Zinc Finger Proteins in the Human Fungal Pathogen Cryptococcus neoformans
Abstract
:1. Introduction
2. Zinc Finger Proteins: An Overview
3. ZNFs Identified in C. neoformans
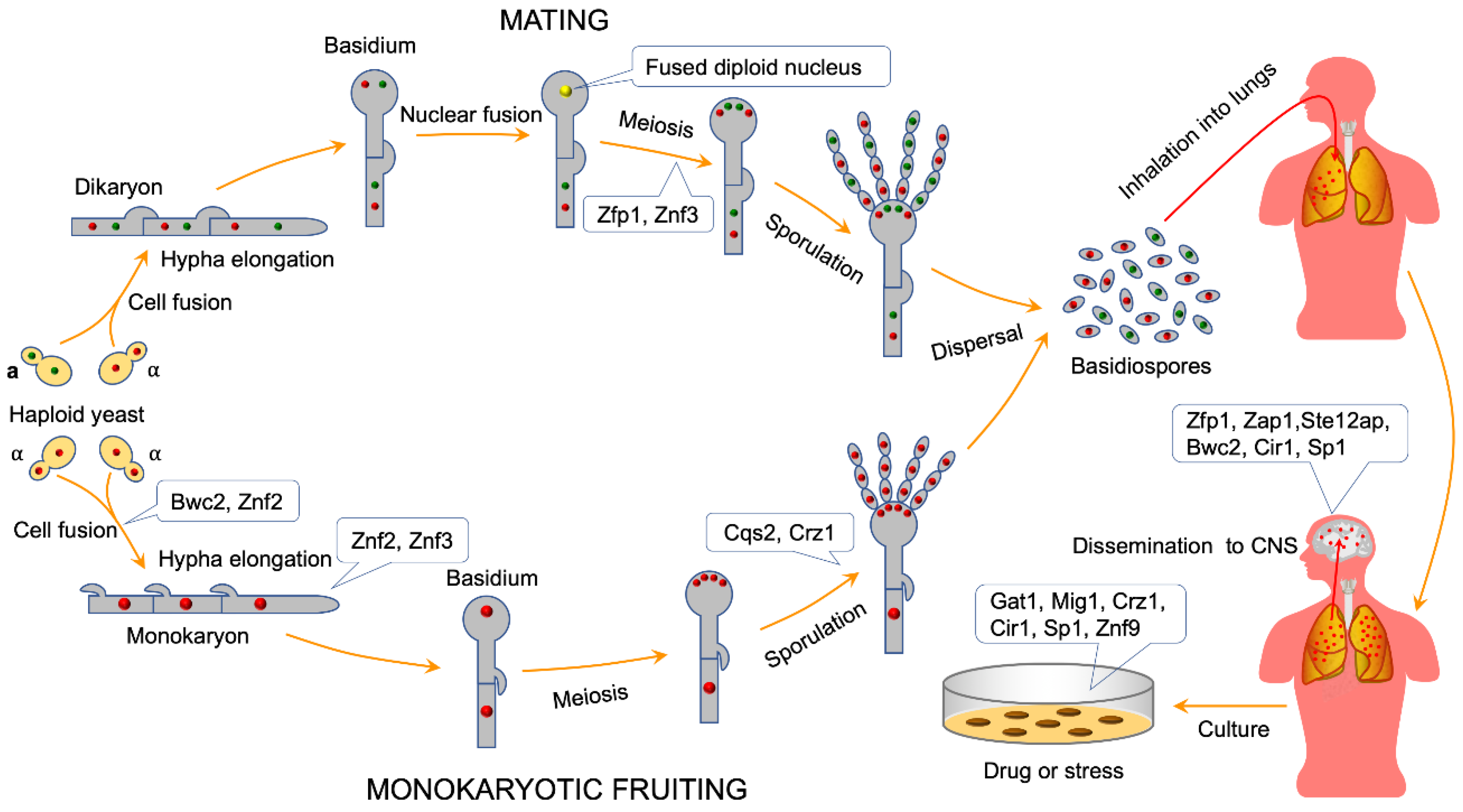
4. ZNFs Regulate Sexual Reproduction in C. neoformans
4.1. ZNFs Regulate Cell Fusion during Mating in C. neoformans
4.2. ZNFs Regulate Mating Hyphal Growth in C. neoformans
4.3. ZNFs Regulate Basidiospore Formation during Mating or Monokaryotic Fruiting in C. neoformans
5. ZNFs Regulate the Virulence in C. neoformans
5.1. ZNFs Regulate Virulence Factors Production in C. neoformans
5.2. ZNFs Mediate Cryptococcus Crossing the Blood-Brain Barrier
5.3. ZNFs Mediate Metal Homeostasis in C. neoformans
6. ZNFs Regulate Stress Resistance in C. neoformans
7. Conclusions and Future Directions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Casadevall, A.; Perfect, J.R. Cryptococcus Neoformans; ASM Press: Washington, DC, USA, 1998. [Google Scholar]
- Lin, X.; Heitman, J. The biology of the Cryptococcus neoformans species complex. Annu. Rev. Microbiol. 2006, 60, 69–105. [Google Scholar] [CrossRef] [PubMed]
- Idnurm, A.; Bahn, Y.S.; Nielsen, K.; Lin, X.; Fraser, J.A.; Heitman, J. Deciphering the model pathogenic fungus Cryptococcus neoformans. Nat. Rev. Microbiol. 2005, 3, 753–764. [Google Scholar] [CrossRef] [PubMed]
- Brown, G.D.; Denning, D.W.; Levitz, S.M. Tackling human fungal infections. Science 2012, 336, 647. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Park, B.J.; Wannemuehler, K.A.; Marston, B.J.; Govender, N.; Pappas, P.G.; Chiller, T.M. Estimation of the current global burden of cryptococcal meningitis among persons living with HIV/AIDS. AIDS 2009, 23, 525–530. [Google Scholar] [CrossRef]
- Rajasingham, R.; Smith, R.M.; Park, B.J.; Jarvis, J.N.; Govender, N.P.; Chiller, T.M.; Denning, D.W.; Loyse, A.; Boulware, D.R. Global burden of disease of HIV-associated cryptococcal meningitis: An updated analysis. Lancet Infect. Dis. 2017, 17, 873–881. [Google Scholar] [CrossRef] [Green Version]
- Kozel, T.R. Virulence factors of Cryptococcus neoformans. Trends Microbiol. 1995, 3, 295–299. [Google Scholar] [CrossRef]
- Kronstad, J.; Jung, W.H.; Hu, G. Beyond the big three: Systematic analysis of virulence factors in Cryptococcus neoformans. Cell Host Microbe 2008, 4, 308–310. [Google Scholar] [CrossRef] [Green Version]
- Lin, X.; Hull, C.M.; Heitman, J. Sexual reproduction between partners of the same mating type in Cryptococcus neoformans. Nature 2005, 434, 1017–1021. [Google Scholar] [CrossRef]
- Belay, T.; Cherniak, R.; ONeill, E.B.; Kozel, T.R. Serotyping of Cryptococcus neoformans by dot enzyme assay. J. Clin. Microbiol. 1996, 34, 466–470. [Google Scholar] [CrossRef] [Green Version]
- Levitz, S.M. The Ecology of Cryptococcus neoformans and the Epidemiology of Cryptococcosis. Rev. Infect. Dis. 1991, 13, 1163–1169. [Google Scholar] [CrossRef]
- Franzot, S.P.; Salkin, I.F.; Casadevall, A. Cryptococcus neoformans var. grubii: Separate varietal status for Cryptococcus neoformans serotype A isolates. J. Clin. Microbiol. 1999, 37, 838–840. [Google Scholar]
- Springer, D.J.; Chaturvedi, V. Projecting Global Occurrence of Cryptococcus gattii. Emerg. Infect. Dis. 2010, 16, 14–20. [Google Scholar] [CrossRef]
- Hagen, F.; Khayhan, K.; Theelen, B.; Kolecka, A.; Polacheck, I.; Sionov, E.; Falk, R.; Parnmen, S.; Lumbsch, H.T.; Boekhout, T. Recognition of seven species in the Cryptococcus gattii/Cryptococcus neoformans species complex. Fungal Genet. Biol. 2015, 78, 16–48. [Google Scholar] [CrossRef] [Green Version]
- Kwon-Chung, K.J.; Bennett, J.E.; Wickes, B.L.; Meyer, W.; Cuomo, C.A.; Wollenburg, K.R.; Bicanic, T.A.; Castaneda, E.; Chang, Y.C.; Chen, J.H.; et al. The Case for Adopting the “Species Complex” Nomenclature for the Etiologic Agents of Cryptococcosis. Msphere 2017, 2, e00357-16. [Google Scholar] [CrossRef] [Green Version]
- Kozubowski, L.; Lee, S.C.; Heitman, J. Signalling pathways in the pathogenesis of Cryptococcus. Cell Microbiol. 2009, 11, 370–380. [Google Scholar] [CrossRef] [Green Version]
- Lengeler, K.B.; Davidson, R.C.; D’Souza, C.; Harashima, T.; Shen, W.C.; Wang, P.; Pan, X.; Waugh, M.; Heitman, J. Signal transduction cascades regulating fungal development and virulence. Microbiol. Mol. Biol. Rev. 2000, 64, 746–785. [Google Scholar] [CrossRef] [Green Version]
- Wang, P.; Heitman, J. Signal transduction cascades regulating mating, filamentation, and virulence in Cryptococcus neoformans. Curr. Opin. Microbiol. 1999, 2, 358–362. [Google Scholar] [CrossRef]
- Liu, T.B.; Wang, Y.; Stukes, S.; Chen, Q.; Casadevall, A.; Xue, C. The F-Box protein Fbp1 regulates sexual reproduction and virulence in Cryptococcus neoformans. Eukaryot. Cell 2011, 10, 791–802. [Google Scholar] [CrossRef] [Green Version]
- Masso-Silva, J.; Espinosa, V.; Liu, T.B.; Wang, Y.; Xue, C.; Rivera, A. The F-Box Protein Fbp1 Shapes the Immunogenic Potential of Cryptococcus neoformans. mBio 2018, 9, e01828-17. [Google Scholar] [CrossRef] [Green Version]
- Laity, J.H.; Lee, B.M.; Wright, P.E. Zinc finger proteins: New insights into structural and functional diversity. Curr. Opin. Struct. Biol. 2001, 11, 39–46. [Google Scholar] [CrossRef]
- MacPherson, S.; Larochelle, M.; Turcotte, B. A fungal family of transcriptional regulators: The zinc cluster proteins. Microbiol. Mol. Biol. Rev. 2006, 70, 583–604. [Google Scholar] [CrossRef] [Green Version]
- Bhat, P.J.; Murthy, T.V. Transcriptional control of the GAL/MEL regulon of yeast Saccharomyces cerevisiae: Mechanism of galactose-mediated signal transduction. Mol. Microbiol. 2001, 40, 1059–1066. [Google Scholar] [CrossRef] [PubMed]
- Miller, J.; Mclachlan, A.D.; Klug, A. Repetitive Zinc-Binding Domains in the Protein Transcription Factor Iiia from Xenopus Oocytes. EMBO J. 1985, 4, 1609–1614. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.S.; Gippert, G.P.; Soman, K.V.; Case, D.A.; Wright, P.E. Three-dimensional solution structure of a single zinc finger DNA-binding domain. Science 1989, 245, 635–637. [Google Scholar] [CrossRef] [PubMed]
- Berg, J.M.; Shi, Y. The galvanization of biology: A growing appreciation for the roles of zinc. Science 1996, 271, 1081–1085. [Google Scholar] [CrossRef] [PubMed]
- Dai, K.S.; Liew, C.C. Characterization of a novel gene encoding zinc finger domains identified from expressed sequence tags (ESTs) of a human heart cDNA database. J. Mol. Cell. Cardiol. 1998, 30, 2365–2375. [Google Scholar] [CrossRef]
- Gupta, S.K.; Rai, A.K.; Kanwar, S.S.; Sharma, T.R. Comparative analysis of zinc finger proteins involved in plant disease resistance. PLoS ONE 2012, 7, e42578. [Google Scholar] [CrossRef] [Green Version]
- Summers, M.F. Zinc finger motif for single-stranded nucleic acids? Investigations by nuclear magnetic resonance. J. Cell. Biochem. 1991, 45, 41–48. [Google Scholar] [CrossRef]
- Bombarda, E.; Grell, E.; Roques, B.P.; Mely, Y. Molecular mechanism of the Zn2+-induced folding of the distal CCHC finger motif of the HIV-1 nucleocapsid protein. Biophys. J. 2007, 93, 208–217. [Google Scholar] [CrossRef] [Green Version]
- Matsui, T.; Kodera, Y.; Endoh, H.; Miyauchi, E.; Komatsu, H.; Sato, K.; Tanaka, T.; Kohno, T.; Maeda, T. RNA recognition mechanism of the minimal active domain of the human immunodeficiency virus type-2 nucleocapsid protein. J. Biochem. 2007, 141, 269–277. [Google Scholar] [CrossRef]
- Narayanan, N.; Gorelick, R.J.; DeStefano, J.J. Structure/function mapping of amino acids in the N-terminal zinc finger of the human immunodeficiency virus type 1 nucleocapsid protein: Residues responsible for nucleic acid helix destabilizing activity. Biochemistry 2006, 45, 12617–12628. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Clay, N.K.; Nelson, T. The recessive epigenetic swellmap mutation affects the expression of two step II splicing factors required for the transcription of the cell proliferation gene STRUWWELPETER and for the timing of cell cycle arrest in the Arabidopsis leaf. Plant Cell 2005, 17, 1994–2008. [Google Scholar] [CrossRef] [Green Version]
- Wang, D.; Guo, Y.; Wu, C.; Yang, G.; Li, Y.; Zheng, C. Genome-wide analysis of CCCH zinc finger family in Arabidopsis and rice. BMC Genom. 2008, 9, 44. [Google Scholar] [CrossRef] [Green Version]
- DuBois, R.N.; McLane, M.W.; Ryder, K.; Lau, L.F.; Nathans, D. A growth factor-inducible nuclear protein with a novel cysteine/histidine repetitive sequence. J. Biol. Chem. 1990, 265, 19185–19191. [Google Scholar] [PubMed]
- Li, W.T.; He, M.; Wang, J.; Wang, Y.P. Zinc finger protein (ZFP) in plants-A review. Plant Omics 2013, 6, 474–480. [Google Scholar]
- Omichinski, J.G.; Clore, G.M.; Schaad, O.; Felsenfeld, G.; Trainor, C.; Appella, E.; Stahl, S.J.; Gronenborn, A.M. NMR structure of a specific DNA complex of Zn-containing DNA binding domain of GATA-1. Science 1993, 261, 438–446. [Google Scholar] [CrossRef] [PubMed]
- Kosarev, P.; Mayer, K.F.; Hardtke, C.S. Evaluation and classification of RING-finger domains encoded by the Arabidopsis genome. Genome Biol. 2002, 3, research0016-1. [Google Scholar] [CrossRef] [PubMed]
- Stone, S.L.; Hauksdottir, H.; Troy, A.; Herschleb, J.; Kraft, E.; Callis, J. Functional analysis of the RING-type ubiquitin ligase family of Arabidopsis. Plant Physiol. 2005, 137, 13–30. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lim, S.D.; Yim, W.C.; Moon, J.C.; Kim, D.S.; Lee, B.M.; Jang, C.S. A gene family encoding RING finger proteins in rice: Their expansion, expression diversity, and co-expressed genes. Plant Mol. Biol. 2010, 72, 369–380. [Google Scholar] [CrossRef] [PubMed]
- Lorick, K.L.; Jensen, J.P.; Fang, S.; Ong, A.M.; Hatakeyama, S.; Weissman, A.M. RING fingers mediate ubiquitin-conjugating enzyme (E2)-dependent ubiquitination. Proc. Natl. Acad. Sci. USA 1999, 96, 11364–11369. [Google Scholar] [CrossRef] [Green Version]
- Deshaies, R.J.; Joazeiro, C.A. RING domain E3 ubiquitin ligases. Annu. Rev. Biochem. 2009, 78, 399–434. [Google Scholar] [CrossRef] [PubMed]
- Freyd, G.; Kim, S.K.; Horvitz, H.R. Novel cysteine-rich motif and homeodomain in the product of the Caenorhabditis elegans cell lineage gene lin-11. Nature 1990, 344, 876–879. [Google Scholar] [CrossRef] [PubMed]
- Marmorstein, R.; Carey, M.; Ptashne, M.; Harrison, S.C. DNA Recognition by Gal4—Structure of a Protein DNA Complex. Nature 1992, 356, 408–414. [Google Scholar] [CrossRef] [PubMed]
- Borden, K.L.; Freemont, P.S. The RING finger domain: A recent example of a sequence-structure family. Curr. Opin. Struct. Biol. 1996, 6, 395–401. [Google Scholar] [CrossRef]
- Gabig, T.G.; Mantel, P.L.; Rosli, R.; Crean, C.D. Requiem: A novel zinc finger gene essential for apoptosis in myeloid cells. J. Biol. Chem. 1994, 269, 29515–29519. [Google Scholar]
- Koken, M.H.; Saib, A.; de The, H. A C4HC3 zinc finger motif. Comptes Rendus Acad. Sci. III 1995, 318, 733–739. [Google Scholar]
- Evans, R.M. The Steroid and Thyroid-Hormone Receptor Superfamily. Science 1988, 240, 889–895. [Google Scholar] [CrossRef]
- Krishna, S.S.; Majumdar, I.; Grishin, N.V. Structural classification of zinc fingers. Nucleic Acids Res. 2003, 31, 532–550. [Google Scholar] [CrossRef] [Green Version]
- Kluska, K.; Adamczyk, J.; Krezel, A. Metal binding properties, stability and reactivity of zinc fingers. Coord. Chem. Rev. 2018, 367, 18–64. [Google Scholar] [CrossRef]
- Gray, K.A.; Yates, B.; Seal, R.L.; Wright, M.W.; Bruford, E.A. Genenames.org: The HGNC resources in 2015. Nucleic Acids Res. 2015, 43, D1079–D1085. [Google Scholar] [CrossRef]
- Loftus, B.J.; Fung, E.; Roncaglia, P.; Rowley, D.; Amedeo, P.; Bruno, D.; Vamathevan, J.; Miranda, M.; Anderson, I.J.; Fraser, J.A.; et al. The genome of the basidiomycetous yeast and human pathogen Cryptococcus neoformans. Science 2005, 307, 1321–1324. [Google Scholar] [CrossRef] [Green Version]
- Janbon, G.; Ormerod, K.L.; Paulet, D.; Byrnes, E.J., 3rd; Yadav, V.; Chatterjee, G.; Mullapudi, N.; Hon, C.C.; Billmyre, R.B.; Brunel, F.; et al. Analysis of the genome and transcriptome of Cryptococcus neoformans var. grubii reveals complex RNA expression and microevolution leading to virulence attenuation. PLoS Genet. 2014, 10, e1004261. [Google Scholar]
- Davidson, R.C.; Nichols, C.B.; Cox, G.M.; Perfect, J.R.; Heitman, J. A MAP kinase cascade composed of cell type specific and non-specific elements controls mating and differentiation of the fungal pathogen Cryptococcus neoformans. Mol. Microbiol. 2003, 49, 469–485. [Google Scholar] [CrossRef] [PubMed]
- Cruz, M.C.; Fox, D.S.; Heitman, J. Calcineurin is required for hyphal elongation during mating and haploid fruiting in Cryptococcus neoformans. EMBO J. 2001, 20, 1020–1032. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jung, K.W.; Bahn, Y.S. The Stress-Activated Signaling (SAS) Pathways of a Human Fungal Pathogen, Cryptococcus neoformans. Mycobiology 2009, 37, 161–170. [Google Scholar] [CrossRef] [Green Version]
- Bahn, Y.S.; Kojima, K.; Cox, G.M.; Heitman, J. Specialization of the HOG pathway and its impact on differentiation and virulence of Cryptococcus neoformans. Mol. Biol. Cell 2005, 16, 2285–2300. [Google Scholar] [CrossRef] [Green Version]
- Lin, X.; Jackson, J.C.; Feretzaki, M.; Xue, C.; Heitman, J. Transcription factors Mat2 and Znf2 operate cellular circuits orchestrating opposite- and same-sex mating in Cryptococcus neoformans. PLoS Genet. 2010, 6, e1000953. [Google Scholar] [CrossRef] [Green Version]
- Feretzaki, M.; Heitman, J. Genetic circuits that govern bisexual and unisexual reproduction in Cryptococcus neoformans. PLoS Genet. 2013, 9, e1003688. [Google Scholar] [CrossRef] [Green Version]
- Lengeler, K.B.; Fox, D.S.; Fraser, J.A.; Allen, A.; Forrester, K.; Dietrich, F.S.; Heitman, J. Mating-type locus of Cryptococcus neoformans: A step in the evolution of sex chromosomes. Eukaryot. Cell 2002, 1, 704–718. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Zhai, B.; Lin, X. The link between morphotype transition and virulence in Cryptococcus neoformans. PLoS Pathog. 2012, 8, e1002765. [Google Scholar] [CrossRef] [Green Version]
- Zhai, B.; Zhu, P.; Foyle, D.; Upadhyay, S.; Idnurm, A.; Lin, X. Congenic strains of the filamentous form of Cryptococcus neoformans for studies of fungal morphogenesis and virulence. Infect. Immun. 2013, 81, 2626–2637. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, J.; Idnurm, A.; Lin, X. Morphology and its underlying genetic regulation impact the interaction between Cryptococcus neoformans and its hosts. Med. Mycol. 2015, 53, 493–504. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, J.; Zhao, Y.; Ferraro, A.R.; Yang, E.; Lewis, Z.A.; Lin, X. Transcription factor Znf2 coordinates with the chromatin remodeling SWI/SNF complex to regulate cryptococcal cellular differentiation. Commun. Biol. 2019, 2, 412. [Google Scholar] [CrossRef]
- Feretzaki, M.; Billmyre, R.B.; Clancey, S.A.; Wang, X.Y.; Heitman, J. Gene Network Polymorphism Illuminates Loss and Retention of Novel RNAi Silencing Components in the Cryptococcus Pathogenic Species Complex. PLoS Genet. 2016, 12, e1005868. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fox, D.S.; Heitman, J. Good fungi gone bad: The corruption of calcineurin. Bioessays 2002, 24, 894–903. [Google Scholar] [CrossRef]
- Steinbach, W.J.; Reedy, J.L.; Cramer, R.A., Jr.; Perfect, J.R.; Heitman, J. Harnessing calcineurin as a novel anti-infective agent against invasive fungal infections. Nat. Rev. Microbiol. 2007, 5, 418–430. [Google Scholar] [CrossRef]
- Fu, C.; Donadio, N.; Cardenas, M.E.; Heitman, J. Dissecting the Roles of the Calcineurin Pathway in Unisexual Reproduction, Stress Responses, and Virulence in Cryptococcus deneoformans. Genetics 2018, 208, 639–653. [Google Scholar] [CrossRef] [Green Version]
- Jung, W.H.; Son, Y.E.; Oh, S.H.; Fu, C.; Kim, H.S.; Kwak, J.H.; Cardenas, M.E.; Heitman, J.; Park, H.S. Had1 Is Required for Cell Wall Integrity and Fungal Virulence in Cryptococcus neoformans. G3 Genes Genomes Genet. 2018, 8, 643–652. [Google Scholar] [CrossRef] [Green Version]
- Lu, Y.K.; Sun, K.H.; Shen, W.C. Blue light negatively regulates the sexual filamentation via the Cwc1 and Cwc2 proteins in Cryptococcus neoformans. Mol. Microbiol. 2005, 56, 480–491. [Google Scholar] [CrossRef]
- Idnurm, A.; Heitman, J. Light controls growth and development via a conserved pathway in the fungal kingdom. PLoS Biol. 2005, 3, 615–626. [Google Scholar] [CrossRef]
- Yeh, Y.L.; Lin, Y.S.; Su, B.J.; Shen, W.C. A screening for suppressor mutants reveals components involved in the blue light-inhibited sexual filamentation in Cryptococcus neoformans. Fungal Genet. Biol. 2009, 46, 42–54. [Google Scholar] [CrossRef] [PubMed]
- Roberts, R.L.; Fink, G.R. Elements of a Single Map Kinase Cascade in Saccharomyces cerevisiae Mediate 2 Developmental Programs in the Same Cell-Type—Mating and Invasive Growth. Genes Dev. 1994, 8, 2974–2985. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wickes, B.L.; Edman, U.; Edman, J.C. The Cryptococcus neoformans STE12alpha gene: A putative Saccharomyces cerevisiae STE12 homologue that is mating type specific. Mol. Microbiol. 1997, 26, 951–960. [Google Scholar] [CrossRef] [PubMed]
- Yue, C.; Cavallo, L.M.; Alspaugh, J.A.; Wang, P.; Cox, G.M.; Perfect, J.R.; Heitman, J. The STE12alpha homolog is required for haploid filamentation but largely dispensable for mating and virulence in Cryptococcus neoformans. Genetics 1999, 153, 1601–1615. [Google Scholar] [PubMed]
- Chang, Y.C.; Penoyer, L.A.; Kwon-Chung, K.J. The second STE12 homologue of Cryptococcus neoformans is MATa-specific and plays an important role in virulence. Proc. Natl. Acad. Sci. USA 2001, 98, 3258–3263. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chang, Y.C.; Wickes, B.L.; Miller, G.F.; Penoyer, L.A.; Kwon-Chung, K.J. Cryptococcus neoformans STE12alpha regulates virulence but is not essential for mating. J. Exp. Med. 2000, 191, 871–882. [Google Scholar] [CrossRef]
- Jonkers, W.; Rep, M. Lessons from fungal F-box proteins. Eukaryot. Cell 2009, 8, 677–695. [Google Scholar] [CrossRef] [Green Version]
- Fan, C.L.; Han, L.T.; Jiang, S.T.; Chang, A.N.; Zhou, Z.Y.; Liu, T.B. The Cys2His2 zinc finger protein Zfp1 regulates sexual reproduction and virulence in Cryptococcus neoformans. Fungal Genet. Biol. 2019, 124, 59–72. [Google Scholar] [CrossRef]
- Abisado, R.G.; Benomar, S.; Klaus, J.R.; Dandekar, A.A.; Chandler, J.R. Bacterial Quorum Sensing and Microbial Community Interactions. mBio 2018, 9, e02331-17. [Google Scholar] [CrossRef] [Green Version]
- Whiteley, M.; Diggle, S.P.; Greenberg, E.P. Progress in and promise of bacterial quorum sensing research. Nature 2017, 551, 313–320. [Google Scholar] [CrossRef]
- Tian, X.; He, G.J.; Hu, P.; Chen, L.; Tao, C.; Cui, Y.L.; Shen, L.; Ke, W.; Xu, H.; Zhao, Y.; et al. Cryptococcus neoformans sexual reproduction is controlled by a quorum sensing peptide. Nat. Microbiol. 2018, 3, 698–707. [Google Scholar] [CrossRef] [PubMed]
- Bahn, Y.S.; Jung, K.W. Stress signaling pathways for the pathogenicity of Cryptococcus. Eukaryot. Cell 2013, 12, 1564–1577. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Choi, J.; Jung, W.H.; Kronstad, J.W. The cAMP/protein kinase A signaling pathway in pathogenic basidiomycete fungi: Connections with iron homeostasis. J. Microbiol. 2015, 53, 579–587. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Roman, E.; Arana, D.M.; Nombela, C.; Alonso-Monge, R.; Pla, J. MAP kinase pathways as regulators of fungal virulence. Trends Microbiol. 2007, 15, 181–190. [Google Scholar] [CrossRef]
- Chang, Y.C.; Wright, L.C.; Tscharke, R.L.; Sorrell, T.C.; Wilson, C.F.; Kwon-Chung, K.J. Regulatory roles for the homeodomain and C2H2 zinc finger regions of Cryptococcus neoformans Ste12alphap. Mol. Microbiol. 2004, 53, 1385–1396. [Google Scholar] [CrossRef]
- Adler, A.; Park, Y.D.; Larsen, P.; Nagarajan, V.; Wollenberg, K.; Qiu, J.; Myers, T.G.; Williamson, P.R. A novel specificity protein 1 (SP1)-like gene regulating protein kinase C-1 (Pkc1)-dependent cell wall integrity and virulence factors in Cryptococcus neoformans. J. Biol. Chem. 2011, 286, 20977–20990. [Google Scholar] [CrossRef] [Green Version]
- Vu, K.; Tham, R.; Uhrig, J.P.; Thompson, G.R., 3rd; Na Pombejra, S.; Jamklang, M.; Bautos, J.M.; Gelli, A. Invasion of the central nervous system by Cryptococcus neoformans requires a secreted fungal metalloprotease. mBio 2014, 5, e01101-14. [Google Scholar] [CrossRef] [Green Version]
- Jung, W.H.; Sham, A.; White, R.; Kronstad, J.W. Iron regulation of the major virulence factors in the AIDS-associated pathogen Cryptococcus neoformans. PLoS Biol. 2006, 4, e410. [Google Scholar] [CrossRef] [Green Version]
- Gerik, K.J.; Bhimireddy, S.R.; Ryerse, J.S.; Specht, C.A.; Lodge, J.K. PKC1 is essential for protection against both oxidative and nitrosative stresses, cell integrity, and normal manifestation of virulence factors in the pathogenic fungus Cryptococcus neoformans. Eukaryot. Cell 2008, 7, 1685–1698. [Google Scholar] [CrossRef] [Green Version]
- Na Pombejra, S.; Jamklang, M.; Uhrig, J.P.; Vu, K.; Gelli, A. The structure-function analysis of the Mpr1 metalloprotease determinants of activity during migration of fungal cells across the blood-brain barrier. PLoS ONE 2018, 13, e0203020. [Google Scholar] [CrossRef]
- Gerwien, F.; Skrahina, V.; Kasper, L.; Hube, B.; Brunke, S. Metals in fungal virulence. FEMS Microbiol. Rev. 2018, 42, fux050. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jung, W.H.; Kronstad, J.W. Iron influences the abundance of the iron regulatory protein Cir1 in the fungal pathogen Cryptococcus neoformans. FEBS Lett. 2011, 585, 3342–3347. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moreno, M.A.; Ibrahim-Granet, O.; Vicentefranqueira, R.; Amich, J.; Ave, P.; Leal, F.; Latge, J.P.; Calera, J.A. The regulation of zinc homeostasis by the ZafA transcriptional activator is essential for Aspergillus fumigatus virulence. Mol. Microbiol. 2007, 64, 1182–1197. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Eide, D.J. Zap1p, a metalloregulatory protein involved in zinc-responsive transcriptional regulation in Saccharomyces cerevisiae. Mol. Cell. Biol. 1997, 17, 5044–5052. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schneider Rde, O.; Fogaca Nde, S.; Kmetzsch, L.; Schrank, A.; Vainstein, M.H.; Staats, C.C. Zap1 regulates zinc homeostasis and modulates virulence in Cryptococcus gattii. PLoS ONE 2012, 7, e43773. [Google Scholar] [CrossRef] [PubMed]
- So, Y.S.; Lee, D.G.; Idnurm, A.; Ianiri, G.; Bahn, Y.S. The TOR Pathway Plays Pleiotropic Roles in Growth and Stress Responses of the Fungal Pathogen Cryptococcus neoformans. Genetics 2019, 212, 1241–1258. [Google Scholar] [CrossRef]
- Dichtl, K.; Samantaray, S.; Wagener, J. Cell wall integrity signalling in human pathogenic fungi. Cell Microbiol. 2016, 18, 1228–1238. [Google Scholar] [CrossRef]
- Moranova, Z.; Virtudazo, E.; Hricova, K.; Ohkusu, M.; Kawamoto, S.; Husickova, V.; Raclavsky, V. The CRZ1/SP1-like gene links survival under limited aeration, cell integrity and biofilm formation in the pathogenic yeast Cryptococcus neoformans. Biomed. Pap. Med. Fac. Univ. Palacky Olomouc 2014, 158, 212–220. [Google Scholar] [CrossRef] [Green Version]
- Karunanithi, S.; Cullen, P.J. The filamentous growth MAPK Pathway Responds to Glucose Starvation Through the Mig1/2 transcriptional repressors in Saccharomyces cerevisiae. Genetics 2012, 192, 869–887. [Google Scholar] [CrossRef] [Green Version]
- Conrad, M.; Schothorst, J.; Kankipati, H.N.; Van Zeebroeck, G.; Rubio-Texeira, M.; Thevelein, J.M. Nutrient sensing and signaling in the yeast Saccharomyces cerevisiae. FEMS Microbiol. Rev. 2014, 38, 254–299. [Google Scholar] [CrossRef] [Green Version]
- Caza, M.; Hu, G.; Price, M.; Perfect, J.R.; Kronstad, J.W. The Zinc Finger Protein Mig1 Regulates Mitochondrial Function and Azole Drug Susceptibility in the Pathogenic Fungus Cryptococcus neoformans. mSphere 2016, 1, e00080-15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Leipheimer, J.; Bloom, A.L.M.; Baumstark, T.; Panepinto, J.C. CNBP Homologues Gis2 and Znf9 Interact with a Putative G-Quadruplex-Forming 3’ Untranslated Region, Altering Polysome Association and Stress Tolerance in Cryptococcus neoformans. mSphere 2018, 3, e00201-18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kmetzsch, L.; Staats, C.C.; Simon, E.; Fonseca, F.L.; Oliveira, D.L.; Joffe, L.S.; Rodrigues, J.; Lourenco, R.F.; Gomes, S.L.; Nimrichter, L.; et al. The GATA-type transcriptional activator Gat1 regulates nitrogen uptake and metabolism in the human pathogen Cryptococcus neoformans. Fungal Genet. Biol. 2011, 48, 192–199. [Google Scholar] [CrossRef] [PubMed]
| Zinc-Binding Domain Type | Consensus Sequence | Function | Examples | References |
|---|---|---|---|---|
| C2H2 | C-X2-4-C-X12-H-X3-5-H | Transcription, nucleic acid binding | TFIIIA | [24] |
| C2HC | C-X2-C-X4-H-X4-C | Genome packaging, single-stranded nucleic binding | Retroviral nucleocapsid | [29,30,31,32,33] |
| C3H | C-X6-14-C-X4-5-C-X3-H | RNA binding | Nup475 | [26,34,35] |
| C4 | C-X2-C-X17-C-X2-C | DNA binding | GATA-1 | [36] |
| C6 | C-X2-C-X6-C-X6-C-X2-C-X6-C | DNA binding | GAL4 | [44] |
| C3HC4 | C-X2-C-X9-39-C-X1-3-H-X2-3-C-X2-C-X4-48-C-X2-C | Protein-protein interaction, nucleic acid binding | RING finger | [38,39,40,45] |
| C2HC5 | C-X2-C-X17-19-H-X2-C-X2-C-X2-C-X16-20-C-X2-3-C-C-X2-C-X17-C-X2-C | Protein-protein interaction, DNA binding | lin-11, isl-1 and mec-3 | [43] |
| C4HC3 | C-X2-C-X11-21-C-X2-C-X4-H-X2-C-X14-17-C-X2-C | Transcription, DNA binding | Requiem, ALL-1 | [46,47] |
| C8 | C-X2-C-X13-C-X2-C-X15-C-X5-C-X12-C-X4-C | Oligomerization, DNA binding | steroid-thyroid receptor | [48] |
| ZNF Type | No. of ZNFs in JEC21 | % | No. of ZNFs in H99 | % |
|---|---|---|---|---|
| C2H2 | 28 | 22.0 | 28 | 21.7 |
| C4 | 28 | 22.0 | 32 | 24.8 |
| C3HC4 | 21 | 16.5 | 23 | 17.8 |
| C3H | 13 | 10.2 | 12 | 9.3 |
| C2HC | 10 | 7.9 | 12 | 9.3 |
| C6 | 5 | 3.9 | 4 | 3.1 |
| C4HC3 | 2 | 1.6 | 1 | 0.8 |
| C2HC5 | 1 | 0.8 | 1 | 0.8 |
| Combination | 6 | 4.7 | 6 | 4.7 |
| Others | 13 | 10.2 | 10 | 7.8 |
| Total | 127 | 100 | 129 | 100 |
| Name | Gene ID | Zinc Finger Type | Description and Function | References |
|---|---|---|---|---|
| Bwc2 | CNE01220 | ZnF_C4 | Sexual filamentation and fungal virulence | [70,71] |
| Cir1 | CNAG_04864CNJ02920 | ZnF_C4 | Iron uptake, iron homeostasis, and virulence | [89,93] |
| Cqs2 | CNF00370 | Znf_C2H2 | Important component of the Qsp1-signaling cascade required for sexual reproduction | [82] |
| Crz1 | CNA01450 | Znf_C2H2 | Regulates the sporulation process | [68] |
| Crz1 | CNAG_00156 | Znf_C2H2 | Cell integrity; Proliferation and survival under reduced aeration; Biofilm formation and susceptibility to fluconazole. | [99] |
| Gat1 | CNAG_00193 | ZnF_C4 | Nitrogen uptake and metabolism | [104] |
| Gis1 | CNAG_02338 | Znf-C2HC | Sterol biosynthesis and fluconazole and oxidative stress sensitivity | [103] |
| Mig1 | CNAG_06327 | Znf_C2H2 | Mitochondrial function and azole drug susceptibility | [102] |
| sp1 | CNAG_00156 | Znf_C2H2 | Cell wall stability, nitrosative stress, extracellular capsule production, and fungal virulence | [87] |
| STE12αp | CND05810 | Znf_C2H2 | Haploid filamentation and fungal virulence | [74,75,77] |
| STE12ap | Znf_C2H2 | Capsule formation and fungal virulence; Capsule-associated genes expression; | [76,86] | |
| Zap1 | CNBG_4460 | Znf_C2H2 | Growth in zinc-limiting conditions Expression of ZRT, IRT-like protein (ZIP) zinc transporters and distinct zinc-binding proteins; Fungal virulence | [96] |
| Zfp1 | CNAG_07329 | Znf_C2H2 | Sexual reproduction and fungal virulence | [79] |
| Znf1 | CND05720 | Znf_C4HC3 | Not essential for Cryptococcus dimorphic hyphal growth | [58] |
| Znf2 | CNG02160 | Znf_C2H2 | Master activator of the yeast to hyphal transition; Key regulator for hyphal growth | [58,61,62,63] |
| Znf3 | CNK01880 | Znf_C2H2 | Hyphal development during unisexual and bisexual reproduction | [59] |
| Znf3 | CNAG_02700 | Znf_C2H2 | Link between RNAi silencing and sexual development | [65] |
| Znf9 | CNAG_01273 | Znf_C2HC | Sterol biosynthesis; Fluconazole and oxidative stress sensitivity | [103] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, Y.-H.; Liu, T.-B. Zinc Finger Proteins in the Human Fungal Pathogen Cryptococcus neoformans. Int. J. Mol. Sci. 2020, 21, 1361. https://doi.org/10.3390/ijms21041361
Li Y-H, Liu T-B. Zinc Finger Proteins in the Human Fungal Pathogen Cryptococcus neoformans. International Journal of Molecular Sciences. 2020; 21(4):1361. https://doi.org/10.3390/ijms21041361
Chicago/Turabian StyleLi, Yuan-Hong, and Tong-Bao Liu. 2020. "Zinc Finger Proteins in the Human Fungal Pathogen Cryptococcus neoformans" International Journal of Molecular Sciences 21, no. 4: 1361. https://doi.org/10.3390/ijms21041361
APA StyleLi, Y.-H., & Liu, T.-B. (2020). Zinc Finger Proteins in the Human Fungal Pathogen Cryptococcus neoformans. International Journal of Molecular Sciences, 21(4), 1361. https://doi.org/10.3390/ijms21041361






