SIRT1 Expression and Regulation in the Primate Testis
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Design
2.2. Tissue Samples of Nonprimate and Non-Human Primates
2.3. Immunohistochemistry on Sectioned Tissues
2.4. Extraction and Quantification of RNA
2.5. Statistics
3. Results
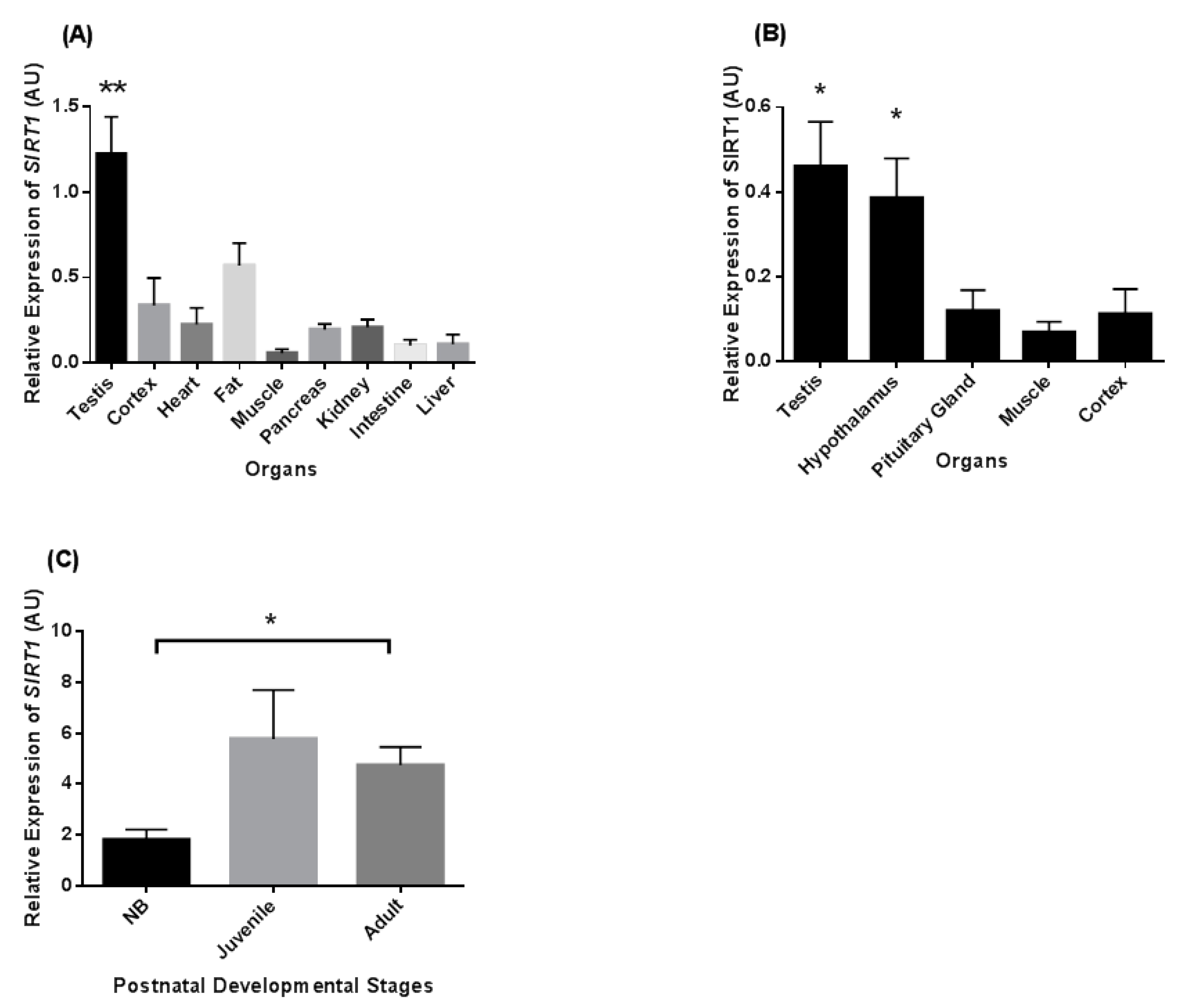
3.1. Tissue Distribution of SIRT1 Transcripts
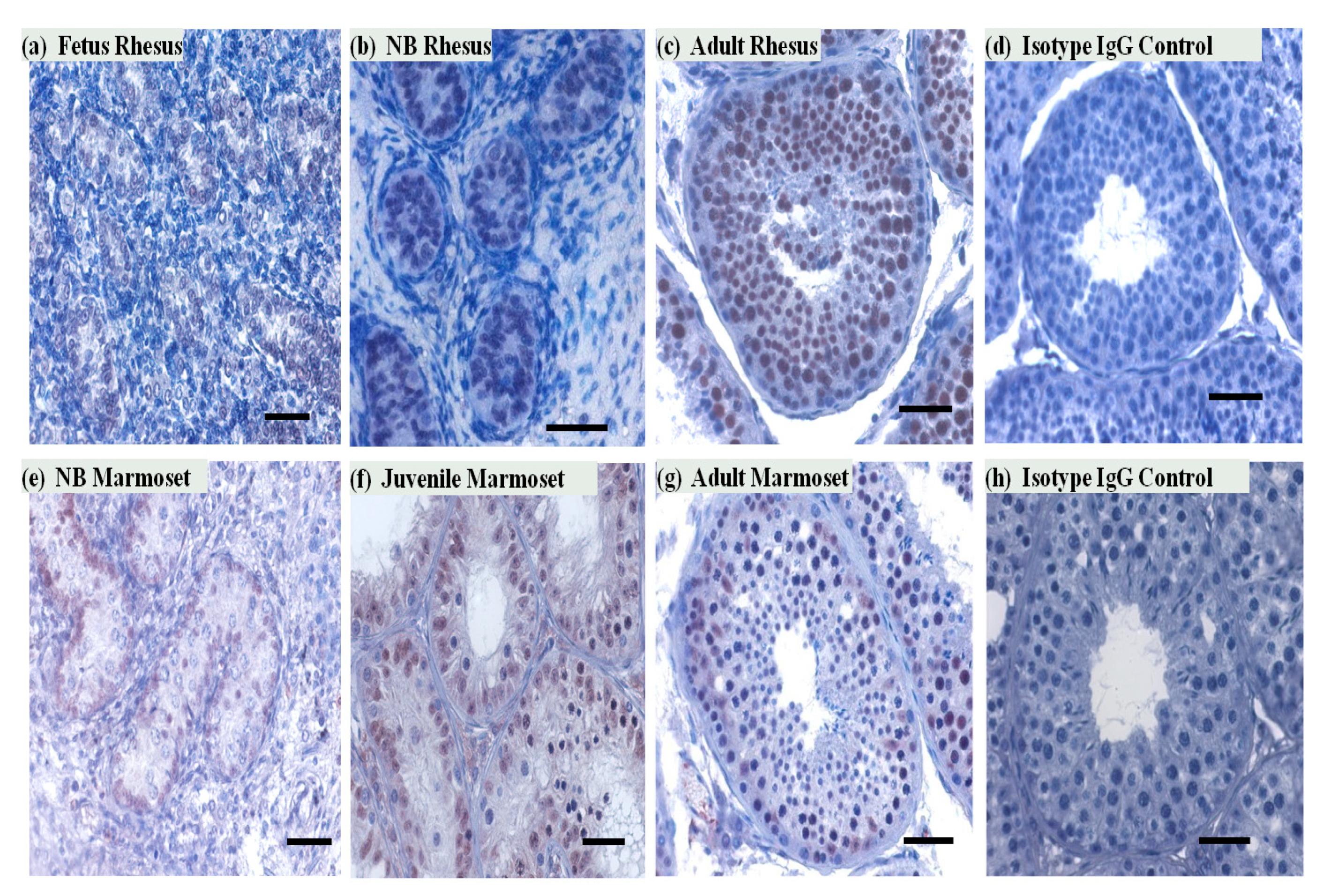
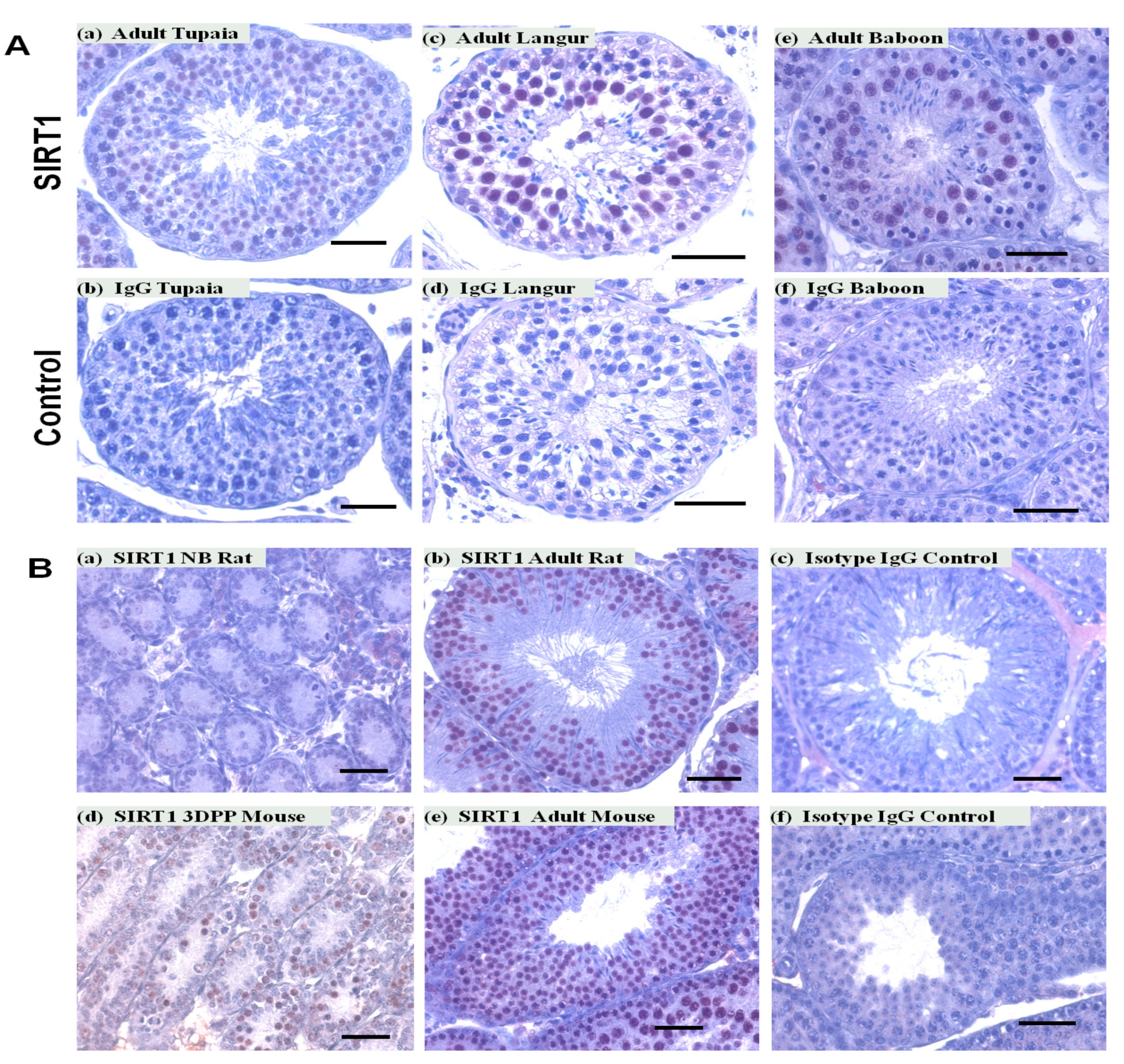
3.2. SIRT1 Expression in the Non-Human Primates Testes
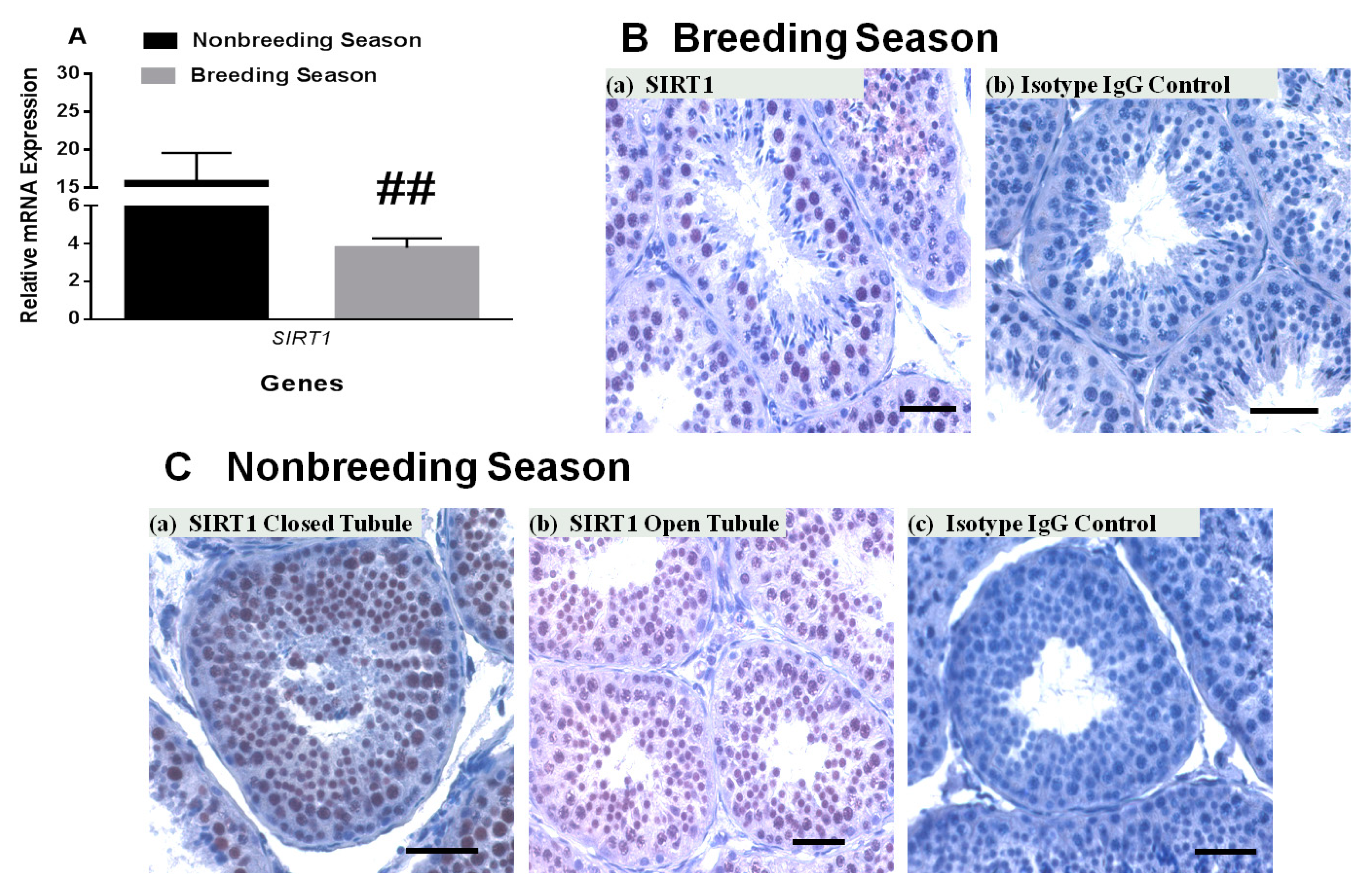
3.3. SIRT1 mRNA and Protein Expression in Rhesus Monkey Testis during Various Seasons
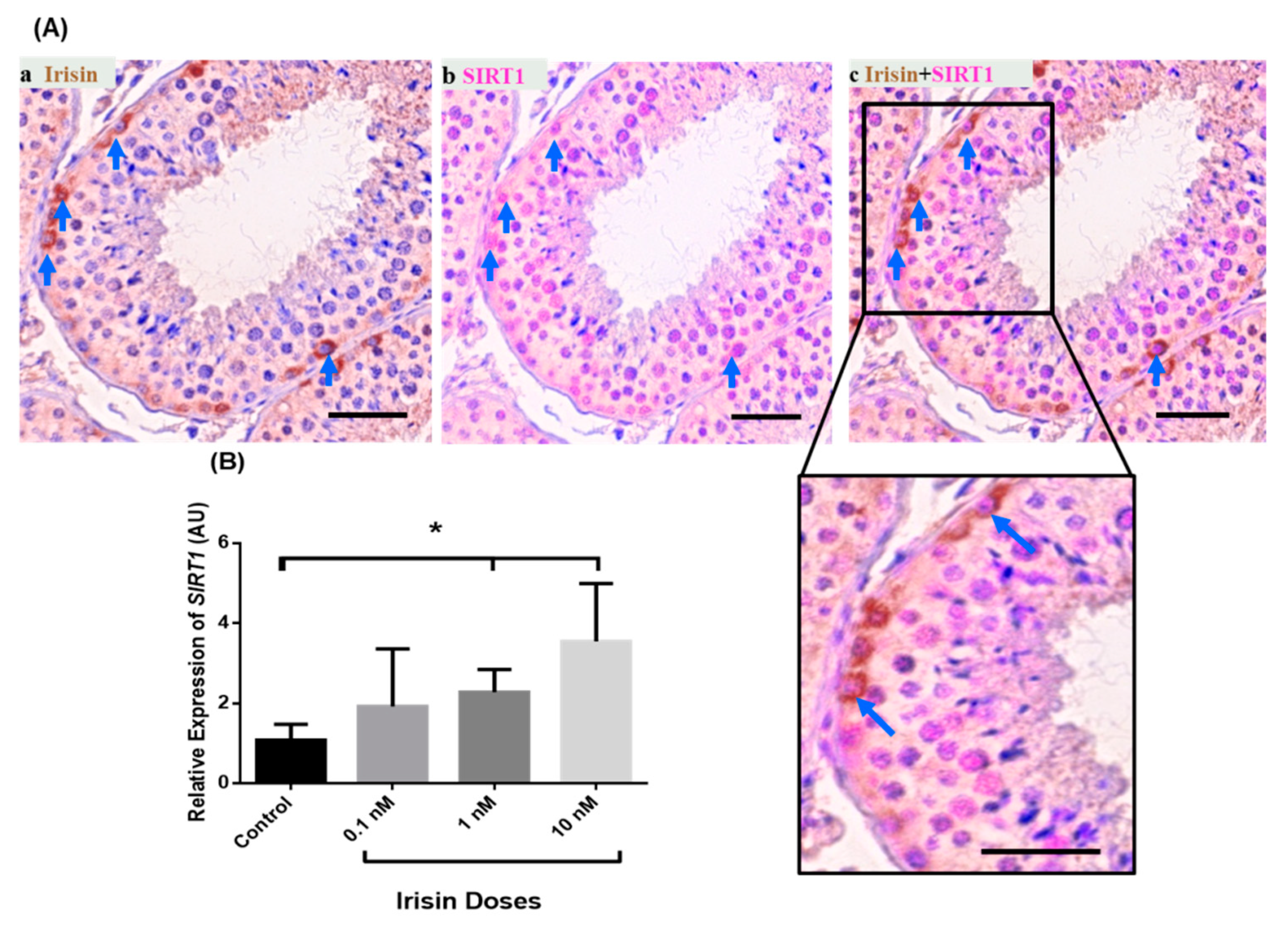
3.4. Exogenous Irisin Effect on SIRT1 Transcript Levels in Testis
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Song, N.; Endo, D.; Koji, T. Roles of epigenome in mammalian spermatogenesis. Reprod. Med. Biol. 2014, 13, 59–69. [Google Scholar] [CrossRef] [PubMed]
- Eun, S.H.; Gan, Q.; Chen, X. Epigenetic regulation of germ cell differentiation. Curr. Opin. Cell Biol. 2010, 22, 737–743. [Google Scholar] [CrossRef] [PubMed]
- Meikar, O.; Da Ros, M.; Kotaja, N. Epigenetic regulation of male germ cell differentiation. Subcell Biochem. 2013, 61, 119–138. [Google Scholar] [CrossRef] [PubMed]
- Bird, A. Perceptions of epigenetics. Nature 2007, 447, 396–398. [Google Scholar] [CrossRef]
- Shahbazian, M.D.; Grunstein, M. Functions of site-specific histone acetylation and deacetylation. Annu. Rev. Biochem. 2007, 76, 75–100. [Google Scholar] [CrossRef]
- Ma, P.; Pan, H.; Montgomery, R.L.; Olson, E.N.; Schultz, R.M. Compensatory functions of histone deacetylase 1 (HDAC1) and HDAC2 regulate transcription and apoptosis during mouse oocyte development. Proc. Natl. Acad. Sci. USA 2012, 109, E481–E489. [Google Scholar] [CrossRef]
- Gu, L.; Wang, Q.; Sun, Q.Y. Histone modifications during mammalian oocyte maturation: Dynamics, regulation and functions. Cell Cycle 2010, 9, 1942–1950. [Google Scholar] [CrossRef] [PubMed]
- Michan, S.; Sinclair, D. Sirtuins in mammals: Insights into their biological function. Biochem. J. 2007, 404, 1–13. [Google Scholar] [CrossRef]
- Pillarisetti, S. A review of Sirt1 and Sirt1 modulators in cardiovascular and metabolic diseases. Recent Pat. Cardiovasc. Drug Discov. 2008, 3, 156–164. [Google Scholar] [CrossRef]
- Yamamoto, H.; Schoonjans, K.; Auwerx, J. Sirtuin functions in health and disease. Mol. Endocrinol. 2007, 21, 1745–1755. [Google Scholar] [CrossRef]
- Haigis, M.C.; Guarente, L.P. Mammalian sirtuins—Emerging roles in physiology, aging, and calorie restriction. Genes Dev. 2006, 20, 2913–2921. [Google Scholar] [CrossRef]
- Kosciuk, T.; Wang, M.; Hong, J.Y.; Lin, H. Updates on the epigenetic roles of sirtuins. Curr. Opin. Chem. Biol. 2019, 51, 18–29. [Google Scholar] [CrossRef]
- Chianese, R.; Viggiano, A.; Urbanek, K.; Cappetta, D.; Troisi, J.; Scafuro, M.; Guida, M.; Esposito, G.; Ciuffreda, L.P.; Rossi, F.; et al. Chronic exposure to low dose of bisphenol A impacts on the first round of spermatogenesis via SIRT1 modulation. Sci. Rep. 2018, 8, 2961. [Google Scholar] [CrossRef] [PubMed]
- D’Angelo, S.; Mele, E.; Di Filippo, F.; Viggiano, A.; Meccariello, R. Sirt1 Activity in the Brain: Simultaneous Effects on Energy Homeostasis and Reproduction. Int. J. Environ. Res. Public Health 2021, 18, 1243. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Song, Z.; Wang, L.; Yu, H.; Liu, W.; Shang, Y.; Xu, Z.; Zhao, H.; Gao, F.; Wen, J.; et al. Sirt1 regulates acrosome biogenesis by modulating autophagic flux during spermiogenesis in mice. Development 2017, 144, 441–451. [Google Scholar] [CrossRef] [PubMed]
- Tatone, C.; Di Emidio, G.; Barbonetti, A.; Carta, G.; Luciano, A.M.; Falone, S.; Amicarelli, F. Sirtuins in gamete biology and reproductive physiology: Emerging roles and therapeutic potential in female and male infertility. Hum. Reprod. Update 2018, 24, 267–289. [Google Scholar] [CrossRef] [PubMed]
- Vazquez, M.J.; Toro, C.A.; Castellano, J.M.; Ruiz-Pino, F.; Roa, J.; Beiroa, D.; Heras, V.; Velasco, I.; Dieguez, C.; Pinilla, L.; et al. SIRT1 mediates obesity- and nutrient-dependent perturbation of pubertal timing by epigenetically controlling Kiss1 expression. Na.t Commun. 2018, 9, 4194. [Google Scholar] [CrossRef]
- Wu, L.; Zhang, A.; Sun, Y.; Zhu, X.; Fan, W.; Lu, X.; Yang, Q.; Feng, Y. Sirt1 exerts anti-inflammatory effects and promotes steroidogenesis in Leydig cells. Fertil. Steril. 2012, 98, 194–199. [Google Scholar] [CrossRef]
- Bell, E.L.; Nagamori, I.; Williams, E.O.; Del Rosario, A.M.; Bryson, B.D.; Watson, N.; White, F.M.; Sassone-Corsi, P.; Guarente, L. SirT1 is required in the male germ cell for differentiation and fecundity in mice. Development 2014, 141, 3495–3504. [Google Scholar] [CrossRef]
- Wahab, F.; Drummer, C.; Matz-Rensing, K.; Fuchs, E.; Behr, R. Irisin is expressed by undifferentiated spermatogonia and modulates gene expression in organotypic primate testis cultures. Mol. Cell. Endocrinol. 2020, 504, 110670. [Google Scholar] [CrossRef] [PubMed]
- Medrano, J.V.; Vilanova-Perez, T.; Fornes-Ferrer, V.; Navarro-Gomezlechon, A.; Martinez-Triguero, M.L.; Garcia, S.; Gomez-Chacon, J.; Povo, I.; Pellicer, A.; Andres, M.M.; et al. Influence of temperature, serum, and gonadotropin supplementation in short- and long-term organotypic culture of human immature testicular tissue. Fertil. Steril. 2018, 110, 1045–1057.e3. [Google Scholar] [CrossRef]
- Kristensen, D.M.; Desdoits-Lethimonier, C.; Mackey, A.L.; Dalgaard, M.D.; De Masi, F.; Munkbol, C.H.; Styrishave, B.; Antignac, J.P.; Le Bizec, B.; Platel, C.; et al. Ibuprofen alters human testicular physiology to produce a state of compensated hypogonadism. Proc. Natl. Acad. Sci. USA 2018, 115, E715–E724. [Google Scholar] [CrossRef] [PubMed]
- Bansode, F.W.; Chowdhury, S.R.; Dhar, J.D. Seasonal changes in the seminiferous epithelium of rhesus and bonnet monkeys. J. Med. Primatol. 2003, 32, 170–177. [Google Scholar] [CrossRef] [PubMed]
- Higashi, Y.; Takahashi, J.; Yoshida, K.; Winters, S.J.; Oshima, H.; Troen, P. Seasonal changes in steroidogenesis in the testis of the rhesus monkey (Macaca mulatta). J. Androl. 1984, 5, 70–79. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wahab, F.; Rodriguez Polo, I.; Behr, R. SIRT1 Expression and Regulation in the Primate Testis. Int. J. Mol. Sci. 2021, 22, 3207. https://doi.org/10.3390/ijms22063207
Wahab F, Rodriguez Polo I, Behr R. SIRT1 Expression and Regulation in the Primate Testis. International Journal of Molecular Sciences. 2021; 22(6):3207. https://doi.org/10.3390/ijms22063207
Chicago/Turabian StyleWahab, Fazal, Ignacio Rodriguez Polo, and Rüdiger Behr. 2021. "SIRT1 Expression and Regulation in the Primate Testis" International Journal of Molecular Sciences 22, no. 6: 3207. https://doi.org/10.3390/ijms22063207
APA StyleWahab, F., Rodriguez Polo, I., & Behr, R. (2021). SIRT1 Expression and Regulation in the Primate Testis. International Journal of Molecular Sciences, 22(6), 3207. https://doi.org/10.3390/ijms22063207





