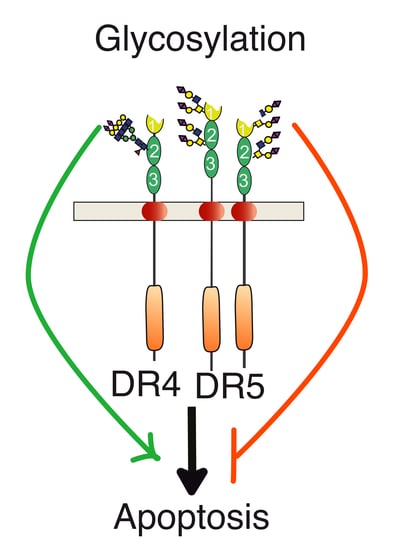
Regulation of TNF-Related Apoptosis-Inducing Ligand Signaling by Glycosylation
Abstract
1. Introduction
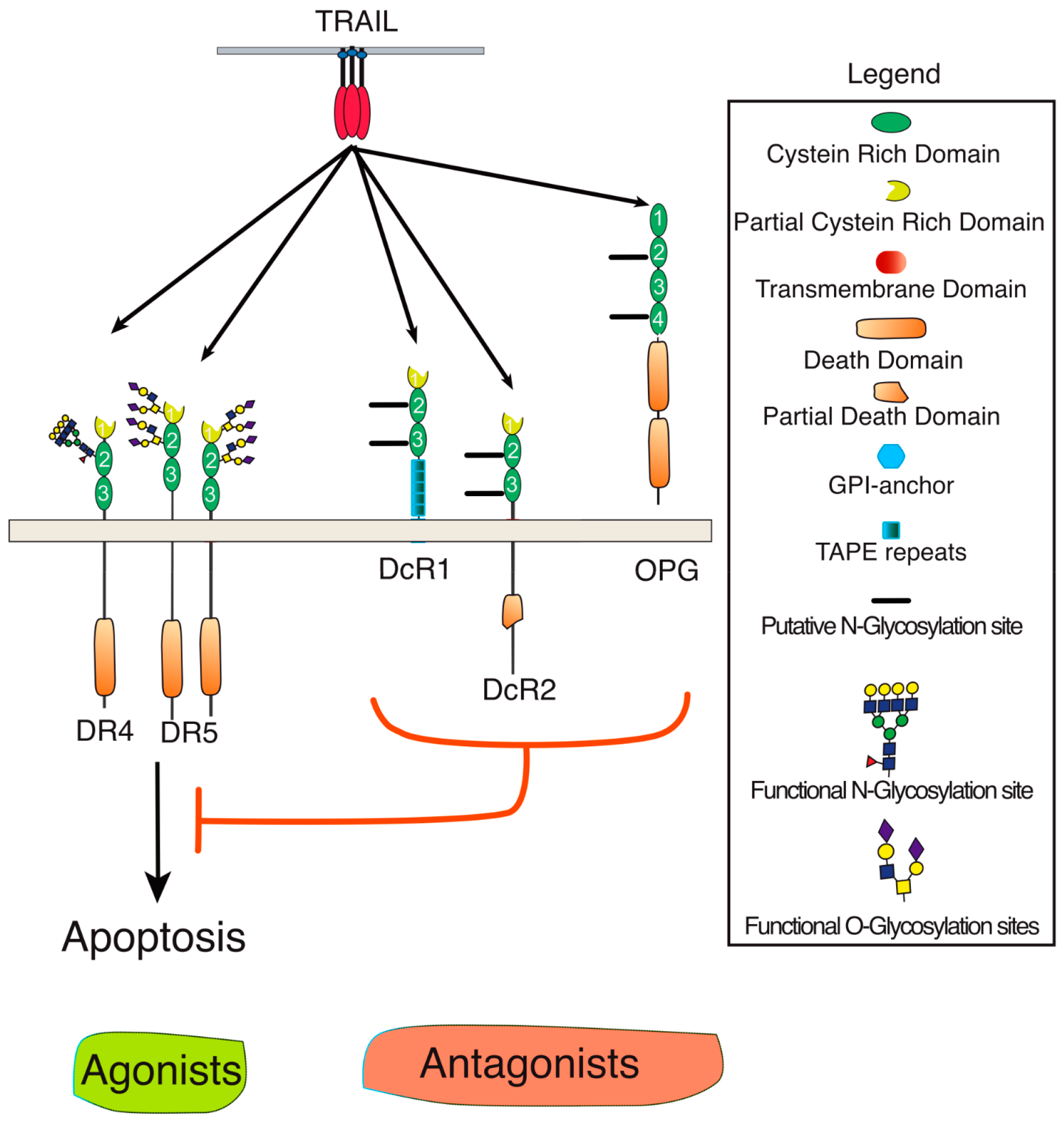
2. Membrane Proximal TRAIL DISC Formation and Signaling Regulation
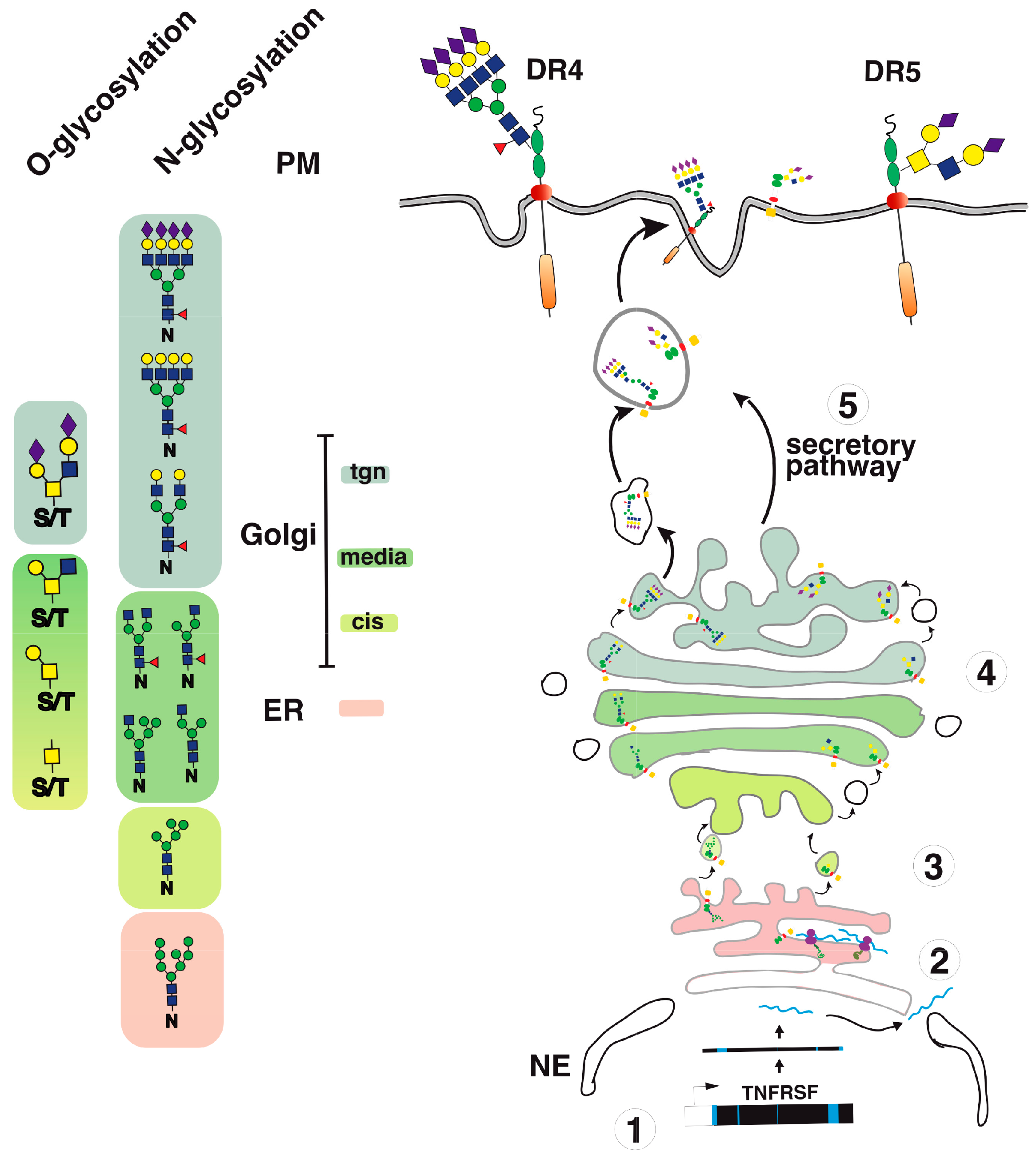
3. Agonist TRAIL Receptors are Glycosylated
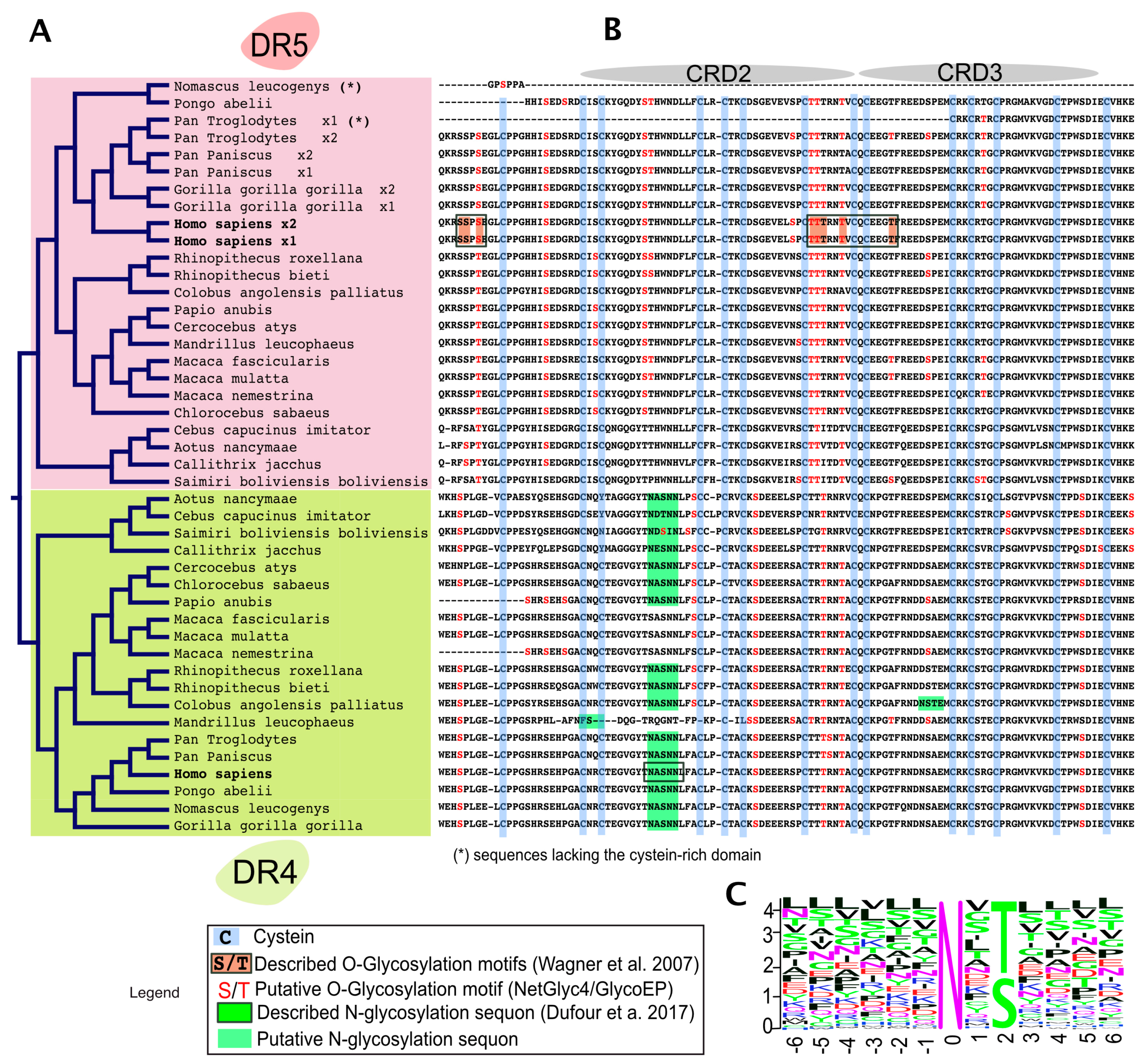
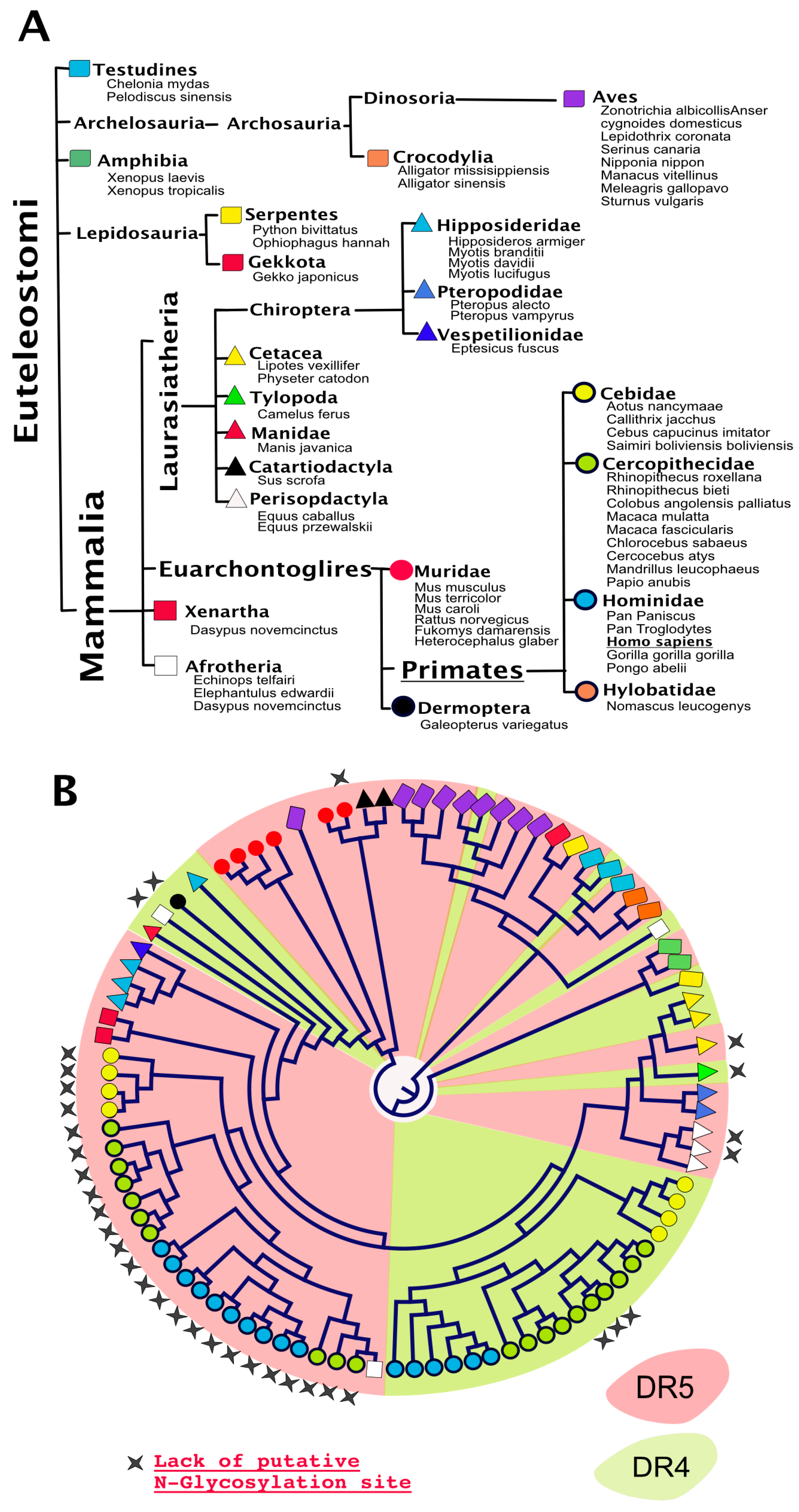
4. TRAIL Receptor Glycosylation during Evolution
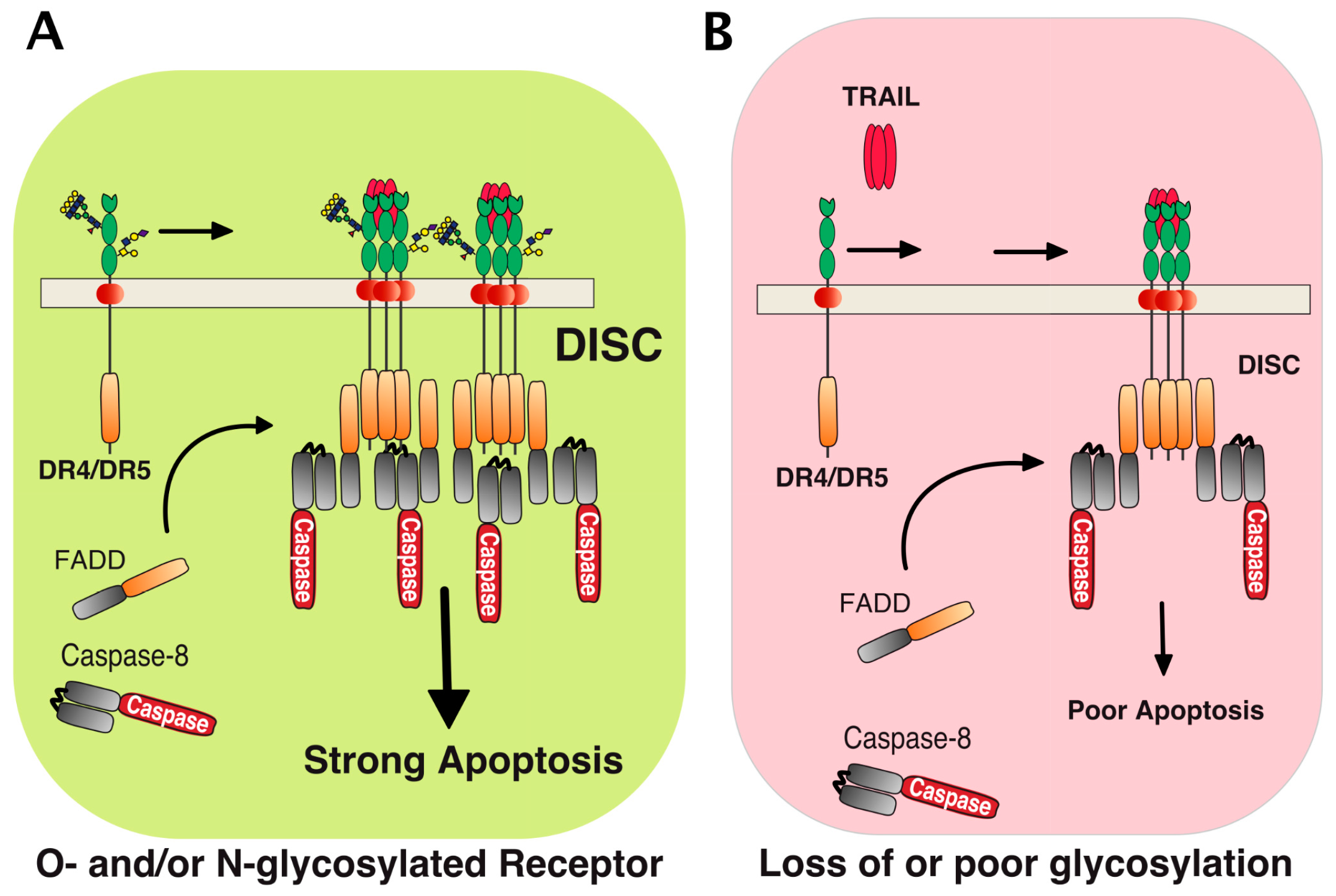
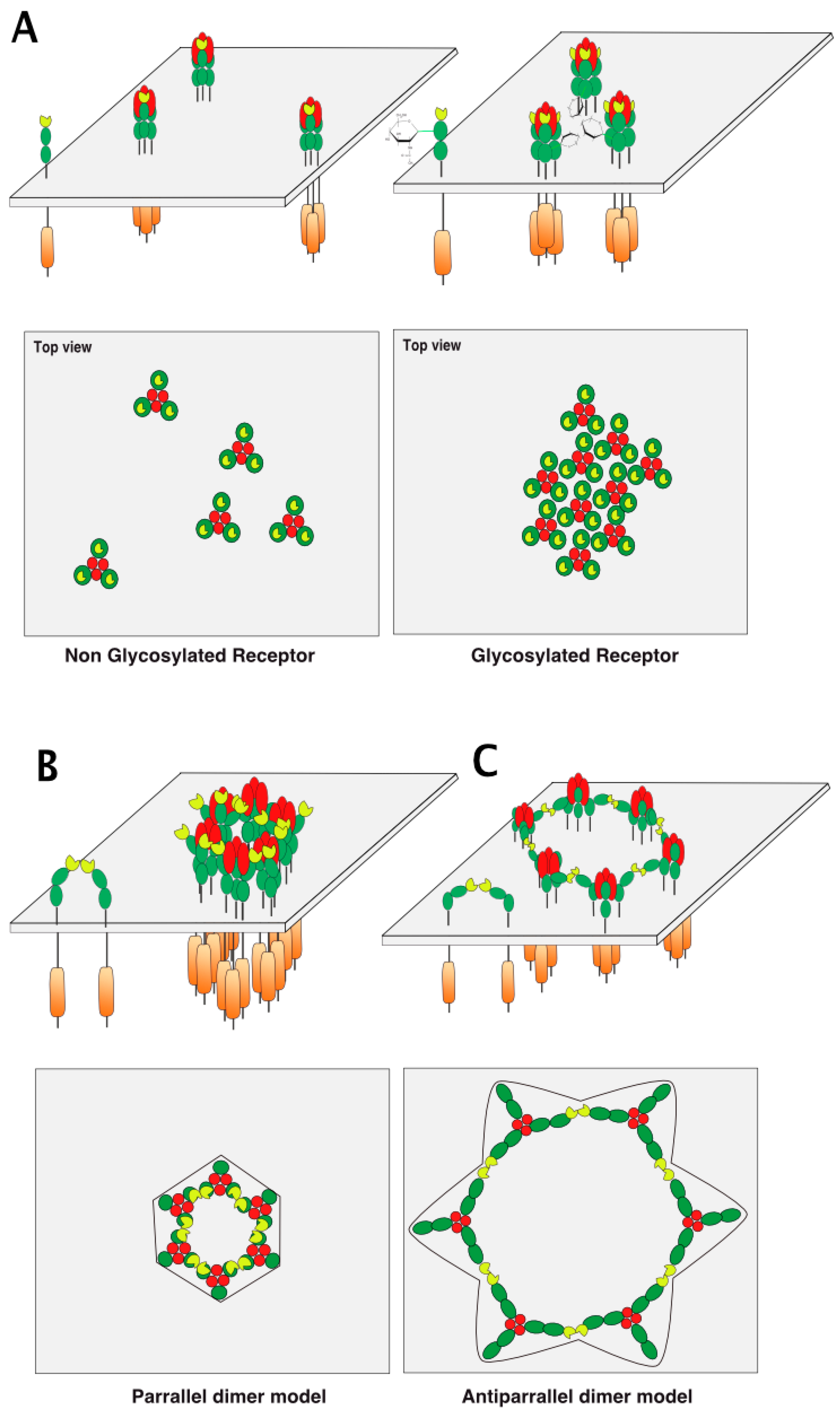
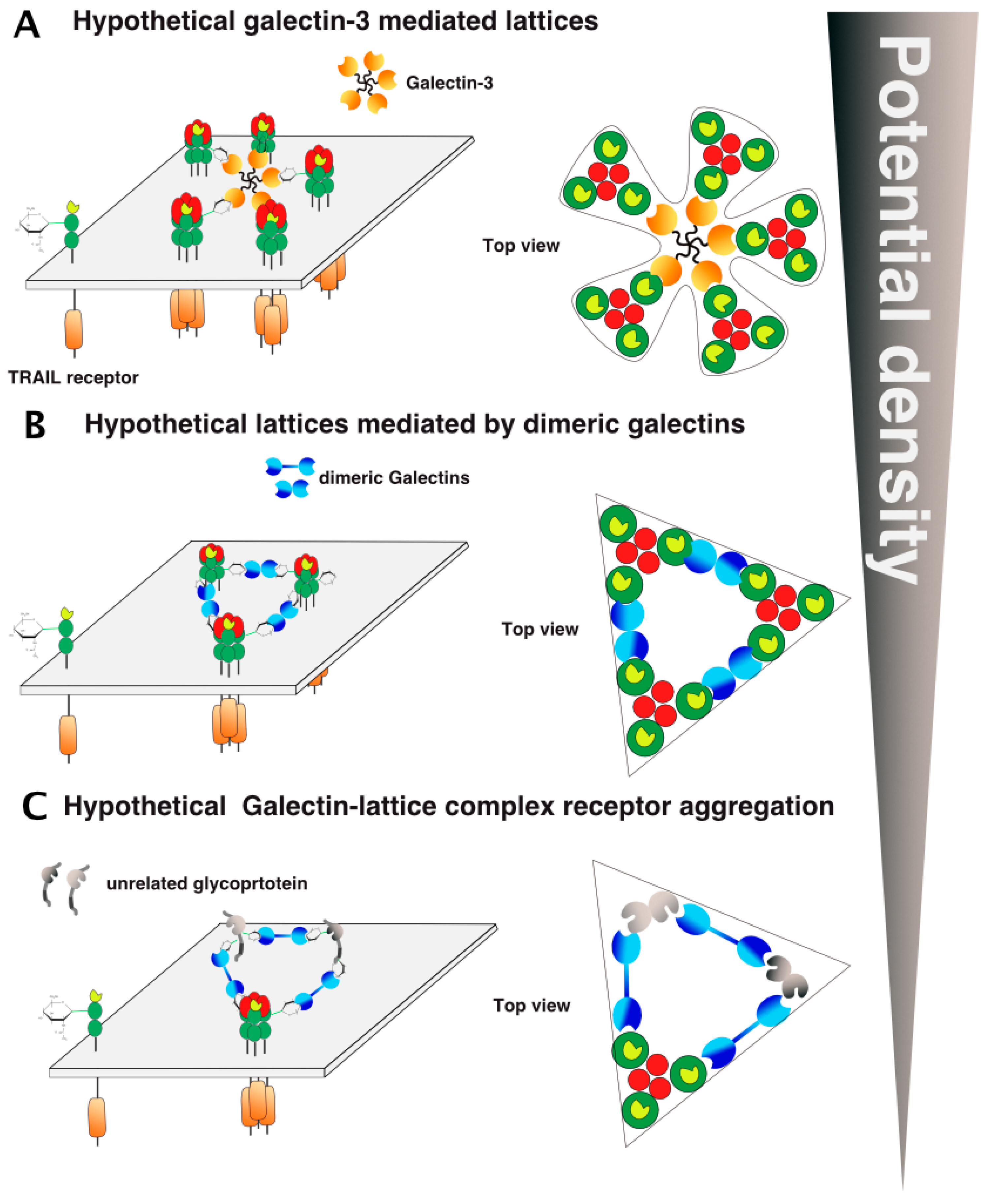
5. TRAIL Receptor Clustering
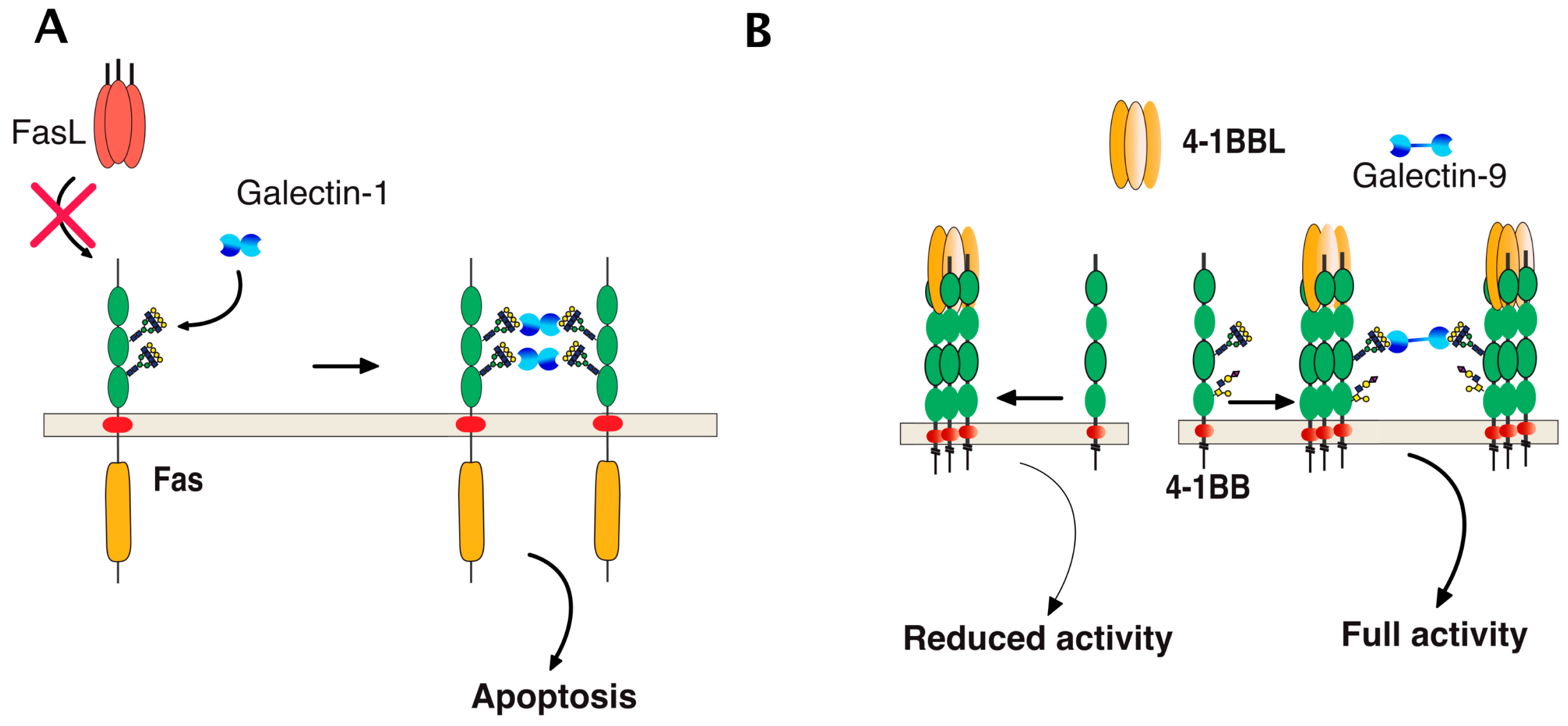
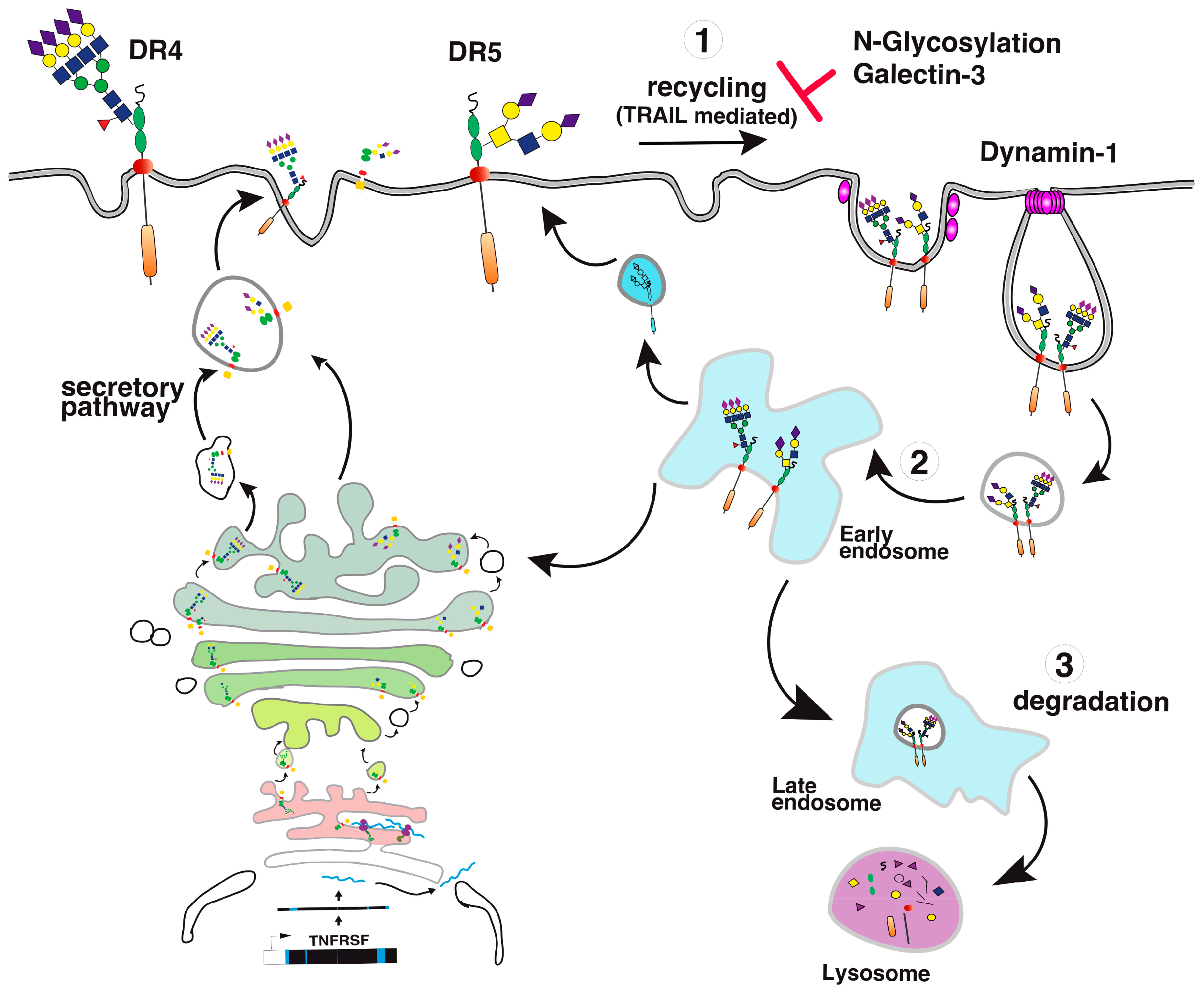
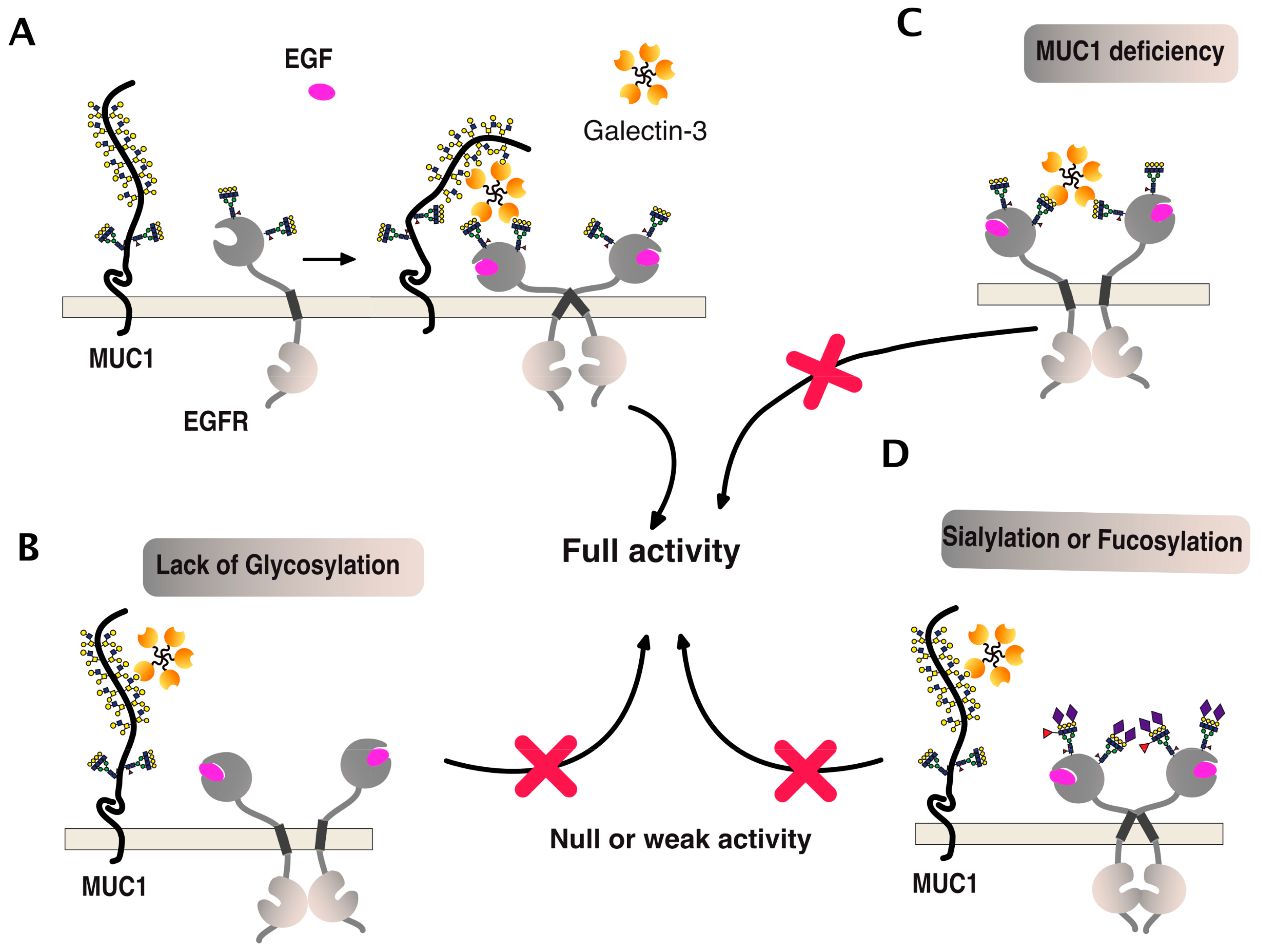
6. TRAIL Signal Transduction Regulation by Carbohydrate-Binding or Modifying Proteins
7. Conclusions
Acknowledgments
Conflicts of Interest
Abbreviations
| AKT | protein kinase B |
| Asn | asparagine |
| c-FLIP | cellular FLICE (FADD-like IL-1β-converting enzyme)-inhibitory protein |
| CRD | cysteine-rich domain |
| DcR | decoy receptor |
| DcR1 | decoy receptor 1, TRAIL-R3, TNFRSF10C |
| DcR2 | decoy receptor 2, TRAIL-R4, TNFRSF10D |
| DD | death domain |
| DISC | death-inducing signaling complex |
| DR | death receptor |
| DR4 | death receptor 4, TRAIL-R1, TNFRSF10A |
| DR5 | death receptor 5, TRAIL-R2, TNFRSF10B |
| EGFR | epithelial growth factor receptor |
| ER | endoplasmic reticulum |
| Fas | fibroblast associated surface antigen, also known as CD95 |
| FADD | Fas associated death domain |
| Fas-L | Fas ligand |
| NE | nuclear envelope |
| NF-κB | nuclear factor-kappa B |
| OPG | osteoprotegerin |
| PLAD | pre-ligand assembly domain |
| PM | plasma membrane |
| Ser | serine |
| SODD | silencer of death domain |
| shRNA | short hairpin RNA |
| siRNA | small interfering RNA |
| Thr | threonine |
| TNF | tumor necrosis factor |
| TNFR1 | tumor necrosis factor receptor 1 |
| TRAIL | TNF related apoptosis inducing ligand, also known as APO2L |
References
- Micheau, O. Posttranslational Modifications and Death Receptor Signalling. In TRAIL, Fas Ligand, TNF and TLR3 in Cancer; Micheau, O., Ed.; Springer International Publishing: Cham, Switzerland, 2017; pp. 247–290. [Google Scholar]
- Varki, A.; Sharon, N. Historical Background and Overview. In Essentials of Glycobiology, 2nd ed.; Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Bertozzi, C.R., Hart, G.W., Etzler, M.E., Eds.; Cold Spring Harbor: Nassau, NY, USA, 2009. [Google Scholar]
- Stanley, P.; Schachter, H.; Taniguchi, N. N-Glycans. In Essentials of Glycobiology, 2nd ed.; Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Bertozzi, C.R., Hart, G.W., Etzler, M.E., Eds.; Cold Spring Harbor: Nassau, NY, USA, 2009. [Google Scholar]
- Brockhausen, I.; Schachter, H.; Stanley, P. O-GalNAc Glycans. In Essentials of Glycobiology, 2nd ed.; Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Bertozzi, C.R., Hart, G.W., Etzler, M.E., Eds.; Cold Spring Harbor: Nassau, NY, USA, 2009. [Google Scholar]
- Matoba, K.; Mihara, E.; Tamura-Kawakami, K.; Miyazaki, N.; Maeda, S.; Hirai, H.; Thompson, S.; Iwasaki, K.; Takagi, J. Conformational Freedom of the LRP6 Ectodomain Is Regulated by N-glycosylation and the Binding of the Wnt Antagonist Dkk1. Cell Rep. 2017, 18, 32–40. [Google Scholar] [CrossRef] [PubMed]
- Emery, J.G.; McDonnell, P.; Burke, M.B.; Deen, K.C.; Lyn, S.; Silverman, C.; Dul, E.; Appelbaum, E.R.; Eichman, C.; DiPrinzio, R.; et al. Osteoprotegerin is a receptor for the cytotoxic ligand TRAIL. J. Biol. Chem. 1998, 273, 14363–14367. [Google Scholar] [CrossRef] [PubMed]
- Truneh, A.; Sharma, S.; Silverman, C.; Khandekar, S.; Reddy, M.P.; Deen, K.C.; McLaughlin, M.M.; Srinivasula, S.M.; Livi, G.P.; Marshall, L.A.; et al. Temperature-sensitive differential affinity of TRAIL for its receptors. DR5 is the highest affinity receptor. J. Biol. Chem. 2000, 275, 23319–23325. [Google Scholar] [CrossRef] [PubMed]
- Micheau, O.; Shirley, S.; Dufour, F. Death receptors as targets in cancer. Br. J. Pharmacol. 2013, 169, 1723–1744. [Google Scholar] [CrossRef] [PubMed]
- Dubuisson, A.; Micheau, O. Antibodies and Derivatives Targeting DR4 and DR5 for Cancer Therapy. Antibodies 2017, 6, 16. [Google Scholar] [CrossRef]
- Boldin, M.P.; Mett, I.L.; Varfolomeev, E.E.; Chumakov, I.; Shemer-Avni, Y.; Camonis, J.H.; Wallach, D. Self-association of the “death domains” of the p55 tumor necrosis factor (TNF) receptor and Fas/APO1 prompts signaling for TNF and Fas/APO1 effects. J. Biol. Chem. 1995, 270, 387–391. [Google Scholar] [CrossRef] [PubMed]
- Bodmer, J.L.; Holler, N.; Reynard, S.; Vinciguerra, P.; Schneider, P.; Juo, P.; Blenis, J.; Tschopp, J. TRAIL receptor-2 signals apoptosis through FADD and caspase-8. Nat. Cell Biol. 2000, 2, 241–243. [Google Scholar] [CrossRef] [PubMed]
- Micheau, O.; Shirley, S.; Morizot, A. TRAIL Receptor-Induced Cell Death Regulation: An Update to Our Deadly Discussion. In Topics in Anti-Cancer Research; Rahman, A., Zaman, K., Eds.; Bentham Science Publishers: Busum, Germany, 2014; Volume 3, pp. 3–36. [Google Scholar]
- Pan, G.; Ni, J.; Yu, G.; Wei, Y.F.; Dixit, V.M. TRUNDD, a new member of the TRAIL receptor family that antagonizes TRAIL signalling. FEBS Lett. 1998, 424, 41–45. [Google Scholar] [CrossRef]
- Pan, G.; Ni, J.; Wei, Y.F.; Yu, G.; Gentz, R.; Dixit, V.M. An antagonist decoy receptor and a death domain-containing receptor for TRAIL. Science 1997, 277, 815–818. [Google Scholar] [CrossRef] [PubMed]
- O’Leary, L.; van der Sloot, A.M.; Reis, C.R.; Deegan, S.; Ryan, A.E.; Dhami, S.P.; Murillo, L.S.; Cool, R.H.; de Sampaio, P.C.; Thompson, K.; et al. Decoy receptors block TRAIL sensitivity at a supracellular level: The role of stromal cells in controlling tumour TRAIL sensitivity. Oncogene 2016, 35, 1261. [Google Scholar] [CrossRef] [PubMed]
- Neumann, S.; Hasenauer, J.; Pollak, N.; Scheurich, P. Dominant negative effects of tumor necrosis factor (TNF)-related apoptosis-inducing ligand (TRAIL) receptor 4 on TRAIL receptor 1 signaling by formation of heteromeric complexes. J. Biol. Chem. 2014, 289, 16576–16587. [Google Scholar] [CrossRef] [PubMed]
- Merino, D.; Lalaoui, N.; Morizot, A.; Schneider, P.; Solary, E.; Micheau, O. Differential inhibition of TRAIL-mediated DR5-DISC formation by decoy receptors 1 and 2. Mol. Cell. Biol. 2006, 26, 7046–7055. [Google Scholar] [CrossRef] [PubMed]
- Anees, M.; Horak, P.; Schiefer, A.I.; Vanhara, P.; El-Gazzar, A.; Perco, P.; Kiesewetter, B.; Mullauer, L.; Streubel, B.; Raderer, M.; et al. The potential evasion of immune surveillance in mucosa associated lymphoid tissue lymphoma by DcR2-mediated up-regulation of nuclear factor-kappaB. Leuk. Lymphoma 2015, 56, 1440–1449. [Google Scholar] [CrossRef] [PubMed]
- Lalaoui, N.; Morle, A.; Merino, D.; Jacquemin, G.; Iessi, E.; Morizot, A.; Shirley, S.; Robert, B.; Solary, E.; Garrido, C.; et al. TRAIL-R4 promotes tumor growth and resistance to apoptosis in cervical carcinoma HeLa cells through AKT. PLoS ONE 2011, 6, e19679. [Google Scholar] [CrossRef] [PubMed]
- Degli-Esposti, M.A.; Dougall, W.C.; Smolak, P.J.; Waugh, J.Y.; Smith, C.A.; Goodwin, R.G. The novel receptor TRAIL-R4 induces NF-kappaB and protects against TRAIL-mediated apoptosis, yet retains an incomplete death domain. Immunity 1997, 7, 813–820. [Google Scholar] [CrossRef]
- Muzio, M.; Stockwell, B.R.; Stennicke, H.R.; Salvesen, G.S.; Dixit, V.M. An induced proximity model for caspase-8 activation. J. Biol. Chem. 1998, 273, 2926–2930. [Google Scholar] [CrossRef] [PubMed]
- Salvesen, G.S.; Dixit, V.M. Caspase activation: The induced-proximity model. Proc. Natl. Acad. Sci. USA 1999, 96, 10964–10967. [Google Scholar] [CrossRef] [PubMed]
- Schleich, K.; Krammer, P.H.; Lavrik, I.N. The chains of death: A new view on caspase-8 activation at the DISC. Cell Cycle 2013, 12, 193–194. [Google Scholar] [CrossRef] [PubMed]
- Dickens, L.S.; Boyd, R.S.; Jukes-Jones, R.; Hughes, M.A.; Robinson, G.L.; Fairall, L.; Schwabe, J.W.; Cain, K.; Macfarlane, M. A death effector domain chain DISC model reveals a crucial role for caspase-8 chain assembly in mediating apoptotic cell death. Mol. Cell 2012, 47, 291–305. [Google Scholar] [CrossRef] [PubMed]
- Berg, D.; Lehne, M.; Muller, N.; Siegmund, D.; Munkel, S.; Sebald, W.; Pfizenmaier, K.; Wajant, H. Enforced covalent trimerization increases the activity of the TNF ligand family members TRAIL and CD95L. Cell Death Differ. 2007, 14, 2021–2034. [Google Scholar] [CrossRef] [PubMed]
- Holler, N.; Tardivel, A.; Kovacsovics-Bankowski, M.; Hertig, S.; Gaide, O.; Martinon, F.; Tinel, A.; Deperthes, D.; Calderara, S.; Schulthess, T.; et al. Two adjacent trimeric Fas ligands are required for Fas signaling and formation of a death-inducing signaling complex. Mol. Cell. Biol. 2003, 23, 1428–1440. [Google Scholar] [CrossRef] [PubMed]
- Shirley, S.; Morizot, A.; Micheau, O. Regulating TRAIL Receptor-Induced Cell Death at the Membrane: A Deadly Discussion. Recent Pat. Anti-Cancer Drug Discov. 2011, 6, 311–323. [Google Scholar] [CrossRef] [PubMed]
- Morizot, A.; Merino, D.; Lalaoui, N.; Jacquemin, G.; Granci, V.; Iessi, E.; Lanneau, D.; Bouyer, F.; Solary, E.; Chauffert, B.; et al. Chemotherapy overcomes TRAIL-R4-mediated TRAIL resistance at the DISC level. Cell Death Differ. 2011, 18, 700–711. [Google Scholar] [CrossRef] [PubMed]
- Lafont, E.; Hartwig, T.; Walczak, H. Paving TRAIL’s Path with Ubiquitin. Trends Biochem. Sci. 2018, 43, 44–60. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.D.; Franco, A.; Myers, K.; Gray, C.; Nguyen, T.; Hersey, P. Relation of TNF-related apoptosis-inducing ligand (TRAIL) receptor and FLICE-inhibitory protein expression to TRAIL-induced apoptosis of melanoma. Cancer Res. 1999, 59, 2747–2753. [Google Scholar] [PubMed]
- Olsson, A.; Diaz, T.; Aguilar-Santelises, M.; Osterborg, A.; Celsing, F.; Jondal, M.; Osorio, L.M. Sensitization to TRAIL-induced apoptosis and modulation of FLICE-inhibitory protein in B chronic lymphocytic leukemia by actinomycin D. Leukemia 2001, 15, 1868–1877. [Google Scholar] [CrossRef] [PubMed]
- Irmler, M.; Thome, M.; Hahne, M.; Schneider, P.; Hofmann, K.; Steiner, V.; Bodmer, J.L.; Schroter, M.; Burns, K.; Mattmann, C.; et al. Inhibition of death receptor signals by cellular FLIP. Nature 1997, 388, 190–195. [Google Scholar] [CrossRef] [PubMed]
- Morle, A.; Garrido, C.; Micheau, O. Hyperthermia restores apoptosis induced by death receptors through aggregation-induced c-FLIP cytosolic depletion. Cell Death Dis. 2015, 6, e1633. [Google Scholar] [CrossRef] [PubMed]
- Kreuz, S.; Siegmund, D.; Scheurich, P.; Wajant, H. NF-kappaB inducers upregulate cFLIP, a cycloheximide-sensitive inhibitor of death receptor signaling. Mol. Cell. Biol. 2001, 21, 3964–3973. [Google Scholar] [CrossRef] [PubMed]
- Haas, T.L.; Emmerich, C.H.; Gerlach, B.; Schmukle, A.C.; Cordier, S.M.; Rieser, E.; Feltham, R.; Vince, J.; Warnken, U.; Wenger, T.; et al. Recruitment of the linear ubiquitin chain assembly complex stabilizes the TNF-R1 signaling complex and is required for TNF-mediated gene induction. Mol. Cell 2009, 36, 831–844. [Google Scholar] [CrossRef] [PubMed]
- Micheau, O. FLIP. In Cancer Therapeutic Targets; Marshall, J.L., Ed.; Springer: New York, NY, USA, 2017; pp. 881–891. [Google Scholar]
- Lavrik, I.N. Systems biology of death receptor networks: Live and let die. Cell Death Dis. 2014, 5, e1259. [Google Scholar] [CrossRef] [PubMed]
- Walczak, H. Death receptor-ligand systems in cancer, cell death, and inflammation. Cold Spring Harb. Perspect. Biol. 2013, 5, a008698. [Google Scholar] [CrossRef] [PubMed]
- Sessler, T.; Healy, S.; Samali, A.; Szegezdi, E. Structural determinants of DISC function: New insights into death receptor-mediated apoptosis signalling. Pharmacol. Ther. 2013, 140, 186–199. [Google Scholar] [CrossRef] [PubMed]
- Estornes, Y.; Toscano, F.; Virard, F.; Jacquemin, G.; Pierrot, A.; Vanbervliet, B.; Bonnin, M.; Lalaoui, N.; Mercier-Gouy, P.; Pacheco, Y.; et al. dsRNA induces apoptosis through an atypical death complex associating TLR3 to caspase-8. Cell Death Differ. 2012, 19, 1482–1494. [Google Scholar] [CrossRef] [PubMed]
- Estornes, Y.; Micheau, O.; Renno, T.; Lebecque, S. Dual role of TLR3 in Inflammation and Cancer. Cell Apoptosis Oncogene Cancer 2013. [Google Scholar] [CrossRef]
- Wagner, K.W.; Punnoose, E.A.; Januario, T.; Lawrence, D.A.; Pitti, R.M.; Lancaster, K.; Lee, D.; von Goetz, M.; Yee, S.F.; Totpal, K.; et al. Death-receptor O-glycosylation controls tumor-cell sensitivity to the proapoptotic ligand Apo2L/TRAIL. Nat. Med. 2007, 13, 1070–1077. [Google Scholar] [CrossRef] [PubMed]
- Dufour, F.; Rattier, T.; Shirley, S.; Picarda, G.; Constantinescu, A.A.; Morle, A.; Zakaria, A.B.; Marcion, G.; Causse, S.; Szegezdi, E.; et al. N-glycosylation of mouse TRAIL-R and human TRAIL-R1 enhances TRAIL-induced death. Cell Death Differ. 2017, 24, 500–510. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, T.; Shiraishi, T.; Horinaka, M.; Wakada, M.; Sakai, T. Glycosylation modulates TRAIL-R1/death receptor 4 protein: Different regulations of two pro-apoptotic receptors for TRAIL by tunicamycin. Oncol. Rep. 2007, 18, 1239–1242. [Google Scholar] [CrossRef] [PubMed]
- Shiraishi, T.; Yoshida, T.; Nakata, S.; Horinaka, M.; Wakada, M.; Mizutani, Y.; Miki, T.; Sakai, T. Tunicamycin enhances tumor necrosis factor-related apoptosis-inducing ligand-induced apoptosis in human prostate cancer cells. Cancer Res. 2005, 65, 6364–6370. [Google Scholar] [CrossRef] [PubMed]
- Jiang, C.C.; Chen, L.H.; Gillespie, S.; Kiejda, K.A.; Mhaidat, N.; Wang, Y.F.; Thorne, R.; Zhang, X.D.; Hersey, P. Tunicamycin sensitizes human melanoma cells to tumor necrosis factor-related apoptosis-inducing ligand-induced apoptosis by up-regulation of TRAIL-R2 via the unfolded protein response. Cancer Res. 2007, 67, 5880–5888. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.Y.; Du, Z.X.; Liu, B.Q.; Gao, Y.Y.; Meng, X.; Guan, Y.; Deng, W.W.; Wang, H.Q. Tunicamycin enhances TRAIL-induced apoptosis by inhibition of cyclin D1 and the subsequent downregulation of survivin. Exp. Mol. Med. 2009, 41, 362–369. [Google Scholar] [CrossRef] [PubMed]
- Jung, Y.H.; Lim, E.J.; Heo, J.; Kwon, T.K.; Kim, Y.H. Tunicamycin sensitizes human prostate cells to TRAIL-induced apoptosis by upregulation of TRAIL receptors and downregulation of cIAP2. Int. J. Oncol. 2012, 40, 1941–1948. [Google Scholar] [PubMed]
- Guo, X.; Meng, Y.; Sheng, X.; Guan, Y.; Zhang, F.; Han, Z.; Kang, Y.; Tai, G.; Zhou, Y.; Cheng, H. Tunicamycin enhances human colon cancer cells to TRAIL-induced apoptosis by JNK-CHOP-mediated DR5 upregulation and the inhibition of the EGFR pathway. Anticancer Drugs 2017, 28, 66–74. [Google Scholar] [CrossRef] [PubMed]
- van Roosmalen, I.A.M.; Reis, C.R.; Setroikromo, R.; Yuvaraj, S.; Joseph, J.V.; Tepper, P.G.; Kruyt, F.A.E.; Quax, W.J. The ER stress inducer DMC enhances TRAIL-induced apoptosis in glioblastoma. Springerplus 2014, 3, 495. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.H.; Jiang, C.C.; Kiejda, K.A.; Wang, Y.F.; Thorne, R.F.; Zhang, X.D.; Hersey, P. Thapsigargin sensitizes human melanoma cells to TRAIL-induced apoptosis by up-regulation of TRAIL-R2 through the unfolded protein response. Carcinogenesis 2007, 28, 2328–2336. [Google Scholar] [CrossRef] [PubMed]
- Martin-Perez, R.; Niwa, M.; Lopez-Rivas, A. ER stress sensitizes cells to TRAIL through down-regulation of FLIP and Mcl-1 and PERK-dependent up-regulation of TRAIL-R2. Apoptosis 2012, 17, 349–363. [Google Scholar] [CrossRef] [PubMed]
- Siegelin, M.D. Utilization of the cellular stress response to sensitize cancer cells to TRAIL-mediated apoptosis. Expert Opin. Ther. Targets 2012, 16, 801–817. [Google Scholar] [CrossRef] [PubMed]
- Munoz-Pinedo, C.; Ruiz-Ruiz, C.; Ruiz de Almodovar, C.; Palacios, C.; Lopez-Rivas, A. Inhibition of glucose metabolism sensitizes tumor cells to death receptor-triggered apoptosis through enhancement of death-inducing signaling complex formation and apical procaspase-8 processing. J. Biol. Chem. 2003, 278, 12759–12768. [Google Scholar] [CrossRef] [PubMed]
- MacFarlane, M.; Robinson, G.L.; Cain, K. Glucose—A sweet way to die: Metabolic switching modulates tumor cell death. Cell Cycle 2012, 11, 3919–3925. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Nam, S.Y.; Amoscato, A.A.; Lee, Y.J. Low glucose-enhanced TRAIL cytotoxicity is mediated through the ceramide-Akt-FLIP pathway. Oncogene 2002, 21, 337–346. [Google Scholar] [CrossRef] [PubMed]
- Qin, J.Z.; Xin, H.; Nickoloff, B.J. 2-deoxyglucose sensitizes melanoma cells to TRAIL-induced apoptosis which is reduced by mannose. Biochem. Biophys. Res. Commun. 2010, 401, 293–299. [Google Scholar] [CrossRef] [PubMed]
- Lu, M.; Lawrence, D.A.; Marsters, S.; Acosta-Alvear, D.; Kimmig, P.; Mendez, A.S.; Paton, A.W.; Paton, J.C.; Walter, P.; Ashkenazi, A. Opposing unfolded-protein-response signals converge on death receptor 5 to control apoptosis. Science 2014, 345, 98–101. [Google Scholar] [CrossRef] [PubMed]
- Dufour, F.; Rattier, T.; Constantinescu, A.A.; Zischler, L.; Morle, A.; Ben Mabrouk, H.; Humblin, E.; Jacquemin, G.; Szegezdi, E.; Delacote, F.; et al. TRAIL receptor gene editing unveils TRAIL-R1 as a master player of apoptosis induced by TRAIL and ER stress. Oncotarget 2017, 8, 9974–9985. [Google Scholar] [CrossRef] [PubMed]
- Iurlaro, R.; Puschel, F.; Leon-Annicchiarico, C.L.; O’Connor, H.; Martin, S.J.; Palou-Gramon, D.; Lucendo, E.; Munoz-Pinedo, C. Glucose Deprivation Induces ATF4-Mediated Apoptosis through TRAIL Death Receptors. Mol. Cell. Biol. 2017, 37, e00479-16. [Google Scholar] [CrossRef] [PubMed]
- Munoz-Pinedo, C.; Lopez-Rivas, A. A role for caspase-8 and TRAIL-R2/DR5 in ER-stress-induced apoptosis. Cell Death Differ. 2018, 25, 226. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Huang, Y.; Li, Y.; Pu, L.; Xia, F.; Jiang, C.; Liu, H.; Jiang, Z. [Glycosylation inhibitor 2-deoxy-d-glucose sensitizes oral cancer cells to TRAIL-induced apoptosis]. J. South. Med. Univ. 2013, 33, 524–527. [Google Scholar]
- Liu, H.; Jiang, C.C.; Lavis, C.J.; Croft, A.; Dong, L.; Tseng, H.Y.; Yang, F.; Tay, K.H.; Hersey, P.; Zhang, X.D. 2-Deoxy-d-glucose enhances TRAIL-induced apoptosis in human melanoma cells through XBP-1-mediated up-regulation of TRAIL-R2. Mol. Cancer 2009, 8, 122. [Google Scholar] [CrossRef] [PubMed]
- Moriwaki, K.; Noda, K.; Furukawa, Y.; Ohshima, K.; Uchiyama, A.; Nakagawa, T.; Taniguchi, N.; Daigo, Y.; Nakamura, Y.; Hayashi, N.; et al. Deficiency of GMDS leads to escape from NK cell-mediated tumor surveillance through modulation of TRAIL signaling. Gastroenterology 2009, 137, 188–198. [Google Scholar] [CrossRef] [PubMed]
- Moriwaki, K.; Shinzaki, S.; Miyoshi, E. GDP-mannose-4,6-dehydratase (GMDS) deficiency renders colon cancer cells resistant to tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) receptor- and CD95-mediated apoptosis by inhibiting complex II formation. J. Biol. Chem. 2011, 286, 43123–43133. [Google Scholar] [CrossRef] [PubMed]
- Sievers, F.; Wilm, A.; Dineen, D.; Gibson, T.J.; Karplus, K.; Li, W.; Lopez, R.; McWilliam, H.; Remmert, M.; Soding, J.; et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 2011, 7, 539. [Google Scholar] [CrossRef] [PubMed]
- Weng, Y.; Sui, Z.; Jiang, H.; Shan, Y.; Chen, L.; Zhang, S.; Zhang, L.; Zhang, Y. Releasing N-glycan from peptide N-terminus by N-terminal succinylation assisted enzymatic deglycosylation. Sci. Rep. 2015, 5, 9770. [Google Scholar] [CrossRef] [PubMed]
- Chan, F.K. The pre-ligand binding assembly domain: A potential target of inhibition of tumour necrosis factor receptor function. Ann. Rheum. Dis. 2000, 59 (Suppl. S1), i50–i53. [Google Scholar] [CrossRef] [PubMed]
- Chan, F.K. Three is better than one: Pre-ligand receptor assembly in the regulation of TNF receptor signaling. Cytokine 2007, 37, 101–107. [Google Scholar] [CrossRef] [PubMed]
- Edmond, V.; Ghali, B.; Penna, A.; Taupin, J.L.; Daburon, S.; Moreau, J.F.; Legembre, P. Precise Mapping of the CD95 Pre-Ligand Assembly Domain. PLoS ONE 2012, 7, e46236. [Google Scholar] [CrossRef] [PubMed]
- Vanamee, E.S.; Faustman, D.L. Structural principles of tumor necrosis factor superfamily signaling. Sci. Signal. 2018, 11, eaao4910. [Google Scholar] [CrossRef] [PubMed]
- Eck, M.J.; Beutler, B.; Kuo, G.; Merryweather, J.P.; Sprang, S.R. Crystallization of trimeric recombinant human tumor necrosis factor (cachectin). J. Biol. Chem. 1988, 263, 12816–12819. [Google Scholar] [PubMed]
- Chan, F.K.; Chun, H.J.; Zheng, L.; Siegel, R.M.; Bui, K.L.; Lenardo, M.J. A domain in TNF receptors that mediates ligand-independent receptor assembly and signaling. Science 2000, 288, 2351–2354. [Google Scholar] [CrossRef] [PubMed]
- Clancy, L.; Mruk, K.; Archer, K.; Woelfel, M.; Mongkolsapaya, J.; Screaton, G.; Lenardo, M.J.; Chan, F.K. Preligand assembly domain-mediated ligand- independent association between TRAIL receptor 4 (TR4) and TR2 regulates TRAIL-induced apoptosis. Proc. Natl. Acad. Sci. USA 2005, 102, 18099–18104. [Google Scholar] [CrossRef] [PubMed]
- Graves, J.D.; Kordich, J.J.; Huang, T.H.; Piasecki, J.; Bush, T.L.; Sullivan, T.; Foltz, I.N.; Chang, W.; Douangpanya, H.; Dang, T.; et al. Apo2L/TRAIL and the Death Receptor 5 Agonist Antibody AMG 655 Cooperate to Promote Receptor Clustering and Antitumor Activity. Cancer Cell 2014, 26, 177–189. [Google Scholar] [CrossRef] [PubMed]
- Valley, C.C.; Lewis, A.K.; Mudaliar, D.J.; Perlmutter, J.D.; Braun, A.R.; Karim, C.B.; Thomas, D.D.; Brody, J.R.; Sachs, J.N. Tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) induces death receptor 5 networks that are highly organized. J. Biol. Chem. 2012, 287, 21265–21278. [Google Scholar] [CrossRef] [PubMed]
- Brandt, B.; Buchse, T.; Abou-Eladab, E.F.; Tiedge, M.; Krause, E.; Jeschke, U.; Walzel, H. Galectin-1 induced activation of the apoptotic death-receptor pathway in human Jurkat T lymphocytes. Histochem. Cell Biol. 2008, 129, 599–609. [Google Scholar] [CrossRef] [PubMed]
- Dorrie, J.; Sapala, K.; Zunino, S.J. Interferon-gamma increases the expression of glycosylated CD95 in B-leukemic cells: An inducible model to study the role of glycosylation in CD95-signalling and trafficking. Cytokine 2002, 18, 98–107. [Google Scholar] [CrossRef] [PubMed]
- Shatnyeva, O.M.; Kubarenko, A.V.; Weber, C.E.; Pappa, A.; Schwartz-Albiez, R.; Weber, A.N.; Krammer, P.H.; Lavrik, I.N. Modulation of the CD95-induced apoptosis: The role of CD95 N-glycosylation. PLoS ONE 2011, 6, e19927. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Kim, S.; Hurtado, J.; Lee, Z.H.; Kim, K.K.; Pollok, K.E.; Kwon, B.S. Characterization of human homologue of 4-1BB and its ligand. Immunol. Lett. 1995, 45, 67–73. [Google Scholar] [CrossRef]
- Madireddi, S.; Eun, S.Y.; Lee, S.W.; Nemcovicova, I.; Mehta, A.K.; Zajonc, D.M.; Nishi, N.; Niki, T.; Hirashima, M.; Croft, M. Galectin-9 controls the therapeutic activity of 4-1BB-targeting antibodies. J. Exp. Med. 2014, 211, 1433–1448. [Google Scholar] [CrossRef] [PubMed]
- Bitra, A.; Doukov, T.; Wang, J.; Picarda, G.; Benedict, C.A.; Croft, M.; Zajonc, D.M. Crystal structure of murine 4-1BB and its interaction with 4-1BBL support a role for galectin-9 in 4-1BB signaling. J. Biol. Chem. 2018, 293, 1317–1329. [Google Scholar] [CrossRef] [PubMed]
- Mazurek, N.; Byrd, J.C.; Sun, Y.; Hafley, M.; Ramirez, K.; Burks, J.; Bresalier, R.S. Cell-surface galectin-3 confers resistance to TRAIL by impeding trafficking of death receptors in metastatic colon adenocarcinoma cells. Cell Death Differ. 2012, 19, 523–533. [Google Scholar] [CrossRef] [PubMed]
- Stewart, S.E.; Menzies, S.A.; Popa, S.J.; Savinykh, N.; Petrunkina Harrison, A.; Lehner, P.J.; Moreau, K. A genome-wide CRISPR screen reconciles the role of N-linked glycosylation in galectin-3 transport to the cell surface. J. Cell Sci. 2017, 130, 3234–3247. [Google Scholar] [CrossRef] [PubMed]
- Reis, C.R.; Chen, P.H.; Bendris, N.; Schmid, S.L. TRAIL-death receptor endocytosis and apoptosis are selectively regulated by dynamin-1 activation. Proc. Natl. Acad. Sci. USA 2017, 114, 504–509. [Google Scholar] [CrossRef] [PubMed]
- Hadari, Y.R.; Arbel-Goren, R.; Levy, Y.; Amsterdam, A.; Alon, R.; Zakut, R.; Zick, Y. Galectin-8 binding to integrins inhibits cell adhesion and induces apoptosis. J. Cell Sci. 2000, 113, 2385–2397. [Google Scholar] [PubMed]
- Perillo, N.L.; Pace, K.E.; Seilhamer, J.J.; Baum, L.G. Apoptosis of T cells mediated by galectin-1. Nature 1995, 378, 736–739. [Google Scholar] [CrossRef] [PubMed]
- Perillo, N.L.; Uittenbogaart, C.H.; Nguyen, J.T.; Baum, L.G. Galectin-1, an endogenous lectin produced by thymic epithelial cells, induces apoptosis of human thymocytes. J. Exp. Med. 1997, 185, 1851–1858. [Google Scholar] [CrossRef] [PubMed]
- Yang, R.Y.; Hsu, D.K.; Liu, F.T. Expression of galectin-3 modulates T-cell growth and apoptosis. Proc. Natl. Acad. Sci. USA 1996, 93, 6737–6742. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.J.; Song, Y.K.; Song, J.J.; Siervo-Sassi, R.R.; Kim, H.R.; Li, L.; Spitz, D.R.; Lokshin, A.; Kim, J.H. Reconstitution of galectin-3 alters glutathione content and potentiates TRAIL-induced cytotoxicity by dephosphorylation of Akt. Exp. Cell Res. 2003, 288, 21–34. [Google Scholar] [CrossRef]
- Oka, N.; Nakahara, S.; Takenaka, Y.; Fukumori, T.; Hogan, V.; Kanayama, H.O.; Yanagawa, T.; Raz, A. Galectin-3 inhibits tumor necrosis factor-related apoptosis-inducing ligand-induced apoptosis by activating Akt in human bladder carcinoma cells. Cancer Res. 2005, 65, 7546–7553. [Google Scholar] [CrossRef] [PubMed]
- Mazurek, N.; Sun, Y.J.; Liu, K.F.; Gilcrease, M.Z.; Schober, W.; Nangia-Makker, P.; Raz, A.; Bresalier, R.S. Phosphorylated galectin-3 mediates tumor necrosis factor-related apoptosis-inducing ligand signaling by regulating phosphatase and tensin homologue deleted on chromosome 10 in human breast carcinoma cells. J. Biol. Chem. 2007, 282, 21337–21348. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.I.; Whang, E.E.; Abramson, M.A.; Donner, D.B.; Bertagnolli, M.M.; Moore, F.D., Jr.; Ruan, D.T. Galectin-3 regulates apoptosis and doxorubicin chemoresistance in papillary thyroid cancer cells. Biochem. Biophys. Res. Commun. 2009, 379, 626–631. [Google Scholar] [CrossRef] [PubMed]
- Mazurek, N.; Byrd, J.C.; Sun, Y.; Ueno, S.; Bresalier, R.S. A galectin-3 sequence polymorphism confers TRAIL sensitivity to human breast cancer cells. Cancer 2011, 117, 4375–4380. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Sun, R.R.; Yu, Z.J.; Liang, H.; Shen, S.; Kan, Q. Galectin-1 Modulates the Survival and Tumor Necrosis Factor-Related Apoptosis-Inducing Ligand (TRAIL) Sensitivity in Human Hepatocellular Carcinoma Cells. Cancer Biother. Radiopharm. 2015, 30, 336–341. [Google Scholar] [CrossRef] [PubMed]
- Fukumori, T.; Takenaka, Y.; Oka, N.; Yoshii, T.; Hogan, V.; Inohara, H.; Kanayama, H.O.; Kim, H.R.; Raz, A. Endogenous galectin-3 determines the routing of CD95 apoptotic signaling pathways. Cancer Res. 2004, 64, 3376–3379. [Google Scholar] [CrossRef] [PubMed]
- Scaffidi, C.; Schmitz, I.; Zha, J.; Korsmeyer, S.J.; Krammer, P.H.; Peter, M.E. Differential modulation of apoptosis sensitivity in CD95 type I and type II cells. J. Biol. Chem. 1999, 274, 22532–22538. [Google Scholar] [CrossRef] [PubMed]
- Mazurek, N.; Jie Sun, Y.; Feng Liu, K.; Gilcrease, M.Z.; Schober, W.; Nangia-Makker, P.; Raz, A.; Bresalier, R.S. Phosphorylated galectin-3 mediates tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) signaling by regulating PTEN in human breast carcinoma cells. J. Biol. Chem. 2007, 282, 21337–21348. [Google Scholar] [CrossRef] [PubMed]
- Lin, T.C.; Chen, S.T.; Huang, M.C.; Huang, J.; Hsu, C.L.; Juan, H.F.; Lin, H.H.; Chen, C.H. GALNT6 expression enhances aggressive phenotypes of ovarian cancer cells by regulating EGFR activity. Oncotarget 2017, 8, 42588–42601. [Google Scholar] [CrossRef] [PubMed]
- Merlin, J.; Stechly, L.; de Beauce, S.; Monte, D.; Leteurtre, E.; van Seuningen, I.; Huet, G.; Pigny, P. Galectin-3 regulates MUC1 and EGFR cellular distribution and EGFR downstream pathways in pancreatic cancer cells. Oncogene 2011, 30, 2514–2525. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Hsu, D.K.; Chen, H.Y.; Yang, R.Y.; Carraway, K.L., 3rd; Isseroff, R.R.; Liu, F.T. Galectin-3 regulates intracellular trafficking of EGFR through Alix and promotes keratinocyte migration. J. Investig. Dermatol. 2012, 132, 2828–2837. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Gu, J.; Ihara, H.; Miyoshi, E.; Honke, K.; Taniguchi, N. Core fucosylation regulates epidermal growth factor receptor-mediated intracellular signaling. J. Biol. Chem. 2006, 281, 2572–2577. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, K.; Yokote, H.; Arao, T.; Maegawa, M.; Tanaka, K.; Fujita, Y.; Shimizu, C.; Hanafusa, T.; Fujiwara, Y.; Nishio, K. N-Glycan fucosylation of epidermal growth factor receptor modulates receptor activity and sensitivity to epidermal growth factor receptor tyrosine kinase inhibitor. Cancer Sci. 2008, 99, 1611–1617. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.C.; Yen, H.Y.; Chen, C.Y.; Chen, C.H.; Cheng, P.F.; Juan, Y.H.; Chen, C.H.; Khoo, K.H.; Yu, C.J.; Yang, P.C.; et al. Sialylation and fucosylation of epidermal growth factor receptor suppress its dimerization and activation in lung cancer cells. Proc. Natl. Acad. Sci. USA 2011, 108, 11332–11337. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira, J.T.; de Matos, A.J.; Santos, A.L.; Pinto, R.; Gomes, J.; Hespanhol, V.; Chammas, R.; Manninen, A.; Bernardes, E.S.; Albuquerque Reis, C.; et al. Sialylation regulates galectin-3/ligand interplay during mammary tumour progression--a case of targeted uncloaking. Int. J. Dev. Biol. 2011, 55, 823–834. [Google Scholar] [CrossRef] [PubMed]
- Piyush, T.; Chacko, A.R.; Sindrewicz, P.; Hilkens, J.; Rhodes, J.M.; Yu, L.G. Interaction of galectin-3 with MUC1 on cell surface promotes EGFR dimerization and activation in human epithelial cancer cells. Cell Death Differ. 2017, 24, 1937–1947. [Google Scholar] [CrossRef] [PubMed]
- Mozzi, A.; Forcella, M.; Riva, A.; Difrancesco, C.; Molinari, F.; Martin, V.; Papini, N.; Bernasconi, B.; Nonnis, S.; Tedeschi, G.; et al. NEU3 activity enhances EGFR activation without affecting EGFR expression and acts on its sialylation levels. Glycobiology 2015, 25, 855–868. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yen, H.Y.; Liu, Y.C.; Chen, N.Y.; Tsai, C.F.; Wang, Y.T.; Chen, Y.J.; Hsu, T.L.; Yang, P.C.; Wong, C.H. Effect of sialylation on EGFR phosphorylation and resistance to tyrosine kinase inhibition. Proc. Natl. Acad. Sci. USA 2015, 112, 6955–6960. [Google Scholar] [CrossRef] [PubMed]
- Lichtenstein, R.G.; Rabinovich, G.A. Glycobiology of cell death: When glycans and lectins govern cell fate. Cell Death Differ. 2013, 20, 976–986. [Google Scholar] [CrossRef] [PubMed]
- Peter, M.E.; Hellbardt, S.; Schwartz-Albiez, R.; Westendorp, M.O.; Walczak, H.; Moldenhauer, G.; Grell, M.; Krammer, P.H. Cell surface sialylation plays a role in modulating sensitivity towards APO-1-mediated apoptotic cell death. Cell Death Differ. 1995, 2, 163–171. [Google Scholar] [PubMed]
- Keppler, O.T.; Peter, M.E.; Hinderlich, S.; Moldenhauer, G.; Stehling, P.; Schmitz, I.; Schwartz-Albiez, R.; Reutter, W.; Pawlita, M. Differential sialylation of cell surface glycoconjugates in a human B lymphoma cell line regulates susceptibility for CD95 (APO-1/Fas)-mediated apoptosis and for infection by a lymphotropic virus. Glycobiology 1999, 9, 557–569. [Google Scholar] [CrossRef] [PubMed]
- Amano, M.; Galvan, M.; He, J.; Baum, L.G. The ST6Gal I sialyltransferase selectively modifies N-glycans on CD45 to negatively regulate galectin-1-induced CD45 clustering, phosphatase modulation, and T cell death. J. Biol. Chem. 2003, 278, 7469–7475. [Google Scholar] [CrossRef] [PubMed]
- Swindall, A.F.; Bellis, S.L. Sialylation of the Fas death receptor by ST6Gal-I provides protection against Fas-mediated apoptosis in colon carcinoma cells. J. Biol. Chem. 2011, 286, 22982–22990. [Google Scholar] [CrossRef] [PubMed]
- Corti, A.; Merli, S.; Bagnasco, L.; D’Ambrosio, F.; Marino, M.; Cassani, G. Identification of two forms (31-33 and 48 kD) of the urinary soluble p55 tumor necrosis factor receptor that are differentially N- and O-glycosylated. J. Interferon Cytokine Res. 1995, 15, 143–152. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Swindall, A.F.; Kesterson, R.A.; Schoeb, T.R.; Bullard, D.C.; Bellis, S.L. ST6Gal-I regulates macrophage apoptosis via alpha2-6 sialylation of the TNFR1 death receptor. J. Biol. Chem. 2011, 286, 39654–39662. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Woronicz, J.D.; Liu, W.; Goeddel, D.V. Prevention of constitutive TNF receptor 1 signaling by silencer of death domains. Science 1999, 283, 543–546. [Google Scholar] [CrossRef] [PubMed]
- Kitson, J.; Raven, T.; Jiang, Y.P.; Goeddel, D.V.; Giles, K.M.; Pun, K.T.; Grinham, C.J.; Brown, R.; Farrow, S.N. A death-domain-containing receptor that mediates apoptosis. Nature 1996, 384, 372–375. [Google Scholar] [CrossRef] [PubMed]
- Walczak, H.; Degli-Esposti, M.A.; Johnson, R.S.; Smolak, P.J.; Waugh, J.Y.; Boiani, N.; Timour, M.S.; Gerhart, M.J.; Schooley, K.A.; Smith, C.A.; et al. TRAIL-R2: A novel apoptosis-mediating receptor for TRAIL. Embo J. 1997, 16, 5386–5397. [Google Scholar] [CrossRef] [PubMed]
- Takada, H.; Chen, N.J.; Mirtsos, C.; Suzuki, S.; Suzuki, N.; Wakeham, A.; Mak, T.W.; Yeh, W.C. Role of SODD in regulation of tumor necrosis factor responses. Mol. Cell. Biol. 2003, 23, 4026–4033. [Google Scholar] [CrossRef] [PubMed]
- Kamili, N.A.; Arthur, C.M.; Gerner-Smidt, C.; Tafesse, E.; Blenda, A.; Dias-Baruffi, M.; Stowell, S.R. Key regulators of galectin-glycan interactions. Proteomics 2016, 16, 3111–3125. [Google Scholar] [CrossRef] [PubMed]

| Target/Compound | Cell Type | TRAIL-Induced Apoptosis | Comment | Reference |
|---|---|---|---|---|
| Carbohydrate-binding proteins | ||||
| Tunicamycin | Colon carcinomas | Enhanced | ⬆ DR5 and ⬇ EGFR | [49] |
| Colon and Lung carcinomas | Enhanced | UPR-mediated ⬆ DR5 | [58] | |
| ER/Golgi Stressor | Multiple Myeloma | |||
| Oral cancer cells | Enhanced | ⬆ DR5 | [62] | |
| 2-deoxy-d-glucose | Melanomas | Enhanced | ⬆ DR5 | [63] |
| Melanomas | Enhanced | ⬆ DR5 | [57] | |
| Glucose deprivation | Leukemias, Breast and Cervical carcinomas | Enhanced | ⬆ TRAIL DISC formation and ⬇ of c-FLIP | [54,60] |
| Colon carcinomas | Enhanced | ATF4-mediated ⬆ of DR4 and DR5 | [61] | |
| Fucosylation (GDMS) | Colon carcinomas | Inhibited | [64,65] | |
| benzyl-a-GalNAc | Pancreatic cancer Melanoma Colon carcinomas | Inhibited | ⬆ TRAIL DISC formation and ⬆ receptor aggregation | |
| siGalnt14 | Inhibited | |||
| siGalnt3 | Inhibited | [42] | ||
| siFUT6 | Inhibited | |||
| GALNT14 overexp. | Enhanced | |||
| Target/Compound | Cell Type | TRAIL-Induced Apoptosis (*FasL) | Comment | Reference |
|---|---|---|---|---|
| Carbohydrate-binding proteins | ||||
| siGal1 | Hepatocellular Carcinomas | Enhanced | ⬇ Survivin and ⬇ Bcl-2 | [95] |
| Gal3 P64→H64 | Breast Carcinomas | Enhanced | [94] | |
| siGal3 | Papillary Thyroid Carcinomas | Enhanced | ⬇ AKT | [93] |
| phospho-Gal3 | Breast Carcinomas | Enhanced | ⬇ AKT | [98] |
| Gal-3 Overexp. | Bladder Carcinomas | Inhibited | ⬆ AKT | [91] |
| Breast Carcinomas | Enhanced | ⬇ AKT | [90] | |
| Leukemias | Enhanced | converts type II cells into type I | [96] | |
| Gal3 extracellular | Colon carcinomas | Inhibited | Inhibition of receptor trafficking (membrane level) | [83] |
© 2018 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Micheau, O. Regulation of TNF-Related Apoptosis-Inducing Ligand Signaling by Glycosylation. Int. J. Mol. Sci. 2018, 19, 715. https://doi.org/10.3390/ijms19030715
Micheau O. Regulation of TNF-Related Apoptosis-Inducing Ligand Signaling by Glycosylation. International Journal of Molecular Sciences. 2018; 19(3):715. https://doi.org/10.3390/ijms19030715
Chicago/Turabian StyleMicheau, Olivier. 2018. "Regulation of TNF-Related Apoptosis-Inducing Ligand Signaling by Glycosylation" International Journal of Molecular Sciences 19, no. 3: 715. https://doi.org/10.3390/ijms19030715
APA StyleMicheau, O. (2018). Regulation of TNF-Related Apoptosis-Inducing Ligand Signaling by Glycosylation. International Journal of Molecular Sciences, 19(3), 715. https://doi.org/10.3390/ijms19030715