Genome-Wide Identification, Expression Profile and Evolution Analysis of Karyopherin β Gene Family in Solanum tuberosum Group Phureja DM1-3 Reveals Its Roles in Abiotic Stresses
Abstract
1. Introduction
2. Results
2.1. Genome-Wide Identification of KPNβ Genes from S. tuberosum
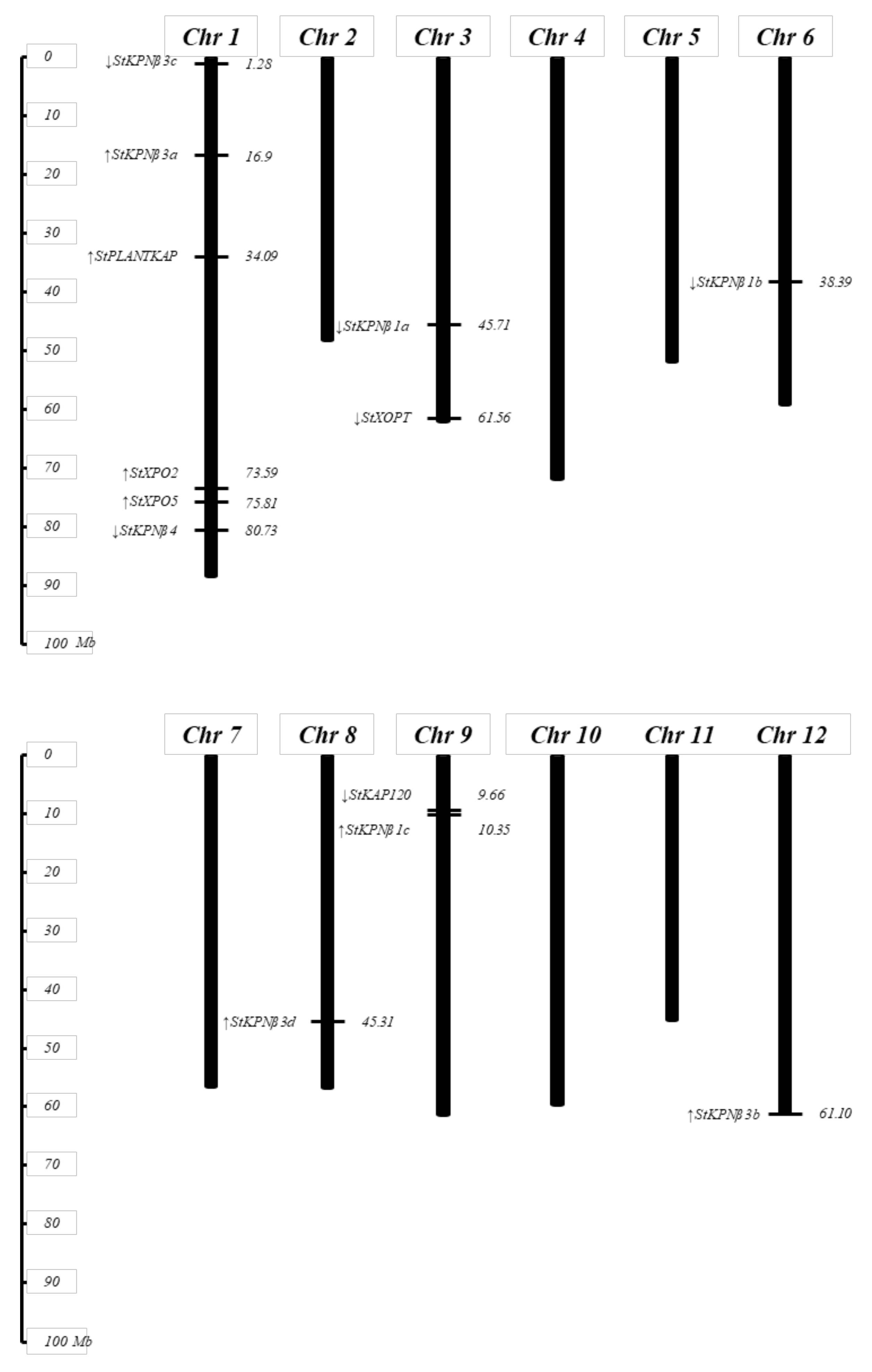
2.2. Chromosomal Distribution and Duplication Events among StKPNβ Genes
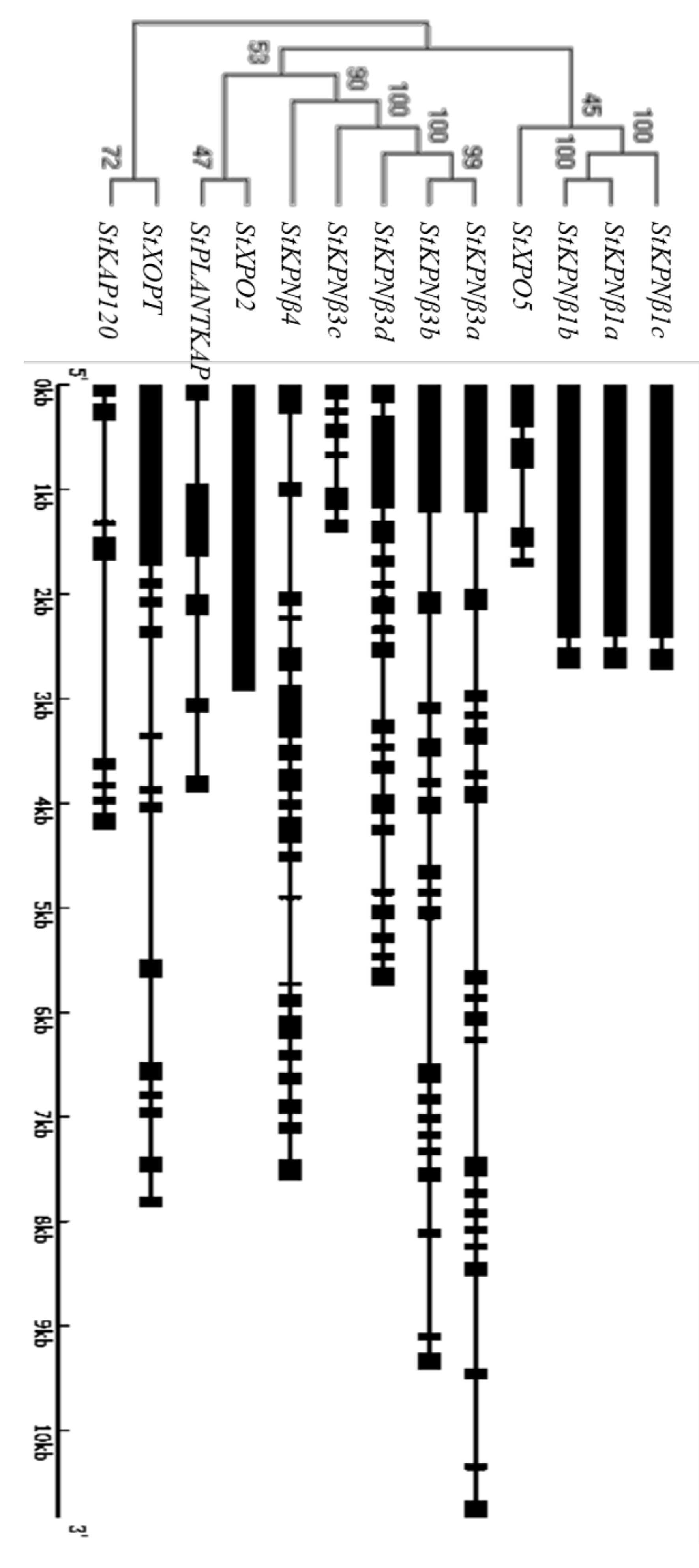
2.3. Gene Structure of StKPNβs
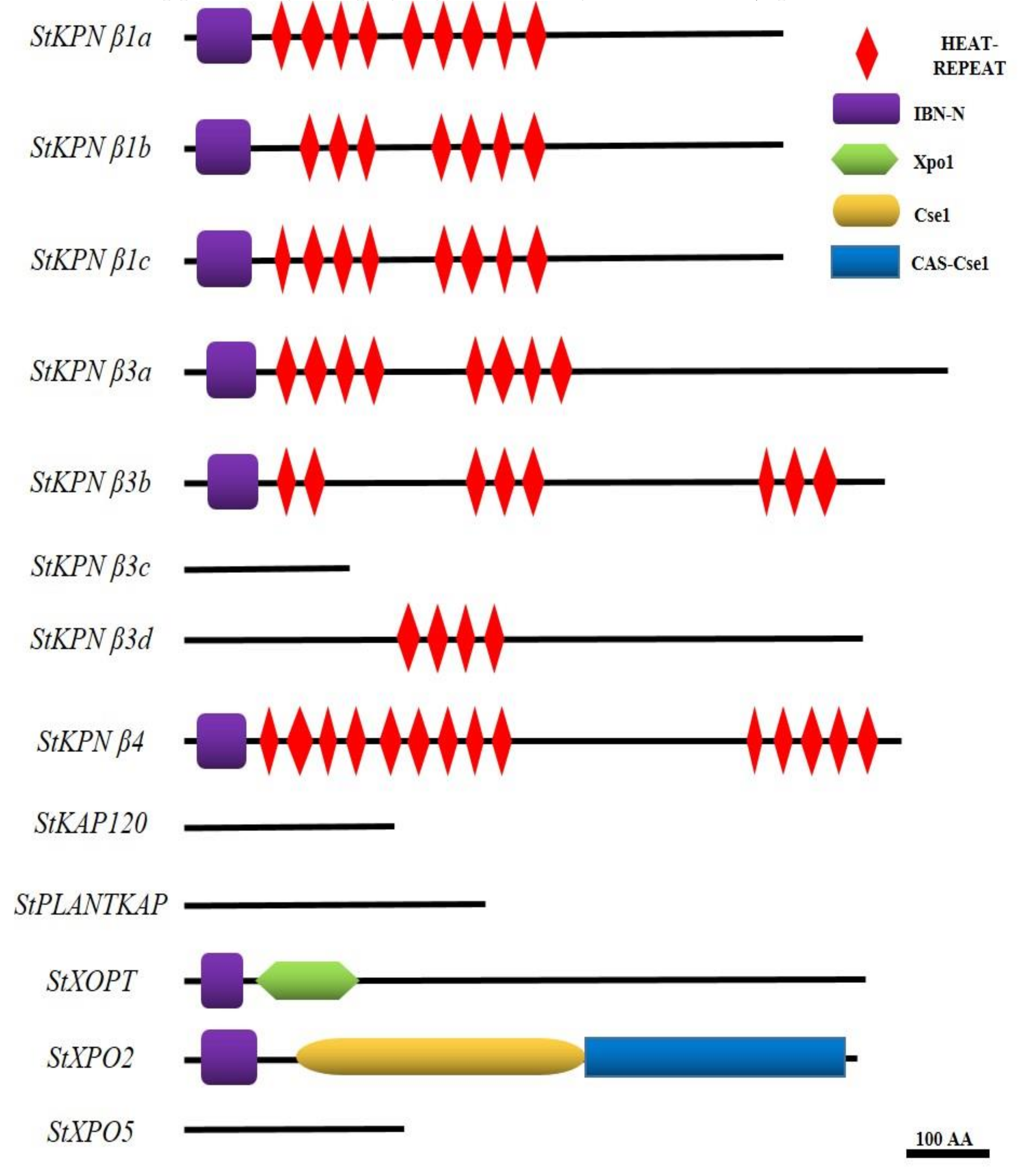
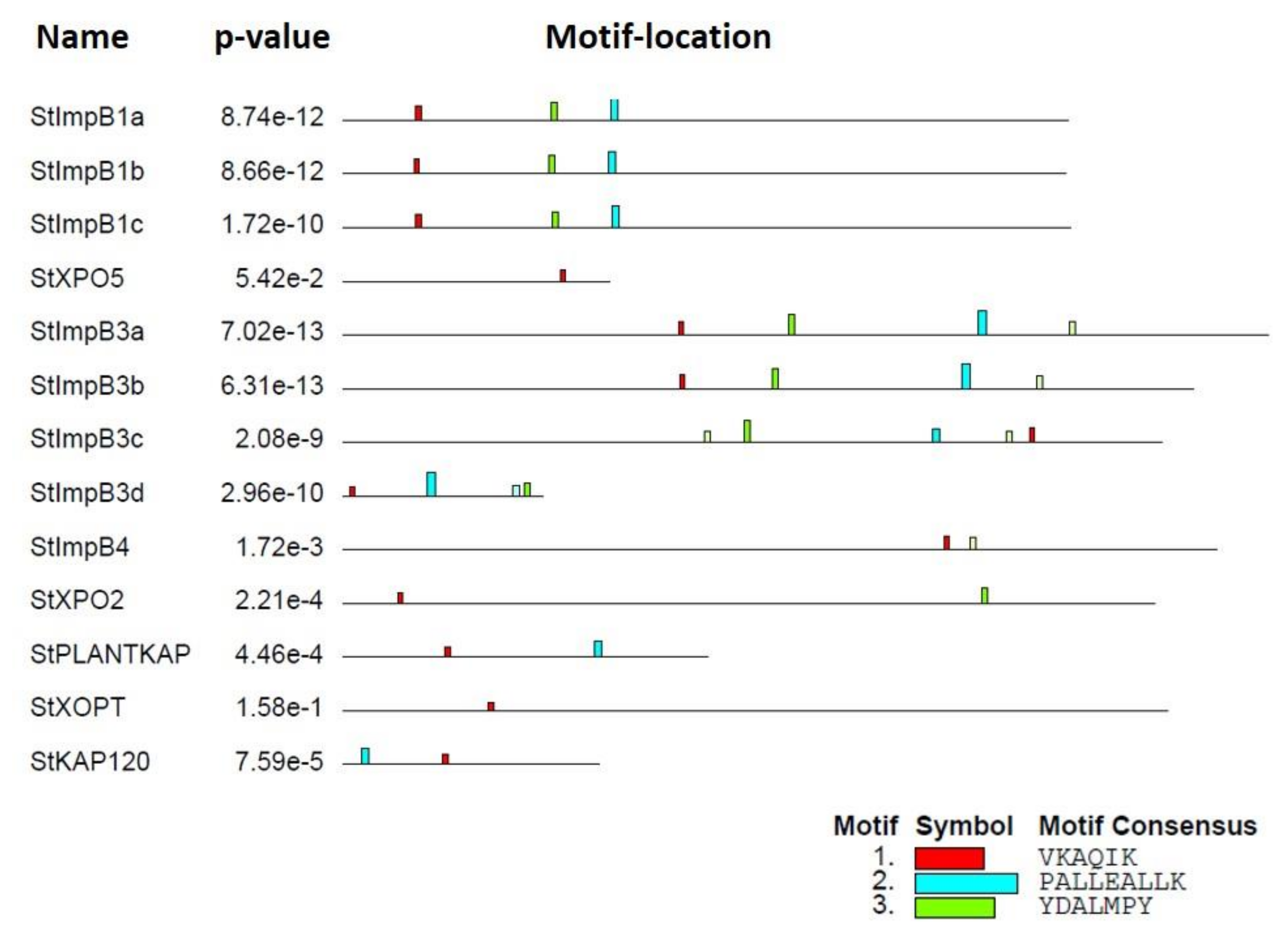
2.4. Conserved Domains and Motif Analysis of StKPNβs
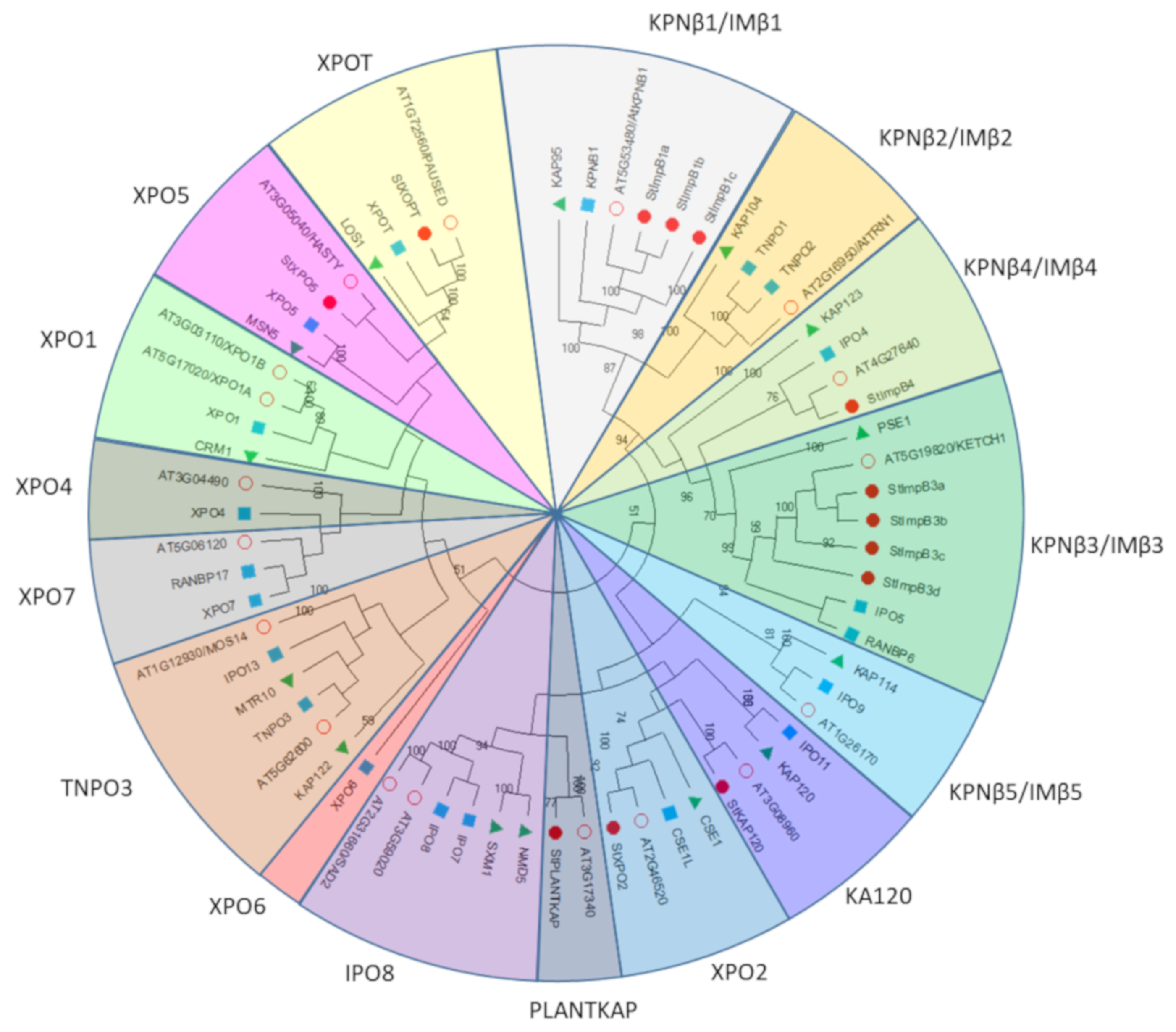
2.5. Phylogenetic Analysis of StKPNβs
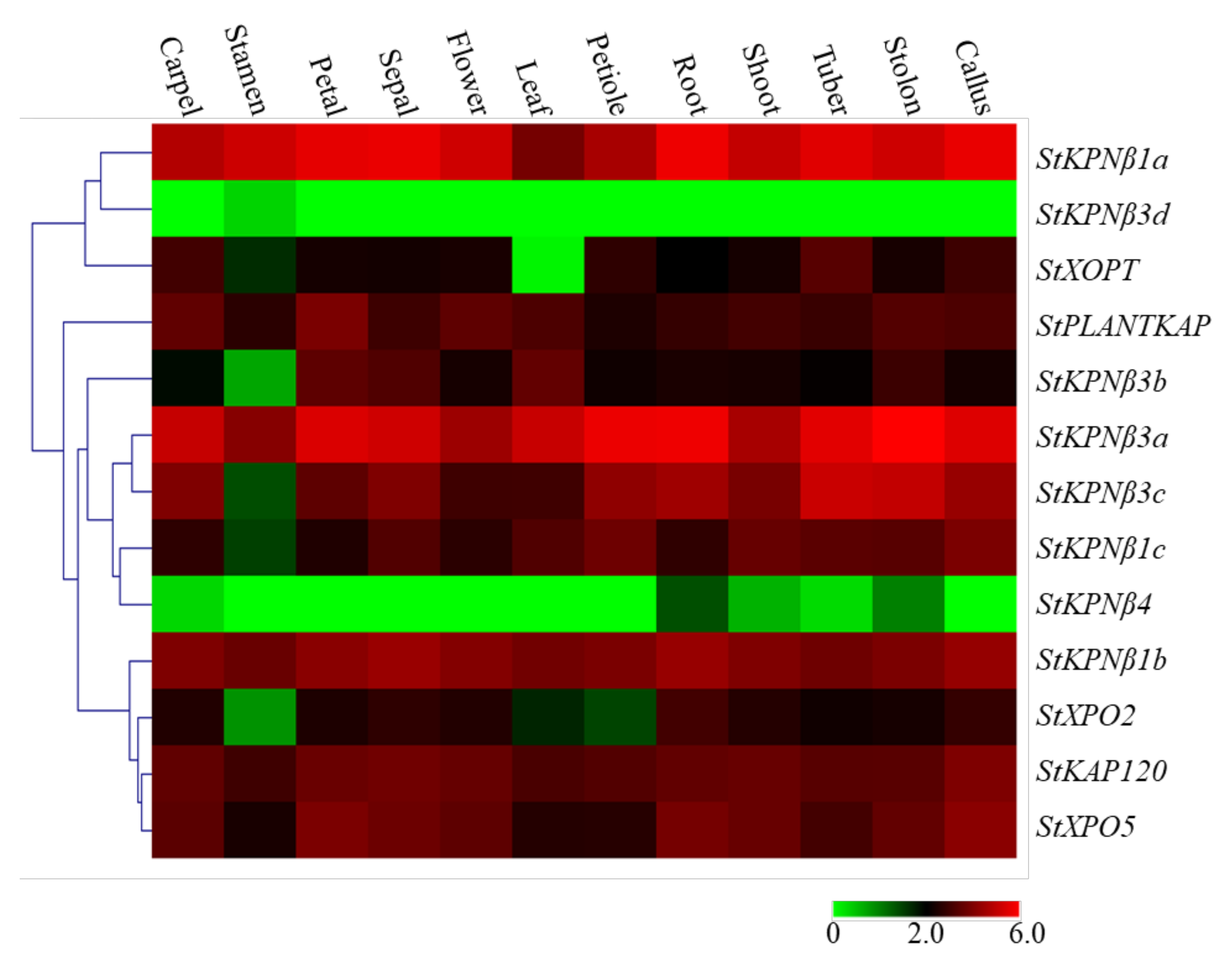
2.6. Expression Profiles of StKPNβs among Various Tissues and Developmental Stages
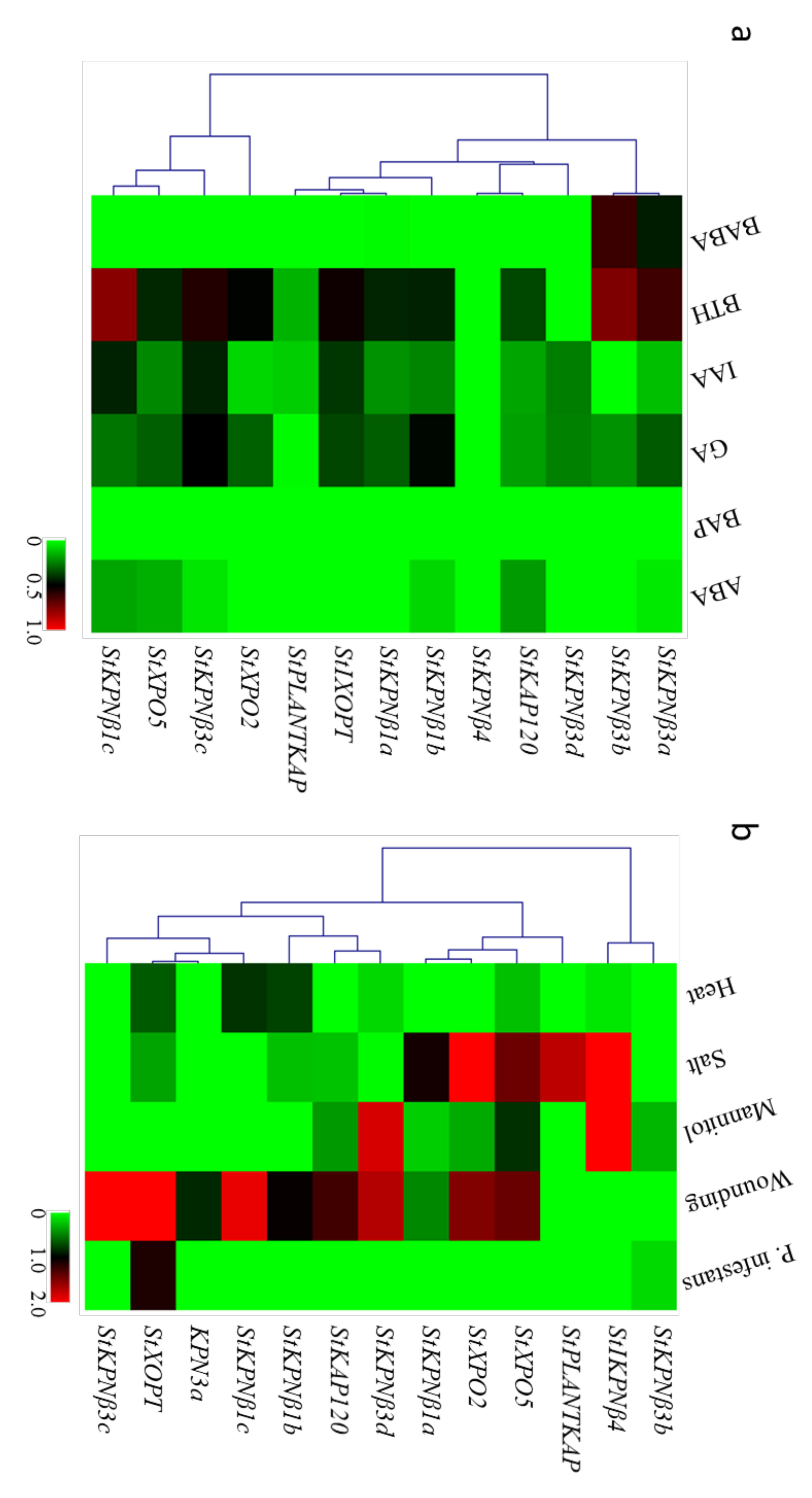
2.7. Expression Profiles of StKPNβs in Response to Biotic and Abiotic Stresses
2.8. StKPNβs Response to Various Phytohormones
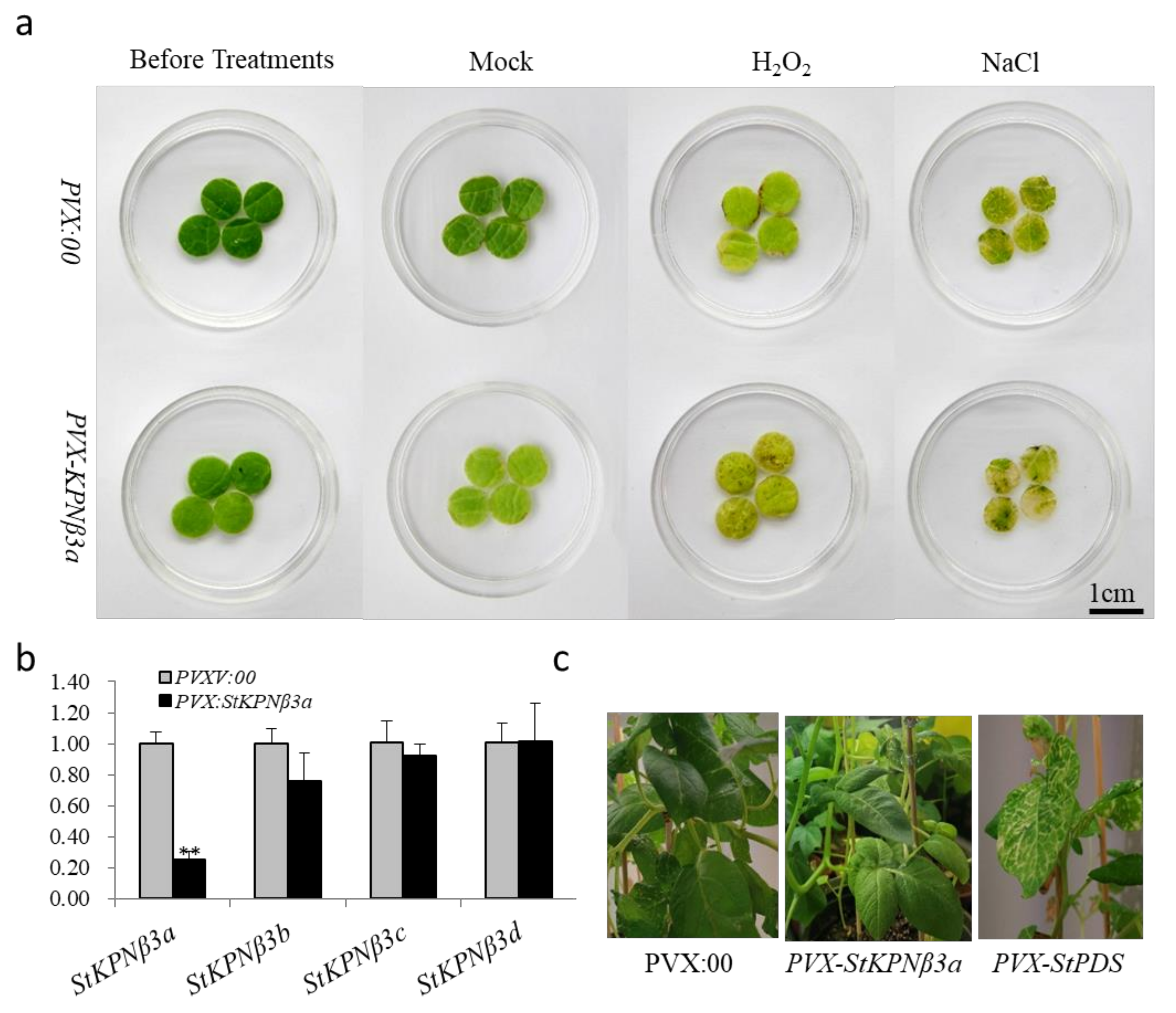
2.9. Knockdown of StKPNβ3a Expression Results in Increased Susceptibility to Environmental Stresses
3. Discussion
4. Materials and Methods
4.1. Plant Material and Treatments
4.2. Identification of KPNβ Genes in S. tuberosum Group Phureja
4.3. Analysis of Gene Structure and Conserved Domains
4.4. Sequence Alignment and Phylogenetic Construction
4.5. Expression Profiling of StKPNβ Genes in Different Tissues or Under Various Stresses
4.6. RNA Extraction and Quantitative Real-Time RT-PCR
4.7. Virus-Induced Gene Silencing (VIGS) of Potato
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Baum, D.A.; Baum, B. An inside-out origin for the eukaryotic cell. BMC Biol. 2014, 12, 76. [Google Scholar] [CrossRef] [PubMed]
- Lane, N.; Martin, W. The energetics of genome complexity. Nature 2010, 467, 929–934. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.M.; Meier, I. The nuclear pore comes to the fore. Trends Plant Sci. 2008, 13, 20–27. [Google Scholar] [CrossRef] [PubMed]
- Fahrenkrog, B.; Aebi, U. The nuclear pore complex: Nucleocytoplasmic transport and beyond. Nat. Rev. Mol. Cell Biol. 2003, 4, 757–766. [Google Scholar] [CrossRef]
- Fried, H.; Kutay, U. Nucleocytoplasmic transport: Taking an inventory. Cell Mol. Life Sci. 2003, 60, 1659–1688. [Google Scholar] [CrossRef] [PubMed]
- Gorlich, D.; Mattaj, I.W. Nucleocytoplasmic transport. Science 1996, 271, 1513–1518. [Google Scholar] [CrossRef]
- Goryaynov, A.; Yang, W. Role of molecular charge in nucleocytoplasmic transport. PLoS ONE 2014, 9, e88792. [Google Scholar] [CrossRef]
- Oka, M.; Yoneda, Y. Importin alpha: Functions as a nuclear transport factor and beyond. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 2018, 94, 259–274. [Google Scholar] [CrossRef]
- Ullman, K.S.; Powers, M.A.; Forbes, D.J. Nuclear export receptors: From importin to exportin. Cell 1997, 90, 967–970. [Google Scholar] [CrossRef]
- Yeon, S.I.; Youn, J.H.; Lim, M.H.; Lee, H.J.; Kim, Y.M.; Choi, J.E.; Lee, J.M.; Shin, J.S. Development of monoclonal antibodies against human IRF-5 and their use in identifying the binding of IRF-5 to nuclear import proteins karyopherin-alpha1 and -beta1. Yonsei Med. J. 2008, 49, 1023–1031. [Google Scholar] [CrossRef][Green Version]
- Yoneda, Y.; Hieda, M.; Nagoshi, E.; Miyamoto, Y. Nucleocytoplasmic protein transport and recycling of Ran. Cell Struct. Funct. 1999, 24, 425–433. [Google Scholar] [CrossRef] [PubMed]
- Kimura, M.; Imamoto, N. Biological significance of the importin-beta family-dependent nucleocytoplasmic transport pathways. Traffic 2014, 15, 727–748. [Google Scholar] [CrossRef] [PubMed]
- Strom, A.C.; Weis, K. Importin-beta-like nuclear transport receptors. Genome Biol. 2001, 2, REVIEWS3008. [Google Scholar] [CrossRef] [PubMed]
- Vetter, I.R.; Arndt, A.; Kutay, U.; Gorlich, D.; Wittinghofer, A. Structural view of the Ran-Importin beta interaction at 2.3 A resolution. Cell 1999, 97, 635–646. [Google Scholar] [CrossRef]
- Tamura, K.; Hara-Nishimura, I. Functional insights of nucleocytoplasmic transport in plants. Front. Plant Sci. 2014, 5, 118. [Google Scholar] [CrossRef]
- O’Reilly, A.J.; Dacks, J.B.; Field, M.C. Evolution of the karyopherin-beta family of nucleocytoplasmic transport factors; ancient origins and continued specialization. PLoS ONE 2011, 6, e19308. [Google Scholar]
- Goldfarb, D.S.; Corbett, A.H.; Mason, D.A.; Harreman, M.T.; Adam, S.A. Importin alpha: A multipurpose nuclear-transport receptor. Trends Cell Biol. 2004, 14, 505–514. [Google Scholar] [CrossRef]
- Huang, J.G.; Yang, M.; Liu, P.; Yang, G.D.; Wu, C.A.; Zheng, C.C. Genome-wide profiling of developmental, hormonal or environmental responsiveness of the nucleocytoplasmic transport receptors in Arabidopsis. Gene 2010, 451, 38–44. [Google Scholar] [CrossRef]
- Yang, Y.; Wang, W.; Chu, Z.; Zhu, J.-K.; Zhang, H. Roles of Nuclear Pores and Nucleo-cytoplasmic Trafficking in Plant Stress Responses. Front. Plant Sci. 2017, 8, 574. [Google Scholar] [CrossRef]
- Luo, Y.; Wang, Z.; Ji, H.; Fang, H.; Wang, S.; Tian, L.; Li, X. An Arabidopsis homolog of importin beta1 is required for ABA response and drought tolerance. Plant J. 2013, 75, 377–389. [Google Scholar] [CrossRef]
- Hunter, C.A.; Aukerman, M.J.; Sun, H.; Fokina, M.; Poethig, R.S. PAUSED encodes the Arabidopsis exportin-t ortholog. Plant Physiol. 2003, 132, 2135–2143. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Chen, X. PAUSED, a putative exportin-t, acts pleiotropically in Arabidopsis development but is dispensable for viability. Plant Physiol. 2003, 132, 1913–1924. [Google Scholar] [CrossRef] [PubMed]
- Musarella, C. Solanum torvum Sw. (Solanaceae): A new alien species for Europe. Genet Resour. Crop Ev. 2020, 67, 515–522. [Google Scholar] [CrossRef]
- FAOSTAT. Available online: http://www.fao.org/faostat/en/\#data/QC (accessed on 6 June 2018).
- Knapp, S.; Vorontsova, M.S.; Särkinen, T. Dichotomous keys to the species of Solanum L. (Solanaceae) in continental Africa, Madagascar (incl. the Indian Ocean islands), Macaronesia and the Cape Verde Islands. PhytoKeys 2019, 127, 39–76. [Google Scholar] [CrossRef]
- The Potato Genome Sequencing Consortium. Genome sequence and analysis of the tuber crop potato. Nature 2011, 475, 189–195. [Google Scholar] [CrossRef]
- Hawkes, J. The Potato: Evolution, Biodiversity and Tenetic Resources; Belhaven Press: London, UK, 1990. [Google Scholar]
- Aversano, R.; Contaldi, F.; Ercolano, M.R.; Grosso, V.; Iorizzo, M.; Tatino, F.; Xumerle, L.; Dal Molin, A.; Avanzato, C.; Ferrarini, A.; et al. The Solanum commersonii Genome Sequence Provides Insights into Adaptation to Stress Conditions and Genome Evolution of Wild Potato Relatives. Plant Cell 2015, 27, 954–968. [Google Scholar] [CrossRef]
- Esposito, S.; Aversano, R.; Bradeen, J.M.; Di Matteo, A.; Villano, C.; Carputo, D. Deep-sequencing of Solanum commersonii small RNA libraries reveals riboregulators involved in cold stress response. Plant Biol. (Stuttg) 2020, 22 Suppl. 1, 133–142. [Google Scholar] [CrossRef]
- Esposito, S.; Aversano, R.; D’Amelia, V.; Villano, C.; Alioto, D.; Mirouze, M.; Carputo, D. Dicer-like and RNA-dependent RNA polymerase gene family identification and annotation in the cultivated Solanum tuberosum and its wild relative S. commersonii. Planta 2018, 248, 729–743. [Google Scholar] [CrossRef]
- Massa, A.N.; Childs, K.L.; Lin, H.; Bryan, G.J.; Giuliano, G.; Buell, C.R. The transcriptome of the reference potato genome Solanum tuberosum Group Phureja clone DM1–3 516R44. PLoS ONE 2011, 6, e26801. [Google Scholar] [CrossRef]
- Harel, A.; Forbes, D.J. Importin beta: Conducting a much larger cellular symphony. Mol. Cell 2004, 16, 319–330. [Google Scholar]
- Bollman, K.M.; Aukerman, M.J.; Park, M.Y.; Hunter, C.; Berardini, T.Z.; Poethig, R.S. HASTY, the Arabidopsis ortholog of exportin 5/MSN5, regulates phase change and morphogenesis. Development 2003, 130, 1493–1504. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Ye, R.; Xin, Y.; Fang, X.; Li, C.; Shi, H.; Zhou, X.; Qi, Y. An importin beta protein negatively regulates MicroRNA activity in Arabidopsis. Plant Cell 2011, 23, 3565–3576. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; Zhang, Z.; Jing, B.; Gannon, P.; Ding, J.; Xu, F.; Li, X.; Zhang, Y. Transportin-SR Is Required for Proper Splicing of Resistance Genes and Plant Immunity. PLoS Genet. 2011, 7, e1002159. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Guo, X.; Ge, C.; Ma, Z.; Jiang, M.; Li, T.; Koiwa, H.; Yang, S.W.; Zhang, X. KETCH1 imports HYL1 to nucleus for miRNA biogenesis in Arabidopsis. Proc. Natl. Acad. Sci. USA 2017, 114, 4011–4016. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Zhan, Q.; Shi, T.; Liu, J.; Zhao, K.; Gao, Y. The nuclear transporter SAD2 plays a role in calcium- and H2 O2 -mediated cell death in Arabidopsis. Plant J. 2020, 101, 324–333. [Google Scholar] [CrossRef] [PubMed]
- Verslues, P.E.; Guo, Y.; Dong, C.H.; Ma, W.; Zhu, J.K. Mutation of SAD2, an importin beta-domain protein in Arabidopsis, alters abscisic acid sensitivity. Plant J. 2006, 47, 776–787. [Google Scholar] [CrossRef]
- Kroukamp, H.; den Haan, R.; van Wyk, N.; van Zyl, W.H. Overexpression of native PSE1 and SOD1 in Saccharomyces cerevisiae improved heterologous cellulase secretion. Appl. Energy 2013, 102, 150–156. [Google Scholar] [CrossRef]
- Makhnevych, T.; Lusk, C.P.; Anderson, A.M.; Aitchison, J.D.; Wozniak, R.W. Cell cycle regulated transport controlled by alterations in the nuclear pore complex. Cell 2003, 115, 813–823. [Google Scholar] [CrossRef]
- Ueta, R.; Fukunaka, A.; Yamaguchi-Iwai, Y. Pse1p mediates the nuclear import of the iron-responsive transcription factor Aft1p in Saccharomyces cerevisiae. J. Biol. Chem. 2003, 278, 50120–50127. [Google Scholar] [CrossRef]
- Mosammaparast, N.; Jackson, K.R.; Guo, Y.; Brame, C.J.; Shabanowitz, J.; Hunt, D.F.; Pemberton, L.F. Nuclear import of histone H2A and H2B is mediated by a network of karyopherins. J. Cell Biol. 2001, 153, 251–262. [Google Scholar] [CrossRef]
- Thiriet, C.; Hayes, J.J. Histone dynamics during transcription: Exchange of H2A/H2B dimers and H3/H4 tetramers during pol II elongation. Results Probl. Cell. Differ. 2006, 41, 77–90. [Google Scholar] [PubMed]
- Yaseen, N.R.; Blobel, G. Cloning and characterization of human karyopherin beta3. Proc. Natl. Acad. Sci. USA 1997, 94, 4451–4456. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [PubMed]
- Liu, N.; Fromm, M.; Avramova, Z. H3K27me3 and H3K4me3 chromatin environment at super-induced dehydration stress memory genes of Arabidopsis thaliana. Mol. Plant 2014, 7, 502–513. [Google Scholar] [CrossRef]
- Song, S.; Hao, L.; Zhao, P.; Xu, Y.; Zhong, N.; Zhang, H.; Liu, N. Genome-wide Identification, Expression Profiling and Evolutionary Analysis of Auxin Response Factor Gene Family in Potato (Solanum tuberosum Group Phureja). Sci Rep. 2019, 9, 1755. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Faivre-Rampant, O.; Gilroy, E.M.; Hrubikova, K.; Hein, I.; Millam, S.; Loake, G.J.; Birch, P.; Taylor, M.; Lacomme, C. Potato virus X-induced gene silencing in leaves and tubers of potato. Plant Physiol. 2004, 134, 1308–1316. [Google Scholar] [CrossRef]
- Lacomme, C.; Chapman, S. Use of potato virus X (PVX)-based vectors for gene expression and virus-induced gene silencing (VIGS). Curr. Protoc. Microbiol. 2008. [Google Scholar] [CrossRef]


| Gene Name a | Locus ID b | Predicted Proteins | Chromosomal Location c | Gene Models d | Putative Proteins e | ||||
|---|---|---|---|---|---|---|---|---|---|
| Chr | Chr_start | Chr_end | Length (aa) | pI | MW (kDa) | ||||
| StKPNβ 1a | PGSC0003DMG400018525 | PGSC0003DMP400032281 | 3 | 45710853 | 45713572 | 1 | 871 | 4.61 | 96.40 |
| StKPNβ 1b | PGSC0003DMG400019597 | PGSC0003DMP400034029 | 6 | 38386746 | 38389457 | 1 | 868 | 4.59 | 96.00 |
| StKPNβ 1c | PGSC0003DMG400026641 | PGSC0003DMP400046282 | 9 | 10350557 | 10353269 | 1 | 873 | 4.62 | 96.22 |
| StKPNβ 3a | PGSC0003DMG400015862 | PGSC0003DMP400027802 | 1 | 16901021 | 16911855 | 1 | 1111 | 4.74 | 123.08 |
| StKPNβ 3b | PGSC0003DMG401004281 | PGSC0003DMP400007618 | 12 | 61091176 | 61100591 | 1 | 1021 | 4.77 | 113.87 |
| StKPNβ 3c | PGSC0003DMG400023766 | PGSC0003DMP400055376 | 1 | 1281756 | 1283167 | 1 | 239 | 4.73 | 27.15 |
| StKPNβ 3d | PGSC0003DMG400013325 | PGSC0003DMP400051493 | 8 | 45306052 | 45311794 | 1 | 983 | 6.10 | 109.56 |
| StKPNβ 4 | PGSC0003DMG400032173 | PGSC0003DMP400041127 | 1 | 80738183 | 80745787 | 5 | 1049 | 4.87 | 115.27 |
| StKAP120 | PGSC0003DMG401000117 | PGSC0003DMP400011095 | 9 | 9658233 | 9662484 | 1 | 307 | 5.28 | 33.77 |
| StPLANTKAP | PGSC0003DMG400006259 | PGSC0003DMP400021286 | 1 | 34095216 | 34099108 | 1 | 438 | 4.22 | 49.23 |
| StXOPT | PGSC0003DMG400012034 | PGSC0003DMP400039669 | 3 | 61569820 | 61577676 | 4 | 990 | 5.44 | 111.51 |
| StXPO2 | PGSC0003DMG400022883 | PGSC0003DMP400000259 | 1 | 73593165 | 73596092 | 1 | 975 | 5.52 | 109.63 |
| StXPO5 | PGSC0003DMG400022491 | PGSC0003DMP400038992 | 1 | 75806958 | 75808695 | 1 | 320 | 6.00 | 35.01 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xu, Y.; Liu, L.; Zhao, P.; Tong, J.; Zhong, N.; Zhang, H.; Liu, N. Genome-Wide Identification, Expression Profile and Evolution Analysis of Karyopherin β Gene Family in Solanum tuberosum Group Phureja DM1-3 Reveals Its Roles in Abiotic Stresses. Int. J. Mol. Sci. 2020, 21, 931. https://doi.org/10.3390/ijms21030931
Xu Y, Liu L, Zhao P, Tong J, Zhong N, Zhang H, Liu N. Genome-Wide Identification, Expression Profile and Evolution Analysis of Karyopherin β Gene Family in Solanum tuberosum Group Phureja DM1-3 Reveals Its Roles in Abiotic Stresses. International Journal of Molecular Sciences. 2020; 21(3):931. https://doi.org/10.3390/ijms21030931
Chicago/Turabian StyleXu, Ya, Lu Liu, Pan Zhao, Jing Tong, Naiqin Zhong, Hongji Zhang, and Ning Liu. 2020. "Genome-Wide Identification, Expression Profile and Evolution Analysis of Karyopherin β Gene Family in Solanum tuberosum Group Phureja DM1-3 Reveals Its Roles in Abiotic Stresses" International Journal of Molecular Sciences 21, no. 3: 931. https://doi.org/10.3390/ijms21030931
APA StyleXu, Y., Liu, L., Zhao, P., Tong, J., Zhong, N., Zhang, H., & Liu, N. (2020). Genome-Wide Identification, Expression Profile and Evolution Analysis of Karyopherin β Gene Family in Solanum tuberosum Group Phureja DM1-3 Reveals Its Roles in Abiotic Stresses. International Journal of Molecular Sciences, 21(3), 931. https://doi.org/10.3390/ijms21030931




