Pilocarpine Induced Behavioral and Biochemical Alterations in Chronic Seizure-Like Condition in Adult Zebrafish
Abstract
1. Introduction
2. Results
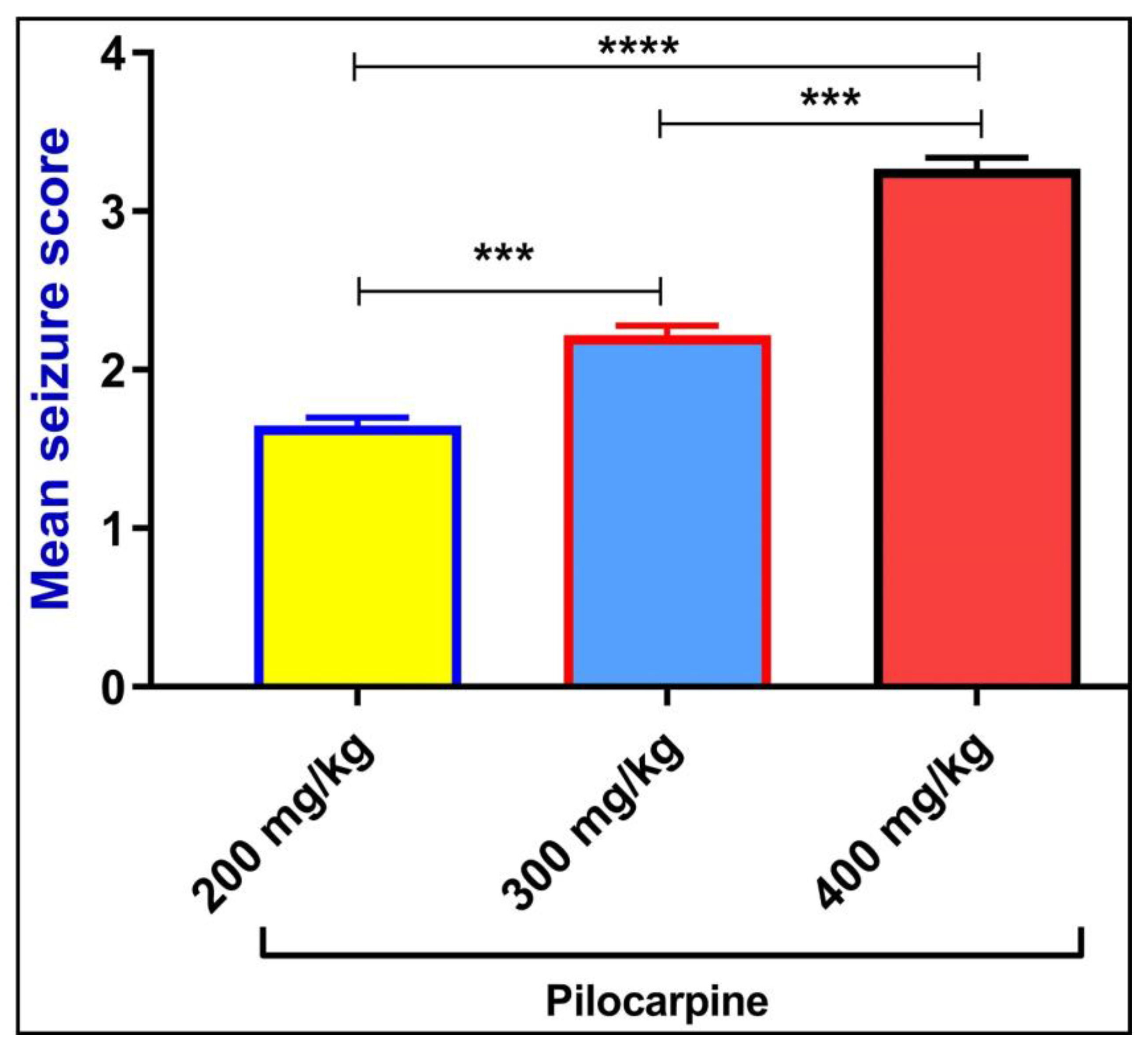
2.1. Pilocarpine-Induced Seizure-Like Behavior
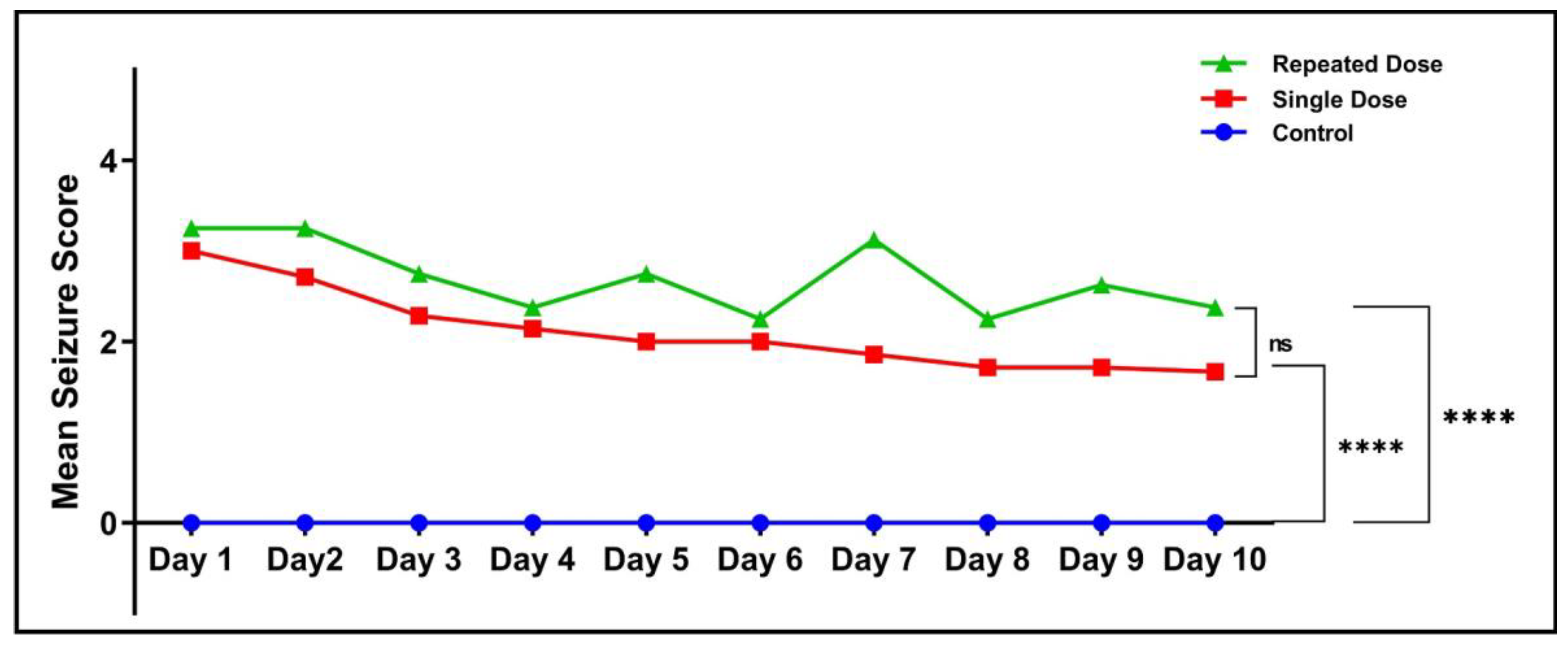
2.2. Mean Seizure Score
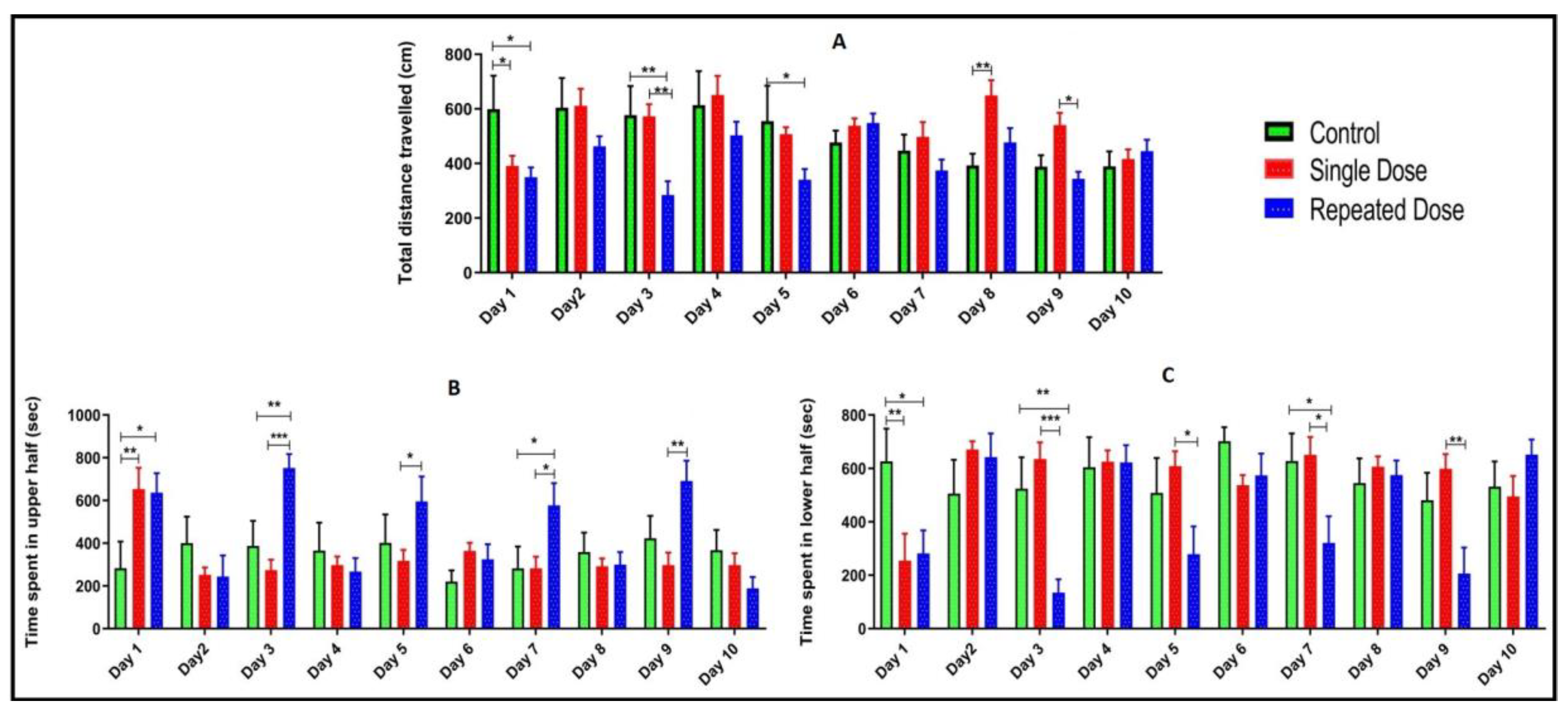
2.3. Total Distance Travelled, Time Spent in Upper and Lower Half of the Tank
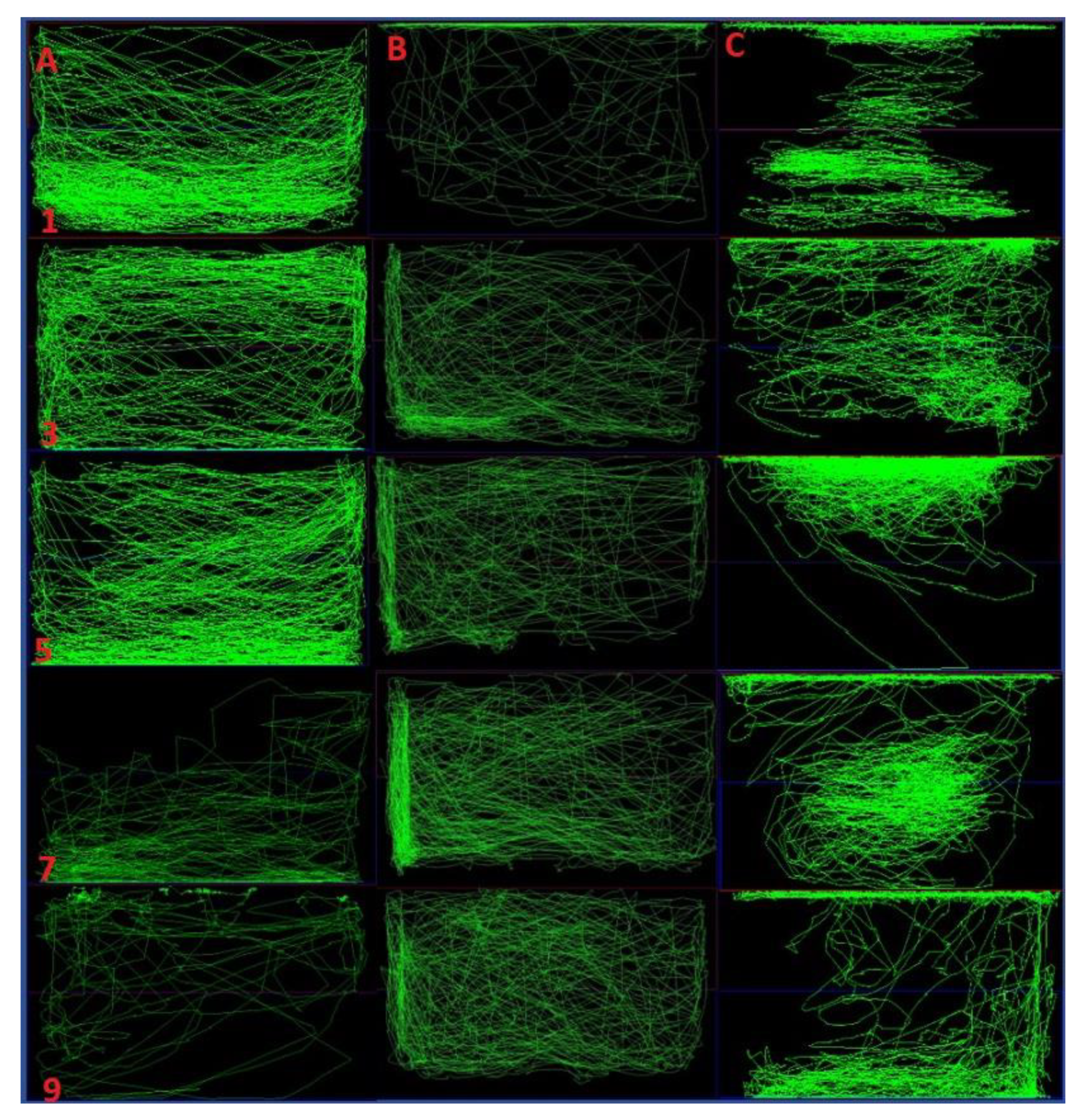
2.4. Comparison of Locomotor Pattern and Behavior
2.5. Modulation of Inflammatory Markers (HMGB1, TLR4, NF-κB, TNF-α, and IL-1), BDNF, CREB-1 and NPY upon Pilocarpine Exposure
2.5.1. HMGB1
2.5.2. TLR4
2.5.3. NF-κB
2.5.4. TNF-α
2.5.5. IL-1
2.5.6. BDNF
2.5.7. CREB-1
2.5.8. NPY
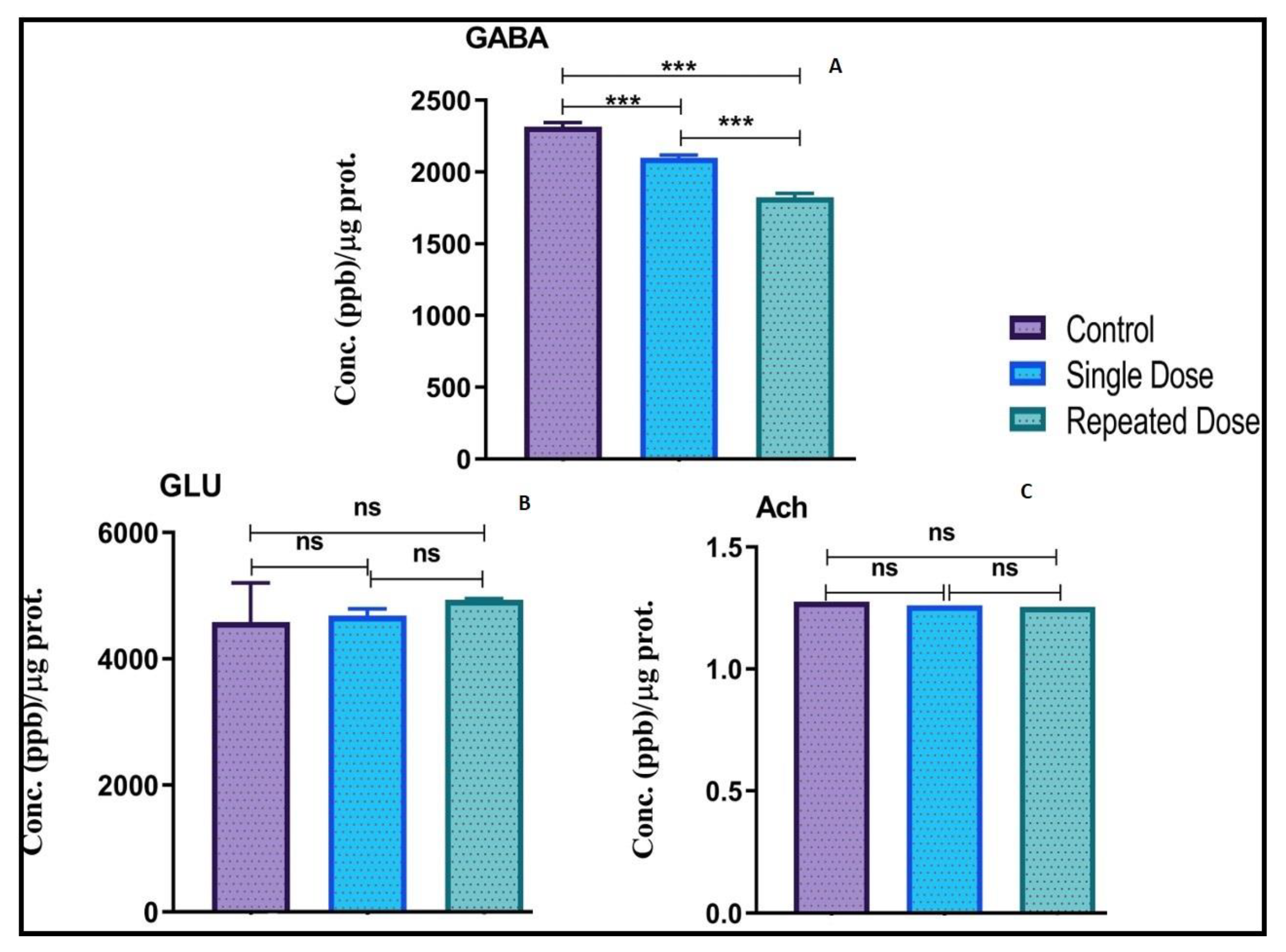
2.6. Modulation of Neurotransmitters Levels in Zebrafish Brain
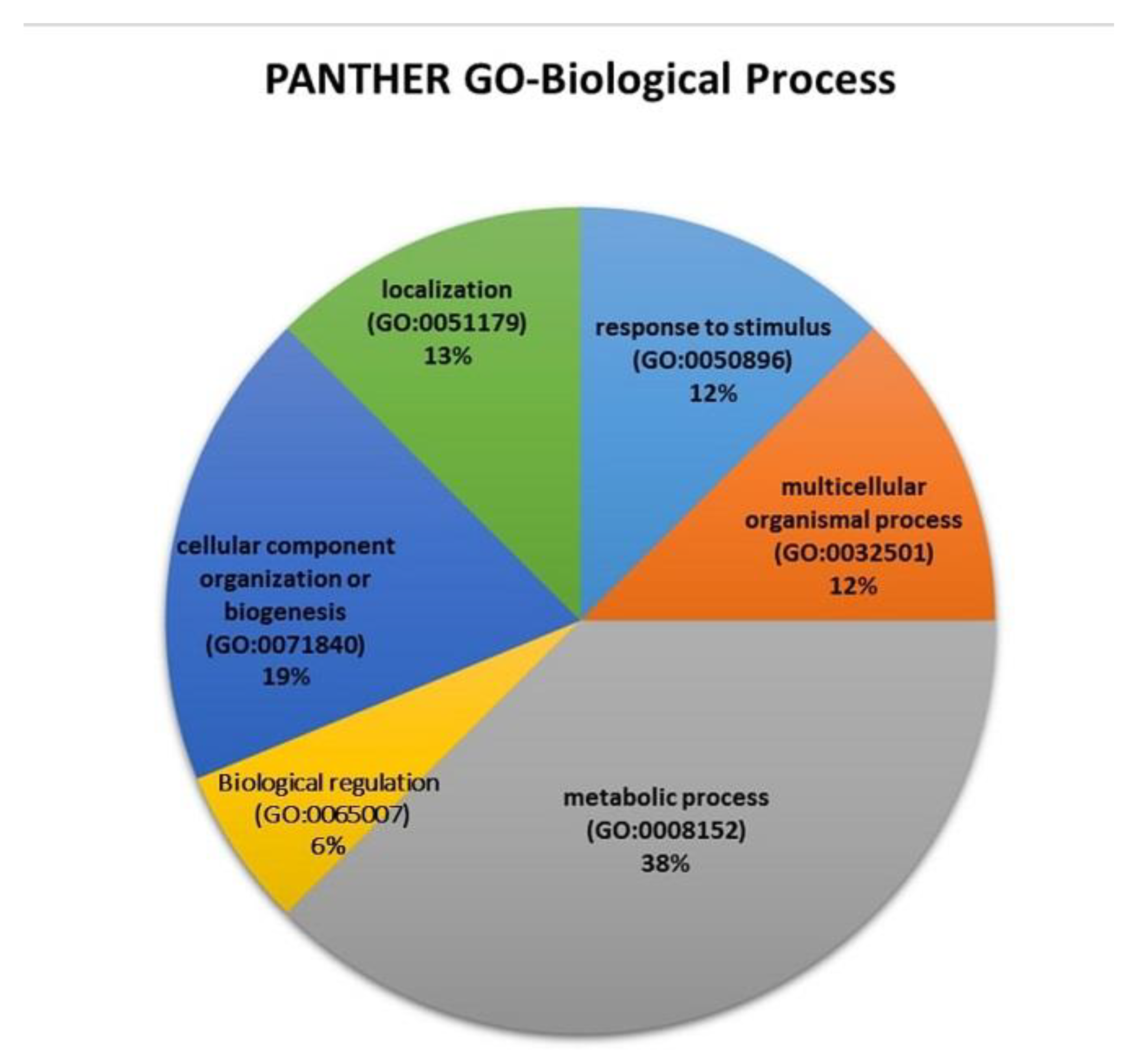
2.7. Proteomic Analysis
3. Discussion
4. Material and Methods
4.1. Experimental Equipment and Chemicals
4.2. Zebrafish (Danio rerio) Care and Maintenance
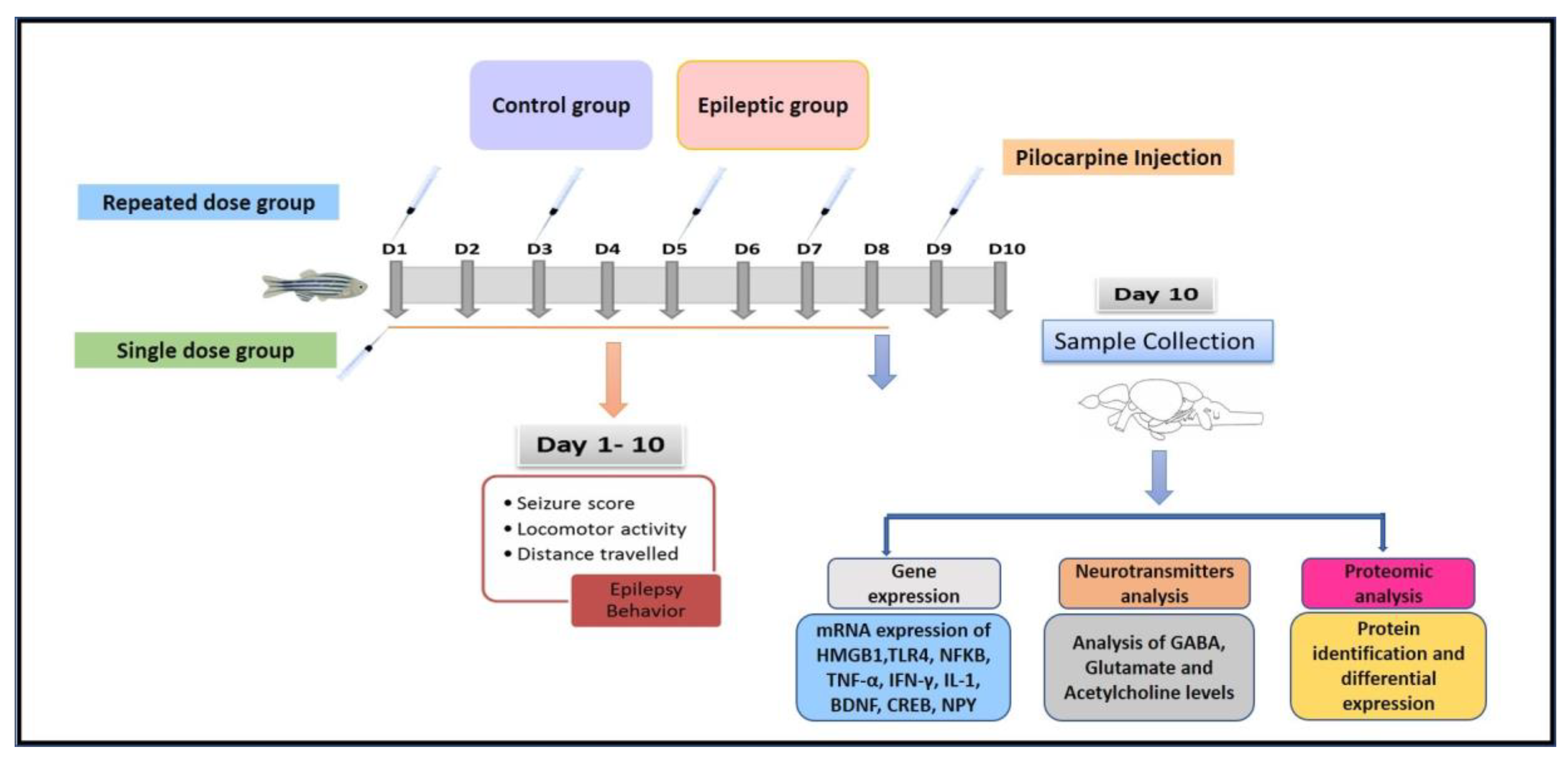
4.3. Pilocarpine-Induced Seizure Behavior
Experimental Design
4.4. Epilepsy Behavior and Seizure Score Recording
4.5. Gene Expression Studies
4.5.1. Brain Harvesting
4.5.2. Isolation of RNA and First-Strand cDNA Synthesis
4.5.3. StepOne®® Real-time PCR
4.6. Brain Neurotransmitter Analysis
4.7. Zebrafish Brain Protein Estimation
4.7.1. In-Solution Tryptic Digestion
4.7.2. Sample Desalting/Cleanup Using Spin Columns
4.7.3. Nanoflow-Ultra High-Performance Liquid Chromatography-Tandem Mass Spectrometry (LC-MS/MS)
4.7.4. Protein Identification and Differential Expression Using PEAKS Bioinformatics Software
4.8. Software and Instrumentation
4.9. Statistical Analysis:
Author Contributions
Funding
Conflicts of Interest
References
- Fisher, R.S.; Acevedo, C.; Arzimanoglou, A.; Bogacz, A.; Cross, J.H.; Elger, C.E.; Engel, J., Jr.; Forsgren, L.; French, J.A.; Glynn, M. ILAE official report: A practical clinical definition of epilepsy. Epilepsia 2014, 55, 475–482. [Google Scholar] [CrossRef] [PubMed]
- Vezzani, A.; Aronica, E.; Mazarati, A.; Pittman, Q.J. Epilepsy and brain inflammation. Exp. Neurol. 2013, 244, 11–21. [Google Scholar] [CrossRef]
- Paudel, Y.N.; Shaikh, M.F.; Shah, S.; Kumari, Y.; Othman, I. Role of inflammation in epilepsy and neurobehavioral comorbidities: Implication for therapy. Eur. J. Pharmacol. 2018. [Google Scholar] [CrossRef]
- Van Vliet, E.A.; Aronica, E.; Vezzani, A.; Ravizza, T. Neuroinflammatory pathways as treatment targets and biomarker candidates in epilepsy: Emerging evidence from preclinical and clinical studies. Neuropathol. Appl. Neurobiol. 2018, 44, 91–111. [Google Scholar] [CrossRef] [PubMed]
- Ravizza, T.; Terrone, G.; Salamone, A.; Frigerio, F.; Balosso, S.; Antoine, D.J.; Vezzani, A. High mobility group box 1 is a novel pathogenic factor and a mechanistic biomarker for epilepsy. Brain Behav. Immun. 2017. [Google Scholar] [CrossRef] [PubMed]
- Paudel, Y.N.; Shaikh, M.; Serrano, Á.A.; Kumari, Y.; Aleksovska, K.; Alvim, M.K.M.; Chakraborti, A.; Othman, I.B. HMGB1: A Common Biomarker and Potential Target for TBI, Neuroinflammation, Epilepsy and Cognitive Dysfunction. Front. Neurosci. 2018, 12, 628. [Google Scholar] [CrossRef] [PubMed]
- Paudel, Y.N.; Angelopoulou, E.; Piperi, C.; Balasubramaniam, V.R.; Othman, I.; Shaikh, M.F. Enlightening the role of high mobility group box 1 (HMGB1) in inflammation: Updates on receptor signalling. Eur. J. Pharmacol. 2019. [Google Scholar] [CrossRef]
- Shams, S.; Rihel, J.; Ortiz, J.G.; Gerlai, R. The zebrafish as a promising tool for modeling human brain disorders: A review based upon an IBNS Symposium. Neurosci. Biobehav. Rev. 2017. [Google Scholar] [CrossRef]
- Copmans, D.l.; Rateb, M.; Tabudravu, J.N.; Pérez-Bonilla, M.; Dirkx, N.; Vallorani, R.; Diaz, C.; Pérez del Palacio, J.; Smith, A.J.; Ebel, R. Zebrafish-Based Discovery of Antiseizure Compounds from the Red Sea: Pseurotin A2 and Azaspirofuran A. ACS Chem. Neurosci. 2018. [Google Scholar] [CrossRef]
- Norton, W.; Bally-Cuif, L. Adult zebrafish as a model organism for behavioural genetics. BMC Neurosci. 2010, 11, 90. [Google Scholar] [CrossRef]
- Mussulini, B.H.M.; Leite, C.E.; Zenki, K.C.; Moro, L.; Baggio, S.; Rico, E.P.; Rosemberg, D.B.; Dias, R.D.; Souza, T.M.; Calcagnotto, M.E. Seizures induced by pentylenetetrazole in the adult zebrafish: A detailed behavioral characterization. PLoS ONE 2013, 8, e54515. [Google Scholar] [CrossRef] [PubMed]
- Gross, A.; Benninger, F.; Madar, R.; Illouz, T.; Griffioen, K.; Steiner, I.; Offen, D.; Okun, E. Toll-like receptor 3 deficiency decreases epileptogenesis in a pilocarpine model of SE-induced epilepsy in mice. Epilepsia 2017, 58, 586–596. [Google Scholar] [CrossRef] [PubMed]
- Lin, W.; Huang, W.; Chen, S.; Lin, M.; Huang, Q.; Huang, H. The Role of 5-HTR6 in Mossy Fiber Sprouting: Activating Fyn and p-ERK1/2 in Pilocarpine-Induced Chronic Epileptic Rats. Cell. Physiol. Biochem. 2017, 42, 231–241. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.; Young Choi, B.; Won Suh, S. Unexpected effects of acetylcholine precursors on pilocarpine seizure-induced neuronal death. Curr. Neuropharmacol. 2018, 16, 51–58. [Google Scholar] [CrossRef]
- Fu, L.; Liu, K.; Wake, H.; Teshigawara, K.; Yoshino, T.; Takahashi, H.; Mori, S.; Nishibori, M. Therapeutic effects of anti-HMGB1 monoclonal antibody on pilocarpine-induced status epilepticus in mice. Sci. Rep. 2017, 7, 1179. [Google Scholar] [CrossRef]
- Mussulini, B.H.M.; Vizuete, A.F.K.; Braga, M.; Moro, L.; Baggio, S.; Santos, E.; Lazzarotto, G.; Zenki, K.C.; Pettenuzzo, L.; da Rocha, J.B.T. Forebrain glutamate uptake and behavioral parameters are altered in adult zebrafish after the induction of Status Epilepticus by kainic acid. Neurotoxicology 2018. [Google Scholar] [CrossRef]
- Vermoesen, K.; Serruys, A.-S.K.; Loyens, E.; Afrikanova, T.; Massie, A.; Schallier, A.; Michotte, Y.; Crawford, A.D.; Esguerra, C.V.; de Witte, P.A. Assessment of the convulsant liability of antidepressants using zebrafish and mouse seizure models. Epilepsy Behav. 2011, 22, 450–460. [Google Scholar] [CrossRef]
- Winter, M.J.; Windell, D.; Metz, J.; Matthews, P.; Pinion, J.; Brown, J.T.; Hetheridge, M.J.; Ball, J.S.; Owen, S.F.; Redfern, W.S. 4-dimensional functional profiling in the convulsant-treated larval zebrafish brain. Sci. Rep. 2017, 7, 1–16. [Google Scholar] [CrossRef]
- Kalueff, A.V.; Gebhardt, M.; Stewart, A.M.; Cachat, J.M.; Brimmer, M.; Chawla, J.S.; Craddock, C.; Kyzar, E.J.; Roth, A.; Landsman, S. Towards a comprehensive catalog of zebrafish behavior 1.0 and beyond. Zebrafish 2013, 10, 70–86. [Google Scholar] [CrossRef]
- Baraban, S.; Taylor, M.; Castro, P.; Baier, H. Pentylenetetrazole induced changes in zebrafish behavior, neural activity and c-fos expression. Neuroscience 2005, 131, 759–768. [Google Scholar] [CrossRef]
- Kundap, U.P.; Kumari, Y.; Othman, I.; Shaikh, M. Zebrafish as a model for epilepsy-induced cognitive dysfunction: A pharmacological, biochemical and behavioral approach. Front. Pharmacol. 2017, 8, 515. [Google Scholar] [CrossRef] [PubMed]
- Mezzomo, N.J.; Fontana, B.D.; Kalueff, A.V.; Barcellos, L.J.; Rosemberg, D.B. Understanding taurine CNS activity using alternative zebrafish models. Neurosci. Biobehav. Rev. 2018, 90, 471–485. [Google Scholar] [CrossRef] [PubMed]
- Choo, B.K.M.; Kundap, U.P.; Kumari, Y.; Hue, S.-M.; Othman, I.; Shaikh, M.F. Orthosiphon stamineus leaf extract affects TNF-α and seizures in a zebrafish model. Front. Pharmacol. 2018, 9, 139. [Google Scholar] [CrossRef] [PubMed]
- Amini, E.; Golpich, M.; Farjam, A.S.; Kamalidehghan, B.; Mohamed, Z.; Ibrahim, N.M.; Ahmadiani, A.; Raymond, A.A. Brain Lipopolysaccharide Preconditioning-Induced Gene Reprogramming Mediates a Tolerance State in Electroconvulsive Shock Model of Epilepsy. Front. Pharmacol. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Vezzani, A.; Granata, T. Brain inflammation in epilepsy: Experimental and clinical evidence. Epilepsia 2005, 46, 1724–1743. [Google Scholar] [CrossRef] [PubMed]
- Vezzani, A.; French, J.; Bartfai, T.; Baram, T.Z. The role of inflammation in epilepsy. Nat. Rev. Neurol. 2011, 7, 31. [Google Scholar] [CrossRef]
- Maroso, M.; Balosso, S.; Ravizza, T.; Liu, J.; Aronica, E.; Iyer, A.M.; Rossetti, C.; Molteni, M.; Casalgrandi, M.; Manfredi, A.A. Toll-like receptor 4 and high-mobility group box-1 are involved in ictogenesis and can be targeted to reduce seizures. Nat. Med. 2010, 16, 413. [Google Scholar] [CrossRef]
- Ying, C.; Ying, L.; Yanxia, L.; Le, W.; Lili, C. High mobility group box 1 antibody represses autophagy and alleviates hippocampus damage in pilocarpine-induced mouse epilepsy model. Acta Histochem. 2019, 151485. [Google Scholar] [CrossRef]
- Shi, Y.; Zhang, L.; Teng, J.; Miao, W. HMGB1 mediates microglia activation via the TLR4/NF-κB pathway in coriaria lactone induced epilepsy. Mol. Med. Rep. 2018, 17, 5125–5131. [Google Scholar] [CrossRef] [PubMed]
- Liu, A.-H.; Wu, Y.-T.; Wang, Y.-P. MicroRNA-129-5p inhibits the development of autoimmune encephalomyelitis-related epilepsy by targeting HMGB1 through the TLR4/NF-kB signaling pathway. Brain Res. Bull. 2017, 132, 139–149. [Google Scholar] [CrossRef] [PubMed]
- Qu, Z.; Jia, L.; Xie, T.; Zhen, J.; Si, P.; Cui, Z.; Xue, Y.; Sun, C.; Wang, W. (−)-Epigallocatechin-3-Gallate Protects Against Lithium-Pilocarpine-Induced Epilepsy by Inhibiting the Toll-Like Receptor 4 (TLR4)/Nuclear Factor-κB (NF-κB) Signaling Pathway. Med. Sci. Mon. 2019, 25, 1749. [Google Scholar] [CrossRef] [PubMed]
- Ashhab, M.U.; Omran, A.; Kong, H.; Gan, N.; He, F.; Peng, J.; Yin, F. Expressions of tumor necrosis factor alpha and microRNA-155 in immature rat model of status epilepticus and children with mesial temporal lobe epilepsy. J. Mol. Neurosci. 2013, 51, 950–958. [Google Scholar] [CrossRef] [PubMed]
- Rijkers, K.; Majoie, H.; Hoogland, G.; Kenis, G.; De Baets, M.; Vles, J. The role of interleukin-1 in seizures and epilepsy: A critical review. Exp. Neurol. 2009, 216, 258–271. [Google Scholar] [CrossRef] [PubMed]
- De Simoni, M.G.; Perego, C.; Ravizza, T.; Moneta, D.; Conti, M.; Marchesi, F.; De Luigi, A.; Garattini, S.; Vezzani, A. Inflammatory cytokines and related genes are induced in the rat hippocampus by limbic status epilepticus. Eur. J. Neurosci. 2000, 12, 2623–2633. [Google Scholar] [CrossRef]
- Gall, C.M. Seizure-induced changes in neurotrophin expression: Implications for epilepsy. Exp. Neurol. 1993, 124, 150–166. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Han, X.; Blendy, J.A.; Porter, B.E. Decreased CREB levels suppress epilepsy. Neurobiol. Dis. 2012, 45, 253–263. [Google Scholar] [CrossRef]
- Kandel, E.R. The molecular biology of memory: cAMP, PKA, CRE, CREB-1, CREB-2, and CPEB. Mol. Brain 2012, 5, 14. [Google Scholar] [CrossRef]
- Noe, F.; Vaghi, V.; Balducci, C.; Fitzsimons, H.; Bland, R.; Zardoni, D.; Sperk, G.; Carli, M.; During, M.; Vezzani, A. Anticonvulsant effects and behavioural outcomes of rAAV serotype 1 vector-mediated neuropeptide Y overexpression in rat hippocampus. Gene Ther. 2010, 17, 643. [Google Scholar] [CrossRef]
- Gøtzsche, C.; Woldbye, D. The role of NPY in learning and memory. Neuropeptides 2016, 55, 79–89. [Google Scholar] [CrossRef]
- Luo, J.; Min, S.; Wei, K.; Li, P.; Dong, J.; Liu, Y.-f. Propofol protects against impairment of learning-memory and imbalance of hippocampal Glu/GABA induced by electroconvulsive shock in depressed rats. J. Anesth. 2011, 25, 657. [Google Scholar] [CrossRef]
- Kaila, K.; Ruusuvuori, E.; Seja, P.; Voipio, J.; Puskarjov, M. GABA actions and ionic plasticity in epilepsy. Curr. Opin. Neurobiol. 2014, 26, 34–41. [Google Scholar] [CrossRef] [PubMed]
- Choo, B.K.M.; Kundap, U.P.; bin Johan Arief, M.F.; Kumari, Y.; Yap, J.L.; Wong, C.P.; Othman, I.; Shaikh, M.F. Effect of newer anti-epileptic drugs (AEDs) on the cognitive status in pentylenetetrazol induced seizures in a zebrafish model. Prog. Neuro-Psychopharmacol. Biol. Psychiatry 2019, 92, 483–493. [Google Scholar] [CrossRef]
- Miller, H.P.; Levey, A.I.; Rothstein, J.D.; Tzingounis, A.V.; Conn, P.J. Alterations in glutamate transporter protein levels in kindling-induced epilepsy. J. Neurochem. 1997, 68, 1564–1570. [Google Scholar] [CrossRef] [PubMed]
- Chapman, A.G. Glutamate and epilepsy. J. Nutr. 2000, 130, 1043S–1045S. [Google Scholar] [CrossRef] [PubMed]
- Holmes, G.L. Seizure-induced neuronal injury: Animal data. Neurology 2002, 59, S3–S6. [Google Scholar] [CrossRef] [PubMed]
- Izquierdo, I.; Medina, J.H. Memory formation: The sequence of biochemical events in the hippocampus and its connection to activity in other brain structures. Neurobiol. Learn. Memory 1997, 68, 285–316. [Google Scholar] [CrossRef]
- Atzori, M.; Kanold, P.; Pineda, J.C.; FLORES-HERNANDEZ, J. Dopamine-Acetylcholine Interactions in the Modulation of Glutamate Release. Ann. N. Y. Acad. Sci. 2003, 1003, 346–348. [Google Scholar] [CrossRef]
- Persike, D.; Marques-Carneiro, J.; Stein, M.; Yacubian, E.; Centeno, R.; Canzian, M.; Fernandes, M. Altered Proteins in the Hippocampus of Patients with Mesial Temporal Lobe Epilepsy. Pharmaceuticals 2018, 11, 95. [Google Scholar] [CrossRef]
- Yang, J.; Czech, T.; Felizardo, M.; Baumgartner, C.; Lubec, G. Aberrant expression of cytoskeleton proteins in hippocampus from patients with mesial temporal lobe epilepsy. Amino Acids 2006, 30, 477–493. [Google Scholar] [CrossRef]
- Yang, Q.; Wang, S.; Karlsson, J.-E.; Hamberger, A.; Haglid, K.G. Phosphorylated and non-phosphorylated neurofilament proteins: Distribution in the rat hippocampus and early changes after kainic acid induced seizures. J. Chem. Neuroanat. 1995, 9, 217–228. [Google Scholar] [CrossRef]
- Kundap, U.P.; Paudel, Y.N.; Kumari, Y.; Othman, I.; Shaikh, M.F. Embelin prevents seizure and associated cognitive impairments in a pentylenetetrazole-induced kindling zebrafish model. Front. Pharmacol. 2019, 10, 315. [Google Scholar] [CrossRef] [PubMed]


| Score 0 | Normal Swimming |
| Score 1 | Jittery movement at the top of the tank |
| Score 2 | Ataxia/Hyperactivity |
| Score 3 | Circular movement, Circling around small area |
| Score 4 | Erratic burst movement with loss of posture/Corkscrew swimming |
| S.N. | Accession | −10lgP | Coverage (%) | #Peptides | #Unique | Avg. Mass | Protein Description | Biological Process (GO) from Uniprot and Protein Analysis Through Evolutionary Relationships (PANTHER) |
|---|---|---|---|---|---|---|---|---|
| 1 | P17561|EPD_DANRE | 89.06 | 15 | 2 | 2 | 24,472 | Ependymin | cell-matrix adhesion (GO:0007160) |
| 2 | tr|Q6DHB6|Q6DHB6_DANRE | 71.15 | 6 | 4 | 3 | 49,994 | Keratin 91 | NA |
| 3 | tr|A0A2R8Q6V3|A0A2R8Q6V3_DANRE | 38.54 | 4 | 2 | 2 | 57,453 | Keratin type II cytoskeletal 8 | NA |
| 4 | tr|A8WGN0|A8WGN0_DANRE | 38.54 | 4 | 2 | 2 | 57,660 | Keratin 8 | NA |
| 5 | tr|Q6P3K5|Q6P3K5_DANRE | 44.56 | 4 | 2 | 2 | 57,828 | Krt5 protein | NA |
| 6 | tr|Q9PUB5|Q9PUB5_DANRE | 44.56 | 4 | 2 | 2 | 58,587 | Type II cytokeratin | fin regeneration (GO:0031101) |
| 7 | tr|F1R5A5|F1R5A5_DANRE | 44.56 | 4 | 2 | 2 | 58,715 | Keratin 5 | |
| 8 | tr|A9JRN9|A9JRN9_DANRE | 61.41 | 3 | 2 | 2 | 95,635 | LOC794362 protein | axon development (GO:0061564) |
| 9 | tr|F1QCR7|F1QCR7_DANRE | 61.41 | 3 | 2 | 2 | 95,577 | Neurofilament medium polypeptide a | |
| 10 | tr|Q32PU7|Q32PU7_DANRE | 83.35 | 8 | 3 | 1 | 49,717 | Tubulin beta chain | microtubule-based process (GO:0007017) |
| 11 | tr|B8A516|B8A516_DANRE | 68.72 | 6 | 2 | 2 | 49,969 | Tubulin alpha chain | microtubule-based process (GO:0007017), microtubule cytoskeleton organization (GO:0000226), mitotic cell cycle (GO:0000278) |
| 12 | tr|Q502J6|Q502J6_DANRE | 79.92 | 25 | 2 | 2 | 11,412 | Gamma1a-synuclein | dopaminergic neuron differentiation (GO:0071542), larval locomotory behavior (GO:0008345) |
| 13 | tr|A0A2R8RP85|A0A2R8RP85_DANRE | 79.92 | 25 | 2 | 2 | 11,442 | Synuclein gamma b (breast cancer-specific protein 1) |
| S.N. | Accession | −10lgP | Coverage (%) | #Peptides | #Unique | Avg. Mass | Protein Description | Biological Process (GO) From Uniprot and Protein Analysis Through Evolutionary Relationships (PANTHER) |
|---|---|---|---|---|---|---|---|---|
| 1 | tr|Q6P5M9|Q6P5M9_DANRE | 153.04 | 50 | 16 | 2 | 49,787 | Tubulin beta chain | Cellular comp. organization or biogenesis (GO:0071840) |
| 2 | Q90486|HBB1_DANRE | 123.66 | 64 | 7 | 7 | 16,389 | Hemoglobin subunit beta-1 | Cellular comp. organization or biogenesis (GO:0071840), localization (GO: 0051179), Metabolic process (GO:0008152), response to stimulus (GO:0050896) |
| 3 | tr|Q6ZM12|Q6ZM12_DANRE | 40.06 | 16 | 2 | 2 | 16,295 | Hemoglobin beta adult 2 | |
| 4 | tr|Q6ZM13|Q6ZM13_DANRE | 80.62 | 29 | 3 | 2 | 15,413 | Hemoglobin alpha adult 2 | |
| 5 | tr|B3DG37|B3DG37_DANRE | 123.66 | 64 | 7 | 7 | 16,389 | Ba1 protein | |
| 6 | tr|Q803Z5|Q803Z5_DANRE | 83.91 | 43 | 5 | 1 | 15,508 | Hbaa1 protein | |
| 7 | tr|Q6DGK4|Q6DGK4_DANRE | 40.06 | 16 | 2 | 2 | 16,279 | Zgc:92880 protein | |
| 8 | tr|Q9DEU2|Q9DEU2_DANRE | 112.71 | 10 | 8 | 2 | 112,703 | Sodium/potassium-transporting ATPase subunit alpha | Biological regulation (GO:0065007), localization (GO: 0051179), |
| 9 | Q7ZVF9|ACTB2_DANRE | 105.86 | 28 | 6 | 6 | 41,753 | Actin cytoplasmic 2 | Cellular process (GO:0009987), localization (GO: 0051179) |
| 10 | Q7ZVI7|ACTB1_DANRE | 116.98 | 27 | 7 | 1 | 41,767 | Actin cytoplasmic 1 | |
| 11 | tr|A8WG05|A8WG05_DANRE | 105.86 | 28 | 6 | 6 | 41,753 | Bactin2 protein | |
| 12 | tr|B2GS08|B2GS08_DANRE | 116.98 | 27 | 7 | 1 | 41,710 | Bactin1 protein | |
| 13 | tr|R4GE02|R4GE02_DANRE | 96.97 | 23 | 4 | 3 | 27,149 | Si:ch211-113a14.11 | Cellular comp. organization or biogenesis (GO:0071840) |
| 14 | tr|Q7ZU04|Q7ZU04_DANRE | 90.53 | 20 | 5 | 5 | 42,916 | Creatine kinase brain b | NA |
| 15 | tr|Q9I8N9|Q9I8N9_DANRE | 81.04 | 29 | 3 | 3 | 14,918 | Brain-type fatty acid-binding protein | NA |
| 16 | tr|A0A2R8Q1X2|A0A2R8Q1X2_DANRE | 78.78 | 12 | 3 | 3 | 47,246 | Enolase 1a (alpha) | Glycolytic process (GO:0006096) |
| 17 | tr|Q08BA1|Q08BA1_DANRE | 77.71 | 7 | 3 | 3 | 59,744 | ATP synthase subunit alpha | Metabolic process (GO:0008152), response to stimulus (GO:0050896) |
| 18 | tr|A8WGC6|A8WGC6_DANRE | 53.73 | 6 | 2 | 2 | 55,130 | ATP synthase subunit beta | |
| 19 | tr|Q6PC77|Q6PC77_DANRE | 61.34 | 29 | 3 | 3 | 18,258 | ATP synthase subunit d mitochondrial | |
| 20 | Q5MJ86|G3P2_DANRE | 76.85 | 8 | 2 | 2 | 36,107 | Glyceraldehyde-3-phosphate dehydrogenase 2 | Metabolic process (GO:0008152) |
| 21 | tr|Q8JHI0|Q8JHI0_DANRE | 67.61 | 12 | 3 | 3 | 32,763 | Solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator) member 5 | Transporter activity (GO:0005215) |
| 22 | tr|F1R5A5|F1R5A5_DANRE | 67.55 | 5 | 4 | 2 | 58,715 | Keratin 5 | Fin regeneration (GO:0031101) |
| 23 | tr|I3IRY2|I3IRY2_DANRE | 46.87 | 10 | 2 | 2 | 15,746 | Keratin type 1 c19e (Fragment) | NA |
| 24 | tr|F1R8U0|F1R8U0_DANRE | 46.87 | 4 | 2 | 2 | 47,785 | Keratin 94 | NA |
| 25 | tr|Q1RLR3|Q1RLR3_DANRE | 46.87 | 3 | 2 | 2 | 50,931 | Keratin 93 | NA |
| 26 | tr|Q1LXJ9|Q1LXJ9_DANRE | 46.87 | 3 | 2 | 2 | 49,934 | Keratin type 1 c19e | NA |
| 27 | tr|Q24JW4|Q24JW4_DANRE | 66.4 | 14 | 3 | 3 | 35,677 | Flj13639 | NA |
| 28 | Q90XG0|TPISB_DANRE | 65.52 | 11 | 2 | 2 | 26,828 | Triosephosphate isomerase B | Metabolic process (GO:0008152) |
| 29 | P17561|EPD_DANRE | 55.6 | 12 | 2 | 2 | 24,472 | Ependymin | Cell-matrix adhesion (GO:0007160) |
| 30 | Q7T356|143BB_DANRE | 44.88 | 10 | 2 | 2 | 27,393 | 14-3-3 protein beta/alpha-B | Cell cycle (GO:0007049), Signal transduction (GO:0007165) |
| 31 | tr|A4FVM3|A4FVM3_DANRE | 44.86 | 7 | 2 | 2 | 36,022 | LOC557717 protein (Fragment) | Exocytosis (GO:0006887), Intracellular protein transport (GO:0006886), lysosomal transport (GO:0007041) |
| 32 | tr|A9JRN9|A9JRN9_DANRE | 92.47 | 8 | 5 | 4 | 95,635 | LOC794362 protein | ND |
| 33 | tr|A0A2R8QP59|A0A2R8QP59_DANRE | 44.86 | 4 | 2 | 2 | 66,189 | Syntaxin binding protein 1b | Localization (GO: 0051179), multicellular organismal process (GO: 0032501), signaling (GO:0023052). |
| 34 | tr|F1QM13|F1QM13_DANRE | 44.86 | 4 | 2 | 2 | 67,075 | Syntaxin-binding protein 1a | |
| 35 | tr|A2BGE0|A2BGE0_DANRE | 44.86 | 3 | 2 | 2 | 68,575 | Si:rp71-10d23.3 | Localization (GO:0051179), multicellular organismal process (GO:0032501), |
| 36 | tr|F1QYN7|F1QYN7_DANRE | 33.39 | 9 | 2 | 2 | 21,828 | Myelin protein zero | Cellular process (GO:0009987) |
| 37 | tr|F8W3W8|F8W3W8_DANRE | 111.06 | 60 | 9 | 9 | 10,776 | Myelin basic protein a | |
| 38 | tr|A0A2R8QC30|A0A2R8QC30_DANRE | 33.39 | 9 | 2 | 2 | 22,667 | Uncharacterized protein | NA |
| 39 | tr|A0A2R8Q6P3|A0A2R8Q6P3_DANRE | 33.39 | 8 | 2 | 2 | 25,013 | Uncharacterized protein | NA |
| 40 | tr|Q0D294|Q0D294_DANRE | 111.32 | 42 | 5 | 5 | 11,351 | Histone H4 | Nucleosome assemble (GO:0006334) |
| 41 | tr|Q6GQM9|Q6GQM9_DANRE | 98.45 | 15 | 4 | 2 | 46,841 | Eno2 protein | Metabolic process (GO:0008152) |
| 42 | Q6PI52|CALM_DANRE | 98.01 | 52 | 6 | 6 | 16,838 | Calmodulin | Biological regulation (GO:0065007), Cellular component organization or biogenesis (GO:0071840), cellular process (GO:0009987) |
| 43 | tr|Q7SZP4|Q7SZP4_DANRE | 94.54 | 15 | 6 | 5 | 64,751 | Zgc:65851 | Nervous system development (GO:0007399) |
| 44 | tr|F1QCR7|F1QCR7_DANRE | 92.47 | 8 | 5 | 4 | 95,577 | Neurofilament medium polypeptide a | Axon development (GO:0061564) |
| 45 | tr|A0A2R8RRA6|A0A2R8RRA6_DANRE | 92.13 | 8 | 4 | 4 | 74,958 | Serotransferrin | Hemoglobin biosynthetic process (GO:0042541) |
| 46 | tr|F1Q8F1|F1Q8F1_DANRE | 87.46 | 9 | 4 | 3 | 54,195 | Internexin neuronal intermediate filament protein alpha b | Neuron projection morphogenesis (GO:0048812) |
| 47 | tr|Q58EH1|Q58EH1_DANRE | 87.46 | 9 | 4 | 3 | 54,223 | Gefiltin | Neuron projection morphogenesis (GO:0048812) |
| 48 | tr|Q5BJC7|Q5BJC7_DANRE | 80.62 | 29 | 3 | 2 | 15,403 | Si:xx-by187g17.5 protein | hydrogen peroxide catabolic process (GO:0042744), protein heterooligomerization (GO:0051291) |
| 49 | tr|A0A0B5JW41|A0A0B5JW41_DANRE | 79.76 | 34 | 3 | 3 | 10,145 | Cd59 (Fragment) | defense response to Gram-positive bacterium (GO:0050830) |
| 50 | tr|B3DFP9|B3DFP9_DANRE | 78.8 | 50 | 4 | 4 | 15,537 | Apolipoprotein A-II | chordate embryonic development (GO:0043009), nuclear division (GO:0000280) |
| 51 | tr|A0A0R4IKF0|A0A0R4IKF0_DANRE | 65.95 | 18 | 4 | 4 | 30,140 | Apolipoprotein A-Ib | biological regulation (GO:0065007), Cellular component organization or biogenesis (GO:0071840), localization (GO:0051179), metabolic process (GO:0008152), multicellular organismal process (GO:0032501) |
| 52 | Q804W2|PRV7_DANRE | 78.02 | 33 | 3 | 3 | 12,029 | Parvalbumin-7 | biological regulation (GO:0065007) |
| 53 | tr|Q6TH32|Q6TH32_DANRE | 72.15 | 21 | 2 | 2 | 18,771 | Cofilin 1 | developmental process (GO:0032502), metabolic process (GO:0008152) |
| 54 | Q9PVK4|LDHBA_DANRE | 71.3 | 11 | 3 | 3 | 36,247 | L-lactate dehydrogenase B-A chain | metabolic process (GO:0008152) |
| 55 | tr|Q7T334|Q7T334_DANRE | 69.4 | 10 | 2 | 2 | 35,420 | Malate dehydrogenase | |
| 56 | tr|U3JAS0|U3JAS0_DANRE | 68.21 | 25 | 3 | 3 | 15,001 | Synaptosomal-associated protein | cellular protein-containing complex assembly (GO:0034622), synaptic vesicle exocytosis (GO:0016079), vesicle fusion to plasma membrane (GO:0099500) |
| 57 | tr|Q7SX92|Q7SX92_DANRE | 63.05 | 21 | 2 | 2 | 13,341 | Beta-synuclein | dopaminergic neuron differentiation (GO:0071542), larval locomotory behavior (GO:0008345) |
| 58 | tr|E7F9E8|E7F9E8_DANRE | 61.67 | 7 | 3 | 2 | 44,183 | Neurofilament light polypeptide a | axon development (GO:0061564) |
| 59 | tr|A0A0N4STS4|A0A0N4STS4_DANRE | 60.07 | 11 | 2 | 2 | 26,174 | Ubiquitin B | modification-dependent protein catabolic process (GO:0019941), protein ubiquitination (GO:0016567) |
| 60 | tr|B3DLH2|B3DLH2_DANRE | 60.07 | 5 | 2 | 2 | 51,501 | Zgc:172187 protein | |
| 61 | tr|A0JPF1|A0JPF1_DANRE | 60.07 | 5 | 2 | 2 | 59,921 | Zgc:153686 | |
| 62 | tr|Q7SXA3|Q7SXA3_DANRE | 60.07 | 16 | 2 | 2 | 17,999 | Ribosomal protein S27a | metabolic process (GO:0008152) |
| 63 | tr|B8JKN6|B8JKN6_DANRE | 59.82 | 12 | 2 | 2 | 17,404 | Peptidyl-prolyl cis-trans isomerase | protein refolding (GO:0042026) |
| 64 | tr|A0A2R8Q9S7|A0A2R8Q9S7_DANRE | 58.58 | 60 | 2 | 2 | 6468 | Si:dkey-46i9.1 | NA |
| 65 | tr|Q0ZBR7|Q0ZBR7_DANRE | 58.26 | 31 | 3 | 3 | 12,412 | Macrophage migration inhibitory factor | auditory receptor cell development (GO:0060117), cell proliferation (GO:0008283), embryonic morphogenesis (GO:0048598), inner ear development (GO:0048839), negative regulation of apoptotic process (GO:0043066) |
| 66 | tr|Q502C8|Q502C8_DANRE | 53.39 | 26 | 3 | 3 | 17,109 | Peroxiredoxin 5 | cell redox homeostasis (GO:0045454), cellular response to oxidative stress (GO:0034599), fin regeneration (GO:0031101), hydrogen peroxide catabolic process (GO:0042744) |
| 67 | tr|A0A0R4IG45|A0A0R4IG45_DANRE | 52.38 | 2 | 2 | 2 | 94,588 | Dynamin 1a | biological regulation (GO:0065007), cellular component organization or biogenesis (GO:0071840), cellular process (GO:0009987), localization (GO:0051179), metabolic process (GO:0008152) |
| 68 | tr|E9QF63|E9QF63_DANRE | 52.38 | 2 | 2 | 2 | 95,446 | Dynamin 1b | |
| 69 | tr|Q6TNV0|Q6TNV0_DANRE | 51.28 | 11 | 2 | 2 | 19,385 | Cytochrome c oxidase subunit 4I1 | metabolic process (GO:0008152) |
| 70 | tr|Q4VBT9|Q4VBT9_DANRE | 51.28 | 11 | 2 | 2 | 19,443 | Cox4i1 protein | metabolic process (GO:0008152), mitochondrial electron transport, cytochrome c to oxygen (GO:0006123) |
| 71 | tr|Q5TZ35|Q5TZ35_DANRE | 50.73 | 13 | 2 | 2 | 22,261 | Visinin-like 1b | NA |
| 72 | tr|A0A2R8RID5|A0A2R8RID5_DANRE | 50.34 | 5 | 2 | 2 | 84,805 | Aconitate hydratase mitochondrial | tricarboxylic acid cycle (GO:0006099) |
| Q6IQM2|CYC_DANRE | 49.91 | 18 | 2 | 2 | 11,456 | Cytochrome c | mitochondrial electron transport, cytochrome c to oxygen (GO:0006123) | |
| 73 | Q8JH70|ALDCB_DANRE | 48.6 | 9 | 2 | 2 | 39,259 | Fructose-bisphosphate aldolase C-B | glycolytic process (GO:0006096) |
| 74 | tr|Q05AL9|Q05AL9_DANRE | 46.87 | 4 | 2 | 2 | 47,758 | Zgc:153629 protein | NA |
| 75 | tr|Q1JQ08|Q1JQ08_DANRE | 46.87 | 3 | 2 | 2 | 51,584 | Si:dkeyp-113d7.4 protein (Fragment) | NA |
| 76 | tr|F1R9V3|F1R9V3_DANRE | 42.3 | 4 | 2 | 2 | 71,094 | Si:dkey-4p15.3 | cellular process (GO:0009987), localization (GO:0051179), response to stimulus (GO:0050896) |
| 77 | tr|A0A0R4IMF8|A0A0R4IMF8_DANRE | 42.3 | 4 | 2 | 2 | 70,145 | Heat shock cognate 71 kDa protein | cellular process (GO:0009987), localization (GO:0051179), response to stimulus (GO:0050896) |
| S.N. | Accession | Group Profile (Ratio, Control: Epileptic) | Description | PANTHER GO-Slim Biological Process (Danio rerio) |
|---|---|---|---|---|
| 1 | tr|F8W3W8|F8W3W8_DANRE | 1.00:0.09 | Myelin basic protein a | No Match |
| 2 | tr|Q8AY63|Q8AY63_DANRE | 1.00:0 | Brain-subtype creatine kinase | No Match |
| 3 | tr|F1R3D3|F1R3D3_DANRE | 1.00:0.02 | Glyceraldehyde-3-phosphate dehydrogenase | No Match |
| 4 | tr|R4GE02|R4GE02_DANRE | 1.00:0.03 | Si:ch211-113a14.11 | Cellular component organization or biogenesis (GO:0071840 |
| 5 | tr|Q9I8N9|Q9I8N9_DANRE | 1.00:0.04 | Brain-type fatty acid-binding protein | No Match |
| 6 | tr|Q6IQP5|Q6IQP5_DANRE | 1.00:0 | Enolase 1 (Alpha) | No Match |
| 7 | tr|Q1LWD7|Q1LWD7_DANRE | 1.00:0.02 | Parvalbumin | No Match |
| 8 | tr|A0A0R4IS04|A0A0R4IS04_DANRE | 1.00:0 | Myelin protein zero | No Match |
| 9 | tr|B3DG37|B3DG37_DANRE | 1.00:0.03 | Ba1 protein | No Match |
| 10 | tr|E9QJ96|E9QJ96_DANRE | 1.00:0.03 | 14-3-3 protein beta/alpha-A (Fragment) | No Match |
| 11 | tr|Q08BA1|Q08BA1_DANRE | 1.00:0.08 | ATP synthase subunit alpha | Metabolic process (GO:0008152), Response to stimulus (GO:0050896) |
| 12 | tr|Q6PC77|Q6PC77_DANRE | 1.00:0 | ATP synthase subunit d mitochondrial | Metabolic process (GO:0008152) |
| 13 | tr|A8WGC6|A8WGC6_DANRE | 1.00:0.04 | ATP synthase subunit | Metabolic process (GO:0008152) |
| 14 | tr|Q24JW4|Q24JW4_DANRE | 1.00:0 | Flj13639 | Multicellular organismal process (GO:0032501) |
| 15 | tr|A3KPR4|A3KPR4_DANRE | 1.00:0.00 | Histone H4 | No Match |
| 16 | tr|Q7T334|Q7T334_DANRE | 1.00:0 | Malate dehydrogenase | Metabolic process (GO:0008152) |
| 17 | tr|Q6ZM12|Q6ZM12_DANRE | 1.00:0 | Hemoglobin beta adult 2 | localization (GO:0051179), Metabolic process (GO:0008152), Response to stimulus (GO:0050896) |
| 18 | tr|A0A0B5JW41|A0A0B5JW41_DANRE | 1.00:0.01 | Cd59 (Fragment) | No Match |
| 19 | tr|A0A0R4IKF0|A0A0R4IKF0_DANRE | 1.00:0 | Apolipoprotein A-Ib | Cellular component organization or biogenesis (GO:0071840), Biological regulation (GO:0065007), localization (GO:0051179), Metabolic process (GO:0008152), Multicellular organismal process (GO:0032501) |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Paudel, Y.N.; Kumari, Y.; Abidin, S.A.Z.; Othman, I.; Shaikh, M.F. Pilocarpine Induced Behavioral and Biochemical Alterations in Chronic Seizure-Like Condition in Adult Zebrafish. Int. J. Mol. Sci. 2020, 21, 2492. https://doi.org/10.3390/ijms21072492
Paudel YN, Kumari Y, Abidin SAZ, Othman I, Shaikh MF. Pilocarpine Induced Behavioral and Biochemical Alterations in Chronic Seizure-Like Condition in Adult Zebrafish. International Journal of Molecular Sciences. 2020; 21(7):2492. https://doi.org/10.3390/ijms21072492
Chicago/Turabian StylePaudel, Yam Nath, Yatinesh Kumari, Syafiq Asnawi Zainal Abidin, Iekhsan Othman, and Mohd. Farooq Shaikh. 2020. "Pilocarpine Induced Behavioral and Biochemical Alterations in Chronic Seizure-Like Condition in Adult Zebrafish" International Journal of Molecular Sciences 21, no. 7: 2492. https://doi.org/10.3390/ijms21072492
APA StylePaudel, Y. N., Kumari, Y., Abidin, S. A. Z., Othman, I., & Shaikh, M. F. (2020). Pilocarpine Induced Behavioral and Biochemical Alterations in Chronic Seizure-Like Condition in Adult Zebrafish. International Journal of Molecular Sciences, 21(7), 2492. https://doi.org/10.3390/ijms21072492







