.; Lian, Y.; Shao, Z.; Yao, X.-S.; Gao, H. New Phenethylamine Derivatives from Arenibacter Nanhaiticus Sp. Nov. NH36AT and Their Antimicrobial Activity. J. Antibiot. 2013, 66, 655–661. [Google Scholar] [CrossRef]
- Wang, L.; Linares-Otoya, V.; Liu, Y.; Mettal, U.; Marner, M.; Armas-Mantilla, L.; Willbold, S.; Kurtán, T.; Linares-Otoya, L.; Schäberle, T.F. Discovery and Biosynthesis of Antimicrobial Phenethylamine Alkaloids from the Marine Flavobacterium Tenacibaculum Discolor Sv11. J. Nat. Prod. 2022, 85, 1039–1051. [Google Scholar] [CrossRef]
- Muchaamba, F.; Stephan, R.; Tasara, T. β-Phenylethylamine as a Natural Food Additive Shows Antimicrobial Activity against Listeria Monocytogenes on Ready-To-Eat Foods. Foods 2020, 9, 1363. [Google Scholar] [CrossRef]
- Pontiki, E.; Hadjipavlou-Litina, D. Synthesis and Pharmacochemical Evaluation of Novel Aryl-Acetic Acid Inhibitors of Lipoxygenase, Antioxidants, and Anti-Inflammatory Agents. Bioorg. Med. Chem. 2007, 15, 5819–5827. [Google Scholar] [CrossRef]
- Shaaban, S.; Abdou, A.; Alhamzani, A.G.; Abou-Krisha, M.M.; Al-Qudah, M.A.; Alaasar, M.; Youssef, I.; Yousef, T.A. Synthesis and in Silico Investigation of Organoselenium-Clubbed Schiff Bases as Potential Mpro Inhibitors for the SARS-CoV-2 Replication. Life 2023, 13, 912. [Google Scholar] [CrossRef] [PubMed]
- Dorafshan Tabatabai, A.S.; Dehghanian, E.; Mansouri-Torshizi, H.; Feizi-Dehnayebi, M. Computational and Experimental Examinations of New Antitumor Palladium(II) Complex: CT-DNA-/BSA-Binding, In-Silico Prediction, DFT Perspective, Docking, Molecular Dynamics Simulation and ONIOM. J. Biomol. Struct. Dyn. 2023, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Nikolova, S.; Milusheva, M.; Gledacheva, V.; Feizi-Dehnayebi, M.; Kaynarova, L.; Georgieva, D.; Delchev, V.; Stefanova, I.; Tumbarski, Y.; Mihaylova, R.; et al. Drug-Delivery Silver Nanoparticles: A New Perspective for Phenindione as an Anticoagulant. Biomedicines 2023, 11, 2201. [Google Scholar] [CrossRef]
- Feizi-Dehnayebi, M.; Dehghanian, E.; Mansouri-Torshizi, H. Biological Activity of Bis-(Morpholineacetato)Palladium(II) Complex: Preparation, Structural Elucidation, Cytotoxicity, DNA-/Serum Albumin-Interaction, Density Functional Theory, In-Silico Prediction and Molecular Modeling. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2022, 281, 121543. [Google Scholar] [CrossRef] [PubMed]
- Alotaibi, H.S.; Momen, A. Anticancer Drugs’ Deoxyribonucleic Acid (DNA) Interactions. In Biophysical Chemistry—Advance Applications; Khalid, M., Ed.; IntechOpen Limited: London, UK, 2020; ISBN 978-1-83880-138-0. [Google Scholar] [CrossRef]
- Kumari, C.S.; Yasmin, N.; Hussain, M.R.; Babuselvam, M. Invitro Anti-Inflammatory and Anti-Artheritic Property of Rhizopora Mucronata Leaves. Int. J. Pharm. Sci. Res. 2015, 6, 482–485. [Google Scholar]
- Ndoye Foe, F.M.C.; Tchinang, T.F.K.; Nyegue, A.M.; Abdou, J.P.; Yaya, A.J.G.; Tchinda, A.T.; Essame, J.O.; Etoa, F.X. Chemical composition, in vitro antioxidant and anti-inflammatory properties of essential oils of four dietary and medicinal plants from Cameroon. BMC Complement. Altern. Med. 2016, 16, 117. [Google Scholar] [CrossRef]
- Kar, B.; Suresh Kumar, R.B.; Karmakar, I.; Dola, N.; Bala, A.; Mazumder, U.K.; Hadar, P.K. Antioxidant and in vitro anti-inflammatory activities of Mimusops elengi leaves. Asian Pac. J. Trop. Biomed. 2012, 2, S976–S980. [Google Scholar] [CrossRef]
- Osman, N.I.; Sidik, N.J.; Awal, A.; Adam, N.A.; Rezali, N.I. In vitro xanthine oxidase and albumin denaturation inhibition assay of Barringtonia racemosa L. and total phenolic content analysis for potential anti-inflammatory use in gouty arthritis. J. Intercult. Ethnopharmacol. 2016, 5, 343–349. [Google Scholar] [CrossRef]
- Ghorai, P.; Saha, R.; Bhuiya, S.; Das, S.; Brandão, P.; Ghosh, D.; Bhaumik, T.; Bandyopadhyay, P.; Chattopadhyay, D.; Saha, A. Syntheses of Zn (II) and Cu (II) Schiff base complexes using N, O donor Schiff base ligand: Crystal structure, DNA binding, DNA cleavage, docking and DFT study. Polyhedron 2018, 141, 153–163. [Google Scholar] [CrossRef]
- Jiang, G.B.; Zhang, W.Y.; He, M.; Gu, Y.Y.; Bai, L.; Wang, Y.J.; Yi, Q.Y.; Du, F. New ruthenium polypyridyl complexes functionalized with fluorine atom or furan: Synthesis, DNA-binding, cytotoxicity and antitumor mechanism studies. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2020, 227, 117534. [Google Scholar] [CrossRef]
- Amend, N.; Horst, T.; Worek, F.; Wille, T. A Pharmacologically Pre-Contracted Smooth Muscle Bowel Model for the Study of Highly-Potent Opioid Receptor Agonists and Antagonists. Toxicol. Lett. 2023, 382, 41–46. [Google Scholar] [CrossRef] [PubMed]
- Jespersen, B.; Tykocki, N.R.; Watts, S.W.; Cobbett, P.J. Measurement of Smooth Muscle Function in the Isolated Tissue Bath-Applications to Pharmacology Research. J. Vis. Exp. 2015, 95, e52324. [Google Scholar] [CrossRef]
- Świt, P.; Pollap, A.; Orzeł, J. Spectroscopic Determination of Acetylcholine (ACh): A Representative Review. Top. Curr. Chem. 2023, 381, 16. [Google Scholar] [CrossRef]
- Tanahashi, Y.; Komori, S.; Matsuyama, H.; Kitazawa, T.; Unno, T. Functions of Muscarinic Receptor Subtypes in Gastrointestinal Smooth Muscle: A Review of Studies with Receptor-Knockout Mice. Int. J. Mol. Sci. 2021, 22, 926. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Khalil, R.A. Evolving Mechanisms of Vascular Smooth Muscle Contraction Highlight Key Targets in Vascular Disease. Biochem. Pharmacol. 2018, 153, 91–122. [Google Scholar] [CrossRef]
- Frisch, M.; Trucks, G.; Schlegel, H.B.; Scuseria, G.E.; Robb, M.A.; Cheeseman, J.R.; Scalmani, G.; Barone, V.; Mennucci, B.; Petersson, G. Gaussian 09, Revision D.01; Gaussian, Inc.: Wallingford, CT, USA, 2009; p. 201. [Google Scholar]
- Garrett, M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comp. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef]
- MGL Tools. 1.5. 6 (ADT)/MGL Tools 1. 6.; The Scripps Research Institute: La Jolla, CA, USA, 2016. [Google Scholar]
- Feizi-Dehnayebi, M.; Effat, D.; Hassan, M.-T. A novel palladium (II) antitumor agent: Synthesis, characterization, DFT perspective, CT-DNA and BSA interaction studies via in-vitro and in-silico approaches. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2021, 249, 119215. [Google Scholar] [CrossRef]
- Revathi, N.; Sankarganesh, M.; Dhaveethu Raja, J.; Vinoth Kumar, G.G.; Sakthivel, A.; Rajasekaran, R. Bio-active mixed ligand Cu (II) and Zn (II) complexes of pyrimidine derivative Schiff base: DFT calculation, antimicrobial, antioxidant, DNA binding, anticancer and molecular docking studies. JBSD 2021, 39, 3012–3024. [Google Scholar] [CrossRef]
- Gaber, M.; Fathalla, S.K.; El-Ghamry, H.A. 2,4-Dihydroxy-5-[(5-mercapto-1H-1, 2, 4-triazole-3-yl) diazenyl] benzaldehyde acetato, chloro and nitrato Cu (II) complexes: Synthesis, structural characterization, DNA binding and anticancer and antimicrobial activity. Appl. Organomet. Chem. 2019, 33, e4707. [Google Scholar] [CrossRef]
- Tumbarski, Y.; Deseva, I.; Mihaylova, D.; Stoyanova, M.; Krastev, L.; Nikolova, R.; Yanakieva, V.; Ivanov, I. Isolation, Characterization and Amino Acid Composition of a Bacteriocin Produced by Bacillus Methylotrophicus Strain BM47. Food Technol. Biotechnol. 2018, 56, 546. [Google Scholar] [CrossRef]
- Lee, S.; Kim, M.-J.; Lee, B.S.; Ryoo, R.; Kim, H.K.; Kim, K.H. Cumulative Effects of Constituents from the Mushroom Calvatia Nipponica on the Contractility of Penile Corpus Cavernosum Smooth Muscle. Mycobiology 2020, 48, 153–156. [Google Scholar] [CrossRef] [PubMed]
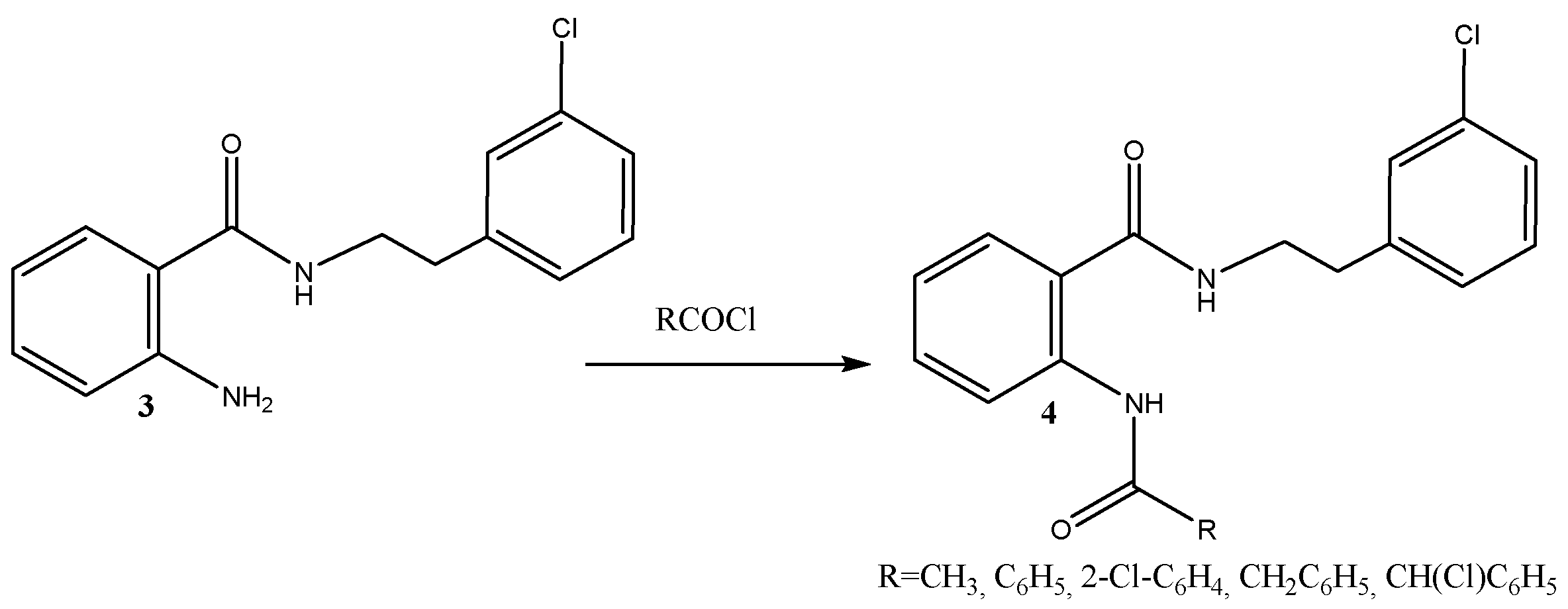

| 4 | R | Yield, % | mp, °C |
|---|---|---|---|
| a | CH3 | 83 | 135–137 |
| b | C6H5 | 79 | 106–108 |
| c | CH2-C6H5 | 79 | 82–83 |
| d | 2-Cl-C6H4 | 80 | 61–63 |
| e | CH(Cl)C6H5 | 81 | 83–84 |
| Anthranilic Acid Hybrid and Its Diamides | 3 | 4a | 4b | 4c | 4d | 4e |
|---|---|---|---|---|---|---|
| logP a | 3.06 | 3.23 | 4.34 | 4.40 | 4.88 | 4.67 |
| TPSA b | 55.12 | 58.20 | 58.20 | 58.20 | 58.20 | 58.20 |
| Nviol c | 0 | 0 | 1:MLogP > 4.15 | 1:MlogP > 4.15 | 1:MlogP > 4.15 | 1:MlogP > 4.15 |
| Nrotb d | 5 | 7 | 8 | 9 | 8 | 9 |
| Water solubility Log S (ESOL) | −4.47 | −4.45 | −5.9 | −5.87 | −6.99 | −6.52 |
| GI absorption | High | High | High | High | High | High |
| Bioavailability score | 0.55 | 0.55 | 0.55 | 0.55 | 0.55 | 0.55 |
| BBB permeability | Yes | Yes | Yes | Yes | Yes | Yes |
| cytochrome P450 inhibition | Yes | Yes | Yes | Yes | Yes | Yes |
| Log Kp (skin permeation) | −5.02 | −5.38 | −4.58 | −4.70 | −3.98 | −4.40 |
| Synth. Access. | 1.81 | 2.28 | 2.59 | 2.71 | 2.84 | 3.34 |
| Inhibition Zones, mm | ||||||
|---|---|---|---|---|---|---|
| Anthranilic Acid Hybrid and Its Diamides | 3 | 4a | 4b | 4c | 4d | 4e |
| Bacillus subtilis ATCC 6633 | 12 | 8 | 11 | 9 | 11 | 10 |
| Bacillus amyloliquefaciens 4BCL-YT | 11 | - | 10 | - | 10 | - |
| Staphylococcus aureus ATCC 25923 | 8 | - | 8 | - | - | - |
| Listeria monocytogenes NBIMCC 8632 | 12 | 8 | 12 | - | 11 | 9 |
| Enterococcus faecalis ATCC 29212 | 12 | 12 | 8 | - | 8 | - |
| Micrococcus luteus 2YC-YT | 19 | 10 | 15 | 12 | 12 | 9 |
| Salmonella enteritidis ATCC 13076 | 12 | 12 | 8 | 8 | 10 | 11 |
| Salmonella typhimurium NBIMCC 1672 | 12 | 8 | 8 | - | 12 | - |
| Klebsiella pneumoniae ATCC 13883 | 8 | 14 | - | - | 8 | 16 |
| Escherichia coli ATCC 25922 | 13 | 9 | 11 | 10 | 12 | 10 |
| Proteus vulgaris ATCC 6380 | 8 | 8 | 8 | - | 8 | 8 |
| Pseudomonas aeruginosa ATCC 9027 | 14 | 9 | 10 | 10 | 10 | 10 |
| Candida albicans NBIMCC 74 | 15 | - | - | - | - | - |
| Saccharomyces cerevisiae ATCC 9763 | 15 | - | - | - | 8 | 8 |
| Aspergillus niger ATCC 1015 | 20 | 10 | 11 | 10 | 10 | 10 |
| Aspergillus flavus | 20 | 8 | 13 | 8 | 11 | 8 |
| Penicillium chrysogenum | 20 | 13 | 15 | 8 | 15 | 8 |
| Rhizopus sp. | 20 | 8 | 11 | 8 | 9 | 9 |
| Fusarium moniliforme ATCC 38932 | 15 | - | - | - | 8 | 8 |
| Mucor sp. | 15 | - | - | - | - | - |
| Cell Line/Compound | LAMA-84 a | K-562 b | T-24 c | CCL-1 d |
|---|---|---|---|---|
| 3 | 114.8 ± 10.5 | 154.4 ± 11.1 | 137.4 ± 12.5 | >200 |
| 4a | 111.5 ± 9.4 | >200 | >200 | >200 |
| 4b | 60.7 ± 5.3 | 55.2 ± 4.7 | 94.9 ± 5.0 | >200 |
| 4c | 31.6 ± 1.1 | 27.4 ± 2.4 | >200 | 69.5 ± 6.1 |
| 4d | 183 ± 13.3 | 99.2 ± 8.8 | >200 | >200 |
| 4e | 193.2 ± 14.7 | 101.5 ± 7.9 | 119.9 ± 9.9 | 69.1 ± 4.6 |
| Compound | IC50, mcg/mL |
|---|---|
| 3 | 660 |
| 4a | 650 |
| 4b | 740 |
| 4c | 670 |
| 4d | 690 |
| 4e | 830 |
| Compound | EHOMO | ELUMO | ΔE | χ | Pi | η | σ | ω | ΔN | ET | Cv | S |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 3 | −6.34 | −1.26 | 5.08 | 3.80 | −3.80 | 2.54 | 0.39 | 2.84 | 1.49 | 181.74 | 66.09 | 140.08 |
| 4a | −6.62 | −1.48 | 5.14 | 4.05 | −4.05 | 2.57 | 0.39 | 3.19 | 1.57 | 207.44 | 77.38 | 161.14 |
| 4b | −6.57 | −1.85 | 4.72 | 4.21 | −4.21 | 2.36 | 0.42 | 3.75 | 1.78 | 242.67 | 90.54 | 176.14 |
| 4c | −6.51 | −1.39 | 5.12 | 3.95 | −3.95 | 2.56 | 0.39 | 3.05 | 1.54 | 261.56 | 94.98 | 183.47 |
| 4d | −6.51 | −1.80 | 4.71 | 4.15 | −4.15 | 2.35 | 0.42 | 3.66 | 1.76 | 237.54 | 94.28 | 181.43 |
| 4e | −6.67 | −1.86 | 4.81 | 4.26 | −4.26 | 2.40 | 0.41 | 3.78 | 1.77 | 256.45 | 98.66 | 188.39 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Milusheva, M.; Gledacheva, V.; Stefanova, I.; Feizi-Dehnayebi, M.; Mihaylova, R.; Nedialkov, P.; Cherneva, E.; Tumbarski, Y.; Tsoneva, S.; Todorova, M.; et al. Synthesis, Molecular Docking, and Biological Evaluation of Novel Anthranilic Acid Hybrid and Its Diamides as Antispasmodics. Int. J. Mol. Sci. 2023, 24, 13855. https://doi.org/10.3390/ijms241813855
Milusheva M, Gledacheva V, Stefanova I, Feizi-Dehnayebi M, Mihaylova R, Nedialkov P, Cherneva E, Tumbarski Y, Tsoneva S, Todorova M, et al. Synthesis, Molecular Docking, and Biological Evaluation of Novel Anthranilic Acid Hybrid and Its Diamides as Antispasmodics. International Journal of Molecular Sciences. 2023; 24(18):13855. https://doi.org/10.3390/ijms241813855
Chicago/Turabian StyleMilusheva, Miglena, Vera Gledacheva, Iliyana Stefanova, Mehran Feizi-Dehnayebi, Rositsa Mihaylova, Paraskev Nedialkov, Emiliya Cherneva, Yulian Tumbarski, Slava Tsoneva, Mina Todorova, and et al. 2023. "Synthesis, Molecular Docking, and Biological Evaluation of Novel Anthranilic Acid Hybrid and Its Diamides as Antispasmodics" International Journal of Molecular Sciences 24, no. 18: 13855. https://doi.org/10.3390/ijms241813855
APA StyleMilusheva, M., Gledacheva, V., Stefanova, I., Feizi-Dehnayebi, M., Mihaylova, R., Nedialkov, P., Cherneva, E., Tumbarski, Y., Tsoneva, S., Todorova, M., & Nikolova, S. (2023). Synthesis, Molecular Docking, and Biological Evaluation of Novel Anthranilic Acid Hybrid and Its Diamides as Antispasmodics. International Journal of Molecular Sciences, 24(18), 13855. https://doi.org/10.3390/ijms241813855












