Dietary AhR Ligands Regulate AhRR Expression in Intestinal Immune Cells and Intestinal Microbiota Composition
Abstract
1. Introduction
2. Results
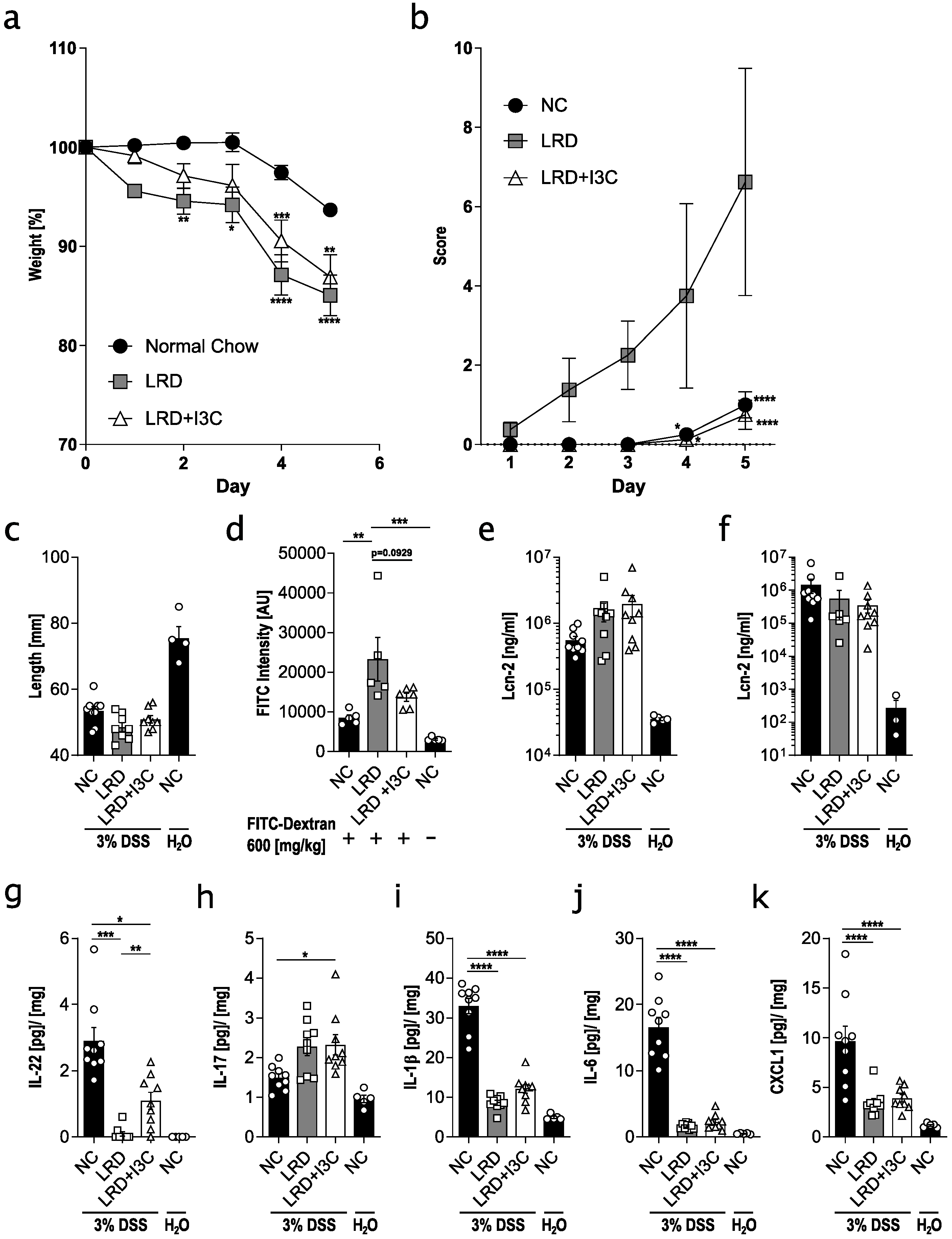
2.1. Dietary AhR Ligands Strengthen Intestinal Barrier Integrity and Lower Susceptibility to Colitis
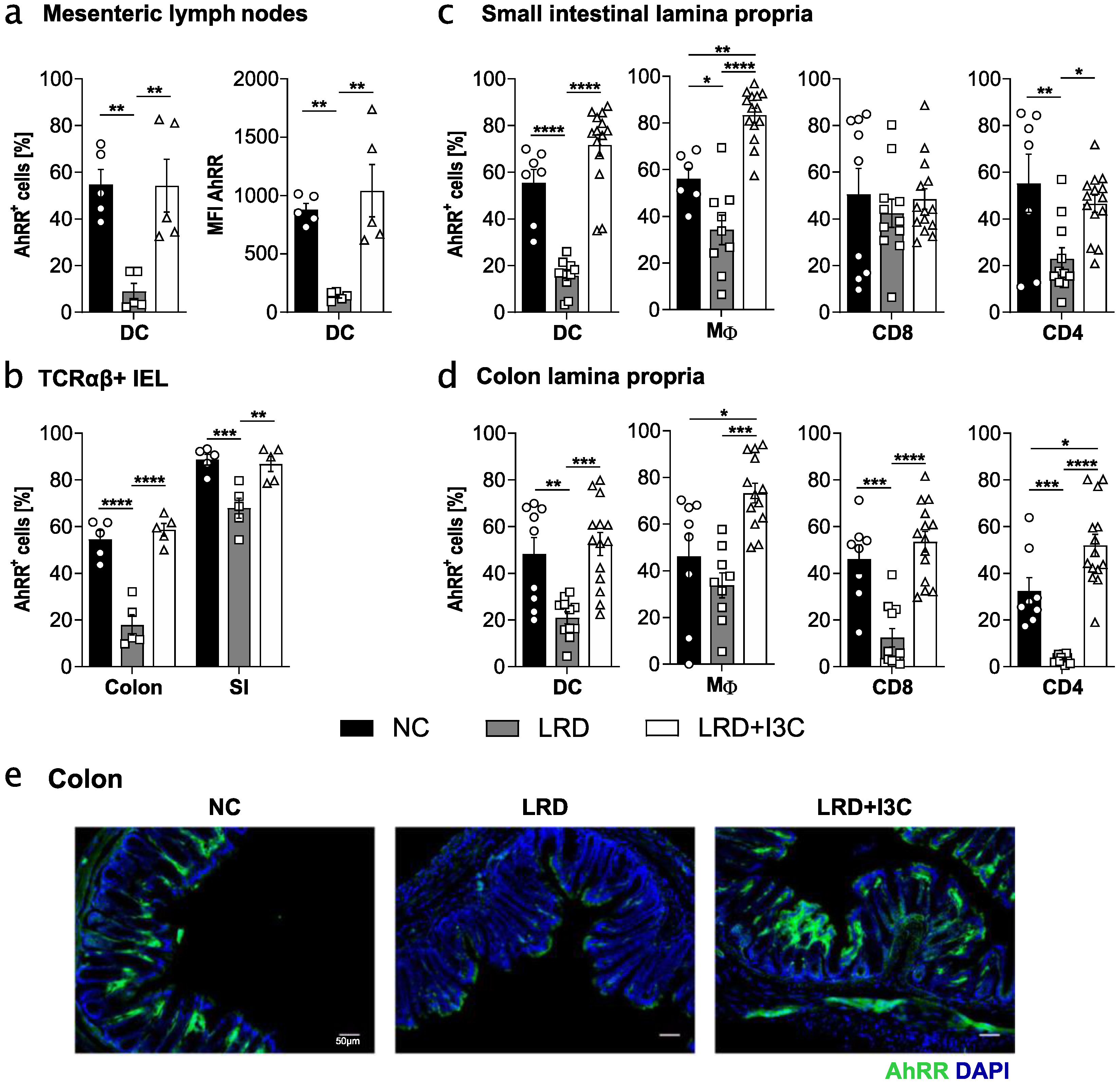
2.2. Dietary AhR Ligands Drive Expression of the AhRR in Intestinal Immune Cells
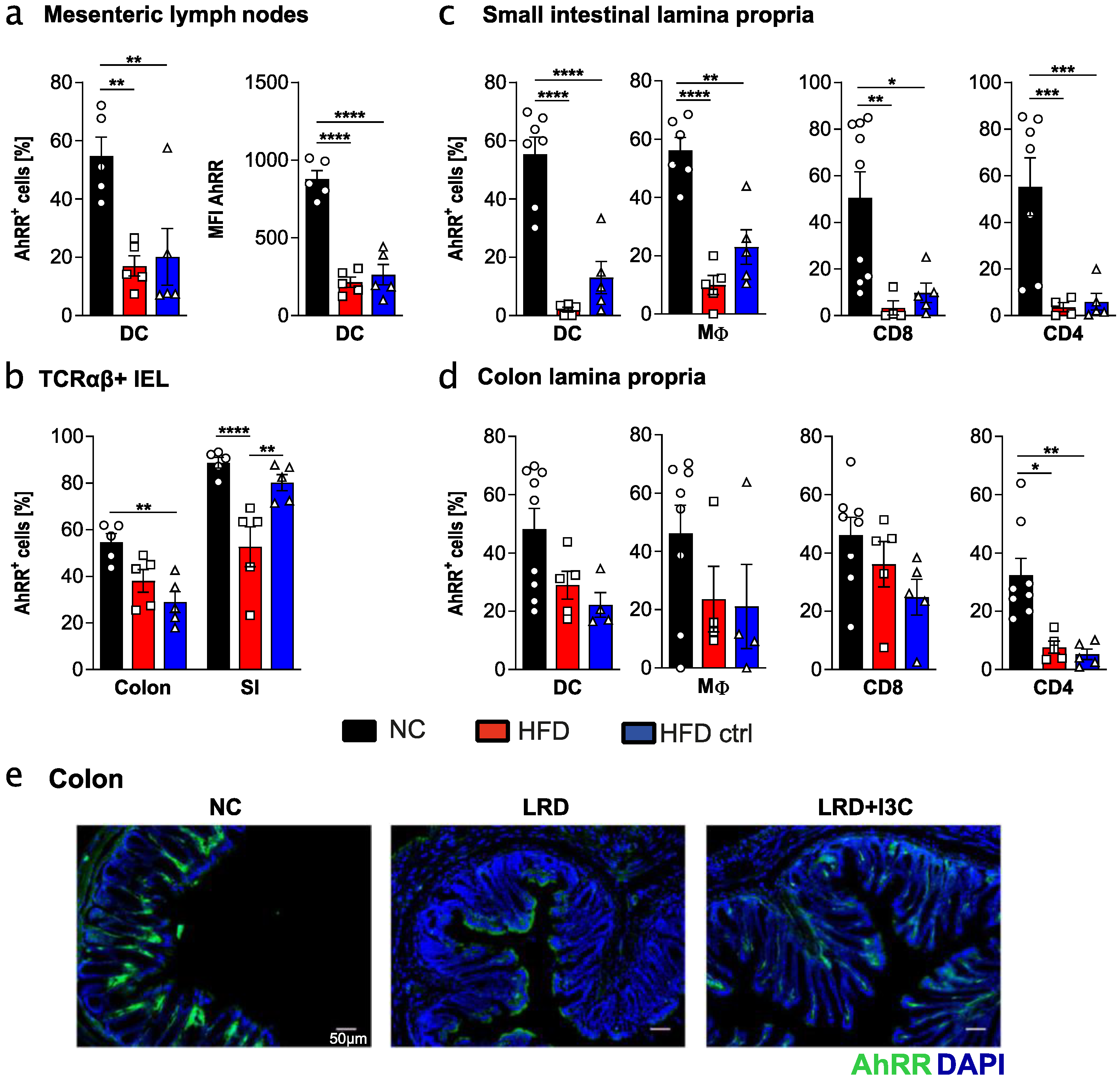
2.3. High Fat Diet and Matching Control Diet Reduce Expression of the AhRR in Intestinal Immune Cells
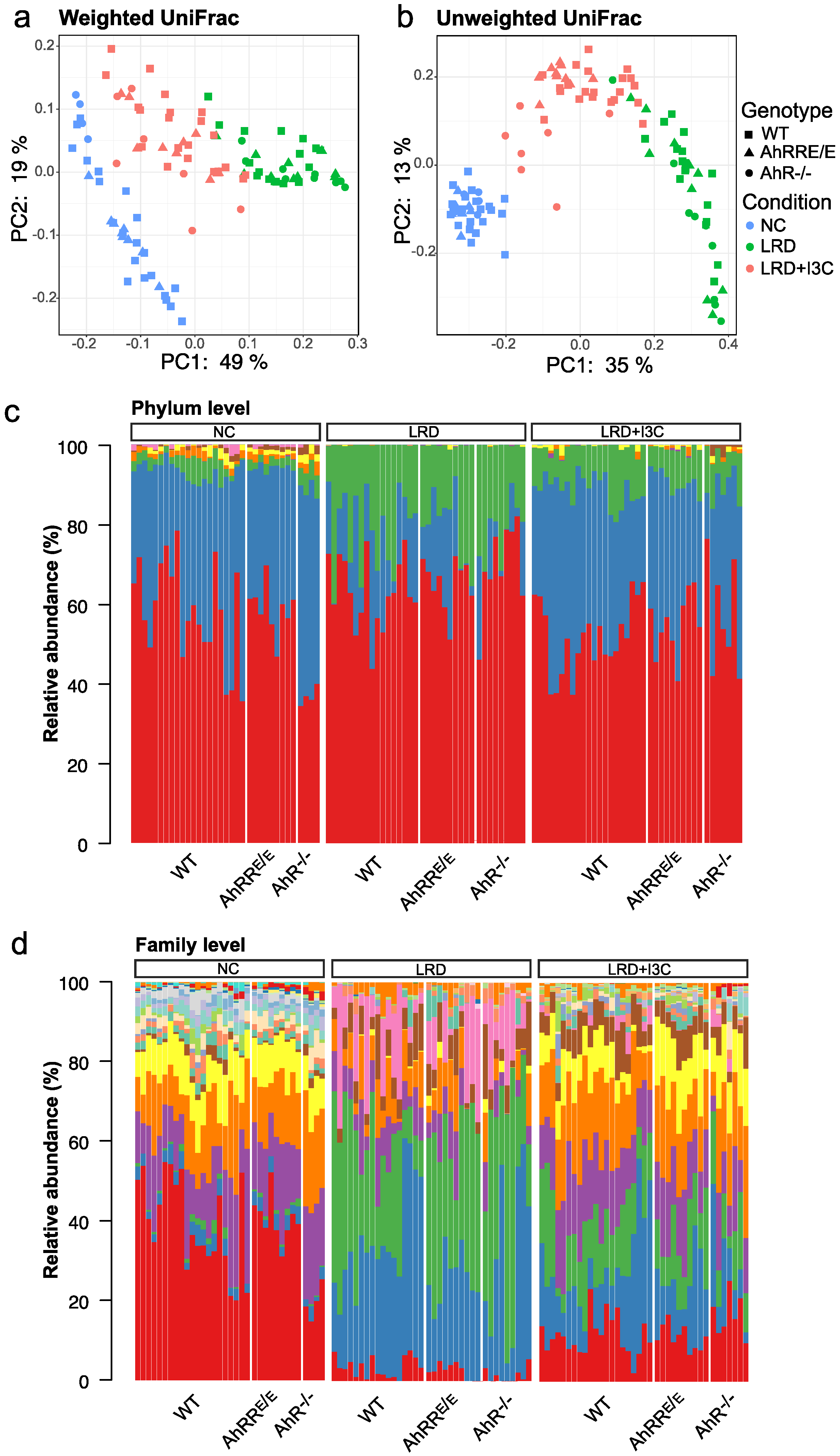
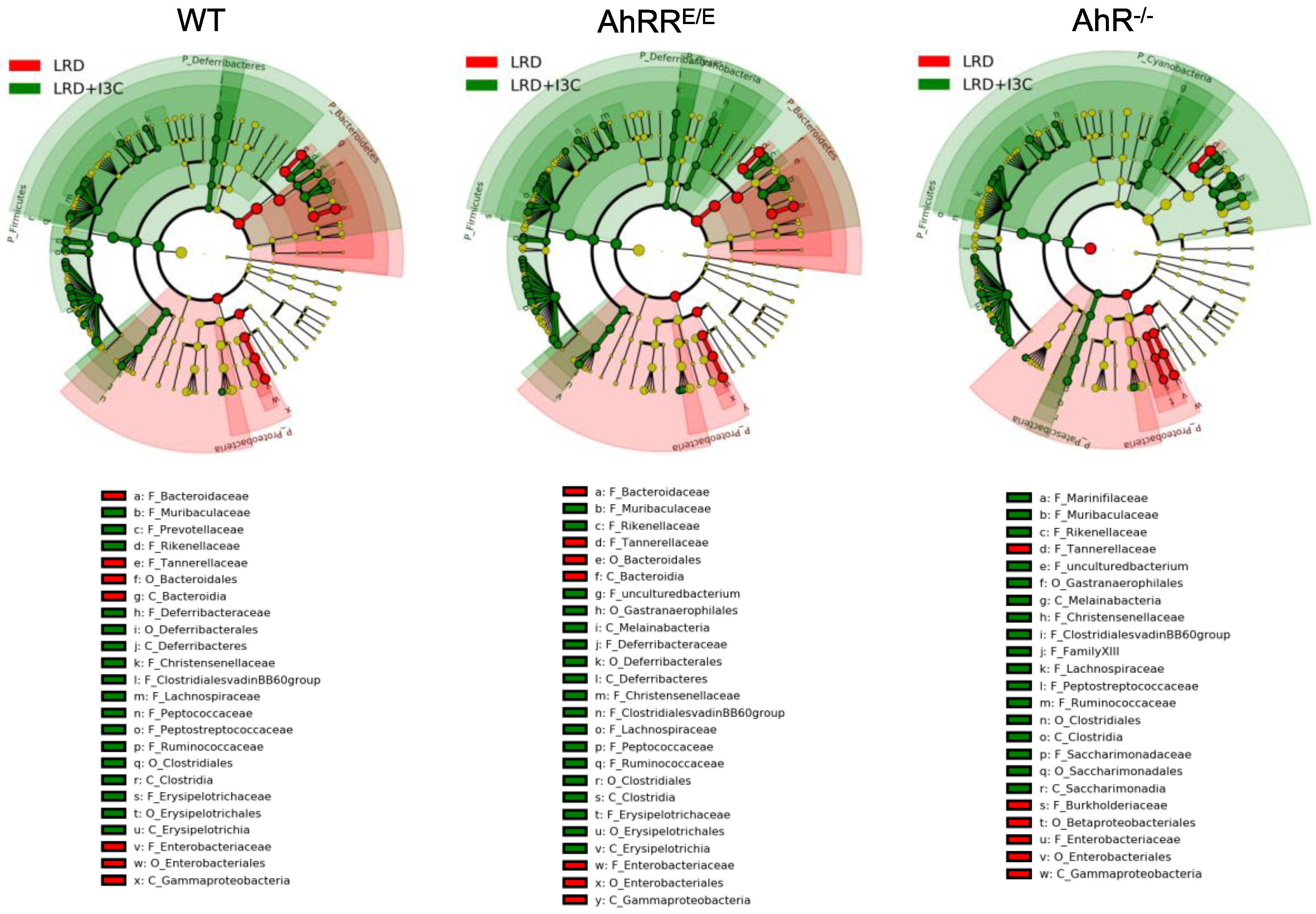
2.4. Influence of Dietary AhR Ligands on the Composition of the Intestinal Microbiota
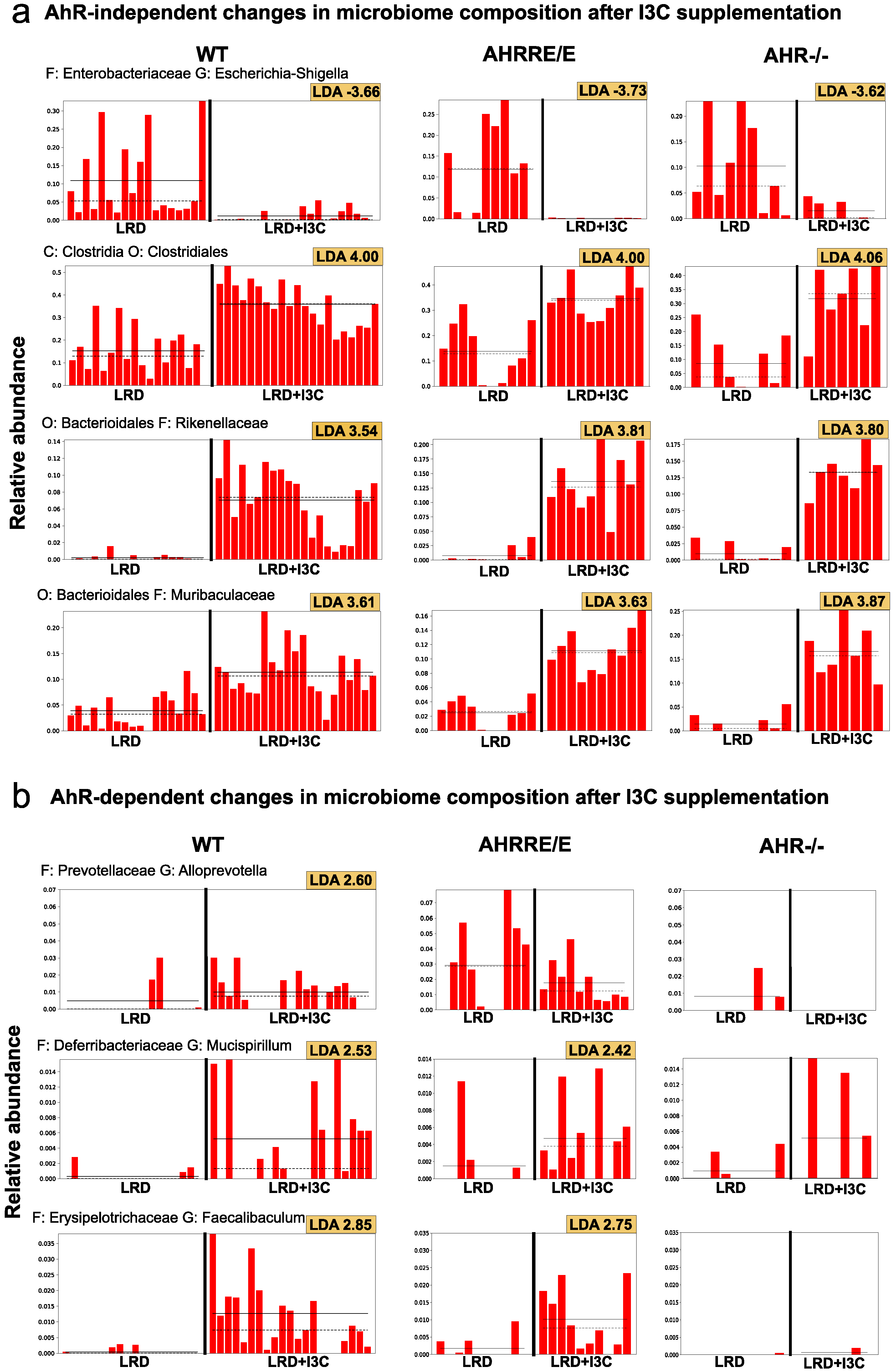
2.4.1. I3C-Induced Changes in the Representation of Bacterial Phyla and Families Are Mostly AhR-Independent
2.4.2. LRD + I3C Also Alters the Frequency of Bacterial Taxa in an AhR-Dependent Manner
3. Discussion
4. Materials and Methods
4.1. Animals
4.2. Feeding of Experimental Diets
4.3. Oral Antibiotic Treatment
4.4. Cytokine Determination in Colon Tissue
4.5. Determination of Intestinal Permeability
4.6. Immunohistology
4.7. Isolation of Intestinal Immune Cell Subsets
4.8. Flow Cytometry
4.9. Real-Time PCR Analysis
4.10. 16S rRNA Gene Data Collection and Sequencing
4.11. Microbiome Sequencing Analysis
4.12. Statistical Analysis
4.13. Data Availability
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| AhR | Aryl hydrocarbon receptor |
| AhRR | Aryl hydrocarbon receptor repressor |
| ARNT | AhR nuclear translocator |
| CFU | Colony forming units |
| DC | Dendritic cell |
| DIM | 3,3′-Diindolylmethane |
| DSS | Dextran sodium sulfate |
| EDTA | Ethylenediaminetetraacetic acid |
| EGFPFCS | Enhanced Green Fluorescent ProteinFetal calf serum |
| FITC | Fluorescein isothiocyanate |
| HBSS | Hank’s balanced salt solution |
| HEPES | 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid |
| HFD | High fat diet |
| HFD ctrl | Matching control diet |
| I3C | Indole-3-carbinol |
| IBD | Inflammatory bowel disease |
| ICZ | indolo[3,2-b]carbazole |
| IEC | Intestinal epithelial cell |
| IEL | Intraepithelial lymphocyte |
| IL | Interleukin |
| ILC | Innate lymphoid cell |
| LCN2 | Lipocalin-2 |
| LDA | Linear discriminant analysis |
| LEfSe | LDA coupled with effect size measurements |
| LPL | Lamina propria lymphocytes |
| LRD | Ligand-reduced diet |
| MFI | Mean fluorescent intensity |
| mLN | Mesenteric lymph node |
| NC | Normal chow |
| PAS-B | Period/ARNT/Single-minded |
| PBS | Phosphate-Buffered Saline |
| PCA | Principal component analysis |
| SI | Small intestine |
| TCR | T cell receptor |
| WT | Wild-type |
References
- Esser, C.; Rannug, A. The aryl hydrocarbon receptor in barrier organ physiology, immunology, and toxicology. Pharmacol. Rev. 2015, 67, 259–279. [Google Scholar] [CrossRef] [PubMed]
- Bernshausen, T.; Jux, B.; Esser, C.; Abel, J.; Fritsche, E. Tissue distribution and function of the Aryl hydrocarbon receptor repressor (AhRR) in C57BL/6 and Aryl hydrocarbon receptor deficient mice. Arch. Toxicol. 2006, 80, 206–211. [Google Scholar] [CrossRef] [PubMed]
- Qiu, J.; Guo, X.; Chen, Z.-M.E.; He, L.; Sonnenberg, G.F.; Artis, D.; Fu, Y.-X.; Zhou, L. Group 3 Innate Lymphoid Cells Inhibit T-Cell-Mediated Intestinal Inflammation through Aryl Hydrocarbon Receptor Signaling and Regulation of Microflora. Immunity 2013, 39, 386–399. [Google Scholar] [CrossRef] [PubMed]
- Kiss, E.A.; Vonarbourg, C.; Kopfmann, S.; Hobeika, E.; Finke, D.; Esser, C.; Diefenbach, A. Natural aryl hydrocarbon receptor ligands control organogenesis of intestinal lymphoid follicles. Science 2011, 334, 1561–1565. [Google Scholar] [CrossRef]
- Zelante, T.; Iannitti, R.G.; Cunha, C.; De Luca, A.; Giovannini, G.; Pieraccini, G.; Zecchi, R.; D’Angelo, C.; Massi-Benedetti, C.; Fallarino, F.; et al. Tryptophan catabolites from microbiota engage aryl hydrocarbon receptor and balance mucosal reactivity via interleukin-22. Immunity 2013, 39, 372–385. [Google Scholar] [CrossRef]
- Li, Y.; Innocentin, S.; Withers, D.R.; Roberts, N.A.; Gallagher, A.R.; Grigorieva, E.F.; Wilhelm, C.; Veldhoen, M. Exogenous Stimuli Maintain Intraepithelial Lymphocytes via Aryl Hydrocarbon Receptor Activation. Cell 2011, 147, 629–640. [Google Scholar] [CrossRef]
- Lee, J.S.; Cella, M.; McDonald, K.G.; Garlanda, C.; Kennedy, G.D.; Nukaya, M.; Mantovani, A.; Kopan, R.; Bradfield, C.A.; Newberry, R.D.; et al. AHR drives the development of gut ILC22 cells and postnatal lymphoid tissues via pathways dependent on and independent of Notch. Nat. Immunol. 2012, 13, 144–151. [Google Scholar] [CrossRef]
- Schiering, C.; Wincent, E.; Metidji, A.; Iseppon, A.; Li, Y.; Potocnik, A.J.; Omenetti, S.; Henderson, C.J.; Wolf, C.R.; Nebert, D.W.; et al. Feedback control of AHR signalling regulates intestinal immunity. Nature 2017, 542, 242–245. [Google Scholar] [CrossRef]
- Singh, N.P.; Singh, U.P.; Rouse, M.; Zhang, J.; Chatterjee, S.; Nagarkatti, P.S.; Nagarkatti, M. Dietary Indoles Suppress Delayed-Type Hypersensitivity by Inducing a Switch from Proinflammatory Th17 Cells to Anti-Inflammatory Regulatory T Cells through Regulation of MicroRNA. J. Immunol. 2016, 196, 1108–1122. [Google Scholar] [CrossRef]
- Chinen, I.; Nakahama, T.; Kimura, A.; Nguyen, N.T.; Takemori, H.; Kumagai, A.; Kayama, H.; Takeda, K.; Lee, S.; Hanieh, H.; et al. The aryl hydrocarbon receptor/microRNA-212/132 axis in T cells regulates IL-10 production to maintain intestinal homeostasis. Int. Immunol. 2015, 27, 405–415. [Google Scholar] [CrossRef]
- Lamas, B.; Natividad, J.M.; Sokol, H. Aryl hydrocarbon receptor and intestinal immunity review-article. Mucosal Immunol. 2018, 11, 1024–1038. [Google Scholar] [CrossRef] [PubMed]
- Monteleone, I.; Rizzo, A.; Sarra, M.; Sica, G.; Sileri, P.; Biancone, L.; MacDonald, T.T.; Pallone, F.; Monteleone, G. Aryl Hydrocarbon Receptor-Induced Signals Up-regulate IL-22 Production and Inhibit Inflammation in the Gastrointestinal Tract. Gastroenterology 2011, 141, 237–248.e1. [Google Scholar] [CrossRef] [PubMed]
- Haarmann-Stemmann, T.; Bothe, H.; Kohli, A.; Sydlik, U.; Abel, J.; Fritsche, E. Analysis of the transcriptional regulation and molecular function of the aryl hydrocarbon receptor repressor in human cell lines. Drug Metab. Dispos. 2007, 35, 2262–2269. [Google Scholar] [CrossRef] [PubMed]
- Baba, T.; Mimura, J.; Gradin, K.; Kuroiwa, A.; Watanabe, T.; Matsuda, Y.; Inazawa, J.; Sogawa, K.; Fujii-Kuriyama, Y. Structure and Expression of the Ah Receptor Repressor Gene. J. Biol. Chem. 2001, 276, 33101–33110. [Google Scholar] [CrossRef] [PubMed]
- Cauchi, S.; Stücker, I.; Cénée, S.; Kremers, P.; Beaune, P.; Massaad-Massade, L. Structure and Polymorphisms of Human Aryl Hydrocarbon Receptor Repressor (AhRR) Gene in a French Population: Relationship With CYP1A1 Inducibility and Lung Cancer. Pharmacogenetics 2003, 13, 339–347. [Google Scholar] [CrossRef]
- Mimura, J.; Ema, M.; Sogawa, K.; Fujii-Kuriyama, Y. Identification of a novel mechanism of regulation of Ah (dioxin) receptor function. Genes Dev. 1999, 13, 20–25. [Google Scholar] [CrossRef]
- Vogel, C.F.A.; Ishihara, Y.; Campbell, C.E.; Kado, S.Y.; Nguyen-Chi, A.; Sweeney, C.; Pollet, M.; Stemmann, T.H.; Tuscano, J.M. A protective role of aryl hydrocarbon receptor repressor in inflammation and tumor growth. Cancers (Basel) 2019, 11, 589. [Google Scholar] [CrossRef]
- Yamamoto, J.; Ihara, K.; Nakayama, H.; Hikino, S.; Satoh, K.; Kubo, N.; Iida, T.; Fujii, Y.; Hara, T. Characteristic expression of aryl hydrocarbon receptor repressor gene in human tissues: Organ-specific distribution and variable induction patterns in mononuclear cells. Life Sci. 2004, 74, 1039–1049. [Google Scholar] [CrossRef]
- Frericks, M.; Meissner, M.; Esser, C. Microarray analysis of the AHR system: Tissue-specific flexibility in signal and target genes. Toxicol. Appl. Pharmacol. 2007, 220, 320–332. [Google Scholar] [CrossRef]
- Tigges, J.; Weighardt, H.; Wolff, S.; Götz, C.; Förster, I.; Kohne, Z.; Huebenthal, U.; Merk, H.F.; Abel, J.; Haarmann-Stemmann, T.; et al. Aryl hydrocarbon receptor repressor (AhRR) function revisited: Repression of CYP1 activity in human skin fibroblasts is not related to AhRR expression. J. Investig. Dermatol. 2013, 133, 87–96. [Google Scholar] [CrossRef]
- Brandstätter, O.; Schanz, O.; Vorac, J.; König, J.; Mori, T.; Maruyama, T.; Korkowski, M.; Haarmann-Stemmann, T.; Von Smolinski, D.; Schultze, J.L.; et al. Balancing intestinal and systemic inflammation through cell type-specific expression of the aryl hydrocarbon receptor repressor. Sci. Rep. 2016, 6, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Busbee, P.B.; Menzel, L.; Alrafas, H.R.; Dopkins, N.; Becker, W.; Miranda, K.; Tang, C.; Chatterjee, S.; Singh, U.P.; Nagarkatti, M.; et al. Indole-3-carbinol prevents colitis and associated microbial dysbiosis in an IL-22–dependent manner. JCI Insight 2020, 5, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Hubbard, T.D.; Murray, I.A.; Nichols, R.G.; Cassel, K.; Podolsky, M.; Kuzu, G.; Tian, Y.; Smith, P.; Kennett, M.J.; Patterson, A.D.; et al. Dietary broccoli impacts microbial community structure and attenuates chemically induced colitis in mice in an Ah receptor dependent manner. J. Funct. Foods 2017, 37, 685–698. [Google Scholar] [CrossRef] [PubMed]
- Moura-Alves, P.; Faé, K.; Houthuys, E.; Dorhoi, A.; Kreuchwig, A.; Furkert, J.; Barison, N.; Diehl, A.; Munder, A.; Constant, P.; et al. AhR sensing of bacterial pigments regulates antibacterial defence. Nature 2014, 512, 387–392. [Google Scholar] [CrossRef]
- Agus, A.; Planchais, J.; Sokol, H. Gut Microbiota Regulation of Tryptophan Metabolism in Health and Disease. Cell Host Microbe 2018, 23, 716–724. [Google Scholar] [CrossRef]
- Takamura, T.; Harama, D.; Fukumoto, S.; Nakamura, Y.; Shimokawa, N.; Ishimaru, K.; Ikegami, S.; Makino, S.; Kitamura, M.; Nakao, A. Lactobacillus bulgaricus OLL1181 activates the aryl hydrocarbon receptor pathway and inhibits colitis. Immunol. Cell Biol. 2011, 89, 817–822. [Google Scholar] [CrossRef]
- Cervantes-Barragan, L.; Chai, J.N.; Tianero, M.D.; Di Luccia, B.; Ahern, P.P.; Merriman, J.; Cortez, V.S.; Caparon, M.G.; Donia, M.S.; Gilfillan, S.; et al. Lactobacillus reuteri induces gut intraepithelial CD4+CD8αα+ T cells. Science 2017, 357, 806–810. [Google Scholar] [CrossRef]
- Stallhofer, J.; Friedrich, M.; Konrad-Zerna, A.; Wetzke, M.; Lohse, P.; Glas, J.; Tillack-Schreiber, C.; Schnitzler, F.; Beigel, F.; Brand, S. Lipocalin-2 Is a Disease Activity Marker in Inflammatory Bowel Disease Regulated by IL-17A, IL-22, and TNF-α and Modulated by IL23R Genotype Status. Inflamm. Bowel Dis. 2015, 21, 2327–2340. [Google Scholar] [CrossRef]
- Alex, P.; Zachos, N.C.; Nguyen, T.; Gonzales, L.; Chen, T.-E.; Conklin, L.S.; Centola, M.; Li, X. Distinct cytokine patterns identified from multiplex profiles of murine DSS and TNBS-induced colitis. Inflamm. Bowel Dis. 2009, 15, 341–352. [Google Scholar] [CrossRef]
- Qiu, J.; Heller, J.J.; Guo, X.; Chen, Z.-M.E.; Fish, K.; Fu, Y.-X.; Zhou, L. The aryl hydrocarbon receptor regulates gut immunity through modulation of innate lymphoid cells. Immunity 2012, 36, 92–104. [Google Scholar] [CrossRef]
- Reeves, P.G.; Nielsen, F.H.; Fahey, G.C. AIN-93 purified diets for laboratory rodents: Final report of the American Institute of Nutrition ad hoc writing committee on the reformulation of the AIN-76A rodent diet. J. Nutr. 1993, 123, 1939–1951. [Google Scholar] [CrossRef] [PubMed]
- Warden, C.H.; Fisler, J.S. Comparisons of Diets Used in Animal Models of High-Fat Feeding. Cell Metab. 2008, 7, 277. [Google Scholar] [CrossRef] [PubMed]
- Segata, N.; Izard, J.; Waldron, L.; Gevers, D.; Miropolsky, L.; Garrett, W.S.; Huttenhower, C. Metagenomic biomarker discovery and explanation. Genome Biol. 2011, 12, R60. [Google Scholar] [CrossRef] [PubMed]
- Lagkouvardos, I.; Lesker, T.R.; Hitch, T.C.A.; Gálvez, E.J.C.; Smit, N.; Neuhaus, K.; Wang, J.; Baines, J.F.; Abt, B.; Stecher, B.; et al. Sequence and cultivation study of Muribaculaceae reveals novel species, host preference, and functional potential of this yet undescribed family. Microbiome 2019, 7, 28. [Google Scholar] [CrossRef]
- Kostic, A.D.; Xavier, R.J.; Gevers, D. The microbiome in inflammatory bowel disease: Current status and the future ahead. Gastroenterology 2014, 146, 1489–1499. [Google Scholar] [CrossRef]
- Korecka, A.; Dona, A.; Lahiri, S.; Tett, A.J.; Al-Asmakh, M.; Braniste, V.; D’Arienzo, R.; Abbaspour, A.; Reichardt, N.; Fujii-Kuriyama, Y.; et al. Bidirectional communication between the Aryl hydrocarbon Receptor (AhR) and the microbiome tunes host metabolism. NPJ Biofilms Microbiomes 2016, 2, 16014. [Google Scholar] [CrossRef]
- Brawner, K.M.; Yeramilli, V.A.; Duck, L.W.; Van Der Pol, W.; Smythies, L.E.; Morrow, C.D.; Elson, C.O.; Martin, C.A. Depletion of dietary aryl hydrocarbon receptor ligands alters microbiota composition and function. Sci. Rep. 2019, 9, 1–12. [Google Scholar] [CrossRef]
- Poland, A.; Knutson, J.C. 2,3,7,8-Tetrachlorodibenzo- p -Dioxin and Related Halogenated Aromatic Hydrocarbons: Examination of the Mechanism of Toxicity. Annu. Rev. Pharmacol. Toxicol. 1982, 22, 517–554. [Google Scholar] [CrossRef]
- Landers, J.P.; Bunce, N.J. The Ah receptor and the mechanism of dioxin toxicity. Biochem. J. 1991, 276, 273–287. [Google Scholar] [CrossRef]
- Lin, Y.H.; Luck, H.; Khan, S.; Schneeberger, P.H.H.; Tsai, S.; Clemente-Casares, X.; Lei, H.; Leu, Y.L.; Chan, Y.T.; Chen, H.Y.; et al. Aryl hydrocarbon receptor agonist indigo protects against obesity-related insulin resistance through modulation of intestinal and metabolic tissue immunity. Int. J. Obes. 2019, 43, 2407–2421. [Google Scholar] [CrossRef]
- Zhang, L.; Nichols, R.G.; Correll, J.; Murray, I.A.; Tanaka, N.; Smith, P.B.; Hubbard, T.D.; Sebastian, A.; Albert, I.; Hatzakis, E.; et al. Persistent organic pollutants modify gut microbiota–host metabolic homeostasis in mice through aryl hydrocarbon receptor activation. Environ. Health Perspect. 2015, 123, 679–688. [Google Scholar] [CrossRef] [PubMed]
- Bjeldanes, L.F.; Kim, J.Y.; Grose, K.R.; Bartholomew, J.C.; Bradfield, C.A. Aromatic hydrocarbon responsiveness-receptor agonists generated from indole-3-carbinol in vitro and in vivo: Comparisons with 2,3,7,8-tetrachlorodibenzo-p-dioxin. Proc. Natl. Acad. Sci. USA 1991, 88, 9543–9547. [Google Scholar] [CrossRef] [PubMed]
- Anderton, M.J.; Manson, M.M.; Verschoyle, R.D.; Gescher, A.; Lamb, J.H.; Farmer, P.B.; Steward, W.P.; Williams, M.L. Pharmacokinetics and tissue disposition of indole-3-carbinol and its acid condensation products after oral administration to mice. Clin. Cancer Res. 2004, 10, 5233–5241. [Google Scholar] [CrossRef]
- Natividad, J.M.; Agus, A.; Planchais, J.; Lamas, B.; Jarry, A.C.; Martin, R.; Michel, M.L.; Chong-Nguyen, C.; Roussel, R.; Straube, M.; et al. Impaired Aryl Hydrocarbon Receptor Ligand Production by the Gut Microbiota Is a Key Factor in Metabolic Syndrome. Cell Metab. 2018, 28, 737–749.e4. [Google Scholar] [CrossRef] [PubMed]
- Lopetuso, L.R.; Scaldaferri, F.; Petito, V.; Gasbarrini, A. Commensal Clostridia: Leading players in the maintenance of gut homeostasis. Gut Pathog. 2013, 5, 1–8. [Google Scholar] [CrossRef]
- Stefka, A.T.; Feehley, T.; Tripathi, P.; Qiu, J.; McCoy, K.; Mazmanian, S.K.; Tjota, M.Y.; Seo, G.Y.; Cao, S.; Theriault, B.R.; et al. Commensal bacteria protect against food allergen sensitization. Proc. Natl. Acad. Sci. USA 2014, 111, 13145–13150. [Google Scholar] [CrossRef]
- Nagano, Y.; Itoh, K.; Honda, K. The induction of Treg cells by gut-indigenous Clostridium. Curr. Opin. Immunol. 2012, 24, 392–397. [Google Scholar] [CrossRef]
- Julliard, W.; De Wolfe, T.J.; Fechner, J.H.; Safdar, N.; Agni, R.; Mezrich, J.D. Amelioration of Clostridium difficile infection in mice by dietary supplementation with indole-3-carbinol. Ann. Surg. 2017, 265, 1183–1191. [Google Scholar] [CrossRef]
- Sung, W.S.; Lee, D.G. In vitro antimicrobial activity and the mode of action of indole-3-carbinol against human pathogenic microorganisms. Biol. Pharm. Bull. 2007, 30, 1865–1869. [Google Scholar] [CrossRef]
- Pillaiyar, T.; Köse, M.; Sylvester, K.; Weighardt, H.; Thimm, D.; Borges, G.; Förster, I.; Von Kügelgen, I.; Müller, C.E. Diindolylmethane Derivatives: Potent Agonists of the Immunostimulatory Orphan G Protein-Coupled Receptor GPR84. J. Med. Chem. 2017, 60, 3636–3655. [Google Scholar] [CrossRef]
- Recio, C.; Lucy, D.; Purvis, G.S.D.; Iveson, P.; Zeboudj, L.; Iqbal, A.J.; Lin, D.; O’Callaghan, C.; Davison, L.; Griesbach, E.; et al. Activation of the immune-metabolic receptor GPR84 enhances inflammation and phagocytosis in macrophages. Front. Immunol. 2018, 9, 1419. [Google Scholar] [CrossRef] [PubMed]
- Gevers, D.; Kugathasan, S.; Denson, L.A.; Vázquez-Baeza, Y.; Van Treuren, W.; Ren, B.; Schwager, E.; Knights, D.; Song, S.J.; Yassour, M.; et al. The treatment-naive microbiome in new-onset Crohn’s disease. Cell Host Microbe 2014, 15, 382–392. [Google Scholar] [CrossRef] [PubMed]
- Loh, G.; Blaut, M. Role of commensal gut bacteria in inflammatory bowel diseases. Gut Microbes 2012, 3, 544–555. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, J.V.; Su, G.H.T.; Reddy, J.K.; Simon, M.C.; Bradfield, C.A. Characterization of a murine Ahr null allele: Involvement of the Ah receptor in hepatic growth and development. Proc. Natl. Acad. Sci. USA 1996, 93, 6731–6736. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Schanz, O.; Chijiiwa, R.; Cengiz, S.C.; Majlesain, Y.; Weighardt, H.; Takeyama, H.; Förster, I. Dietary AhR Ligands Regulate AhRR Expression in Intestinal Immune Cells and Intestinal Microbiota Composition. Int. J. Mol. Sci. 2020, 21, 3189. https://doi.org/10.3390/ijms21093189
Schanz O, Chijiiwa R, Cengiz SC, Majlesain Y, Weighardt H, Takeyama H, Förster I. Dietary AhR Ligands Regulate AhRR Expression in Intestinal Immune Cells and Intestinal Microbiota Composition. International Journal of Molecular Sciences. 2020; 21(9):3189. https://doi.org/10.3390/ijms21093189
Chicago/Turabian StyleSchanz, Oliver, Rieka Chijiiwa, Sevgi Can Cengiz, Yasmin Majlesain, Heike Weighardt, Haruko Takeyama, and Irmgard Förster. 2020. "Dietary AhR Ligands Regulate AhRR Expression in Intestinal Immune Cells and Intestinal Microbiota Composition" International Journal of Molecular Sciences 21, no. 9: 3189. https://doi.org/10.3390/ijms21093189
APA StyleSchanz, O., Chijiiwa, R., Cengiz, S. C., Majlesain, Y., Weighardt, H., Takeyama, H., & Förster, I. (2020). Dietary AhR Ligands Regulate AhRR Expression in Intestinal Immune Cells and Intestinal Microbiota Composition. International Journal of Molecular Sciences, 21(9), 3189. https://doi.org/10.3390/ijms21093189




