HMGA Proteins in Stemness and Differentiation of Embryonic and Adult Stem Cells
Abstract
1. Introduction
2. The Phenotypes of Hmga KO and Transgenic Mice
3. HMGA Functions in Embryonic Stem Cell (ESC) Stemness and Differentiation
4. HMGA Proteins in Adult Stem Cells
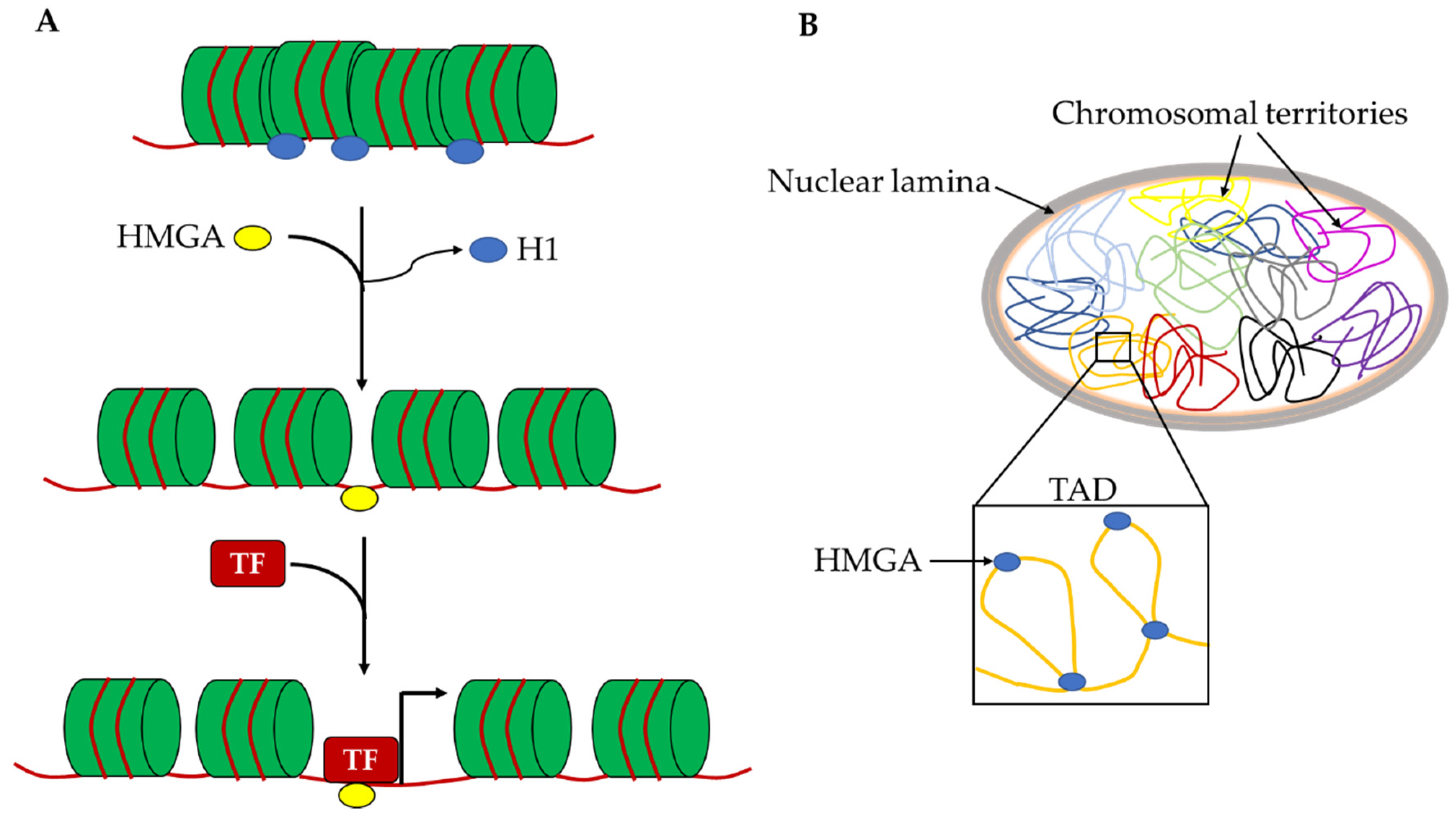
5. HMGA Proteins as Regulators of Chromatin Architecture and Gene Expression
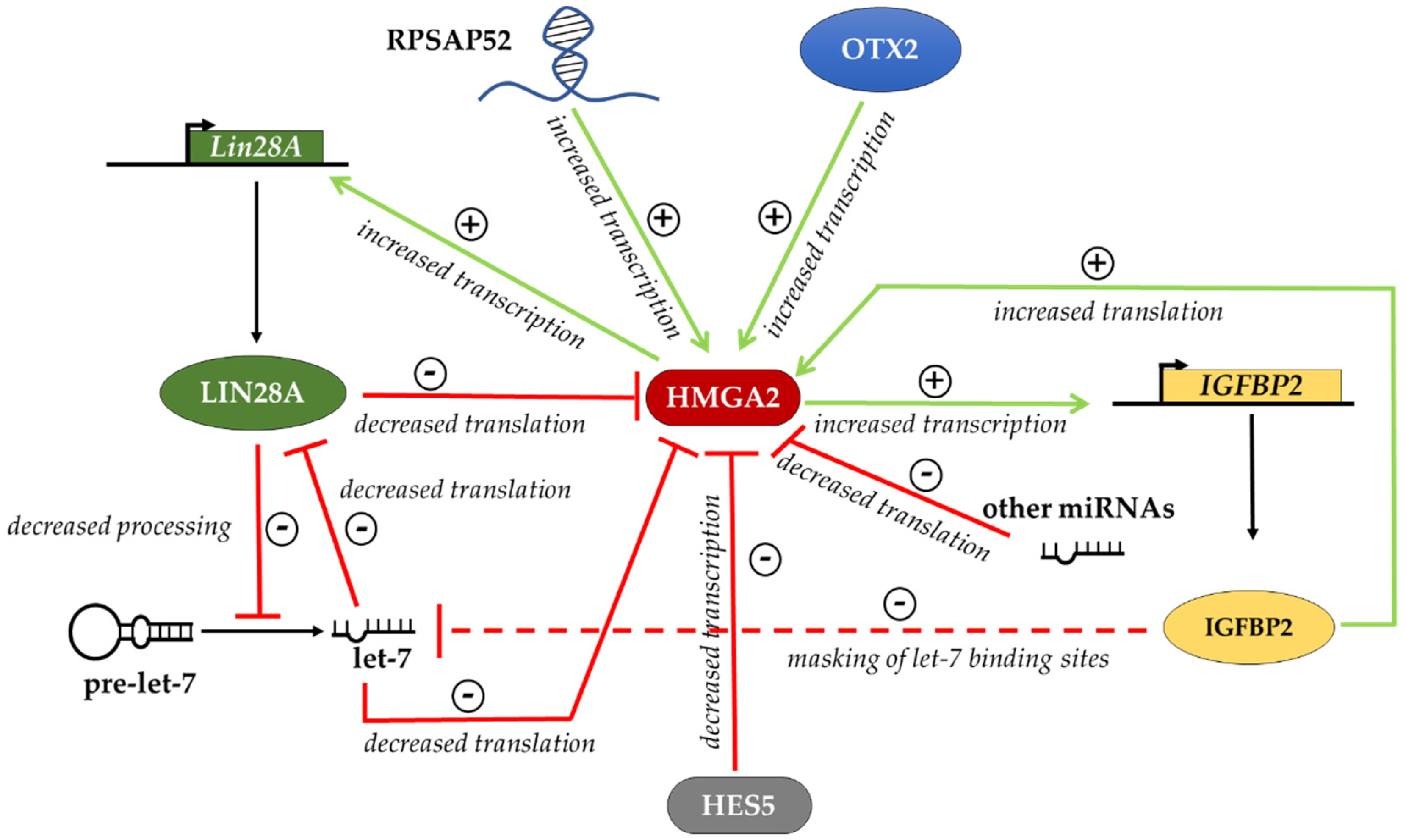
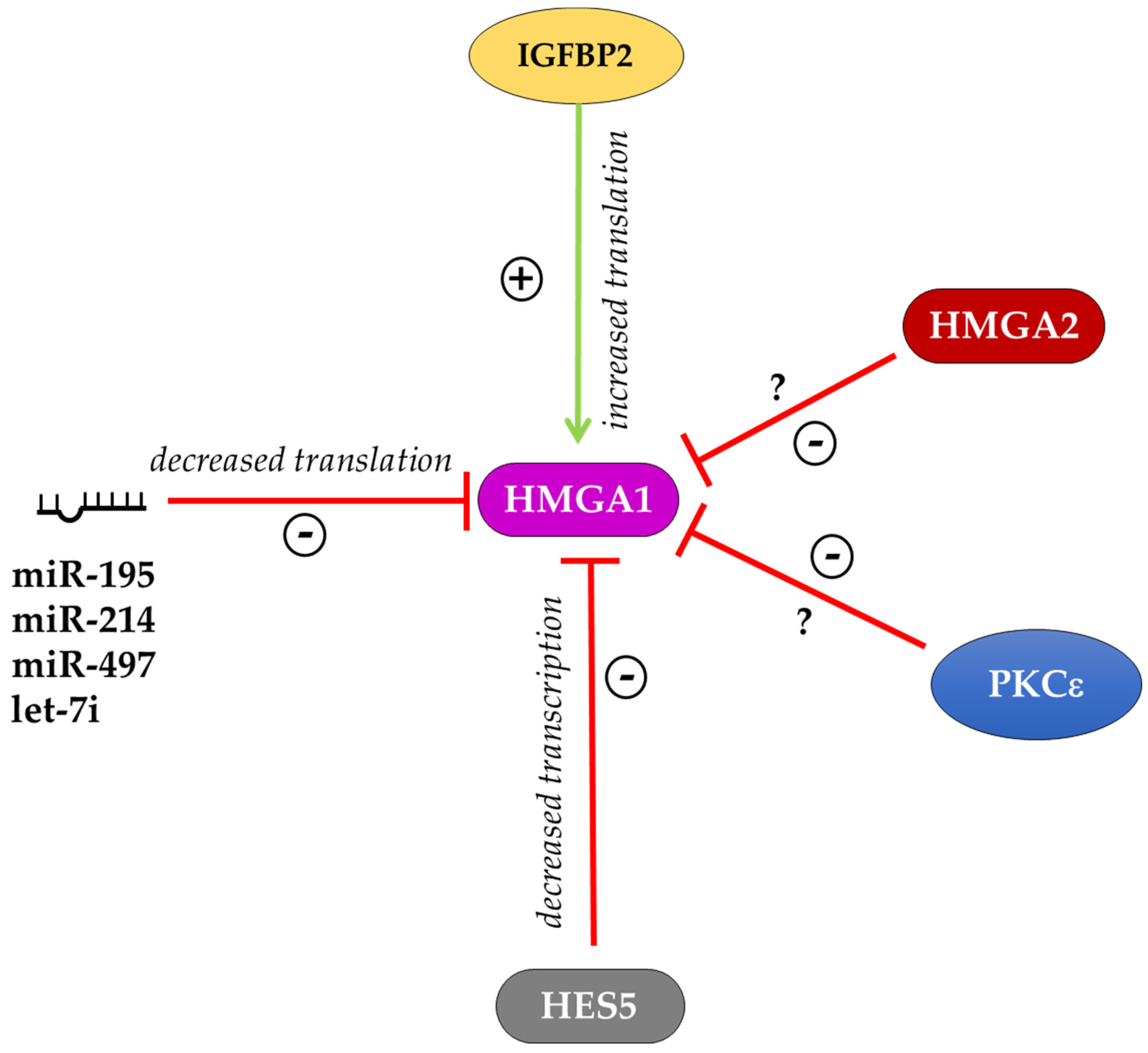
6. Regulation of HMGA Proteins in Different Cellular Contexts
7. Conclusions
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Pallante, P.; Sepe, R.; Puca, F.; Fusco, A. High mobility group a proteins as tumor markers. Front. Med. (Lausanne) 2015, 2, 15. [Google Scholar] [CrossRef] [PubMed]
- Cui, T.; Wei, S.; Brew, K.; Leng, F.J. Energetics of binding the mammalian high mobility group protein HMGA2 to poly(dA-dT)2 and poly(dA)-poly(dT). Mol. Biol. 2005, 352, 629–645. [Google Scholar] [CrossRef] [PubMed]
- Reeves, R.; Beckerbauer, L. HMGI/Y proteins: Flexible regulators of transcription and chromatin structure. Biochim. Biophys. Acta 2001, 1519, 13–29. [Google Scholar] [CrossRef]
- Johnson, K.R.; Lehn, D.A.; Reeves, R. Alternative processing of mRNAs encoding mammalian chromosomal high-mobility-group proteins HMG-I and HMG-Y. Mol. Cell. Biol. 1989, 9, 2114–2123. [Google Scholar] [CrossRef] [PubMed]
- Cesana, M.; Guo, M.H.; Cacchiarelli, D.; Wahlster, L.; Barragan, J.; Doulatov, S.; Vo, L.T.; Salvatori, B.; Trapnell, C.; Clement, K.; et al. A CLK3-HMGA2 Alternative Splicing Axis Impacts Human Hematopoietic Stem Cell Molecular Identity throughout Development. Cell Stem Cell 2018, 22, 575–588. [Google Scholar] [CrossRef]
- Zhou, X.; Benson, K.F.; Ashar, H.R.; Chada, K. Mutation responsible for the mouse pygmy phenotype in the developmentally regulated factor HMGI-C. Nature 1995, 376, 771–774. [Google Scholar] [CrossRef] [PubMed]
- Chiappetta, G.; Avantaggiato, V.; Visconti, R.; Fedele, M.; Battista, S.; Trapasso, F.; Merciai, B.M.; Fidanza, V.; Giancotti, V.; Santoro, M.; et al. High level expression of the HMGI (Y) gene during embryonic development. Oncogene 1996, 13, 2439–2446. [Google Scholar]
- Battista, S.; Pentimalli, F.; Baldassarre, G.; Fedele, M.; Fidanza, V.; Croce, C.M.; Fusco, A. Loss of Hmga1 gene function affects embryonic stem cell lympho-hematopoietic differentiation. FASEB J. 2003, 17, 1496–1498. [Google Scholar] [CrossRef]
- Rohen, C.; Rogalla, P.; Meyer-Bolte, K.; Bartnitzke, S.; Chilla, R.; Bullerdiek, J. Pleomorphic adenomas of the salivary glands: Absence of HMGIY rearrangements. Cancer Genet. Cytogenet. 1999, 111, 178–181. [Google Scholar] [CrossRef]
- Sornberger, K.S.; Weremowicz, S.; Williams, A.J.; Quade, B.J.; Ligon, A.H.; Pedeutour, F.; Vanni, R.; Morton, C.C. Expression of HMGIY in three uterine leiomyomata with complex rearrangements of chromosome 6. Cancer Genet. Cytogenet. 1999, 114, 9–16. [Google Scholar] [CrossRef]
- Battista, S.; Fidanza, V.; Fedele, M.; Klein-Szanto, A.J.; Outwater, E.; Brunner, H.; Santoro, M.; Croce, C.M.; Fusco, A. The expression of a truncated HMGI-C gene induces gigantism associated with lipomatosis. Cancer Res. 1999, 59, 4793–4797. [Google Scholar] [PubMed]
- Arlotta, P.; Tai, A.K.; Manfioletti, G.; Clifford, C.; Jay, G.; Ono, S.J. Transgenic mice expressing a truncated form of the high mobility group I-C protein develop adiposity and an abnormally high prevalence of lipomas. J. Biol. Chem. 2000, 275, 14394–14400. [Google Scholar] [CrossRef]
- Zaidi, M.R.; Okada, Y.; Chada, K.K. Misexpression of full-length HMGA2 induces benign mesenchymal tumors in mice. Cancer Res. 2006, 66, 7453–7459. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Abe, N.; Watanabe, T.; Suzuki, Y.; Matsumoto, N.; Masaki, T.; Mori, T.; Sugiyama, M.; Chiappetta, G.; Fusco, A.; Atomi, Y. An increased high-mobility group A2 expression level is associated with malignant phenotype in pancreatic exocrine tissue. Br. J. Cancer 2003, 89, 2104–2109. [Google Scholar] [CrossRef] [PubMed]
- Flohr, A.M.; Rogalla, P.; Bonk, U.; Puettmann, B.; Buerger, H.; Gohla, G.; Packeisen, J.; Wosniok, W.; Loeschke, S.; Bullerdiek, J. High mobility group protein HMGA1 expression in breast cancer reveals a positive correlation with tumour grade. Histol. Histopathol. 2003, 18, 999–1004. [Google Scholar] [PubMed]
- Narita, M.; Krizhanovsky, V.; Nuñez, S.; Chicas, A.; Hearn, S.A.; Myers, M.P.; Lowe, S.W. A novel role for high-mobility group a proteins in cellular senescence and heterochromatin formation. Cell 2006, 126, 503–514. [Google Scholar] [CrossRef] [PubMed]
- D’Angelo, D.; Mussnich, P.; Arra, C.; Battista, S.; Fusco, A. Critical role of HMGA proteins in cancer cell chemoresistance. J. Mol. Med. 2017, 95, 353–360. [Google Scholar] [CrossRef]
- Sgarra, R.; Pegoraro, S.; Ros, G.; Penzo, C.; Chiefari, E.; Foti, D.; Brunetti, A.; Manfioletti, G. High Mobility Group A (HMGA) proteins: Molecular instigators of breast cancer onset and progression. Biochim Biophys Acta Rev. Cancer 2018, 1869, 216–229. [Google Scholar] [CrossRef]
- Foti, D.; Chiefari, E.; Fedele, M.; Iuliano, R.; Brunetti, L.; Paonessa, F.; Manfioletti, G.; Barbetti, F.; Brunetti, A.; Croce, C.M.; et al. Lack of the architectural factor HMGA1 causes insulin resistance and diabetes in humans and mice. Nat. Med. 2005, 11, 765–773. [Google Scholar] [CrossRef]
- Fedele, M.; Fidanza, V.; Battista, S.; Pentimalli, F.; Klein-Szanto, A.J.; Visone, R.; De Martino, I.; Curcio, A.; Morisco, C.; Del Vecchio, L.; et al. Haploinsufficiency of the Hmga1 gene causes cardiac hypertrophy and myelo-lymphoproliferative disorders in mice. Cancer Res. 2006, 66, 2536–2543. [Google Scholar] [CrossRef]
- Li, Z.; Gilbert, J.A.; Zhang, Y.; Zhang, M.; Qiu, Q.; Ramanujan, K.; Shavlakadze, T.; Eash, J.K.; Scaramozza, A.; Goddeeris, M.M.; et al. An HMGA2-IGF2BP2 axis regulates myoblast proliferation and myogenesis. Dev. Cell 2012, 23, 1176–1188. [Google Scholar] [CrossRef] [PubMed]
- Federico, A.; Forzati, F.; Esposito, F.; Arra, C.; Palma, G.; Barbieri, A.; Palmieri, D.; Fedele, M.; Pierantoni, G.M.; De Martino, I.; et al. Hmga1/Hmga2 double knock-out mice display a “superpygmy” phenotype. Biol. Open 2014, 3, 372–378. [Google Scholar] [CrossRef] [PubMed]
- Ligon, A.H.; Moore, S.D.; Parisi, M.A.; Mealiffe, M.E.; Harris, D.J.; Ferguson, H.L.; Quade, B.J.; Morton, C.C. Constitutional rearrangement of the architectural factor HMGA2: A novel human phenotype including overgrowth and lipomas. Am. J. Hum. Genet. 2005, 76, 340–348. [Google Scholar] [CrossRef] [PubMed]
- Shah, S.N.; Kerr, C.; Cope, L.; Zambidis, E.; Liu, C.; Hillion, J.; Belton, A.; Huso, D.L.; Resar, L.M. HMGA1 reprograms somatic cells into pluripotent stem cells by inducing stem cell transcriptional networks. PLoS ONE 2012, 7, e48533. [Google Scholar] [CrossRef]
- Takahashi, K.; Yamanaka, S. A decade of transcription factor-mediated reprogramming to pluripotency. Nat. Rev. Mol. Cell Biol. 2016, 17, 183–193. [Google Scholar] [CrossRef]
- Li, O.; Vasudevan, D.; Davey, C.A.; Droge, P. High level expression of DNA architectural factor HMGA2 and its association with nucleosomes in human embryonic stem cells. Genesis 2006, 44, 523–529. [Google Scholar] [CrossRef]
- Li, O.; Li, J.; Dröge, P. DNA architectural factor and proto-oncogene HMGA2 regulates key developmental genes in pluripotent human embryonic stem cells. FEBS Lett. 2007, 581, 3533–3537. [Google Scholar] [CrossRef]
- Yu, H.; Lim, H.H.; Tjokro, N.O.; Sathiyanathan, P.; Natarajan, S.; Chew, T.W.; Klonisch, T.; Goodman, S.D.; Surana, U.; Dröge, P. Chaperoning HMGA2 protein protects stalled replication forks in stem and cancer cells. Cell Rep. 2014, 6, 684–697. [Google Scholar] [CrossRef]
- Caron, L.; Bost, F.; Prot, M.; Hofman, P.; Binétruy, B. A new role for the oncogenic high-mobility group A2 transcription factor in myogenesis of embryonic stem cells. Oncogene 2005, 24, 6281–6291. [Google Scholar] [CrossRef]
- Navarra, A.; Musto, A.; Gargiulo, A.; Petrosino, G.; Pierantoni, G.M.; Fusco, A.; Russo, T.; Parisi, S. Hmga2 is necessary for Otx2-dependent exit of embryonic stem cells from the pluripotent ground state. BMC Biol. 2016, 14, 24. [Google Scholar] [CrossRef]
- Parisi, S.; Passaro, F.; Russo, L.; Musto, A.; Navarra, A.; Romano, S.; Petrosino, G.; Russo, T. Lin28 is induced in primed embryonic stem cells and regulates let-7-independent events. FASEB J. 2017, 31, 1046–1058. [Google Scholar] [CrossRef] [PubMed]
- Buecker, C.; Srinivasan, R.; Wu, Z.; Calo, E.; Acampora, D.; Faial, T.; Simeone, A.; Tan, M.; Swigut, T.; Wysocka, J. Reorganization of enhancer patterns in transition from naïve to primed pluripotency. Cell Stem Cell 2014, 14, 838–853. [Google Scholar] [CrossRef] [PubMed]
- Acampora, D.; Di Giovannantonio, L.G.; Simeone, A. Otx2 is an intrinsic determinant of the embryonic stem cell state and is required for transition to a stable epiblast stem cell condition. Development 2013, 140, 43–55. [Google Scholar] [CrossRef] [PubMed]
- Ring, K.L.; Tong, L.M.; Balestra, M.E.; Javier, R.; Andrews-Zwilling, Y.; Li, G.; Walker, D.; Zhang, W.R.; Kreitzer, A.C.; Huang, Y. Direct reprogramming of mouse and human fibroblasts into multipotent neural stem cells with a single factor. Cell Stem Cell 2012, 11, 100–109. [Google Scholar] [CrossRef] [PubMed]
- Yu, K.R.; Shin, J.H.; Kim, J.J.; Koog, M.G.; Lee, J.Y.; Choi, S.W.; Kim, H.S.; Seo, Y.; Lee, S.; Shin, T.H.; et al. Rapid and Efficient Direct Conversion of Human Adult Somatic Cells into Neural Stem Cells by HMGA2/let-7b. Cell Rep. 2015, 10, 441–452. [Google Scholar] [CrossRef]
- Xian, L.; Georgess, D.; Huso, T.; Cope, L.; Belton, A.; Chang, Y.T.; Kuang, W.; Gu, Q.; Zhang, X.; Senger, S.; et al. HMGA1 amplifies Wnt signalling and expands the intestinal stem cell compartment and Paneth cell niche. Nat. Commun. 2017, 8, 15008. [Google Scholar] [CrossRef]
- Resar, L.; Chia, L.; Xian, L. Lessons from the Crypt: HMGA1-Amping up Wnt for Stem Cells and Tumor Progression. Cancer Res. 2018, 78, 1890–1897. [Google Scholar] [CrossRef]
- Nishino, J.; Kim, I.; Chada, K.; Morrison, S.J. Hmga2 promotes neural stem cell self-renewal in young but not old mice by reducing p16Ink4a and p19Arf Expression. Cell 2008, 135, 227–239. [Google Scholar] [CrossRef]
- Kishi, Y.; Fujii, Y.; Hirabayashi, Y.; Gotoh, Y. HMGA regulates the global chromatin state and neurogenic potential in neocortical precursor cells. Nat. Neurosci. 2012, 15, 1127–1133. [Google Scholar] [CrossRef]
- Colombo, D.F.; Burger, L.; Baubec, T.; Schübeler, D. Binding of high mobility group A proteins to the mammalian genome occurs as a function of AT-content. PLoS Genet. 2017, 13, e1007102. [Google Scholar] [CrossRef]
- Shu, P.; Wu, C.; Ruan, X.; Liu, W.; Hou, L.; Fu, H.; Wang, M.; Liu, C.; Zeng, Y.; Chen, P.; et al. Opposing Gradients of MicroRNA Expression Temporally Pattern Layer Formation in the Developing Neocortex. Dev. Cell 2019, 49, 764–785. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, K.; Mason, P.J.; Bessler, M. 3’UTR-truncated Hmga2 cDNA causes MPN-like hematopoiesis by conferring a clonal growth advantage at the level of HSC in mice. Blood 2011, 117, 5860–5869. [Google Scholar] [CrossRef] [PubMed]
- Copley, M.R.; Babovic, S.; Benz, C.; Knapp, D.J.; Beer, P.A.; Kent, D.G.; Wohrer, S.; Treloar, D.Q.; Day, C.; Rowe, K.; et al. The Lin28b-let-7-Hmga2 axis determines the higher self-renewal potential of fetal haematopoietic stem cells. Nat. Cell Biol. 2013, 15, 916–925. [Google Scholar] [CrossRef] [PubMed]
- Rommel, B.; Rogalla, P.; Jox, A.; Kalle, C.V.; Kazmierczak, B.; Wolf, J.; Bullerdiek, J. HMGI-C, a member of the high mobility group family of proteins, is expressed in hematopoietic stem cells and in leukemic cells. Leuk. Lymphoma 1997, 26, 603–607. [Google Scholar] [CrossRef]
- Kumar, P.; Beck, D.; Galeev, R.; Thoms, J.A.I.; Talkhoncheh, M.S.; de Jong, I.; Unnikrishnan, A.; Baudet, A.; Subramaniam, A.; Pimanda, J.E.; et al. HMGA2 promotes long-term engraftment and myeloerythroid differentiation of human hematopoietic stem and progenitor cells. Blood Adv. 2019, 3, 681–691. [Google Scholar] [CrossRef]
- Cavazzana-Calvo, M.; Payen, E.; Negre, O.; Wang, G.; Hehir, K.; Fusil, F.; Down, J.; Denaro, M.; Brady, T.; Westerman, K.; et al. Transfusion independence and HMGA2 activation after gene therapy of human β-thalassaemia. Nature 2010, 467, 318–322. [Google Scholar] [CrossRef]
- Wei, J.; Li, H.; Wang, S.; Li, T.; Fan, J.; Liang, X.; Li, J.; Han, Q.; Zhu, L.; Fan, L.; et al. let-7 enhances osteogenesis and bone formation while repressing adipogenesis of human stromal/mesenchymal stem cells by regulating HMGA2. Stem Cells Dev. 2014, 23, 1452–1463. [Google Scholar] [CrossRef]
- Gao, X.L.; Cao, M.G.; Ai, G.G.; Hu, Y.B. Mir-98 reduces the expression of HMGA2 and promotes osteogenic differentiation of mesenchymal stem cells. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 3311–3317. [Google Scholar]
- Zhang, Y.; Liu, Y.; Wu, M.; Wang, H.; Wu, L.; Xu, B.; Zhou, W.; Fan, X.; Shao, J.; Yang, T. MicroRNA-664a-5p promotes osteogenic differentiation of human bone marrow-derived mesenchymal stem cells by directly downregulating HMGA2. Biochem. Biophys. Res. Commun. 2019, 521, 9–14. [Google Scholar] [CrossRef]
- Yuan, Y.; Xi, Y.; Chen, J.; Zhu, P.; Kang, J.; Zou, Z.; Wang, F.; Bu, S. STAT3 stimulates adipogenic stem cell proliferation and cooperates with HMGA2 during the early stage of differentiation to promote adipogenesis. Biochem. Biophys. Res. Commun. 2017, 482, 1360–1366. [Google Scholar] [CrossRef]
- Puca, F.; Colamaio, M.; Federico, A.; Gemei, M.; Tosti, N.; Bastos, A.U.; Del Vecchio, L.; Pece, S.; Battista, S.; Fusco, A. HMGA1 silencing restores normal stem cell characteristics in colon cancer stem cells by increasing p53 levels. Oncotarget 2014, 5, 3234–3245. [Google Scholar] [CrossRef] [PubMed]
- Colamaio, M.; Tosti, N.; Puca, F.; Mari, A.; Gattordo, R.; Kuzay, Y.; Federico, A.; Pepe, A.; Sarnataro, D.; Ragozzino, E.; et al. HMGA1 silencing reduces stemness and temozolomide resistance in glioblastoma stem cells. Expert Opin. Ther. Targets 2016, 20, 1169–1179. [Google Scholar] [CrossRef] [PubMed]
- Puca, F.; Tosti, N.; Federico, A.; Kuzay, Y.; Pepe, A.; Morlando, S.; Savarese, T.; D’Alessio, F.; Colamaio, M.; Sarnataro, D.; et al. HMGA1 negatively regulates NUMB expression at transcriptional and post transcriptional level in glioblastoma stem cells. Cell Cycle 2019, 18, 1446–1457. [Google Scholar] [CrossRef] [PubMed]
- Reeves, R. Nuclear functions of the HMG proteins. Biochim. Biophys. Acta 2010, 1799, 3–14. [Google Scholar] [CrossRef]
- Yie, J.; Merika, M.; Munshi, N.; Chen, G.; Thanos, D. The role of HMG I(Y) in the assembly and function of the IFN-beta enhanceosome. EMBO J. 1999, 18, 3074–3089. [Google Scholar] [CrossRef]
- Reeves, R.; Leonard, W.J.; Nissen, M.S. Binding of HMG-I(Y) imparts architectural specificity to a positioned nucleosome on the promoter of the human interleukin-2 receptor alpha gene. Mol. Cell. Biol. 2000, 20, 4666–4679. [Google Scholar] [CrossRef]
- Duncan, B.; Zhao, K. Hmga1 mediates the activation of the CRYAB promoter by BRG1. DNA Cell Biol. 2007, 26, 745–752. [Google Scholar] [CrossRef]
- Catez, F.; Yang, H.; Tracey, K.J.; Reeves, R.; Misteli, T.; Bustin, M. Network of dynamic interactions between histone H1 and high-mobility-group proteins in chromatin. Mol. Cell. Biol. 2004, 24, 4321–4328. [Google Scholar] [CrossRef]
- Ozturk, N.; Singh, I.; Mehta, A.; Braun, T.; Barreto, G. HMGA proteins as modulators of chromatin structure during transcriptional activation. Front. Cell Dev. Biol. 2014, 2, 5. [Google Scholar] [CrossRef]
- Cleynen, I.; Brants, J.R.; Peeters, K.; Deckers, R.; Debiec-Rychter, M.; Sciotm, R.; Van de Ven, W.J.; Petit, M.M. HMGA2 regulates transcription of the Imp2 gene via an intronic regulatory element in cooperation with nuclear factor-kappaB. Mol. Cancer Res. 2007, 5, 363–372. [Google Scholar] [CrossRef]
- Schuldenfrei, A.; Belton, A.; Kowalski, J.; Talbot, C.C., Jr.; Di Cello, F.; Poh, W.; Tsai, H.L.; Shah, S.N.; Huso, T.H.; Huso, D.L.; et al. HMGA1 drives stem cell, inflammatory pathway, and cell cycle progression genes during lymphoid tumorigenesis. BMC Genom. 2011, 12, 549. [Google Scholar] [CrossRef] [PubMed]
- Baldassarre, G.; Battista, S.; Belletti, B.; Thakur, S.; Pentimalli, F.; Trapasso, F.; Fedele, M.; Pierantoni, G.; Croce, C.M.; Fusco, A. Negative regulation of BRCA1 gene expression by HMGA1 proteins accounts for the reduced BRCA1 protein levels in sporadic breast carcinoma. Mol. Cell. Biol. 2003, 23, 2225–2238. [Google Scholar] [CrossRef] [PubMed]
- Martinez Hoyos, J.; Fedele, M.; Battista, S.; Pentimalli, F.; Kruhoffer, M.; Arra, C.; Orntoft, T.F.; Croce, C.M.; Fusco, A. Identification of the genes up- and down-regulated by the high mobility group A1 (HMGA1) proteins: Tissue specificity of the HMGA1-dependent gene regulation. Cancer Res. 2004, 64, 5728–5735. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Treiber, T.; Treiber, N.; Meister, G. Regulation of microRNA biogenesis and its crosstalk with other cellular pathways. Nat. Rev. Mol. Cell. Biol. 2019, 20, 5–20. [Google Scholar] [CrossRef]
- Mayr, F.; Heinemann, U. Mechanisms of Lin28-mediated miRNA and mRNA regulation--a structural and functional perspective. Int. J. Mol. Sci. 2013, 14, 16532–16553. [Google Scholar] [CrossRef]
- Hagan, J.P.; Piskounova, E.; Gregory, R.I. Lin28 recruits the TUTase Zcchc11 to inhibit let-7 maturation in mouse embryonic stem cells. Nat. Struct. Mol. Biol. 2009, 16, 1021–1025. [Google Scholar] [CrossRef]
- Xia, X.; Ahmad, I. let-7 microRNA regulates neurogliogenesis in the mammalian retina through Hmga2. Dev. Biol. 2016, 410, 70–85. [Google Scholar] [CrossRef]
- Rowe, R.G.; Wang, L.D.; Coma, S.; Han, A.; Mathieu, R.; Pearson, D.S.; Ross, S.; Sousa, P.; Nguyen, P.T.; Rodriguez, A.; et al. Developmental regulation of myeloerythroid progenitor function by the Lin28b-let-7-Hmga2 axis. J. Exp. Med. 2016, 213, 1497–1512. [Google Scholar] [CrossRef]
- Kuan, I.I.; Lee, C.C.; Chen, C.H.; Lu, J.; Kuo, Y.S.; Wu, H.C. The extracellular domain of epithelial cell adhesion molecule (EpCAM) enhances multipotency of mesenchymal stem cells through EGFR-LIN28-LET7 signaling. J. Biol. Chem. 2019, 294, 7769–7786. [Google Scholar] [CrossRef]
- Dai, N.; Rapley, J.; Angel, M.; Yanik, M.F.; Blower, M.D.; Avruch, J. mTOR phosphorylates IMP2 to promote IGF2 mRNA translation by internal ribosomal entry. Genes Dev. 2011, 25, 1159–1172. [Google Scholar] [CrossRef]
- Hafner, M.; Landthaler, M.; Burger, L.; Khorshid, M.; Hausser, J.; Berninger, P.; Rothballer, A.; Ascano, M., Jr.; Jungkamp, A.C.; Munschauer, M.; et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 2010, 141, 129–141. [Google Scholar] [CrossRef] [PubMed]
- Oliveira-Mateos, C.; Sánchez-Castillo, A.; Soler, M.; Obiols-Guardia, A.; Piñeyro, D.; Boque-Sastre, R.; Calleja-Cervantes, M.E.; Castro de Moura, M.; Martínez-Cardús, A.1; Rubio, T.; et al. The transcribed pseudogene RPSAP52 enhances the oncofetal HMGA2-IGF2BP2-RAS axis through LIN28B-dependent and independent let-7 inhibition. Nat. Commun. 2019, 10, 3979. [Google Scholar] [CrossRef] [PubMed]
- Boque-Sastre, R.; Soler, M.; Oliveira-Mateos, C.; Portela, A.; Moutinho, C.; Sayols, S.; Villanueva, A.; Esteller, M.; Guil, S. Head-to-head antisense transcription and R-loop formation promotes transcriptional activation. Proc. Natl. Acad. Sci. USA 2015, 112, 5785–5790. [Google Scholar] [CrossRef] [PubMed]
- D’Angelo, D.; Mussnich, P.; Sepe, R.; Raia, M.; Del Vecchio, L.; Cappabianca, P.; Pellecchia, S.; Petrosino, S.; Saggio, S.; Solari, D.; et al. RPSAP52 lncRNA is overexpressed in pituitary tumors and promotes cell proliferation by acting as miRNA sponge for HMGA proteins. J. Mol. Med. 2019, 97, 1019–1032. [Google Scholar] [CrossRef]
- Battista, M.; Musto, A.; Navarra, A.; Minopoli, G.; Russo, T.; Parisi, S. miR-125b Regulates the Early Steps of ESC Differentiation through Dies1 in a TGF-Independent Manner. Int. J. Mol. Sci. 2013, 14, 13482–13496. [Google Scholar] [CrossRef]
- Parisi, S.; Battista, M.; Musto, A.; Navarra, A.; Tarantino, C.; Russo, T. A regulatory loop involving Dies1 and miR-125a controls BMP4 signaling in mouse embryonic stem cells. FASEB J. 2012, 10, 3957–3968. [Google Scholar] [CrossRef]
- Musto, A.; Navarra, A.; Vocca, A.; Gargiulo, A.; Minopoli, G.; Romano, S.; Romano, M.F.; Russo, T.; Parisi, S. miR-23a, miR-24 and miR-27a protect differentiating ESCs from BMP4 induced apoptosis. Cell Death Differ. 2015, 22, 1047–1057. [Google Scholar] [CrossRef]
- Xu, N.; Papagiannakopoulos, T.; Pan, G.; Thomson, J.A.; Kosik, K.S. MicroRNA-145 regulates OCT4, SOX2, and KLF4 and represses pluripotency in human embryonic stem cells. Cell 2009, 137, 647–658. [Google Scholar] [CrossRef]
- Zeng, Z.L.; Lin, X.L.; Tan, L.L.; Liu, Y.M.; Qu, K.; Wang, Z. MicroRNAs: Important Regulators of Induced Pluripotent Stem Cell Generation and Differentiation. Stem Cell Rev. Rep. 2018, 14, 71–81. [Google Scholar] [CrossRef]
- Li, N.; Long, B.; Han, W.; Yuan, S.; Wang, K. microRNAs: Important regulators of stem cells. Stem Cell Res. Ther. 2017, 8, 110. [Google Scholar] [CrossRef]
- Bansod, S.; Kageyama, R.; Ohtsuka, T. Hes5 regulates the transition timing of neurogenesis and gliogenesis in mammalian neocortical development. Development 2017, 144, 3156–3167. [Google Scholar] [CrossRef]
- Di Marcantonio, D.; Galli, D.; Carubbi, C.; Gobbi, G.; Queirolo, V.; Martini, S.; Merighi, S.; Vaccarezza, M.; Maffulli, N.; Sykes, S.M.; et al. PKCε as a novel promoter of skeletal muscle differentiation and regeneration. Exp. Cell Res. 2015, 339, 10–19. [Google Scholar] [CrossRef]
- Chandrasekaran, K.S.; Sathyanarayanan, A.; Karunagaran, D. MicroRNA-214 suppresses growth, migration and invasion through a novel target, high mobility group AT-hook 1, in human cervical and colorectal cancer cells. Br. J. Cancer 2016, 115, 741–751. [Google Scholar] [CrossRef]
- Qiu, H.; Zhong, J.; Luo, L.; Tang, Z.; Liu, N.; Kang, K.; Li, L.; Gou, D. Regulatory Axis of miR-195/497 and HMGA1-Id3 Governs Muscle Cell Proliferation and Differentiation. Int. J. Biol. Sci. 2017, 13, 157–166. [Google Scholar] [CrossRef][Green Version]
- Qin, M.M.; Chai, X.; Huang, H.B.; Feng, G.; Li, X.N.; Zhang, J.; Zheng, R.; Liu, X.C.; Pu, C. let-7i inhibits proliferation and migration of bladder cancer cells by targeting HMGA1. BMC Urol. 2019, 19, 53. [Google Scholar] [CrossRef]
- Dai, N.; Ji, F.; Wright, J.; Minichiello, L.; Sadreyev, R.; Avruch, J. IGF2 mRNA binding protein-2 is a tumor promoter that drives cancer proliferation through its client mRNAs IGF2 and HMGA1. Elife 2017, 6. [Google Scholar] [CrossRef]
| HMG | Organism | Cell Type | HMG Level | Phenotype | References |
|---|---|---|---|---|---|
| HMGA1 | mouse | ESCs | KO | Impairment of lymphohematopoietic differentiation | [8] |
| HMGA1 | human | ESCs | overexpression | Block of differentiation, maintenance of the undifferentiated phenotype | [24] |
| HMGA1 | human | ESCs | silencing | Downregulation of pluripotency genes | [24] |
| Hmga1 | human | MSCs | overexpression | Improvement of reprogramming into iPSCs from MSCs | [24] |
| HMGA1 | mouse | ISCs | overexpression | Enhancement of self-renewal | [36] |
| HMGA2 | human | ESCs | silencing | Decrease of self-renewal and mesodermal genes and increase endodermal genes | [27] |
| HMGA2 | human | ESCs | silencing | Increase in the amount of fragmented DNA | [28] |
| HMGA2 | mouse | ESCs | overexpression | Improvement of myogenic differentiation | [29] |
| HMGA2 | mouse | ESCs | Silencing or KO | Block of differentiation, maintenance of the undifferentiated phenotype | [30] |
| HMGA2 | mouse | ESCs | overexpression | Impairment of cell cycle and apoptosis during differentiation | [31] |
| HMGA2 | mouse | Embryonic fibroblasts | KO | Sever impairment of reprogramming into iPSCs | [30] |
| HMGA2 | human | Dermal fibroblasts | overexpression | Improvement of reprogramming into induced neural stem cells | [34] |
| HMGA2 | mouse | Embryonic fibroblasts | overexpression | Improvement of reprogramming into induced neural stem cells | [34] |
| HMGA2 | mouse | NSCs | KO | Decrease of self-renewal | [38] |
| HMGA2 | mouse | HSCs | overexpression | Increase of self-renewal | [42] |
| HMGA2 | mouse | HSCs | KO | Decrease of self-renewal | [43] |
| HMGA2 | human | HSCs | silencing | Decrease of self-renewal, differentiation impairment | [5,45] |
| HMGA2 | human | MSCs | overexpression | Block of differentiation | [47,48,49] |
| HMGA2 | human | MSCs | silencing | Improvement of osteogenic differentiation | [47] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Parisi, S.; Piscitelli, S.; Passaro, F.; Russo, T. HMGA Proteins in Stemness and Differentiation of Embryonic and Adult Stem Cells. Int. J. Mol. Sci. 2020, 21, 362. https://doi.org/10.3390/ijms21010362
Parisi S, Piscitelli S, Passaro F, Russo T. HMGA Proteins in Stemness and Differentiation of Embryonic and Adult Stem Cells. International Journal of Molecular Sciences. 2020; 21(1):362. https://doi.org/10.3390/ijms21010362
Chicago/Turabian StyleParisi, Silvia, Silvia Piscitelli, Fabiana Passaro, and Tommaso Russo. 2020. "HMGA Proteins in Stemness and Differentiation of Embryonic and Adult Stem Cells" International Journal of Molecular Sciences 21, no. 1: 362. https://doi.org/10.3390/ijms21010362
APA StyleParisi, S., Piscitelli, S., Passaro, F., & Russo, T. (2020). HMGA Proteins in Stemness and Differentiation of Embryonic and Adult Stem Cells. International Journal of Molecular Sciences, 21(1), 362. https://doi.org/10.3390/ijms21010362






