Multiplatform Investigation of Plasma and Tissue Lipid Signatures of Breast Cancer Using Mass Spectrometry Tools
Abstract
1. Introduction
2. Results
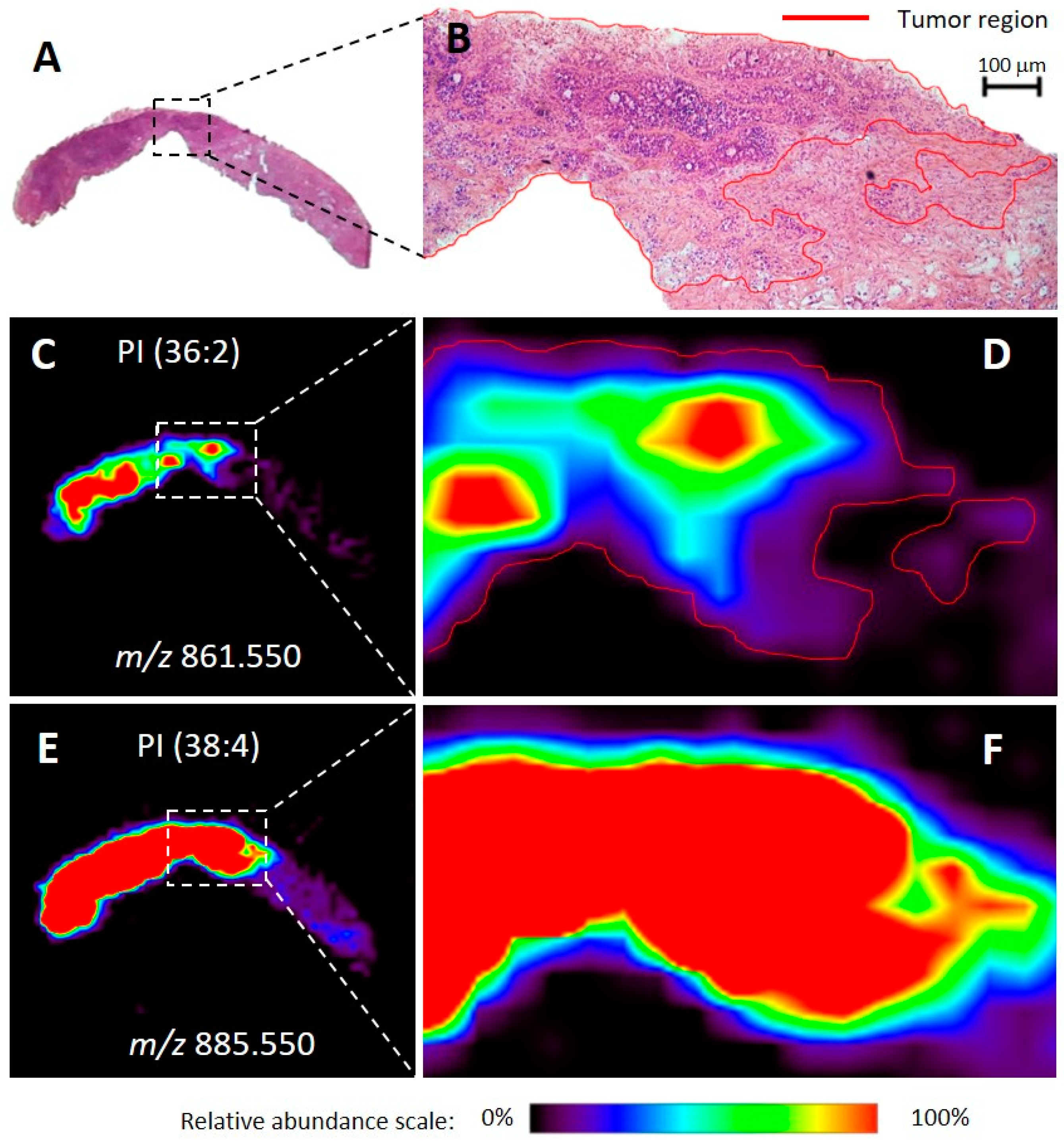
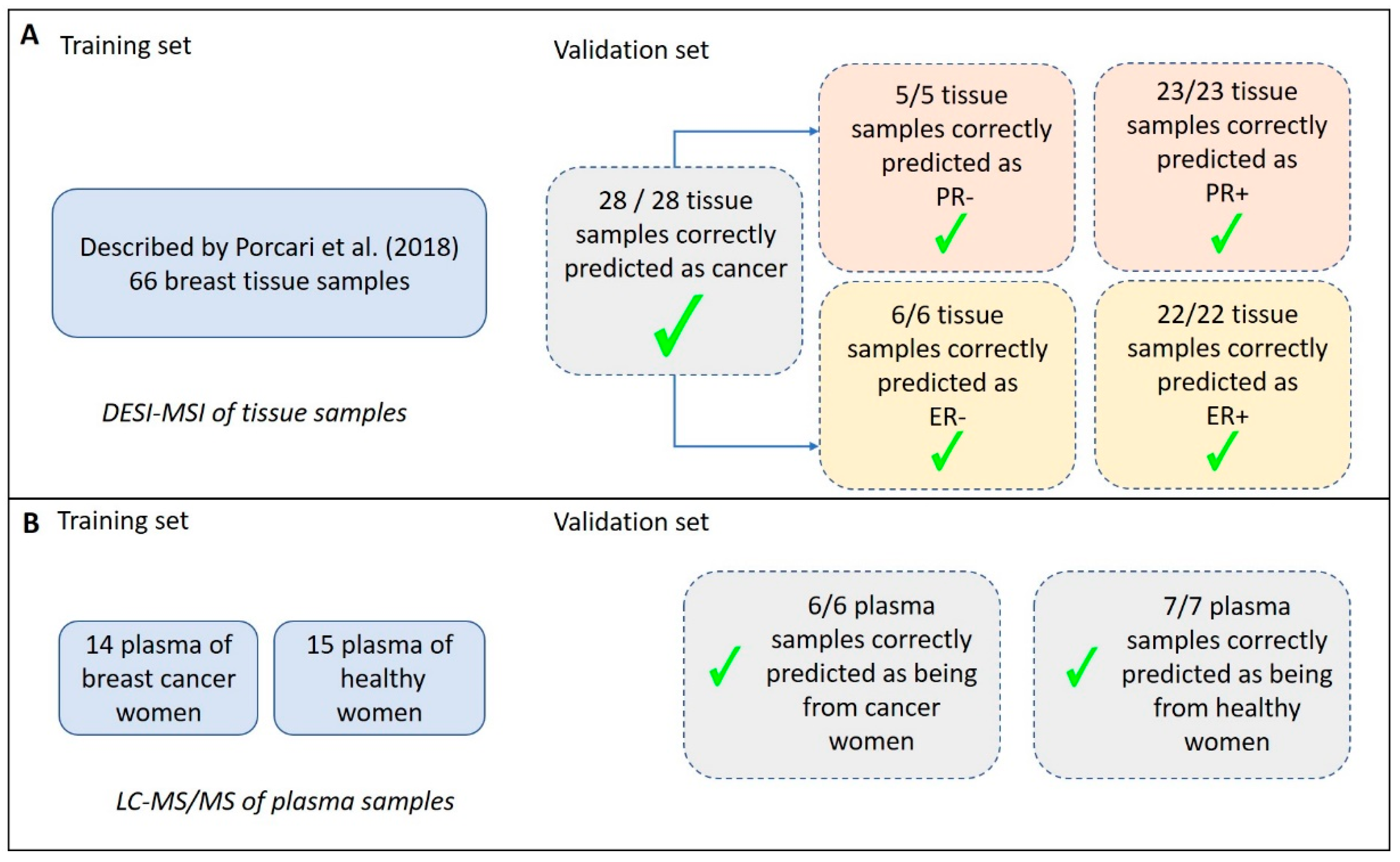
2.1. Molecular Imaging of Breast Tissues by DESI-MSI
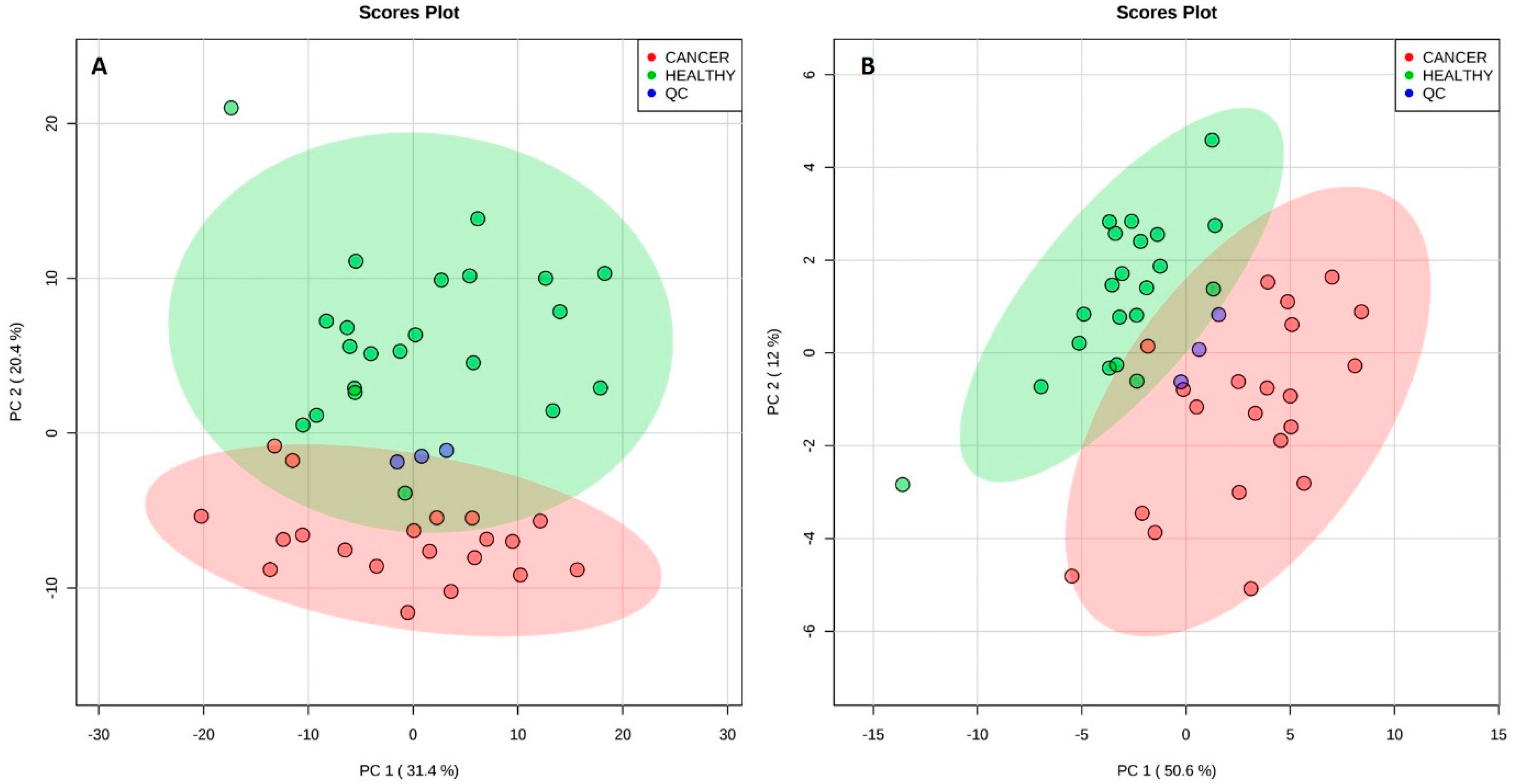
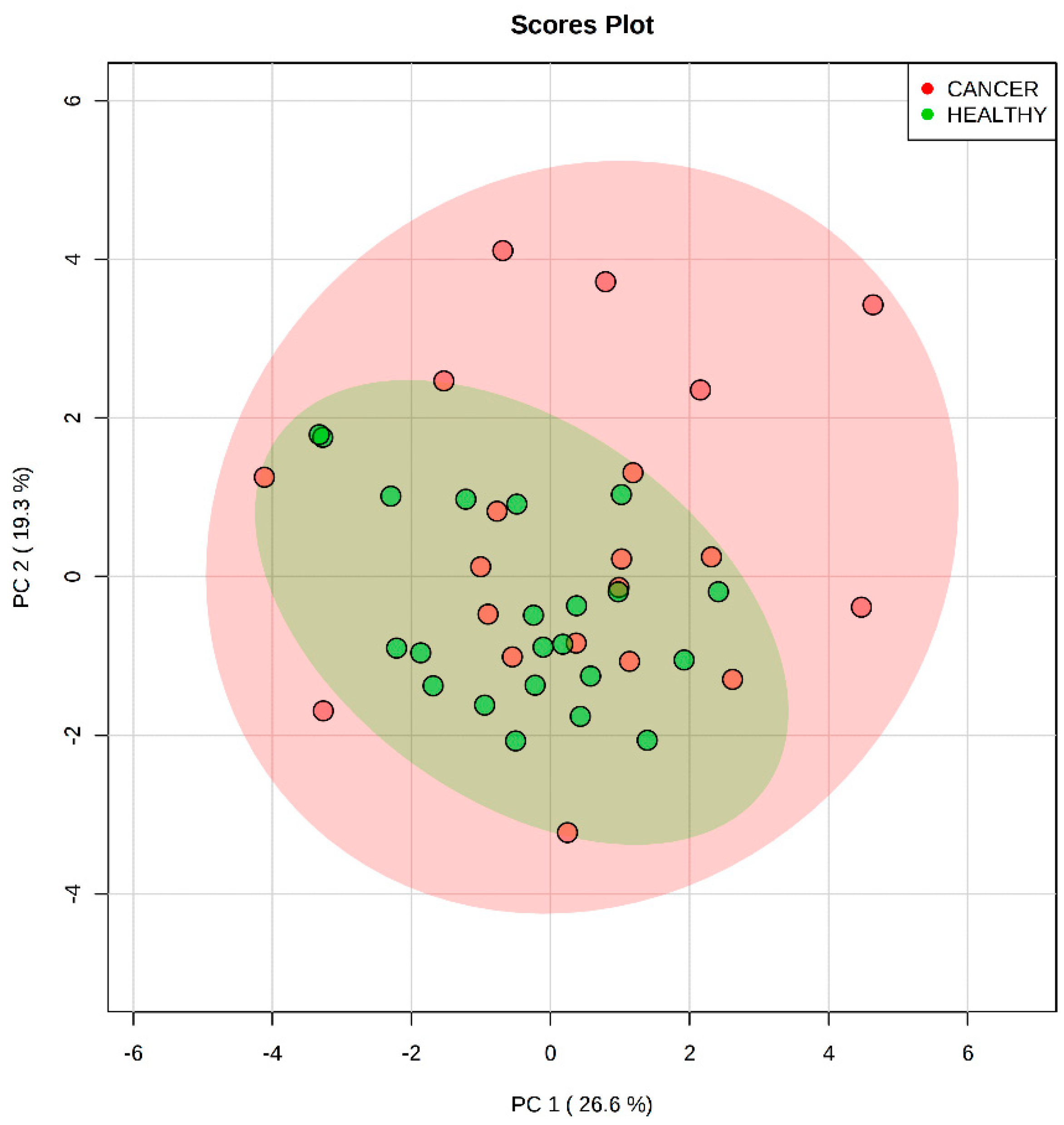
2.2. Analysis of Plasma by LC-MS/MS
2.3. Correspondence of Biomarkers Between Tissue-DESI-MSI and Plasma-LC-MS
3. Discussion
3.1. Molecular Imaging of Breast Tissues by DESI-MSI
3.2. Analysis of Plasma by LC-MS/MS
3.3. Correspondence of Biomarkers Between Tissue and Plasma
4. Materials and Methods
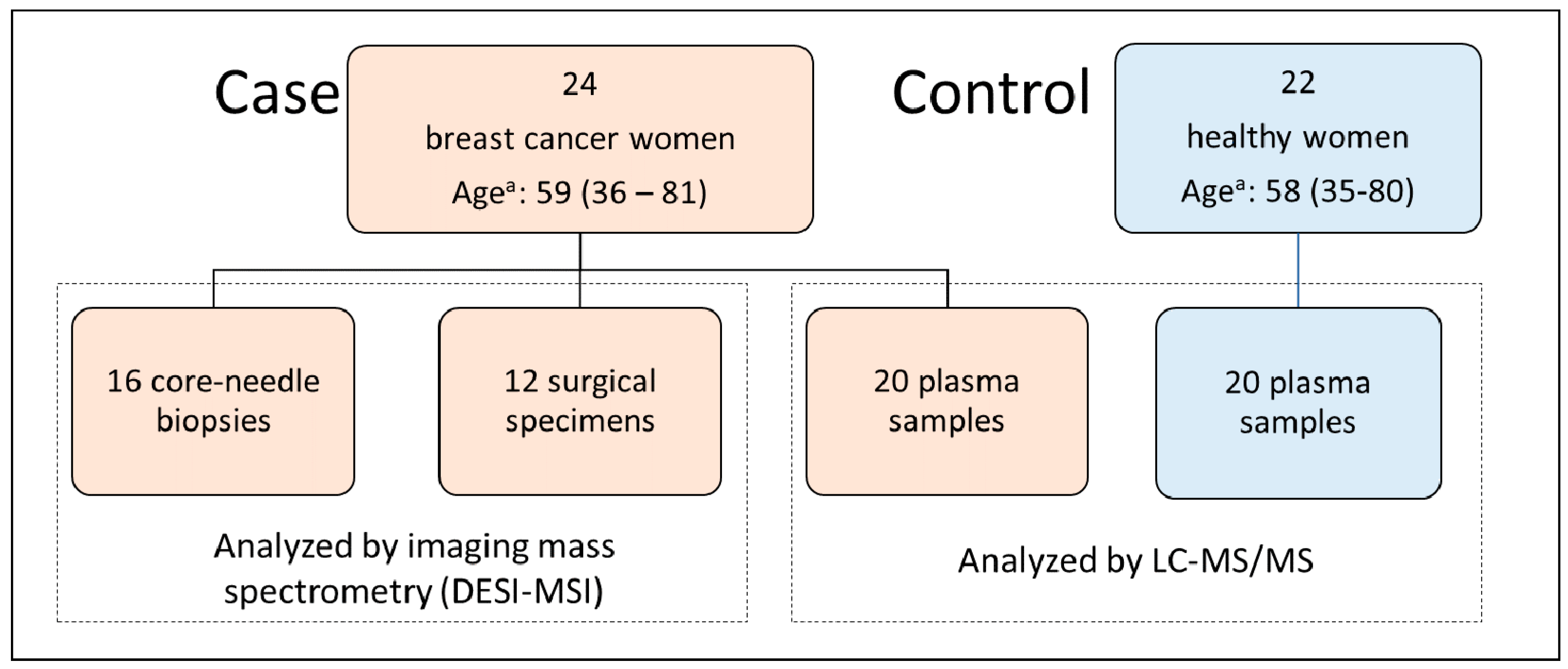
4.1. Subjects and Ethical Consent
4.2. Tissue Samples Analyzed by DESI-MSI
4.3. DESI-MSI Experiments
4.4. Plasma Samples Analyzed by LC-MS
4.4.1. Lipid Extraction
4.4.2. LC-MS Analysis
4.4.3. Data Extraction
4.5. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| AUC | Area under the (ROC) curve |
| CAAE | Brazilian certificate of ethical appreciation approval |
| CAISM-UNICAMP | Center of integrated assistance to women’s health of the University of Campinas |
| Cer | Ceramide |
| CL | Cardiolipin |
| CN:DB | Numbers of fatty acid chain carbons and double bonds in lipid species |
| DC | Ductal carcinoma |
| DCIS | In situ ductal carcinoma |
| DESI-MSI | Desorption-Electrospray-Ionization—Mass Spectrometry |
| ER | Estrogen receptor |
| FA | Fatty acid |
| H&E | Hematoxylin and eosin staining |
| HER2 | Human-epidermal-growth-factor-receptor-2 |
| HR | Hormone receptor status |
| IDC | Invasive ductal carcinoma of the breast |
| Lasso | Least absolute shrinkage and selection operator |
| LC-MS/MS | Liquid chromatography coupled to tandem mass spectrometry |
| LC-MS | Liquid chromatography coupled to mass spectrometry |
| LysoPC | Lysophosphatidylcholine |
| m/z | Mass-to-charge ratio |
| MS | Mass Spectrometry |
| MSE | MS data-independent acquisition |
| MSI | Mass Spectrometry Imaging |
| NST | no special type carcinoma of the breast |
| NPV | Negative predictive value |
| PA | Phosphatidic acid |
| PC | Glycerophosphocholine |
| PC1 | Principal component 1 |
| PC2 | Principal component 2 |
| PCA | Principal Component Analysis |
| PE | Glycerophosphoethanolamine |
| PE | Phosphatidylethanolamine |
| PE-Nme | Methylphosphatidylethanolamine |
| PG | Glycerophosphoglycerol |
| PI | Glycerophosphoinositol |
| ppm | parts per million |
| PPV | Positive predictive value |
| PR | Progesterone receptor |
| PS | Glycerophosphoserine |
| QC | Quality control sample |
| ROC | Receiver operating-characteristic (curves) |
| SVM | Support Vector Machine |
| TG | Triacylglycerol or Triglyceride |
| TIC | Total Ion Current |
References
- Gogiashvili, M.; Nowacki, J.; Hergenröder, R.; Hengstler, J.G.; Lambert, J.; Edlund, K. HR-MAS NMR Based Quantitative Metabolomics in Breast Cancer. Metabolites 2019, 9, 19. [Google Scholar] [CrossRef]
- Hart, C.D.; Tenori, L.; Luchinat, C.; Di Leo, A. Metabolomics in Breast Cancer: Current Status and Perspectives. In Novel Biomarkers in the Continuum of Breast Cancer; Stearns, V., Ed.; Springer International Publishing: Cham, Switzerland, 2016; pp. 217–234. [Google Scholar]
- Wishart, D.S. Emerging applications of metabolomics in drug discovery and precision medicine. Nat. Rev. Drug Discov. 2016, 15, 473. [Google Scholar] [CrossRef]
- Jasbi, P.; Wang, D.; Cheng, S.L.; Fei, Q.; Cui, J.Y.; Liu, L. Breast cancer detection using targeted plasma metabolomics. J. Chromatogr. B. Analyt. Technol. Biomed. Life Sci. 2019, 1105, 26–37. [Google Scholar] [CrossRef] [PubMed]
- Huang, M.; Li, H.-Y.; Liao, H.-W.; Lin, C.-H.; Wang, C.-Y.; Kuo, W.-H.; Kuo, C.-H. Using post-column infused internal standard assisted quantitative metabolomics for establishing prediction models for breast cancer detection. Rapid Commun. Mass Spectrom. 2019, 34. [Google Scholar] [CrossRef] [PubMed]
- Lecuyer, L.; Dalle, C.; Lyan, B.; Demidem, A.; Rossary, A.; Vasson, M.P.; Petera, M.; Lagree, M.; Ferreira, T.; Centeno, D.; et al. Plasma Metabolomic Signatures Associated with Long-term Breast Cancer Risk in the SU.VI.MAX Prospective Cohort. Cancer Epidemiol. Biomarkers Prev. 2019, 28, 300–1307. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Xu, H.; Liu, R.; Gao, P.; Yang, X.; Jin, W. A Novel Strategy for Targeted Lipidomics Based on LC-Tandem-MS Parameters Prediction, Quantification, and Multiple Statistical Data Mining: Evaluation of Lysophosphatidylcholines as Potential Cancer Biomarkers. Anal. Chem. 2019, 91, 3389–3396. [Google Scholar] [CrossRef] [PubMed]
- Cala, M.P.; Aldana, J.; Medina, J.; Sanchez, J.; Guio, J.; Wist, J. Multiplatform plasma metabolic and lipid fingerprinting of breast cancer: A pilot control-case study in Colombian Hispanic women. PLoS ONE 2018, 13. [Google Scholar] [CrossRef]
- Zhang, Q.; Liu, R.; Xu, H.; Yang, X.; Zhang, Y.; Wang, Q.; Gao, P.; Bi, K.; Han, T.; Li, Q. Multifunctional isotopic standards based steroidomics strategy: Exploration of cancer screening model. J. Chromatogr. A 2020, 1614, 460723. [Google Scholar] [CrossRef]
- Luo, X.; Yu, H.; Song, Y.; Sun, T. Integration of metabolomic and transcriptomic data reveals metabolic pathway alteration in breast cancer and impact of related signature on survival. J. Cell Physiol. 2019, 234, 13021–13031. [Google Scholar] [CrossRef]
- Park, J.; Shin, Y.; Kim, T.H.; Kim, D.H.; Lee, A. Plasma metabolites as possible biomarkers for diagnosis of breast cancer. PLoS ONE 2019, 14, e0225129. [Google Scholar] [CrossRef]
- Chen, X.; Chen, H.; Dai, M.; Ai, J.; Li, Y.; Mahon, B.; Dai, S.; Deng, Y. Plasma lipidomics profiling identified lipid biomarkers in distinguishing early-stage breast cancer from benign lesions. Oncotarget 2016, 7, 36622–36631. [Google Scholar] [CrossRef]
- Yamashita, Y.; Nishiumi, S.; Kono, S.; Takao, S.; Azuma, T.; Yoshida, M. Differences in elongation of very long chain fatty acids and fatty acid metabolism between triple-negative and hormone receptor-positive breast cancer. BMC Cancer 2017, 1, 589. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Zheng, X.; Zhou, Q.; Villanueva, N.; Nian, W.; Liu, X.; Huan, T. Metabolomics-Based Discovery of Molecular Signatures for Triple Negative Breast Cancer in Asian Female Population. Sci. Rep. 2020, 10, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.; Xu, R.; Mao, S.; Zhang, Y.; Dai, Y.; Guo, Q.; Song, X.; Zhang, Q.; Li, L.; Chen, Q. Metabolic biomarker signature for predicting the effect of neoadjuvant chemotherapy of breast cancer. Ann. Transl. Med. 2019, 7, 670. [Google Scholar] [CrossRef] [PubMed]
- Silva, C.; Perestrelo, R.; Silva, P.; Tomás, H.; Câmara, J.S. Breast Cancer Metabolomics: From Analytical Platforms to Multivariate Data Analysis. A Review. Metabolites 2019, 9, 102. [Google Scholar] [CrossRef]
- Cardoso, M.R.; Santos, J.C.; Ribeiro, M.L.; Talarico, M.C.R.; Viana, L.R.; Derchain, S.F.M. A Metabolomic Approach to Predict Breast Cancer Behavior and Chemotherapy Response. Int. J. Mol. Sci. 2018, 19, 617. [Google Scholar] [CrossRef]
- McCartney, A.; Vignoli, A.; Biganzoli, L.; Love, R.; Tenori, L.; Luchinat, C.; Di Leo, A. Metabolomics in breast cancer: A decade in review. Cancer Treat. Rev. 2018, 67, 88–96. [Google Scholar] [CrossRef]
- Leung, F.; Eberlin, L.S.; Schwamborn, K.; Heeren, R.M.; Winograd, N.; Cooks, R.G. Mass Spectrometry-Based Tissue Imaging: The Next Frontier in Clinical Diagnostics? Clin. Chem. 2019, 65, 510–513. [Google Scholar] [CrossRef]
- Perez, C.J.; Bagga, A.K.; Prova, S.S.; Taemeh, M.Y.; Ifa, D.R. Review and perspectives on the applications of mass spectrometry imaging under ambient conditions. Rapid Commun. Mass Spectrom. 2019, 33, 27–53. [Google Scholar] [CrossRef]
- Feider, C.L.; Krieger, A.C.; Dehoog, R.J.; Eberlin, L.S. Ambient ionization mass spectrometry: Recent developments and applications. Anal. Chem. 2019, 91, 4266–4290. [Google Scholar] [CrossRef]
- Ifa, D.R.; Eberlin, L.S. Ambient Ionization Mass Spectrometry for Cancer Diagnosis and Surgical Margin Evaluation. Clin. Chem. 2016, 62, 111–123. [Google Scholar] [CrossRef]
- Woolman, M.; Zarrine-Afsar, A. Platforms for rapid cancer characterization by ambient mass spectrometry: Advancements, challenges, and opportunities for improvement towards intrasurgical use. Analyst 2018, 143, 2717–2722. [Google Scholar] [CrossRef] [PubMed]
- Jarmusch, A.K.; Pirro, V.; Baird, Z.; Hattab, E.M.; Cohen-Gadol, A.A.; Cooks, R.G. Lipid, and metabolite profiles of human brain tumors by desorption electrospray ionization-MS. Proc. Natl. Acad Sci. USA 2016, 113, 1486–1491. [Google Scholar] [CrossRef] [PubMed]
- Eberlin, L.S.; Ifa, D.R.; Wu, C.; Cooks, R.G. Three-Dimensional Vizualization of Mouse Brain by Lipid Analysis Using Ambient Ionization Mass Spectrometry. Angew. Chem. Int. Edit. 2010, 49, 873–876. [Google Scholar] [CrossRef] [PubMed]
- Eberlin, L.S.; Gabay, M.; Fan, A.C.; Gouw, A.M.; Tibshirani, R.J.; Felsher, D.W.; Zare, R.N. Alteration of the lipid profile in lymphomas induced by MYC overexpression. Proc. Natl. Acad Sci. USA 2014, 111, 10450–10455. [Google Scholar] [CrossRef]
- Woolman, M.; Tata, A.; Dara, D.; Meens, J.; D’Arcangelo, E.; Perez, C.J.; Saiyara Prova, S.; Bluemke, E.; Ginsberg, H.J.; Ifa, D.; et al. Rapid determination of the tumour stroma ratio in squamous cell carcinomas with desorption electrospray ionization mass spectrometry (DESI-MS): A proof-of-concept demonstration. Analyst 2017, 142, 3250–3260. [Google Scholar] [CrossRef] [PubMed]
- Porcari, A.M.; Zhang, J.; Garza, K.Y.; Rodrigues-Peres, R.M.; Lin, J.Q.; Young, J.H.; Tibshirani, R.; Nagi, C.; Paiva, G.P.; Carter, S.A. Multicenter Study Using Desorption-Electrospray-Ionization-Mass-Spectrometry Imaging for Breast-Cancer Diagnosis. Anal. Chem. 2018, 90, 11324–11332. [Google Scholar] [CrossRef] [PubMed]
- Calligaris, D.; Caragacianu, D.; Liu, X.; Norton, I.; Thompson, C.J.; Richardson, A.L.; Golshan, M.; Easterling, M.L.; Santagata, S.; Dillon, D.A.; et al. Application of desorption electrospray ionization mass spectrometry imaging in breast cancer margin analysis. Proc. Natl. Acad Sci. USA 2014, 111, 15184–15189. [Google Scholar] [CrossRef]
- Guenther, S.; Muirhead, L.J.; Golf, O.; Strittmatter, N.; Ramakrishnan, R.; Goldin, R.D.; Jones, E.A.; Veselkov, K.; Darzi, A.; Takáts, Z.; et al. Spatially Resolved Metabolic Phenotyping of Breast Cancer by Desorption Electrospray Ionization Mass Spectrometry. Cancer Res. 2015, 75, 1828–1837. [Google Scholar] [CrossRef]
- Tata, A.; Gribble, A.; Ventura, M.; Ganguly, M.; Bluemke, E.; Ginsberg, H.J.; Jaffray, D.A.; Ifa, D.R.; Vitkin, A.; Zarrine-Afsar, A. Wide-field tissue polarimetry allows efficient localized mass spectrometry imaging of biological tissues. Chem. Sci. 2016, 7, 2162–2169. [Google Scholar] [CrossRef]
- Woolman, M.; Tata, A.; Bluemke, E.; Dara, D.; Ginsberg, H.J.; Zarrine-Afsar, A. An Assessment of the Utility of Tissue Smears in Rapid Cancer Profiling with Desorption Electrospray Ionization Mass Spectrometry (DESI-MS). J. Am. Soc. Mass Spectrom 2017, 28, 145–153. [Google Scholar] [CrossRef] [PubMed]
- Tata, A.; Woolman, M.; Ventura, M.; Bernards, N.; Ganguly, M.; Gribble, A.; Shrestha, B.; Bluemke, E.; Ginsberg, H.J.; Vitkin, A.; et al. Rapid Detection of Necrosis in Breast Cancer with Desorption Electrospray Ionization Mass Spectrometry. Sci. Rep. 2016, 6, 35374. [Google Scholar] [CrossRef] [PubMed]
- Santoro, A.L.; Drummond, R.D.; Silva, I.T.; Ferreira, S.S.; Juliano, L.; Vendramini, P.H.; Lemos, M.B.D.C.; Eberlin, M.N.; De Andrade, V.P. In Situ DESI-MSI Lipidomic Profiles of Breast Cancer Molecular Subtypes and Precursor Lesions. Cancer Res. 2020, 80, 1246–1257. [Google Scholar] [CrossRef] [PubMed]
- Kertesz, V.; Van Berkel, G.J.; Vavrek, M.; Koeplinger, K.A.; Schneider, B.B.; Covey, T.R. Comparison of drug distribution images from whole-body thin tissue sections obtained using desorption electrospray ionization tandem mass spectrometry and autoradiography. Anal. Chem 2008, 80, 5168–5177. [Google Scholar] [CrossRef]
- Wiseman, J.M.; Ifa, D.R.; Zhu, Y.; Kissinger, C.B.; Manicke, N.E.; Kissinger, P.T. Desorption electrospray ionization mass spectrometry: Imaging drugs and metabolites in tissues. Proc. Natl. Acad Sci. USA 2008, 105, 18120–18125. [Google Scholar] [CrossRef]
- Vismeh, R.; Waldon, D.J.; Teffera, Y.; Zhao, Z. Localization, and quantification of drugs in animal tissues by use of desorption electrospray ionization mass spectrometry imaging. Anal. Chem. 2012, 84, 5439–5445. [Google Scholar] [CrossRef]
- Tata, A.; Perez, C.; Campos, M.L.; Bayfield, M.A.; Eberlin, M.N.; Ifa, D.R. Imprint Desorption Electrospray Ionization Mass Spectrometry Imaging for Monitoring Secondary Metabolites Production during Antagonistic Interaction of Fungi. Anal. Chem. 2015, 87, 12298–12305. [Google Scholar] [CrossRef]
- Abbassi-Ghadi, N.; Jones, E.A.; Romero, M.G.; Golf, O.; Kumar, S.; Huang, J.; Kudo, H.; Goldin, R.D.; Hanna, G.B.; Takáts, Z. A Comparison of DESI-MS and LC-MS for the Lipidomic Profiling of Human Cancer Tissue. J. Am. Soc. Mass Spectrom. 2015, 27, 255–264. [Google Scholar] [CrossRef]
- Gromski, P.S.; Muhamadali, H.; Ellis, D.I.; Xu, Y.; Correa, E.; Turner, M.L.; Goodacre, R. A tutorial review: Metabolomics and partial least squares-discriminant analysis—a marriage of convenience or a shotgun wedding. Anal. Chim. Acta 2015, 879, 10–23. [Google Scholar] [CrossRef]
- Qiu, Y.; Zhou, B.; Su, M.; Baxter, S.; Zheng, X.; Zhao, X.; Yen, Y.; Jia, W. Mass Spectrometry-Based Quantitative Metabolomics Revealed a Distinct Lipid Profile in Breast Cancer Patients. Int. J. Mol. Sci. 2013, 14, 8047–8061. [Google Scholar] [CrossRef]
- Jiang, N.; Zhang, G.; Pan, L.; Yan, C.; Zhang, L.; Weng, Y.; Wang, W.; Chen, X.; Yang, G. Potential plasma lipid biomarkers in early-stage breast cancer. Biotechnol. Lett. 2017, 39, 1657–1666. [Google Scholar] [CrossRef] [PubMed]
- Goodacre, R.; Vaidyanathan, S.; Dunn, W.B.; Harrigan, G.G.; Kell, D.B. Metabolomics by numbers: Acquiring and understanding global metabolite data. Trends Biotechnol. 2004, 22, 245–252. [Google Scholar] [CrossRef] [PubMed]
- Pavlova, N.N.; Thompson, C.B. The Emerging Hallmarks of Cancer Metabolism. Cell Metab. 2016, 23, 27–47. [Google Scholar] [CrossRef]
- Brglez, V.; Pucer, A.; Pungercar, J.; Lambeau, G.; Petan, T. Secreted phospholipases A (2)are differentially expressed and epigenetically silenced in human breast cancer cells. Biochem. Biophys Res. Commun. 2014, 445, 230–235. [Google Scholar] [CrossRef]
- Iorio, E.; Caramujo, M.J.; Cecchetti, S.; Spadaro, F.; Carpinelli, G.; Canese, R.; Podo, F. Key Players in Choline Metabolic Reprograming in Triple-Negative Breast Cancer. Front. Oncol. 2016, 6, 747. [Google Scholar] [CrossRef]
- Taylor, L.A.; Arends, J.; Hodina, A.K.; Unger, C.; Massing, U. Plasma lyso-phosphatidylcholine concentration is decreased in cancer patients with weight loss and activated inflammatory status. Lipids Heal. Dis. 2007, 6, 17. [Google Scholar] [CrossRef]
- Yamashita, S.; Yamashita, J.; Ogawa, M. Overexpression of group II phospholipase A2 in human breast cancer tissues is closely associated with their malignant potency. Br. J. Cancer 1994, 69, 1166–1170. [Google Scholar] [CrossRef]
- Yarla, N.S.; Satyakumar, K.; Srinivasu, D.; Dsvkg, K.; Aliev, G.; Dharmapuri, G. Phospholipase A2: A Potential Therapeutic Target in Inflammation and Cancer (In silico, In vitro, In vivo and Clinical Approach). J. Cancer Sci. Therapy 2015, 7, 249–252. [Google Scholar]
- Lofterød, T.; Mortensen, E.S.; Nalwoga, H.; Wilsgaard, T.; Frydenberg, H.; Risberg, T.; Eggen, A.E.; McTiernan, A.; Aziz, S.; Wist, E.A.; et al. Impact of pre-diagnostic triglycerides and HDL-cholesterol on breast cancer recurrence and survival by breast cancer subtypes. BMC Cancer 2018, 18, 654. [Google Scholar] [CrossRef]
- Zhang, F.; Du, G. Dysregulated lipid metabolism in cancer. World J. Biol. Chem. 2012, 3, 167–174. [Google Scholar] [CrossRef]
- Sailo, B.L.; Banik, K.; Padmavathi, G.; Javadi, M.; Bordoloi, D.; Kunnumakkara, A.B. Tocotrienols: The promising analogues of vitamin E for cancer therapeutics. Pharmacol. Res. 2018, 130, 259–272. [Google Scholar] [CrossRef] [PubMed]
- Sheth, S.A.; Iavarone, A.T.; Liebeskind, D.S.; Won, S.J.; Swanson, R.A. Targeted Lipid Profiling Discovers Plasma Biomarkers of Acute Brain Injury. PLoS ONE 2015, 10, e0129735. [Google Scholar] [CrossRef] [PubMed]
- Bokhart, M.; Nazari, M.; Garrard, K.P.; Muddiman, D.C. MSiReader v1.0: Evolving Open-Source Mass Spectrometry Imaging Software for Targeted and Untargeted Analyses. J. Am. Soc. Mass Spectrom. 2017, 29, 8–16. [Google Scholar] [CrossRef] [PubMed]
| Measured m/z | Ion Mode | Species | Lipid Assignment | Proposed Formula | Exact m/z | Mass Error (ppm) |
|---|---|---|---|---|---|---|
| Characteristic of healthy plasma samples | ||||||
| 496.340 | + | [M + H]+ | LysoPC(16:0) | C24H51NO7P | 496.340 | 0.0 |
| 524.371 | + | [M + H]+ | LysoPC(18:0) | C26H55NO7P | 524.372 | 1.9 |
| 782.569 | + | [M + H]+ | PC(40:4) | C44H81NO8P | 782.570 | 1.3 |
| 810.600 | + | [M + H]+ | PC(38:4) | C46H85NO8P | 810.601 | 1.2 |
| 540.330 | − | [M + FA − H]− | LysoPC(16:0) | C25H51NO9P | 540.330 | 0.0 |
| 568.361 | − | [M + FA − H]− | LysoPC(18:0) | C27H55NO9P | 568.361 | 0.0 |
| 588.330 | − | [M + FA − H]− | LysoPC (20:4) | C29H51NO9P | 588.330 | 0.0 |
| 566.346 | − | [M + FA − H]− | LysoPC(18:1) | C27H53NO9P | 566.346 | 0.0 |
| Characteristic of cancer plasma samples | ||||||
| 786.600 | + | [M + H]+ | PC(36:2) | C44H85NO8P | 786.601 | 1.3 |
| 796.738 | + | [M + NH4]+ | TG (46:0) | C49H98NO6 | 796.739 | 1.3 |
| 758.570 | + | [M + H]+ | PC(34:2) | C42H81NO8P | 758.570 | 0.0 |
| 824.770 | + | [M + NH4]+ | TG(48:0) | C51H102NO6 | 824.771 | 1.2 |
| 407.294 | − | [M − H2O − H]− | 13′-Hydroxy-gamma-tocotrienol | C28H39O2 | 407.295 | 2.5 |
| 409.310 | − | [M − H]− | gamma-tocotrienol | C28H41O2 | 409.311 | 2.4 |
| 802.559 | − | [M + FA − H]− | PC(34:2)/PE-Nme(36:2) | C43H81NO10P | 802.560 | 1.2 |
| 830.590 | − | [M + FA − H]− | PC(36:2) | C45H85NO10P | 830.591 | 1.2 |
| 776.544 | − | [M + FA − H]− | PC(32:1) | C41H79NO10P | 776.544 | 0.0 |
| Tissue Biomarkers for No Special Type (NST) Ductal Carcinoma of the Breast [28] | Prevalence in Plasma Samples of Breast Carcinoma (NST and Special Type) Patients According to LC-MS/MS Results |
|---|---|
| PS(34:1); PE(38:4); PS(38:4); PI(34:1); PS(40:4); PI(36:2); PI(38:3); PE(36:2); PE(O-38:6); PE(O-38:5); PS(36:2); PS(36:1); PC(34:2); PC(34:1); PS(38:1); PI(34:0); PI(38:4) | Yes |
| PG(34:1); PG(36:2); PG(40:7); PS(O-41:0); Cer(t42:1); CL(72:8); CL(72:7); PA(38:2); PS(O-33:0); PE(O-38:4); PG(36:4); PS(P-36:2); PE(39:5); TG(52:3) | No |
| Characteristics | Patients, N | Median Age (Range) |
|---|---|---|
| Core needle biopsy | 16 | 60 (37–80) |
| Surgical specimen | 12 | 61 (36–81) |
| Core needle biopsy + surgical specimen | 5 | 63 (37–80) |
| Plasma | 20 | 58 (36–81) |
| Tumor type | ||
| Ductal NST (no special type) | 16 | 56 (36–81) |
| Special Types | 8 | 65 (37–80) |
| Tumor stage | ||
| I | 10 | 57 (43–77) |
| II | 8 | 59 (36–81) |
| III | 3 | 64 (37–80) |
| IV | 3 | 61 (42–75) |
| Estrogen receptor status | ||
| Positive | 20 | 58 (36–81) |
| Negative | 4 | 65 (42–77) |
| Progesterone receptor status | ||
| Positive | 16 | 56 (36–81) |
| Negative | 8 | 65 (42–80) |
| HER2 receptor status | ||
| Positive | 6 | 47 (36–67) |
| Negative | 18 | 63 (38–81) |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Silva, A.A.R.; Cardoso, M.R.; Rezende, L.M.; Lin, J.Q.; Guimaraes, F.; Silva, G.R.P.; Murgu, M.; Priolli, D.G.; Eberlin, M.N.; Tata, A.; et al. Multiplatform Investigation of Plasma and Tissue Lipid Signatures of Breast Cancer Using Mass Spectrometry Tools. Int. J. Mol. Sci. 2020, 21, 3611. https://doi.org/10.3390/ijms21103611
Silva AAR, Cardoso MR, Rezende LM, Lin JQ, Guimaraes F, Silva GRP, Murgu M, Priolli DG, Eberlin MN, Tata A, et al. Multiplatform Investigation of Plasma and Tissue Lipid Signatures of Breast Cancer Using Mass Spectrometry Tools. International Journal of Molecular Sciences. 2020; 21(10):3611. https://doi.org/10.3390/ijms21103611
Chicago/Turabian StyleSilva, Alex Ap. Rosini, Marcella R. Cardoso, Luciana Montes Rezende, John Q. Lin, Fernando Guimaraes, Geisilene R. Paiva Silva, Michael Murgu, Denise Gonçalves Priolli, Marcos N. Eberlin, Alessandra Tata, and et al. 2020. "Multiplatform Investigation of Plasma and Tissue Lipid Signatures of Breast Cancer Using Mass Spectrometry Tools" International Journal of Molecular Sciences 21, no. 10: 3611. https://doi.org/10.3390/ijms21103611
APA StyleSilva, A. A. R., Cardoso, M. R., Rezende, L. M., Lin, J. Q., Guimaraes, F., Silva, G. R. P., Murgu, M., Priolli, D. G., Eberlin, M. N., Tata, A., Eberlin, L. S., Derchain, S. F. M., & Porcari, A. M. (2020). Multiplatform Investigation of Plasma and Tissue Lipid Signatures of Breast Cancer Using Mass Spectrometry Tools. International Journal of Molecular Sciences, 21(10), 3611. https://doi.org/10.3390/ijms21103611






