Histone Extraction from Human Articular Cartilage for the Study of Epigenetic Regulation in Osteoarthritis
Abstract
:1. Introduction
1.1. The Role of Epigenetics in Osteoarthritis (OA)
1.2. Articular Cartilage: Structure and Composition
1.3. Lack of Articular-Specific Histone Extraction Methods
2. Methods and Results
2.1. Starting Material
2.2. Pulverization of Cryopreserved Articular Cartilage Samples
- Prior to pulverization, place the mortar and the pestle in a foil-covered box, pour liquid nitrogen into it and wait until the tools are cold.
- Introduce the cryogenic tube containing the sample in liquid nitrogen for approximately 5 min. Tissue must be completely frozen, as thawing will hamper its fragmentation and promote protein degradation.
- Take the precooled mortar out of the box and place on it a flat surface.
- Remove the sample from liquid nitrogen and set it in the centre of the mortar.
- Pulverize the tissue by firmly hitting the pestle against the mortar.
- Repeat the process until cartilage acquires a powdery consistency.
- Separate a part of the resulting material with a 5 mm lab spoon into a 1.5 mL microcentrifuge tube. This will be used to obtain the total protein extract of tissue samples.
- Transfer the resulting material to an ice-cold 15 mL tube. Although it is not necessary to use 15 mL tubes, we propose this size because it offers enough space to allow an appropriate homogenization, as the reagents will reach the totality of the sample when vortexing. From now on, all steps must be performed on ice unless otherwise specified.
2.3. Tissue Homogenization
- Add 2 mL of Buffer A to the 15 mL tube and vortex thoroughly.
- Incubate the sample on ice for 10 min (this will allow Buffer A to disrupt cell membranes).
- Vortex the sample, and centrifuge at 4000× g for 10 min at 4 °C. Transfer the supernatant containing the cytosolic fraction (cytosolic fraction 1, CF1) to a fresh 2 mL centrifuge tube and maintain it on ice.
- Due to the low cellularity and ECM abundance in articular cartilage, we recommend performing this homogenization step twice to obtain higher extraction yields. Again, transfer the supernatant containing the cytosolic fraction (cytosolic fraction 2, CF2) to a fresh 2 mL centrifuge tube and keep it on ice. The volume of Buffer A used to homogenize cartilage is considerable, so cytosolic proteins contained in CF1 and CF2 should be concentrated before using them in other assays. Amicon® Ultra-2 Centrifugal Filter Units for concentration and purification of biological solutions (Merck Millipore) can be used for this purpose following manufacturer’s instructions.
- Add 2 mL of Buffer B.
- Vortex the sample to obtain a viscous homogenate and incubate it on ice for 10 min.
- Vortex the sample, centrifuge at 4000× g for 10 min at 4 °C and discard the supernatant.
- Centrifuge at 4000× g for 10 min at 4 °C without adding any buffer to completely remove the remaining Buffer B that could be infiltrated between the tissue fragments of the pellet. Again, discard the supernatant. (Note: Do not expect to obtain a “conventional” pellet, since at this point rest of the ECM will remain, forming a granular pellet.)
2.4. Histone Extraction
- Add 1 mL of 1.2 M H2SO4 to solubilize histones and vortex the sample.
- Centrifuge at 5500× g for 15 min at 4 °C.
- Transfer the supernatant to a 2 mL microcentrifuge tube.
- Add 1 mL of cold acetone (−20 °C).
- Invert the tube a few times to determine whether a whitish precipitate has formed, meaning that histone precipitation is occurring. If the starting material is scarce, you may not see the reaction immediately. However, after an overnight incubation of the samples at −20 °C, a cloudy precipitate should be visible at the bottom of the tube.
- Centrifuge the samples at 10,000× g for 10 min at 4 °C and discard the supernatant.
- Subject the pellet to a second wash with 1 mL of cold acetone (−20 °C).
- Centrifuge at 10,000× g for 10 min at 4 °C and discard the supernatant.
- Let the pellet dry for ~30 min, checking that the acetone has been totally evaporated before resuspension. If not, carefully remove it by pipetting or clean the walls of the tube with tissue paper, avoiding touching the pellet.
- U/T buffer (25 mM ammonium bicarbonate, 6 M urea, 2 M thiourea) facilitates resuspension, so we recommend its use instead of dH2O. Add an appropriate volume of U/T buffer to resuspend the pellet, taking care not to introduce any air bubbles with the pipette. Scrape the wall of the tube with the pipette tip to detach the pellet and patiently dissolve it by vortexing. Verify that no clumps remain and that the whole precipitate has been dissolved.
- Protein concentration can be quantified with the Bradford assay. Histone extracts obtained with this protocol cannot be quantified with PierceTM BCA Protein Assay Kit (Thermo Fisher Scientific) due to the incompatibility with the U/T buffer.
- Store histone extracts at −80 °C or prepare aliquots containing equal amounts of protein in a 1:10 mixture of β-mercaptoethanol and 4× Laemmli Sample Buffer (Bio-Rad) for further procedures.
2.5. Total Protein Extract Obtention
- Add two zirconium oxide grinding balls to each 1.5 mL microcentrifuge tube.
- Add 200 µL of U/S buffer (6 M urea, 2% SDS).
- Vigorously vortex the samples for 2 h at RT.
- Sonicate the samples for 5 min at RT.
- Centrifuge at full speed for 1 min at RT.
- Transfer the supernatant to a fresh 1.5 mL microcentrifuge tube.
- Centrifuge at full speed for 15 min at RT.
- To ensure the complete removal of insoluble material, transfer the supernatant to a fresh 1.5 mL microcentrifuge tube and discard the pellet.
- Protein concentration can be quantified with the Bradford assay.
- Store total protein extracts at −20 °C or prepare aliquots containing equal amounts of protein in a 1:10 mixture of β-mercaptoethanol and 4× Laemmli Sample Buffer (Bio-Rad) for further procedures.
2.6. Electrophoretic Separation of Proteins on Polyacrylamide Gels
- Vortex protein samples before denaturation.
- Boil the samples for 5 min at 95 °C and immediately put them on ice.
- Place the polyacrylamide gels in a vertical electrophoresis system and fill the electrophoresis tank with fresh 10% running buffer (Trizma base, glycine, 20% SDS, dH2O).
- Vortex the samples again.
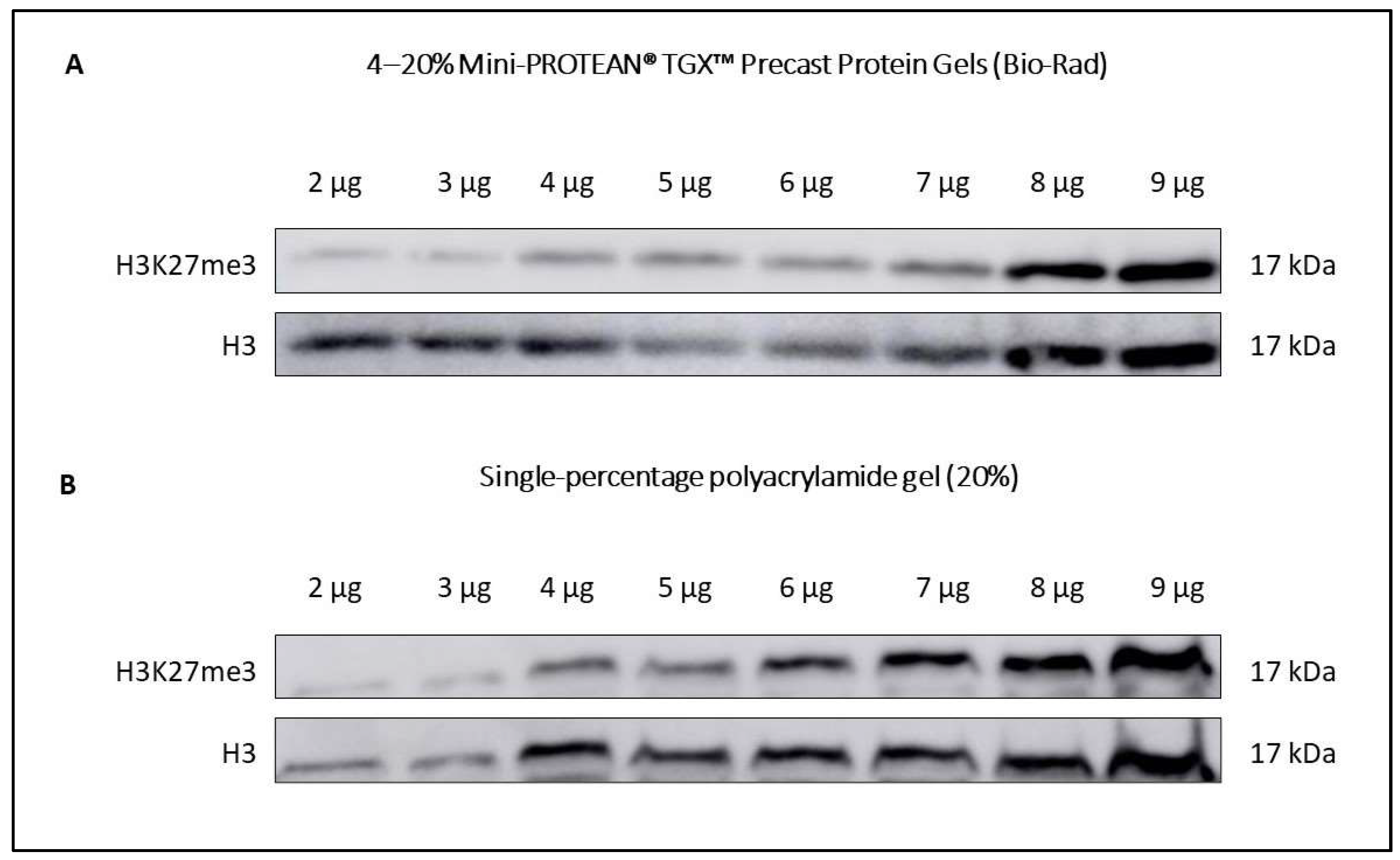
- Load equal sample volumes into the wells of the gel. You can prepare your own gels as long as its polyacrylamide percentage is set between 15–20%, since the molecular weight of histones requires a resolving gel with small pore size (see recipe in Table 4). For better results in Western blotting, we recommend using 4–20% Mini-PROTEAN® TGXTM Precast Protein Gels (Bio-Rad). A comparative example of both types of gels performance in Western blotting is shown in Figure 2.
- Run the SDS-PAGE electrophoresis at 70 V for 20 min or until proteins have reached the stacking gel limit. Then, raise the voltage to 100 V for 1 h 20 min. For Western blotting procedures, soak nitrocellulose membranes of 0.2 µm pore size in transfer buffer (Table 5) with constant shaking while the electrophoresis is running. Although 0.45 µm pore size membranes can be used (whether PVDF or nitrocellulose), we recommend using 0.2 µm nitrocellulose membranes because of their high capacity to retain lower molecular weight proteins. This is especially useful for the study of smaller histones such as H4.
2.7. Silver Staining
- Remove the gel from the cassette and soak it in 50 mL of fixative solution (40% ethanol, 10% glacial acetic acid). Fix the gel for 45 min at RT with smooth constant shaking.
- Wash the gel 2 times × 10 min with dH2O with smooth, constant shaking.
- Sensitize the gel in 0.02% sodium thiosulfate solution for 1 min.
- Wash the gel 2 times × 1 min with dH2O.
- Impregnate the gel with 0.2% silver nitrate and 0.075% formalin solution for 45 min with smooth, constant shaking.
- Wash the gel 2 times × 10 s with dH2O.
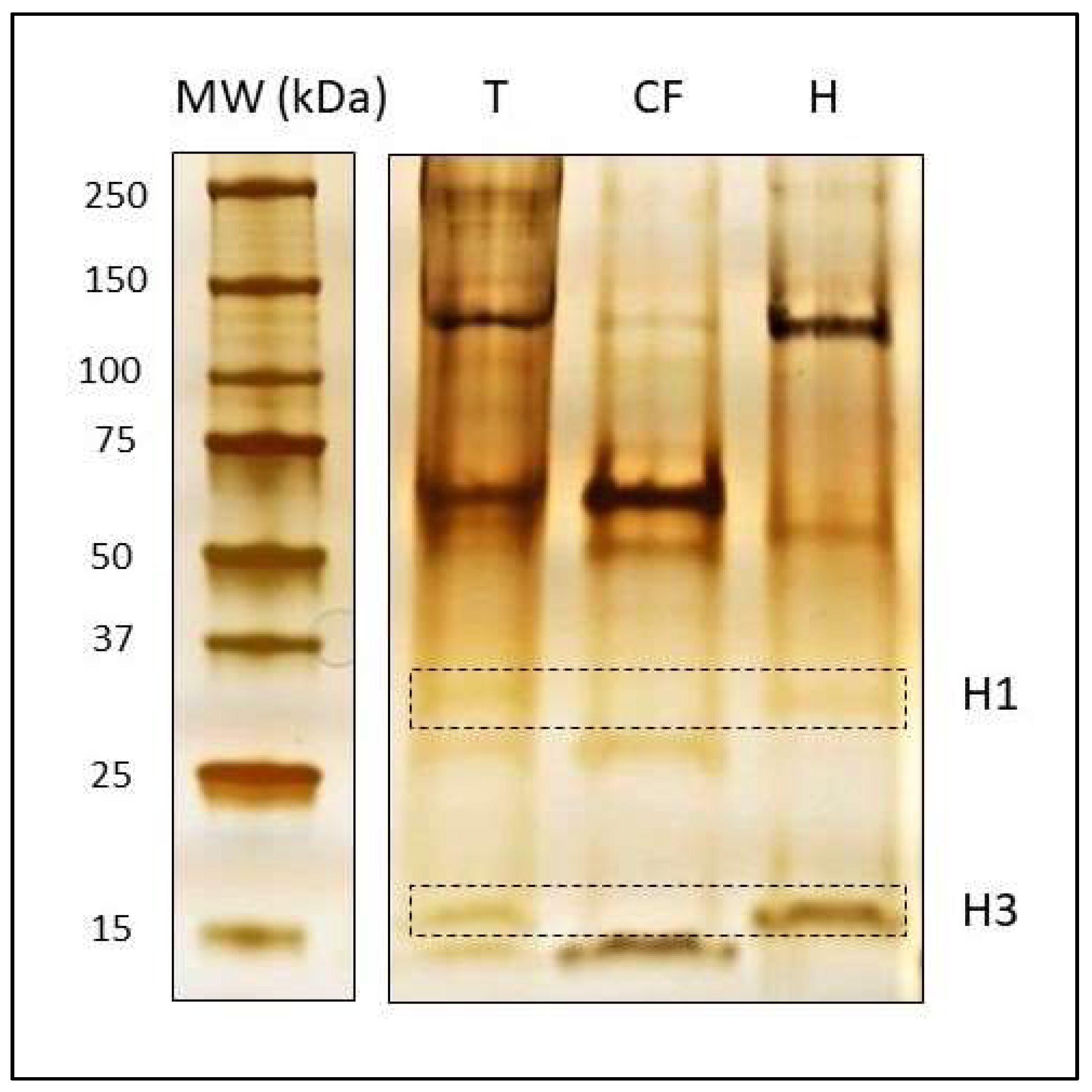
- Detect proteins in the gel by exposing the membrane to 3% sodium carbonate, 12.5 mg/L sodium thiosulfate and 0.025% formalin solution. Shake the gel vigorously for 3–10 min or until a brown precipitate is visible, as shown in Figure 3. Do not extend this step for more than 15 min.
- Stop the image development reaction by immersing the gel in 3% Trizma base and 10% glacial acetic acid solution for 30 min with smooth constant shaking.
- The gel can be conserved in dH2O at 4 °C.
2.8. Western Blot
- When the electrophoresis dye front has reached the bottom of the gel, proceed to transfer the gel to a nitrocellulose membrane using fresh cold transfer buffer (Table 5). Given that the transfer process generates a considerable amount of heat, place the tank into a box filled with ice. Additionally, a cold accumulator can be introduced into the tank next to the transfer cassette. Set the voltage to 100 V for 1 h 30 min.
- When the transfer has finished, immerse the membranes in 5% blocking solution (Table 5) for 1 h at RT with constant shaking.
- Incubate them with specific primary antibodies diluted in 5% blocking solution in a rotator at 4 °C overnight or as long as necessary.
- The following day, wash the membranes 3 times × 10 min at RT with 10% TBST (Table 5) in constant shaking.
- Incubate the membranes with a secondary antibody for 1 h at RT with constant shaking.
- Repeat the 10% TBST washes.
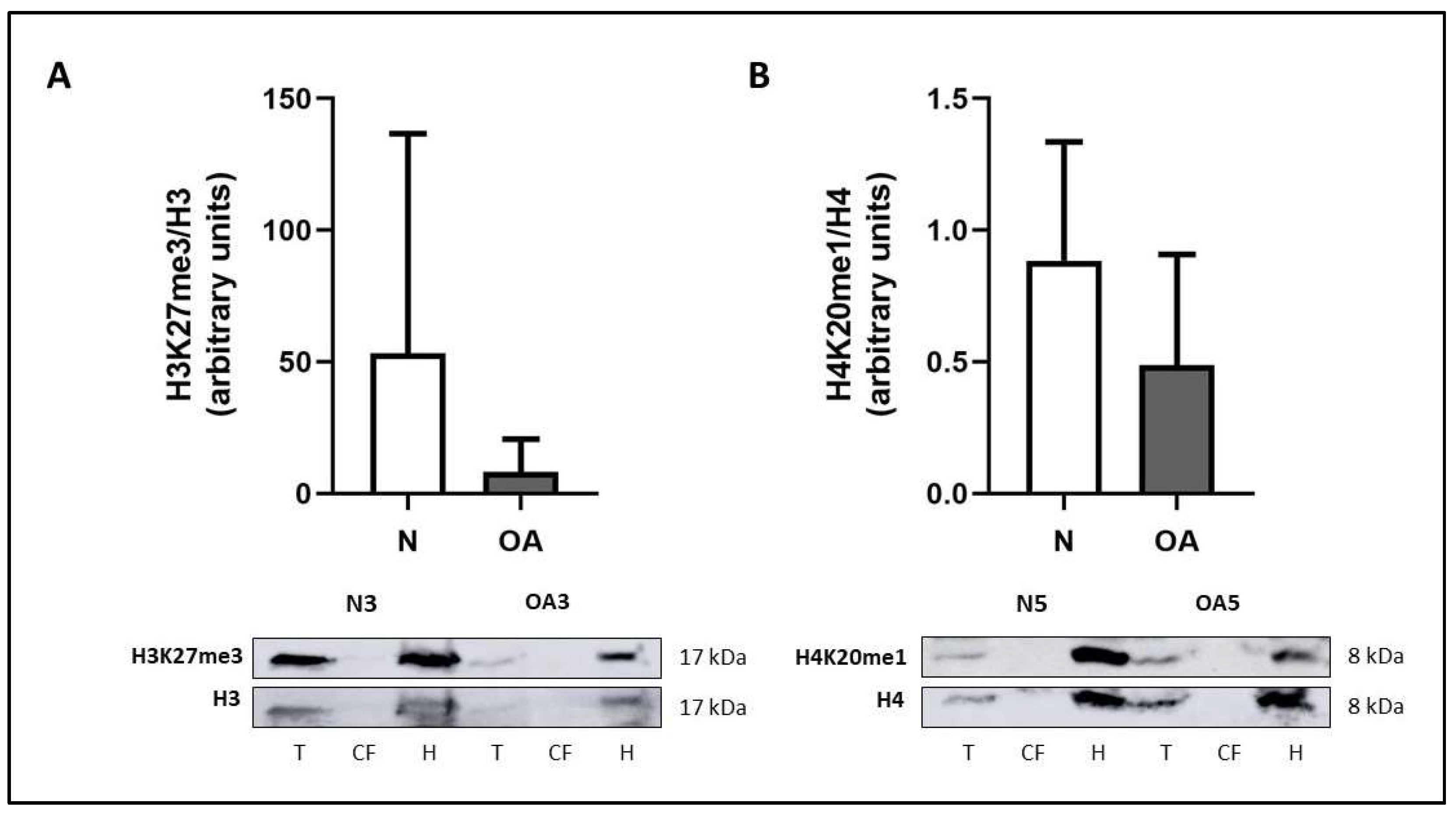
- Detection of the protein signal can be performed using an enhanced chemiluminescent (ECL) system, for instance, Immobilon Classico Western HRP substrate (Merck Millipore). An example with two histone modifications is shown in Figure 4.
2.9. Membrane Stripping
- Immerse the membranes in stripping solution (0.5 M Tris-HCl, 10% SDS, 100 mM β-mercaptoethanol, dH2O) for 45 min at 50 °C with constant shaking.
- Wash the membranes in dH2O for 2 h at RT with constant shaking. Change the dH2O regularly to remove all traces of stripping solution.
- Wash the membranes in 10% TBST for 30 min at RT with constant shaking. At this point, it is advisable to check if protein signal is completely removed.
- Soak the membranes in 5% blocking solution for 1 h at RT with constant shaking.
- Follow steps 3–7 from Section 2.8.
3. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Reynard, L.N.; Loughlin, J. Genetics and epigenetics of osteoarthritis. Maturitas 2012, 71, 200–204. [Google Scholar] [CrossRef] [PubMed]
- Cutter, A.R.; Hayes, J.J. A brief review of nucleosome structure. FEBS Lett. 2015, 589, 2914–2922. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gibney, E.R.; Nolan, C.M. Epigenetics and gene expression. Heredity 2010, 105, 4–13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- He, J.; Cao, W.; Azeem, I.; Shao, Z. Epigenetics of osteoarthritis: Histones and TGF-β1. Clin. Chim. Acta 2020, 510, 593–598. [Google Scholar] [CrossRef] [PubMed]
- Shen, J.; Abu-Amer, Y.; O’Keefe, R.J.; McAlinden, A. Inflammation and epigenetic regulation in osteoarthritis. Connect. Tissue Res. 2017, 58, 49–63. [Google Scholar] [CrossRef]
- Cross, M.; Smith, E.; Hoy, D.; Nolte, S.; Ackerman, I.; Fransen, M.; Bridgett, L.; Williams, S.; Guillemin, F.; Hill, C.L.; et al. The global burden of hip and knee osteoarthritis: Estimates from the Global Burden of Disease 2010 study. Ann. Rheum. Dis. 2014, 73, 1323–1330. [Google Scholar] [CrossRef] [PubMed]
- Goldring, S.R.; Goldring, M.B. Changes in the osteochondral unit during osteoarthritis: Structure, function and cartilage–bone crosstalk. Nat. Rev. Rheumatol. 2016, 12, 632–644. [Google Scholar] [CrossRef] [PubMed]
- Haseeb, A.; Haqqi, T.M. Immunopathogenesis of osteoarthritis. Clin. Immunol. 2013, 146, 185–196. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kapoor, M.; Martel-Pelletier, J.; Lajeunesse, D.; Pelletier, J.-P.; Fahmi, H. Role of proinflammatory cytokines in the pathophysiology of osteoarthritis. Nat. Rev. Rheumatol. 2011, 7, 33–42. [Google Scholar] [CrossRef]
- Chauffier, K.; Laiguillon, M.-C.; Bougault, C.; Gosset, M.; Priam, S.; Salvat, C.; Mladenovic, Z.; Nourissat, G.; Jacques, C.; Houard, X.; et al. Induction of the chemokine IL-8/Kc by the articular cartilage: Possible influence on osteoarthritis. Joint Bone Spine 2012, 79, 604–609. [Google Scholar] [CrossRef]
- Hashimoto, K.; Oreffo, R.O.C.; Gibson, M.B.; Goldring, M.B.; Roach, H.I. DNA demethylation at specific CpG sites in the IL1B promoter in response to inflammatory cytokines in human articular chondrocytes. Arthritis Rheum. 2009, 60, 3303–3313. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Takahashi, A.; de Andrés, M.C.; Hashimoto, K.; Itoi, E.; Oreffo, R.O.C. Epigenetic regulation of interleukin-8, an inflammatory chemokine, in osteoarthritis. Osteoarthritis Cartilage 2015, 23, 1946–1954. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Roach, H.I.; Yamada, N.; Cheung, K.S.C.; Tilley, S.; Clarke, N.M.P.; Oreffo, R.O.C.; Kokubun, S.; Bronner, F. Association between the abnormal expression of matrix-degrading enzymes by human osteoarthritic chondrocytes and demethylation of specific CpG sites in the promoter regions. Arthritis Rheum. 2005, 52, 3110–3124. [Google Scholar] [CrossRef] [PubMed]
- Cheung, K.S.C.; Hashimoto, K.; Yamada, N.; Roach, H.I. Expression of ADAMTS-4 by chondrocytes in the surface zone of human osteoarthritic cartilage is regulated by epigenetic DNA de-methylation. Rheumatol. Int. 2009, 29, 525–534. [Google Scholar] [CrossRef] [PubMed]
- de Andrés, M.C.; Imagawa, K.; Hashimoto, K.; Gonzalez, A.; Roach, H.I.; Goldring, M.B.; Oreffo, R.O.C. Loss of methylation in CpG sites in the NF-κB enhancer elements of inducible nitric oxide synthase is responsible for gene induction in human articular chondrocytes. Arthritis Rheum. 2013, 65, 732–742. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Imagawa, K.; de Andrés, M.C.; Hashimoto, K.; Itoi, E.; Otero, M.; Roach, H.I.; Goldring, M.B.; Oreffo, R.O.C. Association of Reduced Type IX Collagen Gene Expression in Human Osteoarthritic Chondrocytes With Epigenetic Silencing by DNA Hypermethylation: Collagen Expression and DNA Methylation in OA Chondrocytes. Arthritis Rheumatol. 2014, 66, 3040–3051. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Roach, H.I.; Aigner, T. DNA methylation in osteoarthritic chondrocytes: A new molecular target. Osteoarthritis Cartilage 2007, 15, 128–137. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fuks, F. DNA methylation and histone modifications: Teaming up to silence genes. Curr. Opin. Genet. Dev. 2005, 15, 490–495. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Jordan, J.M. Epidemiology of Osteoarthritis. Clin. Geriatr. Med. 2010, 26, 355–369. [Google Scholar] [CrossRef] [Green Version]
- Gabay, O.; Clouse, K.A. Epigenetics of cartilage diseases. Joint Bone Spine 2016, 83, 491–494. [Google Scholar] [CrossRef]
- Alford, J.W.; Cole, B.J. Cartilage Restoration, Part 1: Basic Science, Historical Perspective, Patient Evaluation, and Treatment Options. Am. J. Sports Med. 2005, 33, 295–306. [Google Scholar] [CrossRef] [PubMed]
- Buckwalter, J.A.; Mankin, H.J.; Grodzinsky, A.J. Articular Cartilage and Osteoarthritis. Instr. Course Lect. 2005, 54, 465–480. [Google Scholar] [PubMed]
- Sophia Fox, A.J.; Bedi, A.; Rodeo, S.A. The Basic Science of Articular Cartilage: Structure, Composition, and Function. Sports Health Multidiscip. Approach 2009, 1, 461–468. [Google Scholar] [CrossRef] [PubMed]
- Kiani, C.; Chen, L.; Wu, Y.J.; Yee, A.J.; Yang, B.B. Structure and function of aggrecan. Cell Res. 2002, 12, 19–32. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Carballo, C.B.; Nakagawa, Y.; Sekiya, I.; Rodeo, S.A. Basic Science of Articular Cartilage. Clin. Sports Med. 2017, 36, 413–425. [Google Scholar] [CrossRef] [PubMed]
- Ulrich-Vinther, M.; Maloney, M.D.; Schwarz, E.M.; Rosier, R.; O’Keefe, R.J. Orthopaedic Research Society Special Article. J. Am. Acad. Orthop. Surg. 2003, 11, 10. [Google Scholar]
- Kim, H.; Blanco, F. Cell Death and Apoptosis in Ostearthritic Cartilage. Curr. Drug Targets 2007, 8, 333–345. [Google Scholar] [CrossRef] [PubMed]
- Rivera-Casas, C.; Gonzalez-Romero, R.; Garduño, R.A.; Cheema, M.S.; Ausio, J.; Eirin-Lopez, J.M. Molecular and Biochemical Methods Useful for the Epigenetic Characterization of Chromatin-Associated Proteins in Bivalve Molluscs. Front. Physiol. 2017, 8, 490. [Google Scholar] [CrossRef] [Green Version]
- Rumbaugh, G.; Miller, C.A. Epigenetic Changes in the Brain: Measuring Global Histone Modifications. In Alzheimer’s Disease and Frontotemporal Dementia; Roberson, E.D., Ed.; Methods in Molecular Biology; Humana Press: Totowa, NJ, USA, 2010; Volume 670, pp. 263–274. ISBN 978-1-60761-743-3. [Google Scholar]
- Blum, H.; Beier, H.; Gross, H.J. Improved silver staining of plant proteins, RNA and DNA in polyacrylamide gels. Electrophoresis 1987, 8, 93–99. [Google Scholar] [CrossRef]

| ECM | Chondrocytes | |||||
|---|---|---|---|---|---|---|
| Zone | Function | Collagen | Proteoglycans | Shape | Abundance | |
| Fibres | Distribution | Abundance | ||||
| Superficial | Protecting the underlying layers and providing elasticity | Packed | Parallel | Low | Flattened | Relatively high |
| Middle | First defence line against compressive forces | Thick | Oblique | High | Spherical | Low |
| Deep | Resistance against compressive forces | Very thick | Perpendicular | Very high | Spherical, columnar arrangement | Relatively high |
| Calcified | Anchoring of collagen fibres to the bone | Anchored to the subchondral bone | - | Hypertrophied | ||
| Sample | Weight | Protein Yield | Quantification Method | Resuspension Solvent |
|---|---|---|---|---|
| OA1 | 1.20 g | 5.30 µg/µL | BCA | H2O |
| OA2 | 350 mg | 1.94 µg/µL | BCA | H2O |
| OA3 | 556 mg | 4.04 µg/µL | Bradford | U/T |
| OA4 | 797 mg | 2.86 µg/µL | Bradford | U/T |
| OA5 | 952 mg | 2.60 µg/µL | Bradford | U/T |
| N1 | 1.20 g | 2.39 µg/µL | BCA | H2O |
| N2 | 140 mg | 0.82 µg/µL | BCA | H2O |
| N3 | 1.64 g | 1.24 µg/µL | Bradford | U/T |
| N4 | 1.98 g | 1.29 µg/µL | Bradford | U/T |
| N5 | 1.28 g | 0.67 µg/µL | Bradford | U/T |
| Buffer A | Buffer B |
|---|---|
| 0.15 M NaCl | 0.1 M KCl |
| 10 mM Tris-HCl pH = 7.5 | 50 mM Tris-HCl pH = 7.5 |
| 0.5% Triton X-100 | 1 mM MgCl2 |
| 2 mM sodium butyrate | |
| 1 mM sodium orthovanadate | |
| 0.5 mM PMSF | |
| 1× protease inhibitor cocktail (Sigma) | |
| Stacking Gel | Running Gel | |||
|---|---|---|---|---|
| 4% | 15% | 20% | ||
| dH2O | 4.8 mL | 4.7 mL | 1.4 mL | |
| 0.5 M Tris-HCl pH = 6.8 | 2.0 mL | – | – | |
| 1.5 M Tris-HCl pH = 8.8 | – | 5.0 mL | 5.0 mL | |
| 30% Acrylamide/Bis | 1.1 mL | 10 mL | 13.3 mL | |
| 10% Sodium dodecyl sulfate (SDS) | 80 µL | 200 µL | 200 µL | |
| 10% Ammonium persulfate (APS) | 40 µL | 100 µL | 100 µL | |
| Tetramethylethylenediamine (TEMED) | 8.0 µL | 10 µL | 10 µL | |
| Reagent | Composition | Storage |
|---|---|---|
| Transfer buffer | 100% running buffer | 4 °C until use |
| 100% ethanol | ||
| dH2O | ||
| 100% wash buffer (TBST) | Trizma base | RT |
| NaCl | ||
| Tween-20 | ||
| dH2O | ||
| 5% blocking solution | Bovine Serum Albumin (BSA) | 4 °C |
| 10% TBST |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Núñez-Carro, C.; Blanco-Blanco, M.; Montoya, T.; Villagrán-Andrade, K.M.; Hermida-Gómez, T.; Blanco, F.J.; de Andrés, M.C. Histone Extraction from Human Articular Cartilage for the Study of Epigenetic Regulation in Osteoarthritis. Int. J. Mol. Sci. 2022, 23, 3355. https://doi.org/10.3390/ijms23063355
Núñez-Carro C, Blanco-Blanco M, Montoya T, Villagrán-Andrade KM, Hermida-Gómez T, Blanco FJ, de Andrés MC. Histone Extraction from Human Articular Cartilage for the Study of Epigenetic Regulation in Osteoarthritis. International Journal of Molecular Sciences. 2022; 23(6):3355. https://doi.org/10.3390/ijms23063355
Chicago/Turabian StyleNúñez-Carro, Carmen, Margarita Blanco-Blanco, Tatiana Montoya, Karla M. Villagrán-Andrade, Tamara Hermida-Gómez, Francisco J. Blanco, and María C. de Andrés. 2022. "Histone Extraction from Human Articular Cartilage for the Study of Epigenetic Regulation in Osteoarthritis" International Journal of Molecular Sciences 23, no. 6: 3355. https://doi.org/10.3390/ijms23063355
APA StyleNúñez-Carro, C., Blanco-Blanco, M., Montoya, T., Villagrán-Andrade, K. M., Hermida-Gómez, T., Blanco, F. J., & de Andrés, M. C. (2022). Histone Extraction from Human Articular Cartilage for the Study of Epigenetic Regulation in Osteoarthritis. International Journal of Molecular Sciences, 23(6), 3355. https://doi.org/10.3390/ijms23063355






