Modification of Serine 1040 of SIBRI1 Increases Fruit Yield by Enhancing Tolerance to Heat Stress in Tomato
Abstract
1. Introduction
2. Results
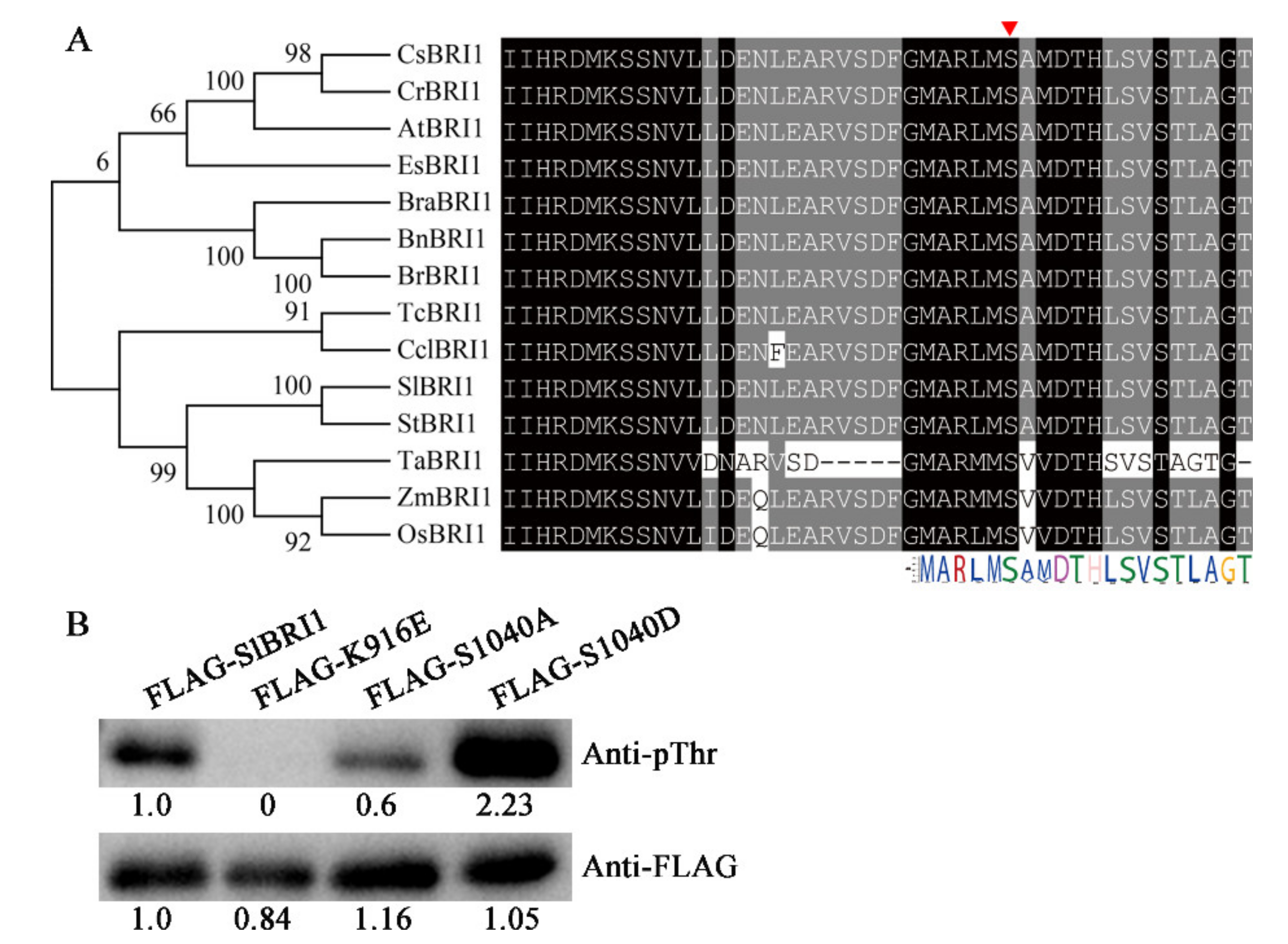
2.1. SlBRI1 Ser-1040 Influences Autophosphorylation of SlBRI1
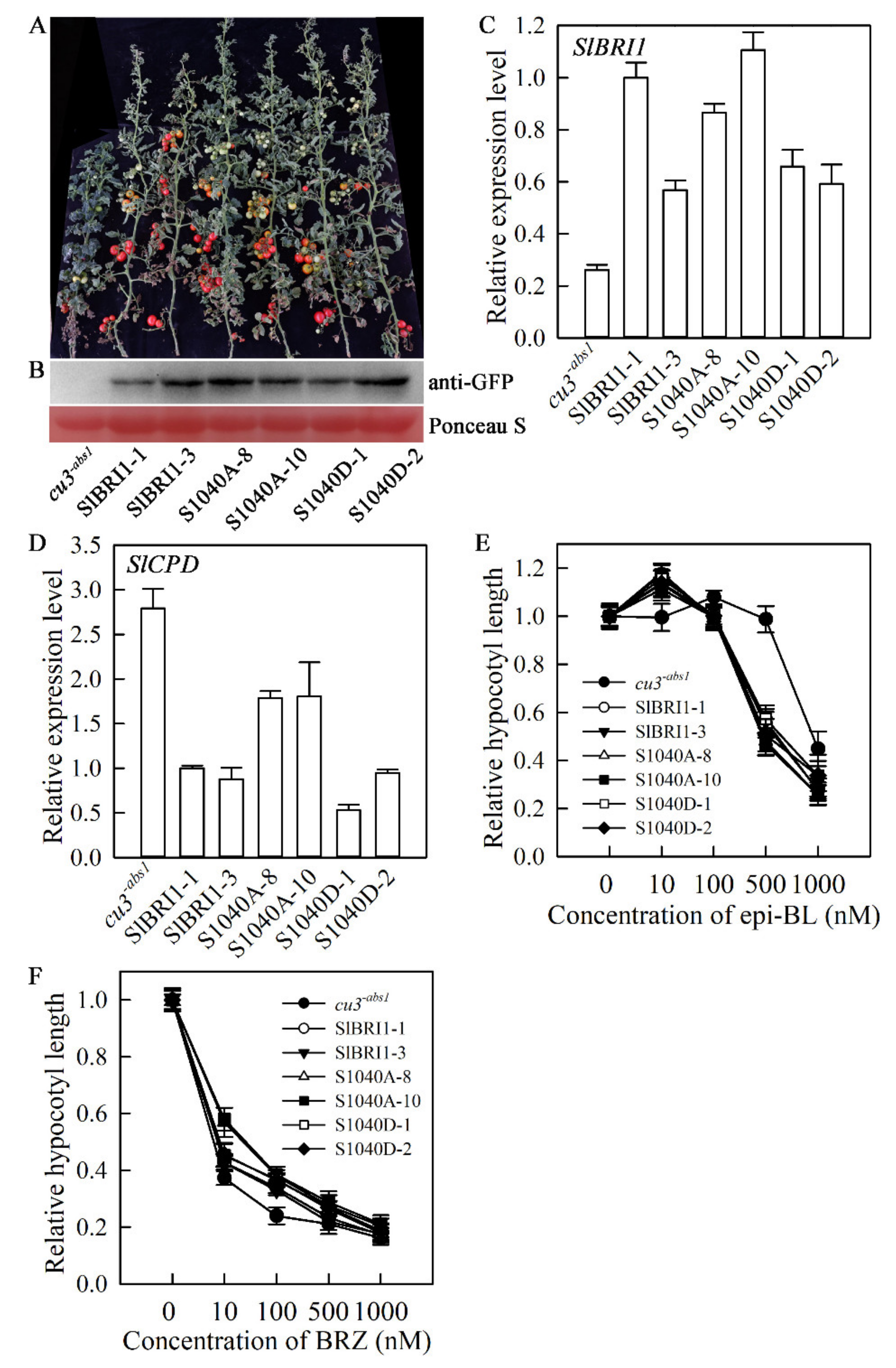
2.2. SlBRI1 Ser-1040 Slightly Affects BR Signalling in Tomato
2.3. Dephosphorylation of Ser-1040 Improves Tomato Yield under Heat Stress
2.4. Dephosphorylation of Ser-1040 Promotes Germination and Seedling Growth under Heat Stress
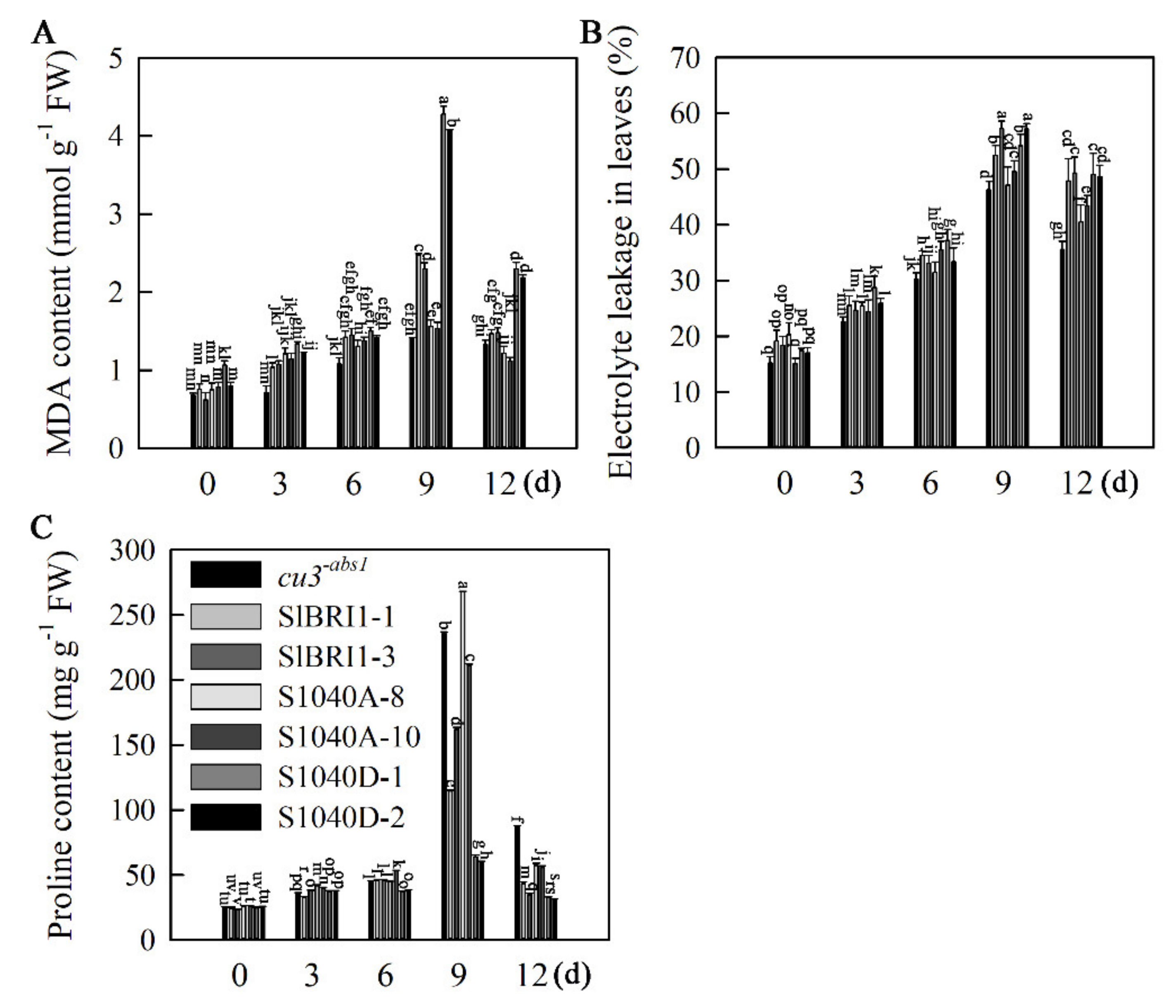
2.5. Dephosphorylation of Ser-1040 Promotes Heat Stress Tolerance of Seedlings
2.6. Dephosphorylation of Ser-1040 Promotes ROS Detoxification of Seedlings under Heat Stress
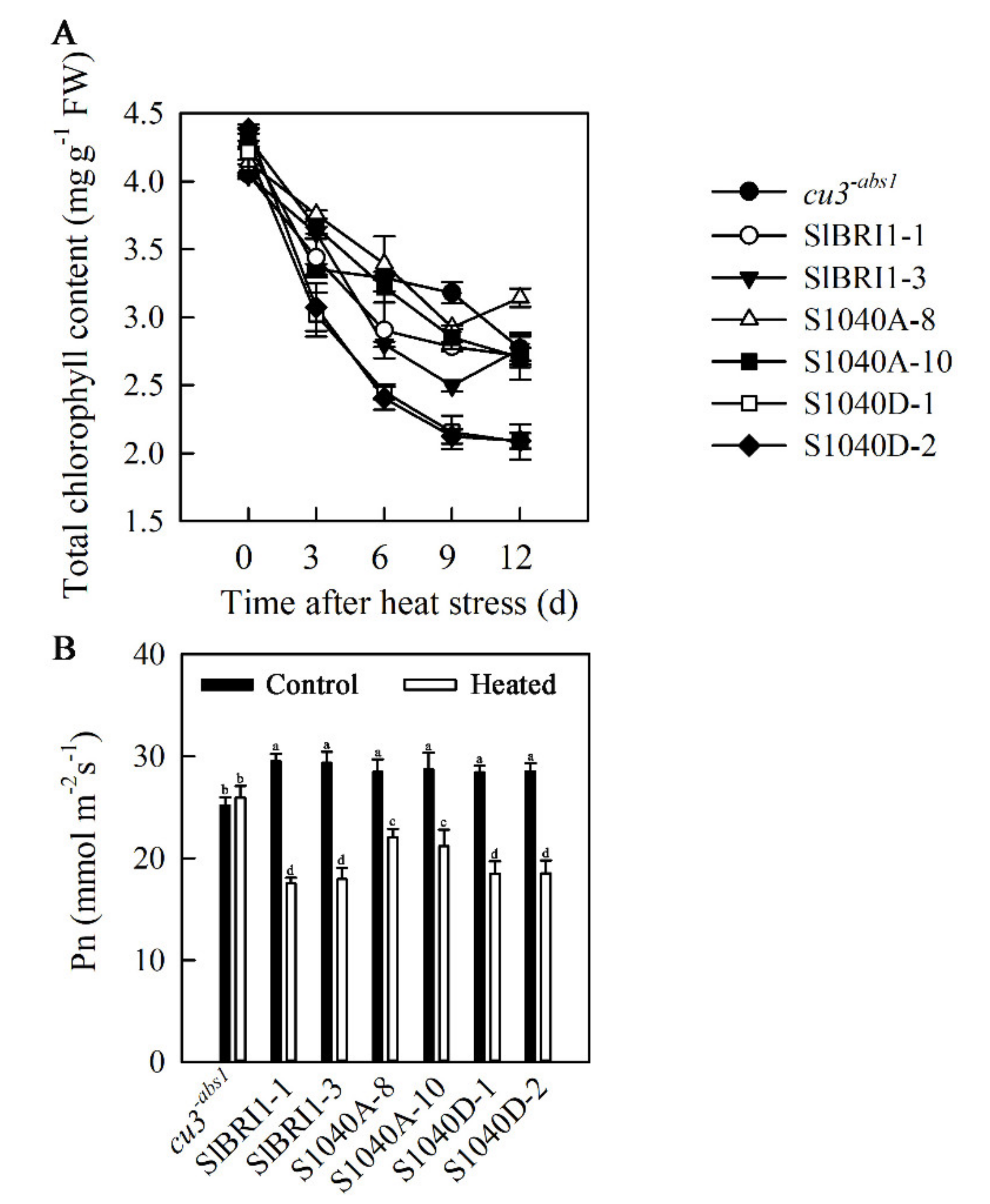
2.7. Dephosphorylation of Ser-1040 Promotes Photosynthesis in Seedlings under Heat Stress
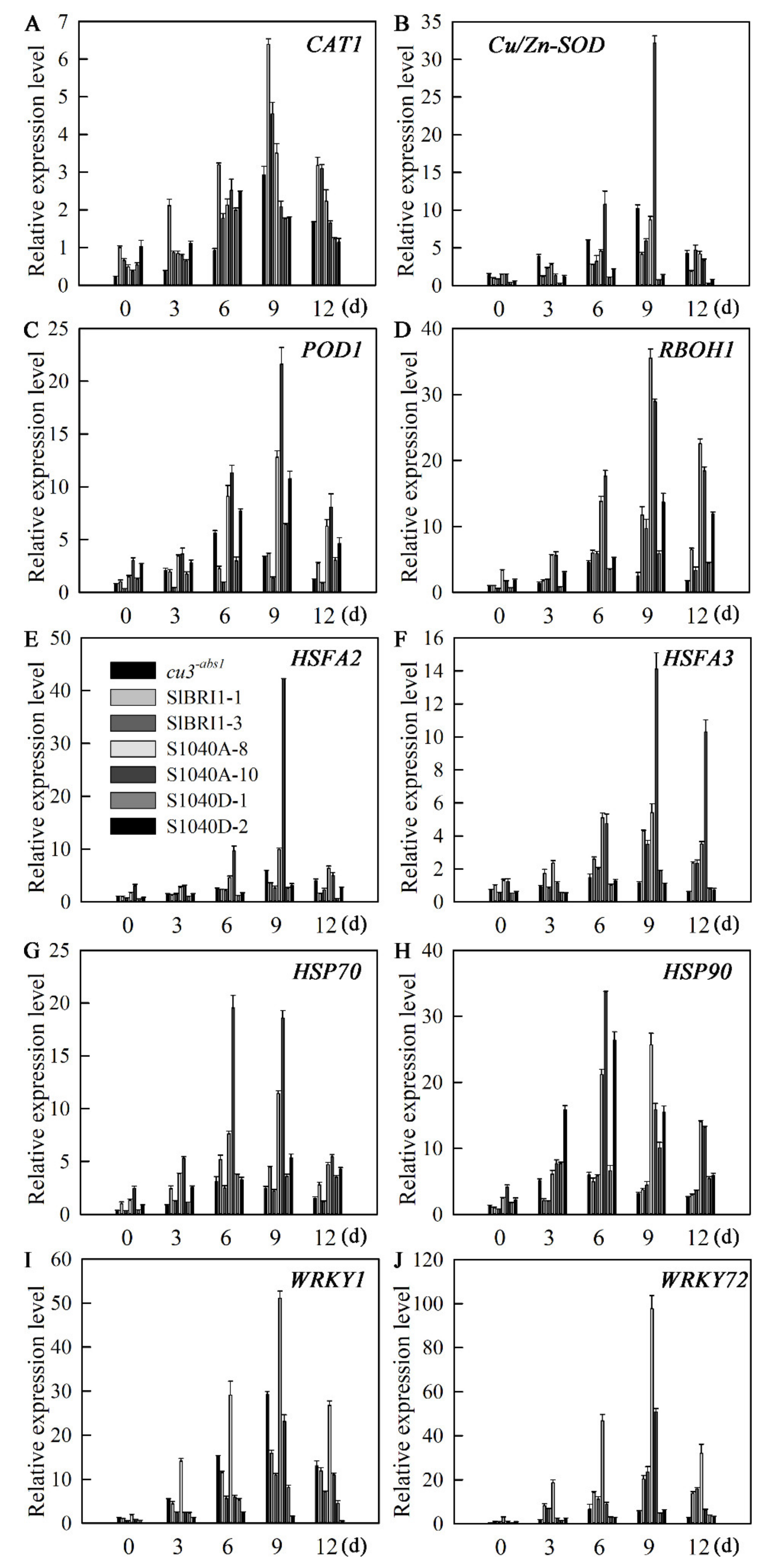
2.8. Dephosphorylation of Ser-1040 Enhances the Expression of Tomato Defence-Related Genes under Heat Stress
3. Discussion
4. Materials and Methods
4.1. Sequence Alignment and Phylogenetic Analysis
4.2. Generation of Transgenic Plants by Site-Directed Mutagenesis
4.3. Agronomic Trait Characterization
4.4. Growth Response of Hypocotyls to Exogenous BRZ and BL
4.5. Germination Analysis of Plants under Heat Stress
4.6. Heat Treatment of Seedlings
4.7. Growth Performances of Seedlings under Heat Stress
4.8. Physiological Parameter Measurements
4.9. DAB Staining
4.10. Autophosphorylation Analysis
4.11. Western Blot Analysis
4.12. Quantitative Real-time PCR Analysis
4.13. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| BR | Brassinosteroid |
| BRI1 | Brassinosteroid Insensitive1 |
| CPD | Constitutive Photomorphogenesis and Dwarf |
| epi-BL | 24-Epibrassinolide |
| BRZ | Brassinazole |
| K916E | Lysine-916-Glutamic acid |
| GFP | Green fluorescent protein |
| Ser | Serine |
| S1040A | Serine-1040-alanine |
| S1040D | Serine-1040-aspartic acid |
| Thr | Threonine |
| Tyr | Tyrosine |
| ROS | Reactive oxygen species |
| DAB | 3,3′-Diaminobenzidine |
| CAT | Catalase |
| SOD | Superoxide dismutase |
| RBOH1 | RESPIRATORY BURST OXIDASE HOMOLOG1 |
| HSF | Heat shock transcription factor |
| HSP | Heat shock protein |
| POD | Peroxidase |
| MDA | Malondialdehyde |
| Pn | Net photosynthetic rate |
| qRT-PCR | Quantitative real-time PCR analysis |
| MS | Murashige and Skoog medium |
| PPFD | Photosynthetic photon flux density |
References
- Bita, C.E.; Gerats, T. Plant tolerance to high temperature in a changing environment: Scientific fundamentals and production of heat stress-tolerant crops. Front. Plant. Sci. 2013, 4, 273. [Google Scholar] [CrossRef]
- Suzuki, N.; Mittler, R. Reactive oxygen species and temperature stresses: A delicate balance between signaling and destruction. Physiol. Plant. 2006, 126, 45–51. [Google Scholar] [CrossRef]
- Kim, S.Y.; Kim, B.H.; Lim, C.J.; Lim, C.O.; Nam, K.H. Constitutive activation of stress-inducible genes in a brassinosteroid-insensitive 1 (bri1) mutant results in higher tolerance to cold. Physiol. Plant 2010, 138, 191–204. [Google Scholar] [CrossRef]
- Xia, X.J.; Wang, Y.J.; Zhou, Y.H.; Mao, W.H.; Shi, K.; Asami, T.; Chen, Z.; Yu, J.Q. Reactive oxygen species are involved in brassinosteroid-induced stress tolerance in cucumber. Plant Physiol. 2009, 150, 801–814. [Google Scholar] [CrossRef]
- Fariduddin, Q.; Yusuf, M.; Ahmad, I.; Ahmad, A. Brassinosteroids and their role in response of plants to abiotic stresses. Biol. Plantarum 2014, 58, 9–17. [Google Scholar] [CrossRef]
- Niu, J.H.; Anjum, S.A.; Wang, R.; Li, J.H.; Liu, M.R.; Song, J.X.; Zohaib, A.; Lv, J.; Wang, S.G.; Zong, X.F. Exogenous application of brassinolide can alter morphological and physiological traits of Leymus chinensis (Trin.) Tzvelev under room and high temperatures. Chil. J. Agr. Res. 2016, 76, 27–33. [Google Scholar] [CrossRef]
- Singh, A.; Mittal, D.; Lavania, D.; Agarwal, M.; Mishra, R.C.; Grover, A. OsHsfA2c and OsHsfB4b are involved in the transcriptional regulation of cytoplasmic OsClpB (Hsp100) gene in rice (Oryza sativa L.). Cell Stress Chaperones 2012, 17, 243–254. [Google Scholar] [CrossRef] [PubMed]
- Planas-Riverola, A.; Gupta, A.; Betegon-Putze, I.; Bosch, N.; Ibanes, M.; Cano-Delgado, A.I. Brassinosteroid signaling in plant development and adaptation to stress. Development 2019, 146, 151894. [Google Scholar] [CrossRef]
- Nawaz, F.; Naeem, M.; Zulfiqar, B.; Akram, A.; Ashraf, M.Y.; Raheel, M.; Shabbir, R.N.; Hussain, R.A.; Anwar, I.; Aurangzaib, M. Understanding brassinosteroid-regulated mechanisms to improve stress tolerance in plants: A critical review. Environ. Sci. Pollut. Res. 2017, 24, 15959–15975. [Google Scholar] [CrossRef]
- Yuan, G.F.; Jia, C.G.; Li, Z.; Sun, B.; Zhang, L.P.; Liu, N.; Wang, Q.M. Effect of brassinosteroids on drought resistance and abscisic acid concentration in tomato under water stress. Sci. Hortic. 2010, 126, 103–108. [Google Scholar] [CrossRef]
- Wang, X.F.; Kota, U.; He, K.; Blackburn, K.; Li, J.; Goshe, M.B.; Huber, S.C.; Clouse, S.D. Sequential transphosphorylation of the BRI1/BAK1 receptor kinase complex impacts early events in brassinosteroid signaling. Dev. Cell. 2008, 15, 220–235. [Google Scholar] [CrossRef]
- Clouse, S.D. Brassinosteroid signal transduction: From receptor kinase activation to transcriptional networks regulating plant development. Plant Cell 2011, 23, 1219–1230. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.W.; Wang, Z.Y. Brassinosteroid signal transduction from receptor kinases to transcription factors. Annu. Rev. Plant. Biol. 2010, 61, 681–704. [Google Scholar] [CrossRef]
- Zhang, X.W.; Zhou, L.Y.; Qin, Y.K.; Chen, Y.W.; Liu, X.L.; Wang, M.Y.; Mao, J.; Zhang, J.J.; He, Z.H.; Liu, L.C.; et al. A temperature-sensitive misfolded bri1-301 receptor requires its kinase activity to promote growth. Plant Physiol. 2018, 178, 1704–1719. [Google Scholar] [CrossRef] [PubMed]
- Gruszka, D.; Szarejko, I.; Maluszynski, M. New allele of HvBRI1 gene encoding brassinosteroid receptor in barley. J. Appl. Genetics 2011, 52, 257–268. [Google Scholar] [CrossRef] [PubMed]
- Nie, S.M.; Huang, S.H.; Wang, S.F.; Cheng, D.D.; Liu, J.W.; Lv, S.Q.; Li, Q.; Wang, X.F. Enhancing brassinosteroid signaling via overexpression of tomato (Solanum lycopersicum) SlBRI1 improves major agronomic traits. Front. Plant. Sci. 2017, 8, 1386. [Google Scholar] [CrossRef]
- Nie, S.M.; Huang, S.H.; Wang, S.F.; Mao, Y.J.; Liu, J.W.; Ma, R.L.; Wang, X.F. Enhanced brassinosteroid signaling intensity via SlBRI1 overexpression negatively regulates drought resistance in a manner opposite of that via exogenous BR application in tomato. Plant Physiol. Biochem. 2019, 138, 36–47. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.; Breja, P.; Khurana, J.P.; Khurana, P. Wheat Brassinosteroid-Insensitive1 (TaBRI1) interacts with members of TaSERK gene family and cause early flowering and seed yield enhancement in Arabidopsis. PLoS ONE 2016, 11, e0153273. [Google Scholar] [CrossRef]
- Morinaka, Y.; Sakamoto, T.; Inukai, Y.; Agetsuma, M.; Kitano, H.; Ashikari, M.; Matsuoka, M. Morphological alteration caused by brassinosteroid insensitivity increases the biomass and grain production of rice. Plant. Physiol. 2006, 141, 924–931. [Google Scholar] [CrossRef]
- Zheng, B.W.; Bai, Q.W.; Wu, L.; Liu, H.; Liu, Y.P.; Xu, W.J.; Li, G.S.; Ren, H.Y.; She, X.P.; Wu, G. EMS1 and BRI1 control separate biological processes via extracellular domain diversity and intracellular domain conservation. Nat. Commun. 2019, 10, 4165. [Google Scholar] [CrossRef]
- Wang, X.F.; Goshe, M.B.; Soderblom, E.J.; Phinney, B.S.; Kuchar, J.A.; Li, J.; Asami, T.; Yoshida, S.; Huber, S.C.; Clouse, S.D. Identification and functional analysis of in vivo phosphorylation sites of the Arabidopsis BRASSINOSTEROID-INSENSITIVE1 receptor kinase. Plant Cell 2005, 17, 1685–1703. [Google Scholar] [CrossRef]
- Wang, Q.N.; Wang, S.F.; Gan, S.F.; Wang, X.; Liu, J.W.; Wang, X.F. Role of specific phosphorylation sites of Arabidopsis Brassinosteroid-Insensitive 1 receptor kinase in plant growth and development. J. Plant. Growth Regul. 2016, 35, 755–769. [Google Scholar] [CrossRef]
- Oh, M.H.; Sun, J.D.; Oh, D.H.; Zielinski, R.E.; Clouse, S.D.; Huber, S.C. Enhancing Arabidopsis leaf growth by engineering the BRASSINOSTEROID INSENSITIVE1 receptor kinase. Plant. Physiol. 2011, 157, 120–131. [Google Scholar] [CrossRef] [PubMed]
- Bajwa, V.S.; Wang, X.F.; Blackburn, R.K.; Goshe, M.B.; Mitra, S.K.; Williams, E.L.; Bishop, G.J.; Krasnyanski, S.; Allen, G.; Huber, S.C.; et al. Identification and functional analysis of tomato BRI1 and BAK1 receptor kinase phosphorylation sites. Plant Physiol. 2013, 163, 30–42. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.F.; Liu, J.W.; Zhao, T.; Du, C.X.; Nie, S.M.; Zhang, Y.Y.; Lv, S.Q.; Huang, S.H.; Wang, X.F. Modification of Threonine-1050 of SlBRI1 regulates BR Signalling and increases fruit yield of tomato. BMC Plant Biol. 2019, 19, 256. [Google Scholar] [CrossRef] [PubMed]
- Mathur, J.; Molnar, G.; Fujioka, S.; Takatsuto, S.; Sakurai, A.; Yokota, T.; Adam, G.; Voigt, B.; Nagy, F.; Maas, C.; et al. Transcription of the Arabidopsis CPD gene, encoding a steroidogenic cytochrome P450, is negatively controlled by brassinosteroids. Plant J. 1998, 14, 593–602. [Google Scholar] [CrossRef]
- Nie, W.F.; Wang, M.M.; Xia, X.J.; Zhou, Y.H.; Shi, K.; Chen, Z.; Yu, J.Q. Silencing of tomato RBOH1 and MPK2 abolishes brassinosteroid-induced H2O2 generation and stress tolerance. Plant Cell Environ. 2013, 36, 789–803. [Google Scholar] [CrossRef]
- Hussain, M.; Khan, T.A.; Yusuf, M.; Fariduddin, Q. Silicon-mediated role of 24-epibrassinolide in wheat under high-temperature stress. Environ. Sci. Pollut. Res. Int. 2019, 26, 17163–17172. [Google Scholar] [CrossRef]
- Mazorra, L.M.; Holton, N.; Bishop, G.J.; Nunez, M. Heat shock response in tomato brassinosteroid mutants indicates that thermotolerance is independent of brassinosteroid homeostasis. Plant Physiol. Biochem. 2011, 49, 1420–1428. [Google Scholar] [CrossRef]
- Miller, G.; Schlauch, K.; Tam, R.; Cortes, D.; Torres, M.A.; Shulaev, V.; Dangl, J.L.; Mittler, R. The plant NADPH oxidase RBOHD mediates rapid systemic signaling in response to diverse stimuli. Sci. Signal. 2009, 2, 45. [Google Scholar] [CrossRef]
- Marino, D.; Dunand, C.; Puppo, A.; Pauly, N. A burst of plant NADPH oxidases. Trends Plant Sci. 2012, 17, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.T.; Chen, T.H.; Lo, K.F.; Chiu, C.Y. Effects of proline on copper transport in rice seedlings under excess copper stress. Plant Sci. 2004, 166, 103–111. [Google Scholar] [CrossRef]
- Yin, Y.L.; Qin, K.Z.; Song, X.W.; Zhang, Q.H.; Zhou, Y.H.; Xia, X.J.; Yu, J.Q. BZR1 transcription factor regulates heat stress tolerance through FERONIA receptor-like kinase-mediated reactive oxygen species signaling in tomato. Plant Cell Physiol. 2018, 59, 2239–2254. [Google Scholar] [CrossRef]
- Farhad, M.S.; Babak, A.M.; Reza, Z.M.; Hassan, R.S.M.; Afshin, T. Response of proline, soluble sugars, photosynthetic pigments and antioxidant enzymes in potato (Solanum tuberosum L.) to different irrigation regimes in greenhouse condition. Aust. J. Crop. Sci. 2011, 5, 55–60. [Google Scholar]
- Al-Whaibi, M.H. Plant heat-shock proteins: A mini review. J. King Saud Univ. Sci. 2011, 23, 139–150. [Google Scholar] [CrossRef]
- Wang, W.X.; Vinocur, B.; Shoseyov, O.; Altman, A. Role of plant heat-shock proteins and molecular chaperones in the abiotic stress response. Trends Plant Sci. 2004, 9, 244–252. [Google Scholar] [CrossRef] [PubMed]
- Usman, M.G.; Rafii, M.Y.; Martini, M.Y.; Yusuff, O.A.; Ismail, M.R.; Miah, G. Molecular analysis of Hsp70 mechanisms in plants and their function in response to stress. Biotechnol. Genet. Eng. Rev. 2017, 33, 26–39. [Google Scholar] [CrossRef]
- Sadura, I.; Libik-Konieczny, M.; Jurczyk, B.; Gruszka, D.; Janeczko, A. HSP transcript and protein accumulation in brassinosteroid barley mutants acclimated to low and high temperatures. Int. J. Mol. Sci. 2020, 21, 1889. [Google Scholar] [CrossRef] [PubMed]
- Neta-Sharir, I.; Isaacson, T.; Lurie, S.; Weiss, D. Dual role for tomato heat shock protein 21: Protecting photosystem II from oxidative stress and promoting color changes during fruit maturation. Plant Cell 2005, 17, 1829–1838. [Google Scholar] [CrossRef]
- Ogawa, D.; Yamaguchi, K.; Nishiuchi, T. High-level overexpression of the Arabidopsis HsfA2 gene confers not only increased themotolerance but also salt/osmotic stress tolerance and enhanced callus growth. J. Exp. Bot. 2007, 58, 3373–3383. [Google Scholar] [CrossRef]
- Zhu, B.G.; Ye, C.J.; Lu, H.Y.; Chen, X.J.; Chai, G.H.; Chen, J.N.; Wang, C. Identification and characterization of a novel heat shock transcription factor gene, GmHsfA1, in soybeans (Glycine max). J. Plant Res. 2006, 119, 247–256. [Google Scholar] [CrossRef]
- Liu, J.G.; Qin, Q.L.; Zhang, Z.; Peng, R.H.; Xiong, A.S.; Chen, J.M.; Yao, Q.H. OsHSF7 gene in rice, Oryza sativa L., encodes a transcription factor that functions as a high temperature receptive and responsive factor. BMB Rep. 2009, 42, 16–21. [Google Scholar] [CrossRef]
- Li, S.J.; Fu, Q.T.; Huang, W.D.; Yu, D.Q. Functional analysis of an Arabidopsis transcription factor WRKY25 in heat stress. Plant Cell Rep. 2009, 28, 683–693. [Google Scholar] [CrossRef] [PubMed]
- Li, S.J.; Zhou, X.; Chen, L.G.; Huang, W.D.; Yu, D.Q. Functional characterization of Arabidopsis thaliana WRKY39 in heat stress. Mol. Cells 2010, 29, 475–483. [Google Scholar] [CrossRef]
- Wu, X.L.; Shiroto, Y.; Kishitani, S.; Ito, Y.; Toriyama, K. Enhanced heat and drought tolerance in transgenic rice seedlings overexpressing OsWRKY11 under the control of HSP101 promoter. Plant Cell Rep. 2009, 28, 21–30. [Google Scholar] [CrossRef]
- Casal, J.J.; Balasubramanian, S. Thermomorphogenesis. Annu. Rev. Plant Biol. 2019, 70, 321–346. [Google Scholar] [CrossRef] [PubMed]
- Khan, A.R.; Hui, C.Z.; Ghazanfar, B.; Khan, M.A.; Ahmad, S.S.; Ahmad, I. Acetyl salicylic acid and 24-epibrassinolide attenuate decline in photosynthesis, chlorophyll contents and membrane thermo-stability in tomato (Lycopersicon Esculentum Mill.) under heat stress. Pak. J. Bot. 2015, 47, 63–70. [Google Scholar]
- Li, J.; Chory, J. A putative leucine-rich repeat receptor kinase involved in brassinosteroid signal transduction. Cell 1997, 90, 929–938. [Google Scholar] [CrossRef]
- Park, S.H.; Morris, J.L.; Park, J.E.; Hirschi, K.D.; Smith, R.H. Efficient and genotype-independent Agrobacterium-mediated tomato transformation. J. Plant Physiol. 2003, 160, 1253–1257. [Google Scholar] [CrossRef]
- Qian, C.M.; Wu, X.J.; Chen, L.; Jiang, C.H. Effects of high temperature stress on the germination of tomato seed. Seed 2002, 20, 89. [Google Scholar]
- Liu, H.; Yu, C.Y.; Li, H.X.; Ouyang, B.; Wang, T.T.; Zhang, J.H.; Wang, X.; Ye, Z.B. Overexpression of ShDHN, a dehydrin gene from Solanum habrochaites enhances tolerance to multiple abiotic stresses in tomato. Plant Sci. 2015, 231, 198–211. [Google Scholar] [CrossRef] [PubMed]
- Kong, F.Y.; Deng, Y.S.; Zhou, B.; Wang, G.D.; Wang, Y.; Meng, Q.W. A chloroplast-targeted DnaJ protein contributes to maintenance of photosystem II under chilling stress. J. Exp. Bot. 2014, 65, 143–158. [Google Scholar] [CrossRef]
- Li, J.; Wen, J.Q.; Lease, K.A.; Doke, J.T.; Tax, F.E.; Walker, J.C. BAK1, an Arabidopsis LRR receptor-like protein kinase, interacts with BRI1 and modulates brassinosteroid signaling. Cell 2002, 110, 213–222. [Google Scholar] [CrossRef]
- Zhou, J.; Wang, J.; Li, X.; Xia, X.J.; Zhou, Y.H.; Shi, K.; Chen, Z.; Yu, J.Q. H2O2 mediates the crosstalk of brassinosteroid and abscisic acid in tomato responses to heat and oxidative stresses. J. Exp. Bot. 2014, 65, 4371–4383. [Google Scholar] [CrossRef] [PubMed]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, S.; Hu, T.; Tian, A.; Luo, B.; Du, C.; Zhang, S.; Huang, S.; Zhang, F.; Wang, X. Modification of Serine 1040 of SIBRI1 Increases Fruit Yield by Enhancing Tolerance to Heat Stress in Tomato. Int. J. Mol. Sci. 2020, 21, 7681. https://doi.org/10.3390/ijms21207681
Wang S, Hu T, Tian A, Luo B, Du C, Zhang S, Huang S, Zhang F, Wang X. Modification of Serine 1040 of SIBRI1 Increases Fruit Yield by Enhancing Tolerance to Heat Stress in Tomato. International Journal of Molecular Sciences. 2020; 21(20):7681. https://doi.org/10.3390/ijms21207681
Chicago/Turabian StyleWang, Shufen, Tixu Hu, Aijuan Tian, Bote Luo, Chenxi Du, Siwei Zhang, Shuhua Huang, Fei Zhang, and Xiaofeng Wang. 2020. "Modification of Serine 1040 of SIBRI1 Increases Fruit Yield by Enhancing Tolerance to Heat Stress in Tomato" International Journal of Molecular Sciences 21, no. 20: 7681. https://doi.org/10.3390/ijms21207681
APA StyleWang, S., Hu, T., Tian, A., Luo, B., Du, C., Zhang, S., Huang, S., Zhang, F., & Wang, X. (2020). Modification of Serine 1040 of SIBRI1 Increases Fruit Yield by Enhancing Tolerance to Heat Stress in Tomato. International Journal of Molecular Sciences, 21(20), 7681. https://doi.org/10.3390/ijms21207681




