Methylation: An Ineluctable Biochemical and Physiological Process Essential to the Transmission of Life
Abstract
1. Introduction
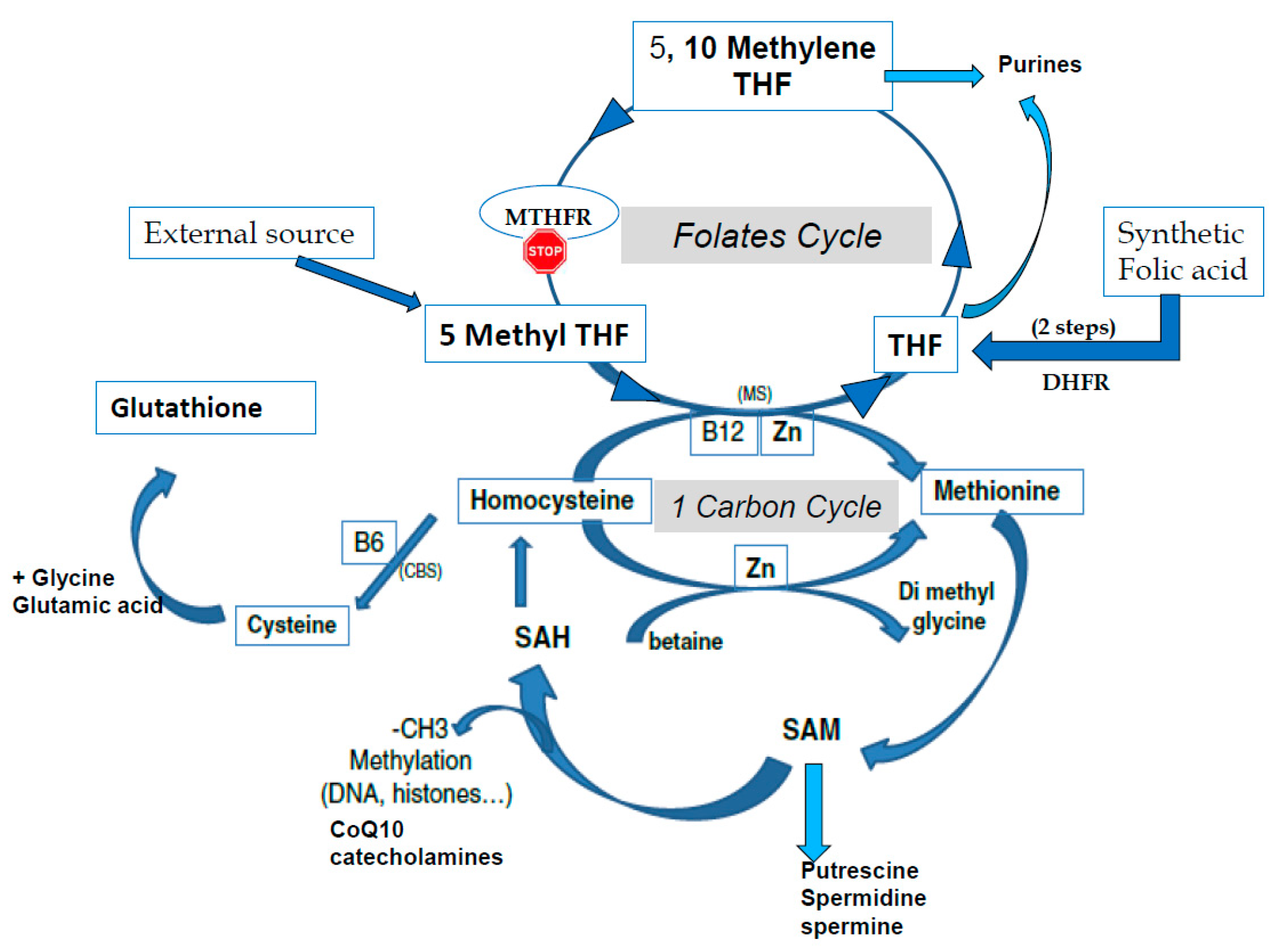
1.1. A Reminder of the Methylation Process
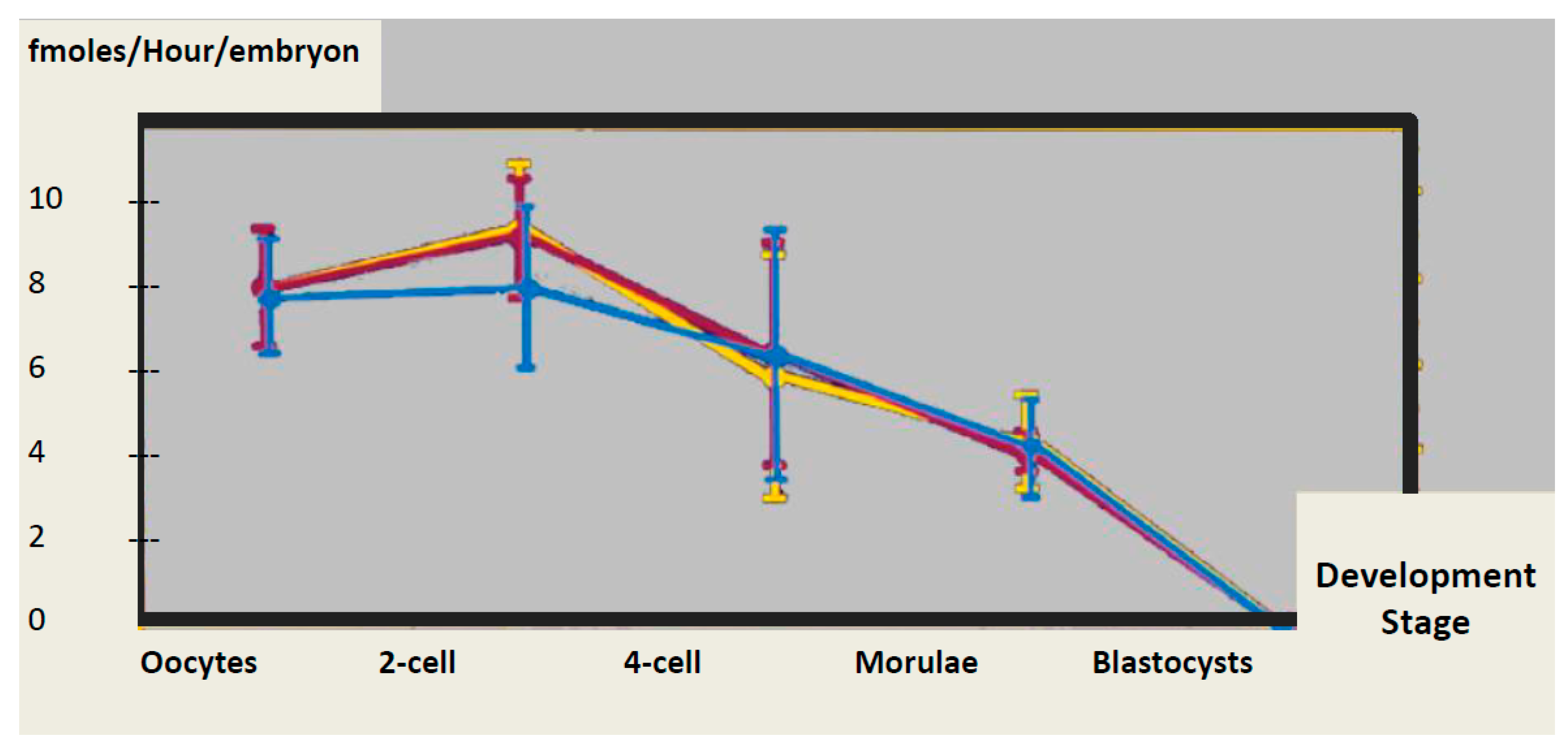
1.2. The Beginning of Life: Fertilization and Immediately Postfertilization
1.3. Problems Associated with Assisted Reproductive Technology
1.4. Blastocyst Formation, Implantation, and Placental Development
1.5. Embryo Growth, DNA Methylation, and Fetal Testis and Ovary
1.6. Methylation Errors in Gametogenesis, Postnatal Life, and Adults
1.7. Methylation Errors in the Male
1.8. Methylation in Female Gametes
2. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Berger, S.L.; Kouzarides, T.; Shiekhattar, R.; Shilatifard, A. An operational definition of epigenetics. Genes Dev. 2009, 23, 781–783. [Google Scholar] [CrossRef]
- Ménézo, Y.; Elder, K. Epigenetic remodeling of chromatin in human ART: Addressing deficiencies in culture media. J. Assist. Reprod. Genet. 2020, 37, 1781–1788. [Google Scholar] [CrossRef]
- Guérin, P.; El Mouatassim, S.; Ménézo, Y. Oxidative stress and protection against reactive oxygen species in the pre-implantation embryo and its surroundings. Hum. Reprod. Update 2001, 7, 175–189. [Google Scholar]
- Guérin, P.; Ménézo, Y. Hypotaurine and taurine in gamete and embryo environments: De novo synthesis via the cysteine sulfinic acid pathway in oviduct cells. Zygote 1995, 3, 333–337. [Google Scholar]
- Ménézo, Y.; Lichtblau, I.; Elder, K. New insights into human pre-implantation metabolism in vivo and in vitro. J. Assist. Reprod. Genet. 2013, 30, 293–303. [Google Scholar]
- Denomme, M.M.; McCallie, B.R.; Parks, J.C.; Schoolcraft, W.B.; Katz-Jaffe, M.G. Alterations in the sperm histone-retained epigenome are associated with unexplained male factor infertility and poor blastocyst development in donor oocyte IVF cycles. Hum. Reprod. 2017, 32, 2443–2450. [Google Scholar] [CrossRef] [PubMed]
- Ward, W.S. Function of sperm chromatin structural elements in fertilization and development. Mol. Hum. Reprod. 2010, 16, 30–36. [Google Scholar] [CrossRef]
- Kutchy, N.A.; Menezes, E.S.B.; Ugur, M.R.; Ul Husna, A.; ElDebaky, H.; Evans, H.C.; Beaty, E.; Santos, F.C.; Tan, W.; Wills, R.W.; et al. Sperm cellular and nuclear dynamics associated with bull fertility. Anim. Reprod. Sci. 2019, 211, 106203. [Google Scholar] [CrossRef]
- Ihara, M.; Meyer-Ficca, M.L.; Leu, N.A.; Rao, S.; Li, F.; Gregory, B.D.; Zalenskaya, I.A.; Schultz, R.M.; Meyer, R.G. Paternal poly (ADP-ribose) metabolism modulates retention of inheritable sperm histones and early embryonic gene expression. PLoS Genet. 2014, 10, e1004317. [Google Scholar] [CrossRef]
- Park, J.S.; Jeong, Y.S.; Shin, S.T.; Lee, K.K.; Kang, Y.K. Dynamic DNA methylation reprogramming: Active demethylation and immediate re-methylation in the male pronucleus of bovine zygotes. Dev. Dyn. 2007, 236, 2523–2533. [Google Scholar] [CrossRef]
- Menezo, Y.; Clement, P.; Dale, B. DNA Methylation Patterns in the Early Human Embryo and the Epigenetic/Imprinting Problems: A Plea for a More Careful Approach to Human Assisted Reproductive Technology (ART). Int. J. Mol. Sci. 2019, 20, 1342. [Google Scholar] [CrossRef] [PubMed]
- Uysal, F.; Akkoyunlu, G.; Ozturk, S. Dynamic expression of DNA methyltransferases (DNMTs) in oocytes and early embryos. Biochimie 2015, 116, 103–113. [Google Scholar] [CrossRef] [PubMed]
- Hara-Isono, K.; Matsubara, K.; Mikami, M.; Arima, T.; Ogata, T.; Fukami, M.; Kagami, M. Assisted reproductive technology represents a possible risk factor for development of epimutation-mediated imprinting disorders for mothers aged ≥30 years. Clin. Epigenet. 2020, 12, 111. [Google Scholar] [CrossRef] [PubMed]
- Enciso, M.; Sarasa, J.; Xanthopoulou, L.; Bristow, S.; Bowles, M.F.; Fragouli, E.; Delhanty, J.; Wells, D. Polymorphisms in the MTHFR gene influence embryo viability and the incidence of aneuploidy. Hum. Genet. 2016, 135, 555–568. [Google Scholar] [CrossRef] [PubMed]
- Servy, E.J.; Jacquesson-Fournols, L.; Cohen, M.; Menezo, Y. MTHFR isoform carriers. 5-MTHF (5-methyl tetrahydrofolate) vs. folic acid: A key to pregnancy outcome: A case series. J. Assist. Reprod. Genet. 2018, 35, 1431–1435. [Google Scholar] [CrossRef] [PubMed]
- Ishitan, H.; Ikeda, S.; Egashira, K.; Sugimoto, M.; Kume, S.; Minami, N.; Ohta, T.J. Embryonic MTHFR contributes to blastocyst development. J. Assist. Reprod. Genet. 2020, 37, 1807–1814. [Google Scholar] [CrossRef]
- Marshall, K.L.; Rivera, R.M. The effects of superovulation and reproductive aging on the epigenome of the oocyte and embryo. Mol. Reprod. Dev. 2018, 85, 90–105. [Google Scholar] [CrossRef]
- Hiura, H.; Okae, H.; Miyauchi, N.; Sato, F.; Sato, A.; Van de Pette, M.; John, R.M.; Kagami, M.; Nakai, K.; Soejima, H.; et al. Characterization of DNA methylation errors in patients with disorders conceived by assisted reproduction technologies. Hum. Reprod. 2012, 8, 2541–2548. [Google Scholar] [CrossRef]
- Hattori, H.; Hiura, H.; Kitamura, A.; Miyauchi, N.; Kobayashi, N.; Takahashi, S.; Okae, H.; Kyono, K.; Kagami, M.; Ogat, T.; et al. Association of four imprinting disorders and ART. Clin. Epigenet. 2019, 11, 21. [Google Scholar] [CrossRef]
- Choux, C.; Binquet, C.; Carmignac, V.; Bruno, C.; Chapusot, C.; Barberet, J.; Lamotte, M.; Sagot, P.; Bourc’his, D.; Fauque, P. The epigenetic control of transposable elements and imprinted genes in newborns is affected by the mode of conception: ART versus spontaneous conception without underlying infertility. Hum. Reprod. 2018, 33, 331–340. [Google Scholar] [CrossRef]
- Senapati, S.; Wang, F.; Ord, T.; Coutifaris, C.; Feng, R.; Mainigi, M. Superovulation alters the expression of endometrial genes critical to tissue remodeling and placentation. J. Assist. Reprod. Genet. 2018, 35, 1799–1808. [Google Scholar] [CrossRef] [PubMed]
- Smith, Z.D.; Chan, M.M.; Humm, K.C.; Karnik, R.; Mekhoubad, S.; Regev, A.; Eggan, K.; Meissner, A. DNA methylation dynamics of the human pre-implantation embryo. Nature 2014, 511, 611–615. [Google Scholar] [CrossRef] [PubMed]
- Luke, B.; Brown, M.B.; Nichols, H.B.; Schymura, M.J.; Browne, M.L.; Fisher, S.C.; Forestieri, N.E.; Rao, C.; Yazdy, M.M.; Gershman, S.T.; et al. Assessment of Birth Defects and Cancer Risk in Children Conceived via In Vitro Fertilization in the US. JAMA Netw. Open 2020, 3, e2022927. [Google Scholar] [CrossRef] [PubMed]
- Market-Velker, B.A.; Fernandes, A.D.; Mann, M.R. Side-by-side comparison of five commercial media systems in a mouse model: Suboptimal in vitro culture interferes with imprint maintenance. Biol. Reprod. 2010, 83, 938–950. [Google Scholar] [CrossRef]
- Berker, B.; Kaya, C.; Aytac, R.; Satiroglu, H. Homocysteine concentrations in follicular fluid are associated with poor oocyte and embryo qualities in polycystic ovary syndrome patients undergoing assisted reproduction. Hum. Reprod. 2009, 24, 2293–2302. [Google Scholar] [CrossRef]
- Menezo, Y.; Khatchadourian, C.; Gharib, A.; Hamidi, J.; Greenland, T.; Sarda, N. Regulation of S-adenosyl methionine synthesis in the mouse embryo. Life Sci. 1989, 44, 1601–1609. [Google Scholar] [CrossRef]
- Morbeck, D.E.; Krisher, R.L.; Herrick, J.R.; Baumann, N.A.; Matern, D.; Moyer, T. Composition of commercial media used for human embryo culture. Fertil. Steril. 2014, 102, 749–766. [Google Scholar] [CrossRef]
- Martın-Romero, F.J.; Miguel-Lasobras, E.M.; Domınguez-Arroyo, J.A.; Gonzalez-Carrera, E.; Alvarez, L.S. Contribution of culture media to oxidative stress and its effect on human oocytes. Reprod. Biomed. Online 2008, 17, 652–661. [Google Scholar] [CrossRef]
- Okamoto, Y.; Yoshida, N.; Suzuki, T.; Shimozawa, N.; Asami, M.; Matsuda, T.; Kojima, N.; Perry, A.C.; Takada, T. DNA methylation dynamics in mouse preimplantation embryos revealed by mass spectrometry. Sci. Rep. 2016, 6, 19134. [Google Scholar] [CrossRef]
- de Waal, E.; Mak, W.; Calhoun, S.; Stein, P.; Ord, T.; Krapp, C.; Coutifaris, C.; Schultz, R.M.; Bartolomei, M.S. In vitro culture increases the frequency of stochastic epigenetic errors at imprinted genes in placental tissues from mouse concepti produced through assisted reproductive technologies. Biol. Reprod. 2014, 90, 22. [Google Scholar] [CrossRef]
- Chen, S.; Wang, J.; Wang, M.; Lu, J.; Cai, Y.; Li, B. In vitro fertilization alters phospholipid profiles in mouse placenta. J. Assist. Reprod. Genet. 2019, 36, 557–567. [Google Scholar] [CrossRef] [PubMed]
- Croteau, S.; Menezo, Y. Methylation in fertilised and parthenogenetic preimplantation mouse embryos. Zygote 1994, 2, 47–52. [Google Scholar] [CrossRef] [PubMed]
- Branco, M.R.; King, M.; Perez-Garcia, V.; Bogutz, A.B.; Caley, M.; Fineberg, E.; Fineberg, E.; Lefebvre, L.; Cook, S.J.; Dean, W.; et al. Maternal DNA methylation regulates early trophoblast development. Dev. Cell 2016, 36, 152–163. [Google Scholar] [CrossRef] [PubMed]
- Georgiades, P.; Watkins, M.; Burton, G.J.; Ferguson-Smith, A.C. Rolesfor genomic imprinting and the zygotic genome in placental development. Proc. Natl. Acad. Sci. USA 2001, 98, 4522–4527. [Google Scholar] [CrossRef]
- Serman, L.; Dodig, D. Impact of DNA methylation on trophoblast function. Clin. Epigenet. 2011, 3, 7. [Google Scholar] [CrossRef]
- Koukoura, O.; Sifakis, S.; Spandidos, D.A. DNA methylation in the human placenta and fetal growth. Mol. Med. Rep. 2012, 5, 883–889. [Google Scholar] [CrossRef]
- Koukoura, O.; Sifakis, S.; Soufla, G.; Zaravinos, A.; Apostolidou, S.; Jones, A.; Widschwendter, M.; Spandidos, D.A. Loss of imprinting and aberrant methylation of IGF2 in placentas from pregnancies complicated with fetal growth restriction. Int. J. Mol. Med. 2011, 28, 481–487. [Google Scholar]
- Novakovic, B.; Yuen, R.K.; Gordon, L.; Penaherrera, M.S.; Sharkey, A.; Moffett, A.; Craig, J.M.; Robinson, W.P.; Saffery, R. Evidence for widespread changes in promoter methylation profile in human placenta in response to increasing gestational age and environmental/stochastic factors. BMC Genom. 2011, 12, 529. [Google Scholar] [CrossRef]
- Dasarathy, J.; Gruca, L.L.; Bennett, C.; Parimi, P.S.; Duenas, C.; Marczewski, S.; Fierro, J.L.; Kalhan, S.C. Methionine metabolism in human pregnancy. Am. J. Clin. Nutr. 2010, 91, 357–365. [Google Scholar] [CrossRef]
- Kamrani, A.; Alipourfard, I.; Ahmadi-Khiavi, H.; Yousefi, M.; Rostamzadeh, D.; Izadi, M.; Ahmadi, M. The role of epigenetic changes in preeclampsia. Biofactors 2019, 45, 712–724. [Google Scholar] [CrossRef]
- Yeung, K.R.; Chiu, C.L.; Pidsley, R.; Makris, A.; Hennessy, A.; Lind, J.M. DNA methylation profiles in preeclampsia and healthy control placentas. Am. J. Physiol. Heart Circ. Physiol. 2016, 310, H1295–H1303. [Google Scholar] [CrossRef] [PubMed]
- Ruano, C.S.M.; Méhats, C.; Miralles, F.; Vaiman, D. The Role of Epigenetics in Placental Development and the Etiology of Preeclampsia. Int. J. Mol. Sci. 2019, 20, 2837. [Google Scholar]
- Hoffman, M. Hypothesis: Hyperhomocysteinemia is an indicator of oxidant stress. Med. Hypothesis 2011, 77, 1088–1093. [Google Scholar] [CrossRef] [PubMed]
- Lorente-Pozo, S.; Parra-Llorca, A.; Núñez-Ramiro, A.; Cernada, M.; Hervás, D.; Boronat, N.; Sandoval, J.; Vento, M.J. The Oxygen Load Supplied during Delivery Room Stabilization of Preterm Infants Modifies the DNA Methylation. Profile Pediatr. 2018, 202, 70–76.e2. [Google Scholar] [CrossRef] [PubMed]
- Bianco-Miotto, T.; Mayne, B.T.; Buckberry, S.; Breen, J.; Rodriguez Lopez, C.M.; Roberts, C.T. Recent progress towards understanding the role of DNA methylation in human placental development. Reproduction 2016, 152, R23–R30. [Google Scholar] [CrossRef]
- Yuen, R.K.; Avila, L.; Peñaherrera, M.S.; von Dadelszen, P.; Lefebvre, L.; Kobor, M.S.; Robinson, W.P. Human placental-specific epipolymorphism and its association with adverse pregnancy outcomes. PLoS ONE 2009, 4, e7389. [Google Scholar] [CrossRef][Green Version]
- Manikkam, M.; Tracey, R.; Guerrero-Bosagna, C.; Skinner, M.K. Plastics derived endocrine disruptors (BPA, DEHP and DBP) induce epigenetic transgenerational inheritance of obesity, reproductive disease and sperm epimutations. PLoS ONE 2013, 8, e55387. [Google Scholar] [CrossRef]
- Skinner, M.K. Environment, epigenetics and reproduction. Mol. Cell. Endocrinol. 2014, 398, 1–3. [Google Scholar] [CrossRef]
- Skinner, M.K. Endocrine disruptor induction of epigenetic transgenerational inheritance of disease. Mol. Cell. Endocrinol. 2014, 398, 4–12. [Google Scholar] [CrossRef]
- Lane, M.; McPherson, N.O.; Fullston, T.; Spillane, M.; Sandeman, L.; Kang, W.X.; Zander-Fox, D.L. Oxidative stress in mouse sperm impairs embryo development, fetal growth and alters adiposity and glucose regulation in female offspring. PLoS ONE 2014, 9, e100832. [Google Scholar] [CrossRef]
- Dolinoy, D.C.; Huang, D.; Jirtle, R. Maternal nutrient supplementation counteracts bisphenol A-induced DNA hypomethylation in early development. Proc. Natl. Acad. Sci. USA 2007, 104, 13056–13061. [Google Scholar] [CrossRef]
- Cooney, C.A.; Dave, A.A.; Wolff, G.L. Maternal methyl supplements in mice affect epigenetic variation and DNA methylation of offspring. J. Nutr. 2002, 32 (Suppl. 8), 2393S–2400S. [Google Scholar] [CrossRef]
- Dattilo, M.; D’Amato, G.; Caroppo, E.; Menezo, Y. Improvement of gamete quality by stimulating and feeding the endogenous antioxidant system: Mechanisms, clinical results, insights on gene-environment interactions and the role of diet. J. Assist. Reprod. Genet. 2016, 33, 1633–1648. [Google Scholar] [CrossRef]
- Bonde, J.P.; Flachs, E.M.; Rimborg, S.; Glazer, C.H.; Giwercman, A.; Ramlau-Hansen, C.H.; Bräuner, E.V. The epidemiologic evidence linking prenatal and postnatal exposure to endocrine disrupting chemicals with male reproductive disorders: A systematic review and meta-analysis. Hum. Reprod. Update 2016, 23, 104–125. [Google Scholar] [CrossRef]
- Menezo, Y.; Dale, B.; Elder, K. The negative impact of the environment on methylation/epigenetic marking in gametes and embryos: A plea for action to protect the fertility of future generation’s. Mol. Reprod. Dev. 2019, 86, 1273–1282. [Google Scholar] [CrossRef]
- Menezo, Y.J.; Silvestris, E.; Dale, B.; Elder, K. Oxidative stress and alterations in DNA methylation: Two sides of the same coin in reproduction. Reprod. Biomed. Online 2016, 33, 668–683. [Google Scholar] [CrossRef]
- Menezo, Y.; Dale, B.; Cohen, M. DNA damage and repair in human oocytes and embryos: A review. Zygote 2010, 18, 357–365. [Google Scholar] [CrossRef]
- Yao, C.; Liu, Y.; Sun, M.; Niu, M.; Yuan, Q.; Hai, Y.; Guo, Y.; Chen, Z.; Hou, J.; Liu, Y.; et al. MicroRNAs and DNA methylation as epigenetic regulators of mitosis, meiosis and spermiogenesis. Reproduction 2015, 150, R25–R34. [Google Scholar] [CrossRef]
- Aston, K.I.; Uren, P.J.; Jenkins, T.G.; Horsager, A.; Cairns, B.R.; Smith, A.D.; Carrel, D.T. Aberrant sperm DNA methylation predicts male fertility status and embryo quality. Fertil. Steril. 2015, 104, 1388–1397. [Google Scholar] [CrossRef]
- Illum, L.R.H.; Bak, S.T.; Lund, S.; Nielsen, A.L.J. DNA methylation in epigenetic inheritance of metabolic diseases through the male germ line. Mol. Endocrinol. 2018, 60, R39–R56. [Google Scholar] [CrossRef]
- McSwiggin, H.M.; O’Doherty, A.M. Epigenetic reprogramming during spermatogenesis and male factor infertility. Reproduction 2018, 156, R9–R21. [Google Scholar] [CrossRef] [PubMed]
- Jenkins, T.G.; Aston, K.I.; Pflueger, C.; Cairns, B.R.; Carrell, D.T. Age associated sperm DNA methylation alterations: Possible implications in offspring disease susceptibility. PLoS Genet. 2014, 10, e1004458. [Google Scholar] [CrossRef] [PubMed]
- Ou, X.H.; Zhu, C.C.; Sun, S.C. Effects of obesity and diabetes on the epigenetic modification of mammalian gametes. Cell Physiol. 2019, 234, 7847–7855. [Google Scholar] [CrossRef] [PubMed]
- Jaroudi, S.; Kakourou, G.; Cawood, S.; Doshi, A.; Ranieri, D.M.; Serhal, P.; Harper, J.C.; SenGupta, S.B. Expression profiling of DNA repair genes in human oocytes and blastocysts using microarrays. Hum. Reprod. 2009, 24, 2649–2655. [Google Scholar] [CrossRef] [PubMed]
- Wyck, S.; Herrera, C.; Requena, C.E.; Bittner, L.; Hajkova, P.; Bollwein, H.; Santoro, R. Oxidative stress in sperm affects the epigenetic reprogramming in early embryonic development. Epigenet. Chromatin 2018, 11, 60. [Google Scholar] [CrossRef]
- Ben Maamar, M.; Nilsson, E.; Sadler-Riggleman, I.; Beck, D.; McCarrey, J.R.; Skinner, M.K. Developmental origins of transgenerational sperm DNA methylation epimutations following ancestral DDT exposure. Dev. Biol. 2019, 445, 280–293. [Google Scholar] [CrossRef]
- Jacquesson-Fournols, L.; Alvarez, S.; Cohen, M.; Clement, P.; Menezo, Y.J. A paternal effect of MTHFR SNPs on gametes and embryos should not be overlooked: Case reports. J. Assist. Reprod. Genet. 2019, 36, 1351–1353. [Google Scholar] [CrossRef]
- El Aarabi, M.; Christensen, K.E.; Chan, D.; Leclerc, D.; Landry, M.; Ly, L.; Rozen, R.; Trasler, J. Testicular MTHFR deficiency may explain sperm DNA hypomethylation associated with high dose folic acid supplementation. Hum. Mol. Genet. 2018, 27, 1123–1135. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Wu, Y.; Yang, P.J. High glucose-induced oxidative stress represses sirtuin deacetylase expression and increases histone acetylation leading to neural tube defects. J. Neurochem. 2016, 137, 371–383. [Google Scholar] [CrossRef]
- Fan, L.H.; Wang, Z.B.; Li, Q.N.; Meng, T.G.; Dong, M.Z.; Hou, Y.; Ouyang, Y.C.; Schatten, H.; Sun, Q.Y. Absence of mitochondrial DNA methylation in mouse oocyte maturation, aging and early embryo development. Biochem. Biophys. Res. Commun. 2019, 513, 912–918. [Google Scholar] [CrossRef]
- Yu, B.; Dong, X.; Gravina, S.; Kartal, Ö.; Schimmel, T.; Cohen, J.; Tortoriello, D.; Zody, R.; Hawkins, R.D.; Vijg, J. Genome-wide, Single-Cell DNA Methylomics Reveals Increased Non-CpG Methylation during Human Oocyte Maturation. Stem Cell Rep. 2017, 9, 397–407. [Google Scholar] [CrossRef]
- Franzago, M.; La Rovere, M.; Franchi, P.G.; Vitacolonna, E.; Stuppia, L. Epigenetics and human reproduction: The primary prevention of the non-communicable diseases. Epigenomics 2019, 11, 1441–1460. [Google Scholar] [CrossRef]
- Ebisch, I.M.; Thomas, C.M.; Peters, W.H.M.; Braat, D.D.M.; Steegers-Theunissen, R.P.M. The importance of folate, zinc and antioxidants in the pathogenesis and prevention of subfertility. Hum. Reprod. Update 2007, 13, 163–174. [Google Scholar] [CrossRef]
- Ziv-Gal, A.; Flaws, J.A. Evidence for bisphenol A–induced female infertility: A review (2007–2016). Fertil. Steril. 2016, 106, 827–856. [Google Scholar] [CrossRef]
- Lopes, S.; Jurisicova, A.; Casper, R.F. Gamete-specific DNA fragmentation in unfertilized human oocytes after intracytoplasmic sperm injection. Hum. Reprod. 1998, 13, 703–708. [Google Scholar] [CrossRef]
- Vázquez-Martínez, E.R.; Gómez-Viais, Y.I.; García-Gómez, E.; Reyes-Mayoral, C.; Reyes-Muñoz, E.; Camacho-Arroyo, I.; Cerbón, M. DNA methylation in the pathogenesis of polycystic ovary syndrome. Reproduction 2019, 158, R27–R40. [Google Scholar] [CrossRef]
- Concha, C.F.; Sir, P.T.; Recabarren, S.E.; Pérez, B.F. Epigenetics of polycystic ovary syndrome. Rev. Med. Chil. 2017, 145, 907–915. [Google Scholar]
- Koukoura, O.; Sifakis, S.; Spandidos, D.A. DNA methylation in endometriosis (Review). Mol. Med. Rep. 2016, 13, 2939–2948. [Google Scholar] [CrossRef] [PubMed]
- Greathouse, K.L.; Bredfeldt, T.; Everitt, J.I.; Lin, K.; Berry, T.; Kannan, K.; Mittelstadt, M.L.; Ho, S.; Walker, C.L. Environmental estrogens differentially engage the histone methyltransferase EZH2 to increase risk of uterine tumorigenesis. Mol. Cancer Res. 2012, 10, 546–557. [Google Scholar] [CrossRef]
- Khanna, P.; Ong, C.; Bay, B.H.; Baeg, G.H. Nanotoxicity: An Interplay of Oxidative Stress, Inflammation and Cell Death. Nanomaterials 2015, 5, 1163–1180. [Google Scholar] [CrossRef]
- Bartosch, C.; Lopes, J.M.; Jerónimo, C. Epigenetics in endometrial carcinogenesis—Part 1: DNA methylation. Epigenomics 2017, 9, 737–755. [Google Scholar] [CrossRef] [PubMed]
- Natanzon, Y.; Goode, E.L.; Cunningham, J.M. Epigenetics in ovarian cancer. Semin. Cancer. Biol. 2018, 51, 160–169. [Google Scholar] [CrossRef]
- Kalhan, S.C. One carbon metabolism in pregnancy: Impact on maternal, fetal and neonatal health. Mol. Cell. Endocrinol. 2016, 435, 48–60. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Menezo, Y.; Clement, P.; Clement, A.; Elder, K. Methylation: An Ineluctable Biochemical and Physiological Process Essential to the Transmission of Life. Int. J. Mol. Sci. 2020, 21, 9311. https://doi.org/10.3390/ijms21239311
Menezo Y, Clement P, Clement A, Elder K. Methylation: An Ineluctable Biochemical and Physiological Process Essential to the Transmission of Life. International Journal of Molecular Sciences. 2020; 21(23):9311. https://doi.org/10.3390/ijms21239311
Chicago/Turabian StyleMenezo, Yves, Patrice Clement, Arthur Clement, and Kay Elder. 2020. "Methylation: An Ineluctable Biochemical and Physiological Process Essential to the Transmission of Life" International Journal of Molecular Sciences 21, no. 23: 9311. https://doi.org/10.3390/ijms21239311
APA StyleMenezo, Y., Clement, P., Clement, A., & Elder, K. (2020). Methylation: An Ineluctable Biochemical and Physiological Process Essential to the Transmission of Life. International Journal of Molecular Sciences, 21(23), 9311. https://doi.org/10.3390/ijms21239311





