Comparative Transcriptome Profiling of Rice Near-Isogenic Line Carrying Xa23 under Infection of Xanthomonas oryzae pv. oryzae
Abstract
1. Introduction
2. Results
2.1. RNA Sequencing of Different Samples and Data Analysis
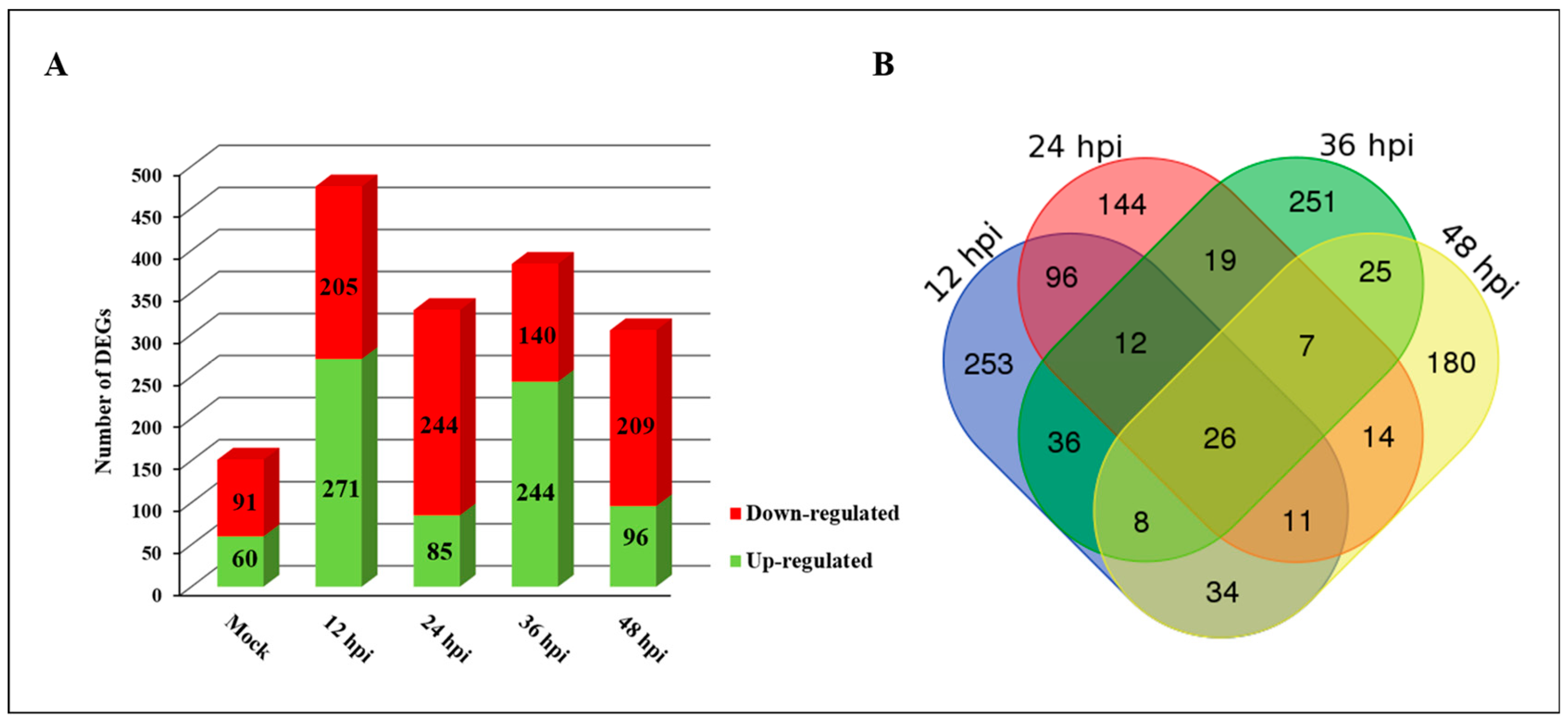
2.2. Identification of DEGs in Response to PXO99A in CBB23 and JG30
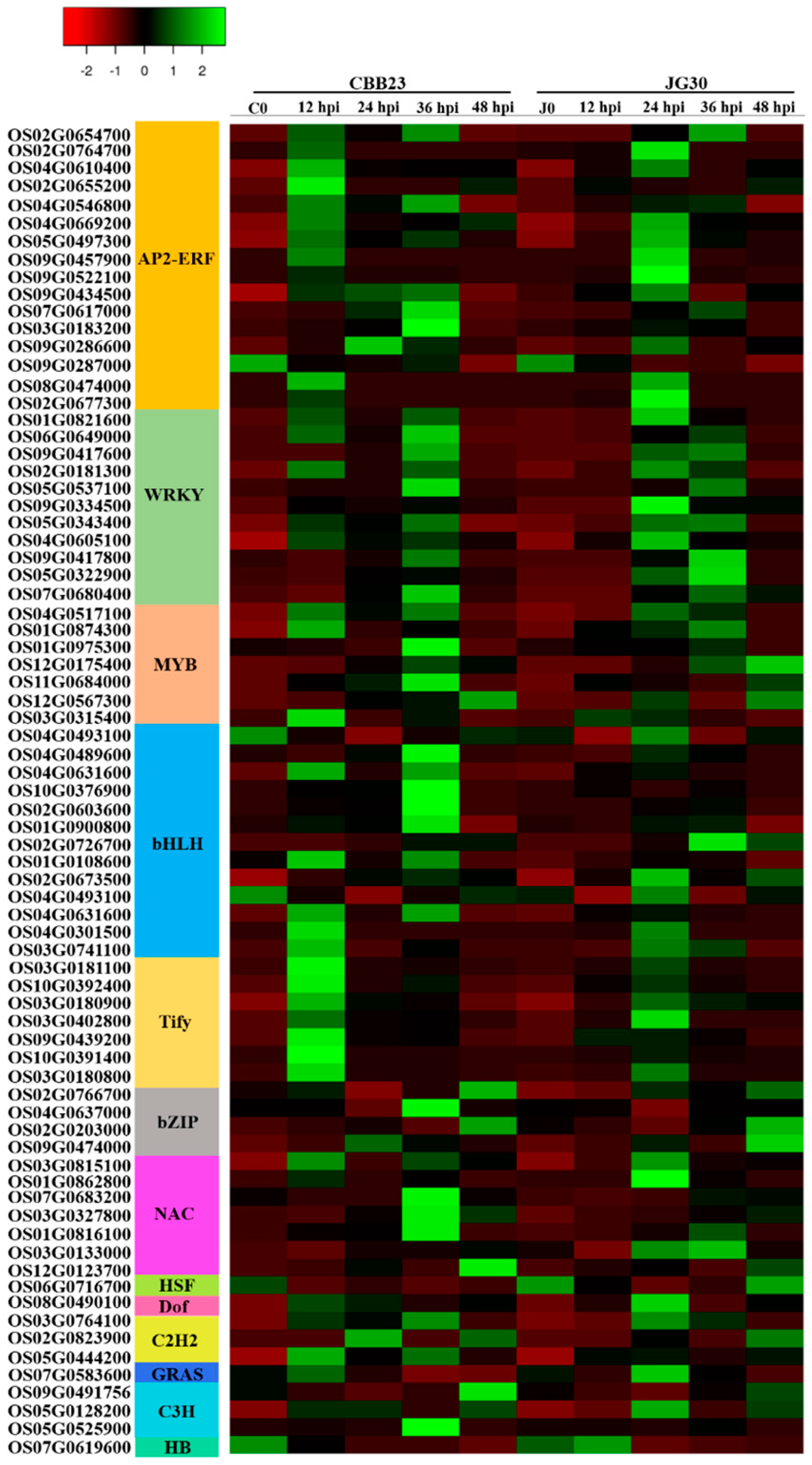
2.3. Response of Differentially Expressed Transcription Factors to PXO99A Infection
2.4. Response of Different Kinase Responsive Genes to PXO99A Infection
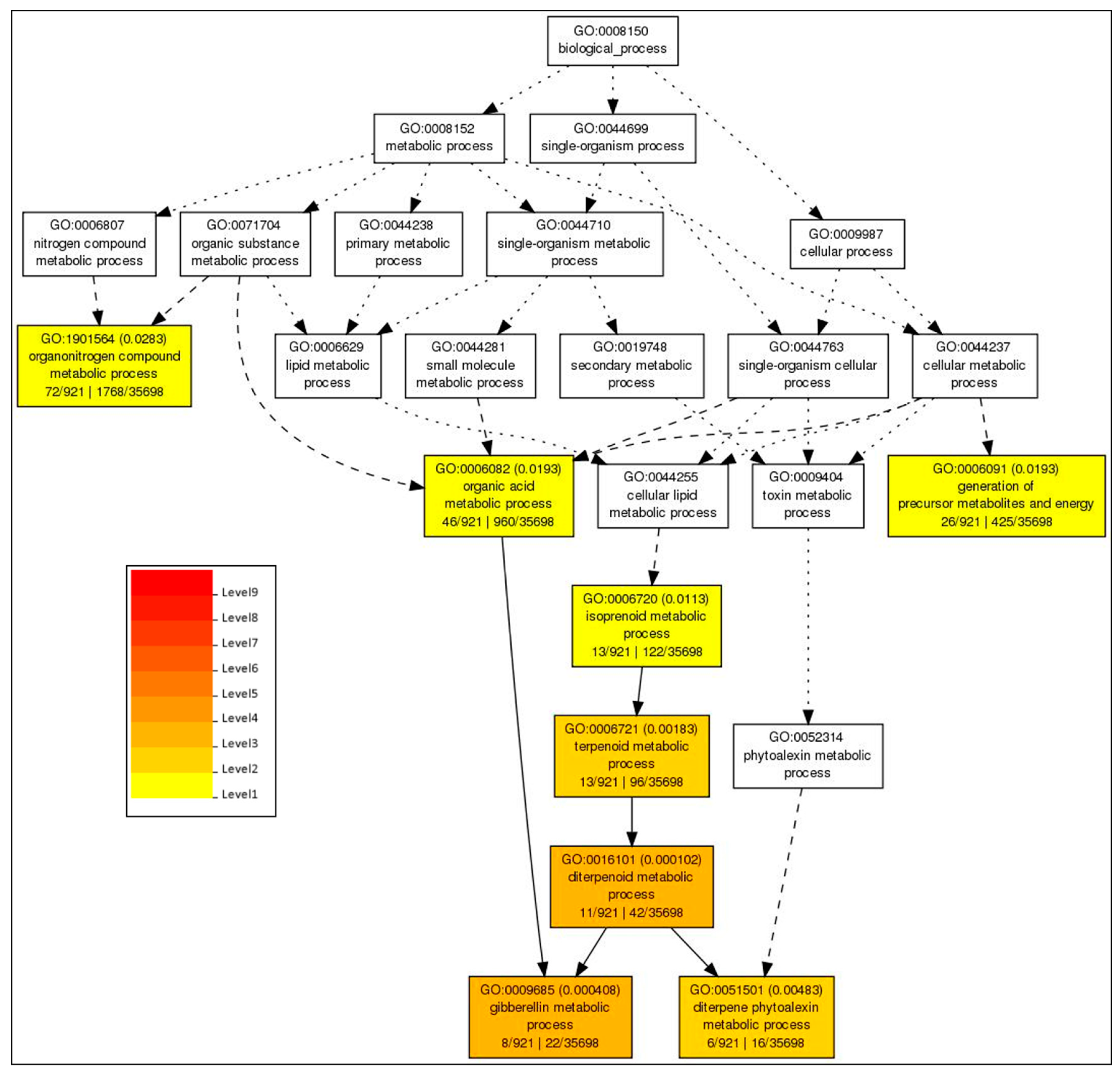
2.5. GO Based Analysis of the DEGs
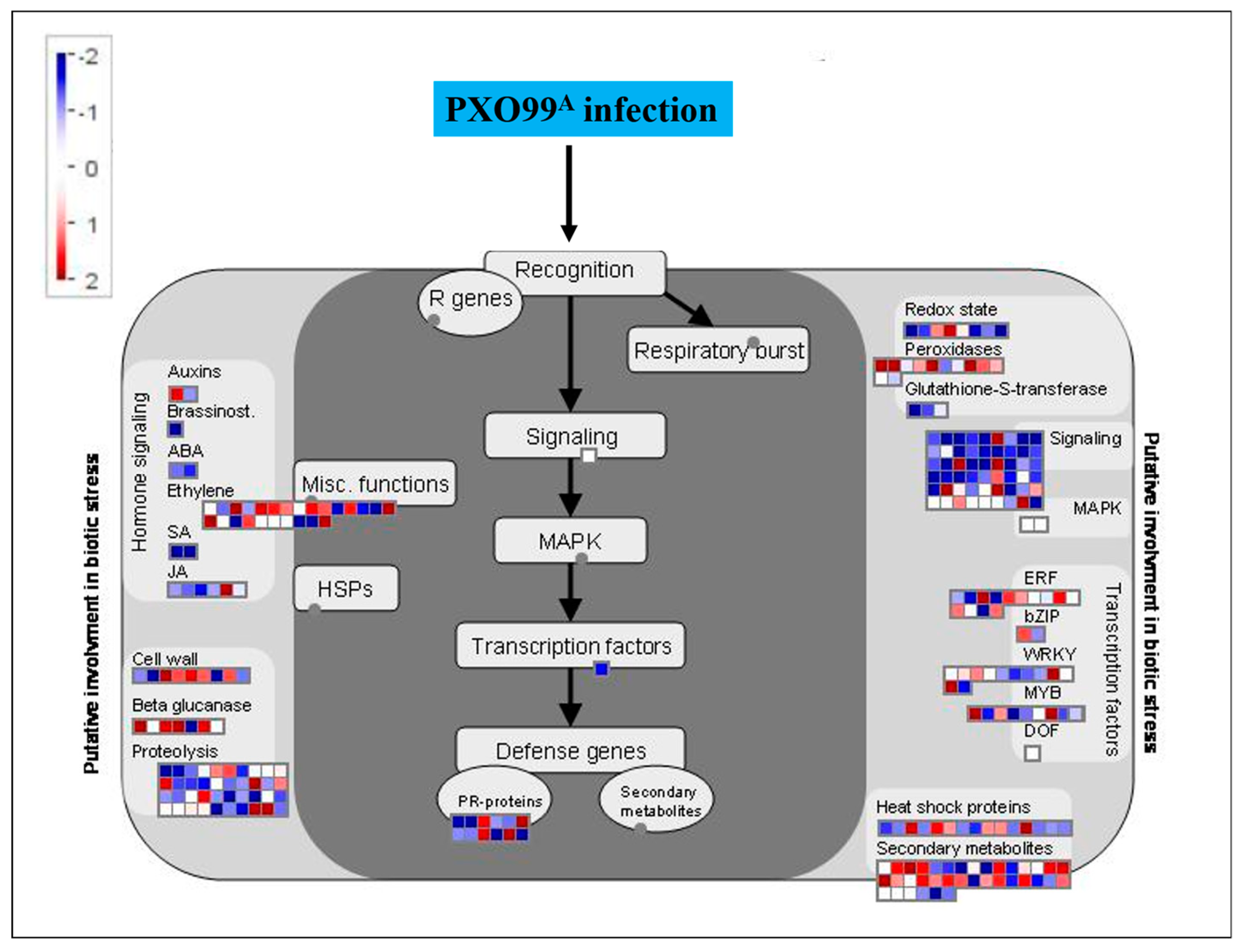
2.6. MapMan Overview of DEGs Related to Plant–Pathogen Interaction
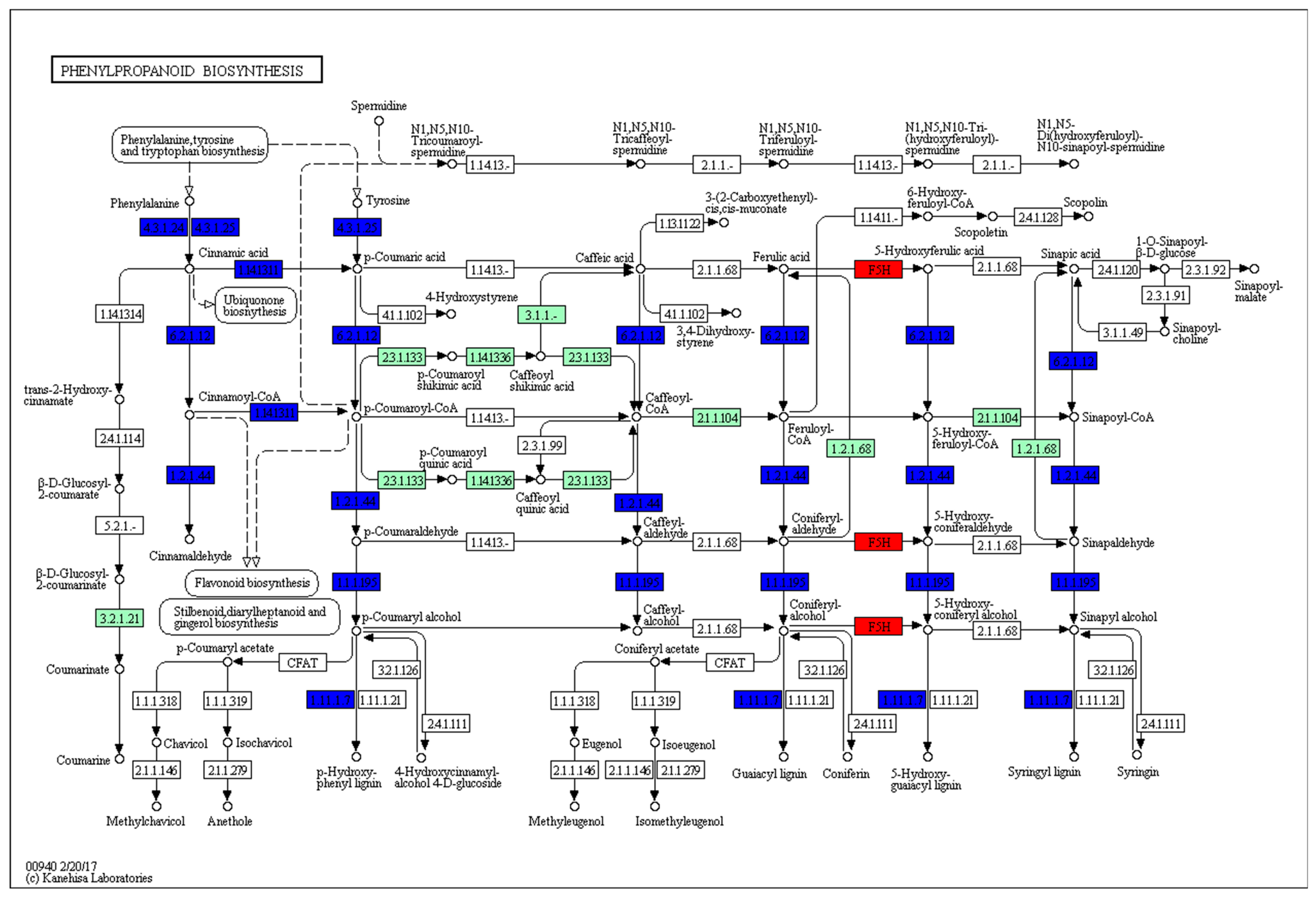
2.7. KEGG Pathway
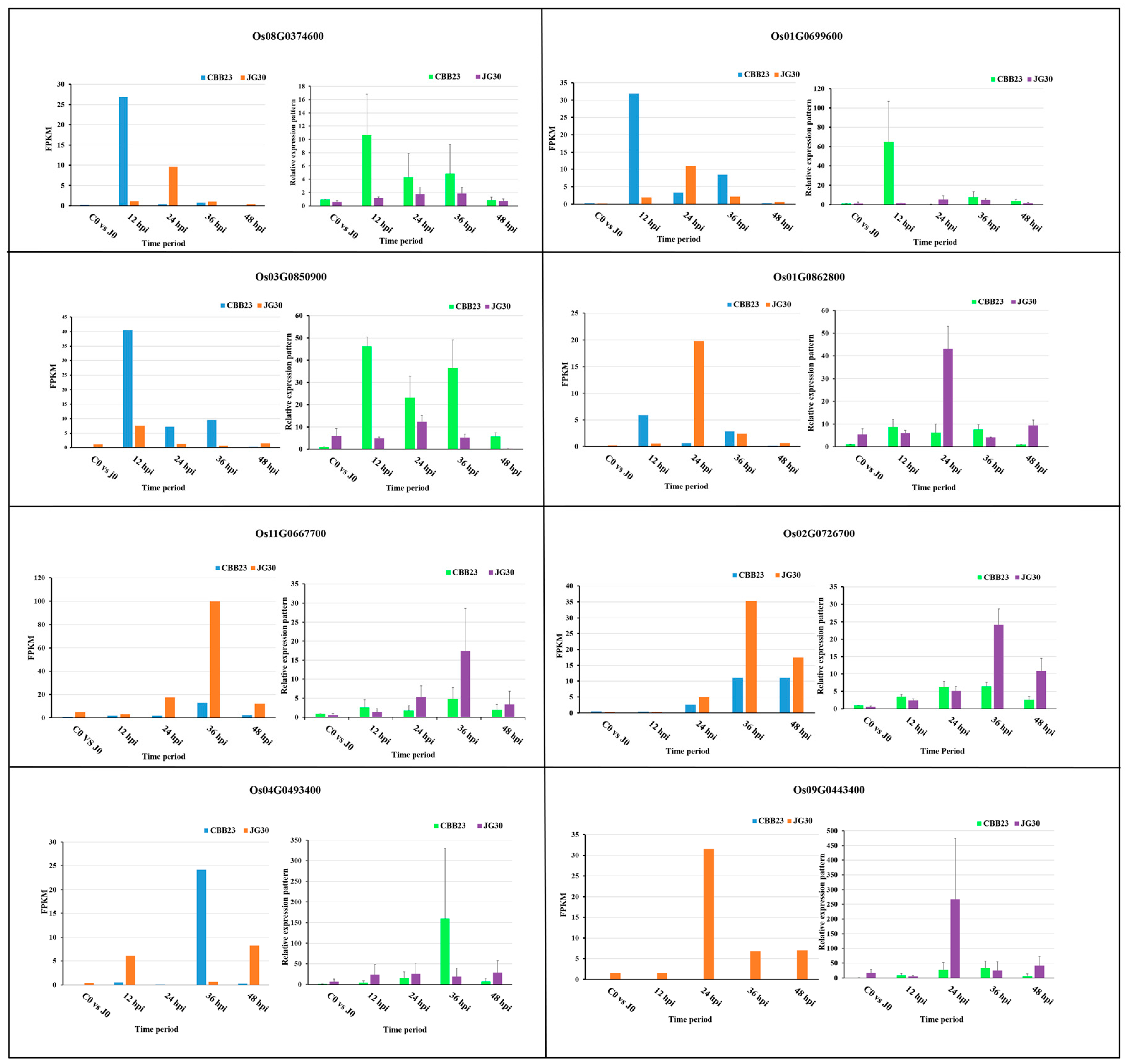
2.8. Validation of RNA-Seq Results by qRT-PCR
3. Discussion
4. Materials and Methods
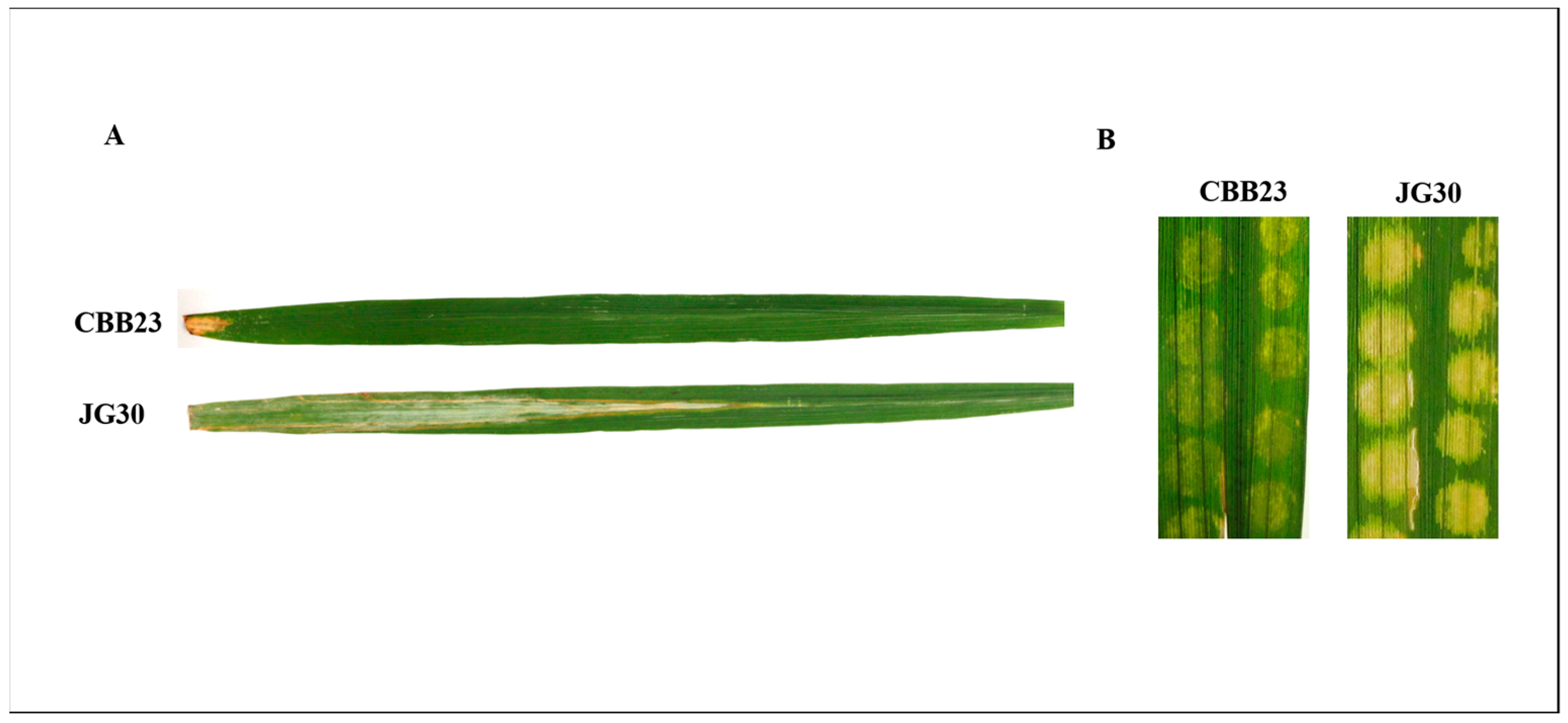
4.1. Plant Material and Growth Conditions
4.2. Inoculation of Rice Plants with PXO99A and Collection of Leaf Samples
4.3. RNA Extraction and Library Preparation for Illumina Sequencing
4.4. Data Analysis
4.5. Gene Enrichment Analysis
4.6. Validation of RNA-Seq Data
Supplementary Materials
Availability of Supporting Data
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Heckman, D.S.; Geiser, D.M.; Eidell, B.R.; Stauffer, R.L.; Kardos, N.L.; Hedges, S.B. Molecular evidence for the early colonization of land by fungi and plants. Science 2001, 293, 1129–1133. [Google Scholar] [CrossRef] [PubMed]
- Zuluaga, P.; Szurek, B.; Koebnik, R.; Kroj, T.; Morel, J.-B. Effector mimics and integrated decoys, the never-ending arms race between rice and Xanthomonas oryzae. Front. Plant Sci. 2017, 8, 431. [Google Scholar] [CrossRef] [PubMed]
- Anderson, J.P.; Gleason, C.A.; Foley, R.C.; Thrall, P.H.; Burdon, J.B.; Singh, K.B. Plants versus pathogens: An evolutionary arms race. Funct. Plant Biol. 2010, 37, 499–512. [Google Scholar] [CrossRef] [PubMed]
- Webster, R.K. Compendium of Rice Diseases; The American Phytopathological Society (APS): St. Paul, MN, USA, 1992. [Google Scholar]
- Gnanamanickam, S.; Priyadarisini, V.B.; Narayanan, N.; Vasudevan, P.; Kavitha, S. An overview of bacterial blight disease of rice and strategies for its management. Curr. Sci. 1999, 1435–1444. [Google Scholar]
- Tagami, Y.; Mizukami, T. Historical review of the researches on bacterial blight of rice caused by Xanthomonas oryzae (Uyede and Ishiyama) dowson. Spec. Rep. Plant Dis. Insect Pests Forecast. Serv. 1962, 10, 1–112. [Google Scholar]
- Mizukami, T.; Wakimoto, S. Epidemiology and control of bacterial leaf blight of rice. Annu. Rev. Phytopathol. 1969, 7, 51–72. [Google Scholar] [CrossRef]
- Mew, T.W. Current status and future prospects of research on bacterial blight of rice. Annu. Rev. Phytopathol. 1987, 25, 359–382. [Google Scholar] [CrossRef]
- Noda, T.; Kaku, H. Growth of Xanthomonas oryzae pv. oryzae in planta and in guttation fluid of rice. Jpn. J. Phytopathol. 1999, 65, 9–14. [Google Scholar] [CrossRef]
- Boch, J.; Scholze, H.; Schornack, S.; Landgraf, A.; Hahn, S.; Kay, S.; Lahaye, T.; Nickstadt, A.; Bonas, U. Breaking the code of DNA binding specificity of TAL-type III effectors. Science 2009, 326, 1509–1512. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Rinaldi, F.C.; Singh, P.; Doyle, E.L.; Dubrow, Z.E.; Tran, T.T.; Pérez-Quintero, A.L.; Szurek, B.; Bogdanove, A.J. TAL effectors drive transcription bidirectionally in plants. Mol. Plant 2017, 10, 285–296. [Google Scholar] [CrossRef] [PubMed]
- Busungu, C.; Taura, S.; Sakagami, J.-I.; Ichitani, K. Identification and linkage analysis of a new rice bacterial blight resistance gene from Xm14, a mutant line from IR24. Breed. Sci. 2016, 66, 636–645. [Google Scholar] [CrossRef] [PubMed]
- Vikal, Y.; Bhatia, D. Genetics and genomics of bacterial blight resistance in rice. In Advances in International Rice Research; InTech: London, UK, 2017. [Google Scholar]
- Wang, C.; Zhang, X.; Fan, Y.; Gao, Y.; Zhu, Q.; Zheng, C.; Qin, T.; Li, Y.; Che, J.; Zhang, M. Xa23 is an executor R protein and confers broad-spectrum disease resistance in rice. Mol. Plant 2015, 8, 290–302. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Wang, C.; Zhao, K.; Zhao, Y.; Caslana, V.; Zhu, X.; Li, D.; Jiang, Q.X. The Effectiveness of Advanced Rice Lines with New Resistance Gene Xa23 to Rice Bacterial Blight. Rice Genet. Newsl. 2001, 18, 71–72. [Google Scholar]
- Han, Y.; Gao, S.; Muegge, K.; Zhang, W.; Zhou, B. Advanced applications of RNA sequencing and challenges. Bioinform. Biol. Insights 2015, 9, BBI-S28991. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Gerstein, M.; Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 2009, 10, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Vogel, C.; Bodenhausen, N.; Gruissem, W.; Vorholt, J.A. The arabidopsis leaf transcriptome reveals distinct but also overlapping responses to colonization by phyllosphere commensals and pathogen infection with impact on plant health. New Phytol. 2016, 212, 192–207. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Xia, H.; Liu, Y.; Wei, H.; Zheng, X.; Song, C.; Chen, L.; Liu, H.; Luo, L. Transcriptomic and metabolomic studies disclose key metabolism pathways contributing to well-maintained photosynthesis under the drought and the consequent drought-tolerance in rice. Front. Plant Sci. 2016, 7, 1886. [Google Scholar] [CrossRef] [PubMed]
- Yadav, I.S.; Sharma, A.; Kaur, S.; Nahar, N.; Bhardwaj, S.C.; Sharma, T.R.; Chhuneja, P. Comparative temporal transcriptome profiling of wheat near isogenic line carrying Lr57 under compatible and incompatible interactions. Front. Plant Sci. 2016, 7, 1943. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhou, Z.; Gao, J.; Wu, Y.; Xia, Z.; Zhang, H.; Wu, J. The mechanisms of maize resistance to Fusarium verticillioides by comprehensive analysis of RNA-Seq data. Front. Plant Sci. 2016, 7, 1654. [Google Scholar] [CrossRef] [PubMed]
- Eulgem, T.; Rushton, P.J.; Robatzek, S.; Somssich, I.E. The wrky superfamily of plant transcription factors. Trends Plant Sci. 2000, 5, 199–206. [Google Scholar] [CrossRef]
- Chisholm, S.T.; Coaker, G.; Day, B.; Staskawicz, B.J. Host-microbe interactions: Shaping the evolution of the plant immune response. Cell 2006, 124, 803–814. [Google Scholar] [CrossRef] [PubMed]
- Dodds, P.N.; Rathjen, J.P. Plant immunity: Towards an integrated view of plant–pathogen interactions. Nat. Rev. Genet. 2010, 11, 539–548. [Google Scholar] [CrossRef] [PubMed]
- Bonas, U.; Stall, R.E.; Staskawicz, B. Genetic and structural characterization of the avirulence gene avrBs3 from Xanthomonas campestris pv. Vesicatoria. Mol. Gen. Genet. 1989, 218, 127–136. [Google Scholar] [CrossRef] [PubMed]
- Kono, Y.; Kojima, A.; Nagai, R.; Watanabe, M.; Kawashima, T.; Onizawa, T.; Teraoka, T.; Watanab, M.; Koshino, H.; Uzawa, J. Antibacterial diterpenes and their fatty acid conjugates from rice leaves. Phytochemistry 2004, 65, 1291–1298. [Google Scholar] [CrossRef] [PubMed]
- Schmelz, E.A.; Huffaker, A.; Sims, J.W.; Christensen, S.A.; Lu, X.; Okada, K.; Peters, R.J. Biosynthesis, elicitation and roles of monocot terpenoid phytoalexins. Plant J. 2014, 79, 659–678. [Google Scholar] [CrossRef] [PubMed]
- Xie, K.; Chen, J.; Wang, Q.; Yang, Y. Direct phosphorylation and activation of a mitogen-activated protein kinase by a calcium-dependent protein kinase in rice. Plant Cell 2014, 26, 3077–3089. [Google Scholar] [CrossRef] [PubMed]
- Fu, L.; Yu, X.; An, C. Overexpression of constitutively active OsCPK10 increases Arabidopsis resistance against Pseudomonas syringae pv. tomato and rice resistance against Magnaporthe grisea. Plant Physiol. Biochem. 2013, 73, 202–210. [Google Scholar] [CrossRef] [PubMed]
- Almagro, L.; Gómez Ros, L.; Belchi-Navarro, S.; Bru, R.; Ros Barceló, A.; Pedreno, M. Class III peroxidases in plant defence reactions. J. Exp. Bot. 2008, 60, 377–390. [Google Scholar] [CrossRef] [PubMed]
- Jiang, G.; Yin, D.; Zhao, J.; Chen, H.; Guo, L.; Zhu, L.; Zhai, W. The rice thylakoid membrane-bound ascorbate peroxidase OsAPX8 functions in tolerance to bacterial blight. Sci. Rep. 2016, 6, 26104. [Google Scholar] [CrossRef] [PubMed]
- Zheng, W.; Ma, L.; Zhao, J.; Li, Z.; Sun, F.; Lu, X. Comparative transcriptome analysis of two rice varieties in response to rice stripe virus and small brown planthoppers during early interaction. PLoS ONE 2013, 8, e82126. [Google Scholar] [CrossRef] [PubMed]
- Foley, R.C.; Kidd, B.N.; Hane, J.K.; Anderson, J.P.; Singh, K.B. Reactive oxygen species play a role in the infection of the necrotrophic fungi, Rhizoctonia solani in wheat. PLoS ONE 2016, 11, e0152548. [Google Scholar] [CrossRef] [PubMed]
- Ralph, J.; Bunzel, M.; Marita, J.M.; Hatfield, R.D.; Lu, F.; Kim, H.; Schatz, P.F.; Grabber, J.H.; Steinhart, H. Peroxidase-dependent cross-linking reactions of p-hydroxycinnamates in plant cell walls. Phytochem. Rev. 2004, 3, 79–96. [Google Scholar] [CrossRef]
- Passardi, F.; Longet, D.; Penel, C.; Dunand, C. The class III peroxidase multigenic family in rice and its evolution in land plants. Phytochemistry 2004, 65, 1879–1893. [Google Scholar] [CrossRef] [PubMed]
- Leng, Y.; Yang, Y.; Ren, D.; Huang, L.; Dai, L.; Wang, Y.; Chen, L.; Tu, Z.; Gao, Y.; Li, X. A rice pectate lyase-like gene is required for plant growth and leaf senescence. Plant Physiol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Rodiuc, N.; Barlet, X.; Hok, S.; Perfusfusi, L.; Allasia, V.; Engler, G.; Seassau, A.; Marteu, N.; Almeida-Engler, J.; Panabierer, F.; et al. Evolution distant pathogens require the Arabidopsis phytosulfokine signaling pathway to establish disease. Plant Cell Environ. 2016, 39, 1396–1407. [Google Scholar] [CrossRef] [PubMed]
- Matsubayashi, Y.; Ogawa, M.; Morita, A.; Sakagami, Y. An LRR receptor kinase involved in perception of a peptide plant hormone, phytosulfokine. Science 2002, 296, 1470–1472. [Google Scholar] [CrossRef] [PubMed]
- Lin, N.-C.; Martin, G.B. Pto-and prf-mediated recognition of AvrPto and AvrPtoB restricts the ability of diverse Pseudomonas syringae pathovars to infect tomato. Mol. Plant Microbe Interact. 2007, 20, 806–815. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Loh, Y.-T.; Bressan, R.A.; Martin, G.B. The tomato gene pti1 encodes a serine/threonine kinase that is phosphorylated by Pto and is involved in the hypersensitive response. Cell 1995, 83, 925–935. [Google Scholar] [CrossRef]
- Großkinsky, D.K.; van der Graaff, E.; Roitsch, T. Regulation of abiotic and biotic stress responses by plant hormones. In Plant Pathogen Resistance Biotechnology; John Wiley & Sons: Hoboken, NJ, USA, 2016; pp. 131–154. [Google Scholar]
- Sun, X.; Cao, Y.; Yang, Z.; Xu, C.; Li, X.; Wang, S.; Zhang, Q. Xa26, a gene conferring resistance to Xanthomonas oryzae pv. oryzae in rice, encodes an LRR receptor kinase-like protein. Plant J. 2004, 37, 517–527. [Google Scholar] [PubMed]
- Balagué, C.; Gouget, A.; Bouchez, O.; Souriac, C.; Haget, N. Boutettet gene conferring resistance to Xanthomonas oryzae pv. oryzae in rice, encodes an an d leafLecRK-I. 9 is required for full resistance to Pseudomonas syringae and affects jasmonate signalling. Mol. Plant Pathol. 2017, 18, 937–948. [Google Scholar]
- Tsuda, K.; Katagiri, F. Comparing signaling mechanisms engaged in pattern-triggered and effector-triggered immunity. Curr. Opin. Plant Biol. 2010, 13, 459–465. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, P.; Rabara, R.C.; Rushton, P.J. A systems biology perspective on the role of WRKY transcription factors in drought responses in plants. Planta 2014, 239, 255–266. [Google Scholar] [CrossRef] [PubMed]
- Berri, S.; Abbruscato, P.; Faivre-Rampant, O.; Brasileiro, A.C.; Fumasoni, I.; Satoh, K.; Kikuchi, S.; Mizzi, L.; Morandini, P.; Pè, M.E. Characterization of WRKY co-regulatory networks in rice and arabidopsis. BMC Plant Biol. 2009, 9, 120. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.; Bartley, L.E.; Canlas, P.; Ronald, P.C. OsWRKY IIa transcription factors modulate rice innate immunity. Rice 2010, 3, 36–42. [Google Scholar] [CrossRef] [PubMed]
- Park, C.Y.; Lee, J.H.; Yoo, J.H.; Moon, B.C.; Choi, M.S.; Kang, Y.H.; Lee, S.M.; Kim, H.S.; Kang, K.Y.; Chung, W.S. WRKY group IId transcription factors interact with calmodulin. FEBS Lett. 2005, 579, 1545–1550. [Google Scholar] [CrossRef] [PubMed]
- Chujo, T.; Miyamoto, K.; Ogawa, S.; Masuda, Y.; Shimizu, T.; Kishi-Kaboshi, M.; Takahashi, A.; Nishizawa, Y.; Minami, E.; Nojiri, H. Overexpression of phosphomimic mutated OsWRKY53 leads to enhanced blast resistance in rice. PLoS ONE 2014, 9, e98737. [Google Scholar] [CrossRef] [PubMed]
- Licausi, F.; Ohmehmeblast Resistance in Rice; Shimizu, T.; Kishi-Kaboshi, M. AP2/ERF transcription factors: Mediators of stress responses and developmental programs. New Phytol. 2013, 199, 639–649. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Guo, Z. Tobacco OPBP1 enhances salt tolerance and disease resistance of transgenic rice. Int. J. Mol. Sci. 2008, 9, 2601–2613. [Google Scholar] [CrossRef] [PubMed]
- Bernoux, M.; Moncuquet, P.; Kroj, T.; Dodds, P.N. A novel conserved mechanism for plant NLR protein pairs: The ‘integrated decoy’ hypothesis. Front. Plant Sci. 2014, 5, 606. [Google Scholar] [CrossRef]
- Liu, D.; Chen, X.; Liu, J.; Ye, J.; Guo, Z. The rice ERF transcription factor OsERF922 negatively regulates resistance to Magnaporthe oryzae and salt tolerance. J. Exp. Bot. 2012, 63, 3899–3911. [Google Scholar] [CrossRef] [PubMed]
- Pooja, S.; Sweta, K.; Mohanapriya, A.; Sudandiradoss, C.; Siva, R.; Gothandam, K.M.; Babu, S. Homotypic clustering of OsMYB4 binding site motifs in promoters of the rice genome and cellular-level implications on sheath blight disease resistance. Gene 2015, 561, 209–218. [Google Scholar] [CrossRef] [PubMed]
- Soler, M.; Camargo, E.L.O.; Carocha, V.; Cassan-Wang, H.; San, C.H.; Savelli, B.; Hefer, C.A.; Paiva, J.A.; Myburg, A.A.; Grima-Pettenati, J. The Eucalyptus grandis R2R3-MYB transcription factor family: Evidence for woody growth-related evolution and function. New Phytol. 2015, 206, 1364–1377. [Google Scholar] [CrossRef] [PubMed]
- Kaneda, T.; Taga, Y.; Takai, R.; Iwano, M.; Matsui, H.; Takayama, S.; Isogai, A.; Che, F.S. The transcription factor OsNAC4 is a key positive regulator of plant hypersensitive cell death. EMBO J. 2009, 28, 926–936. [Google Scholar] [CrossRef] [PubMed]
- Park, S.R.; Kim, H.S.; Lee, K.S.; Hwang, D.-J.; Bae, S.-C.; Ahn, I.-P.; Lee, S.H.; Kim, S.T. Overexpression of rice NAC transcription factor OsNAC58 on increased resistance to bacterial leaf blight. J. Plant Biotechnol. 2017, 44, 149–155. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhao, J.; Li, Y.; Yuan, Z.; He, H.; Yang, H.; Qu, H.; Ma, C.; Qu, S. Transcriptome analysis highlights defense and signaling pathways mediated by rice Pi21 gene with partial resistance to Magnaporthe oryzae. Front. Plant Sci. 2016, 7, 1834. [Google Scholar] [CrossRef] [PubMed]
- Hamberger, B.; Ellis, M.; Friedmann, M.; de Azevedo Souza, C.; Barbazuk, B.; Douglas, C.J. Genome-wide analyses of phenylpropanoid-related genes in Populus trichocarpa, Arabidopsis thaliana, and Oryza sativa: The populus lignin toolbox and conservation and diversification of angiosperm gene families. Botany 2007, 85, 1182–1201. [Google Scholar]
- Vogt, T. Phenylpropanoid biosynthesis. Mol. Plant 2010, 3, 2–20. [Google Scholar] [CrossRef] [PubMed]
- Miedes, E.; Vanholme, R.; Boerjan, W.; Molina, A. The role of the secondary cell wall in plant resistance to pathogens. Front. Plant Sci. 2014, 5, 358. [Google Scholar] [CrossRef] [PubMed]
- Bari, R.; Jones, J.D. Role of plant hormones in plant defence responses. Plant Mol. Biol. 2009, 69, 473–488. [Google Scholar] [CrossRef] [PubMed]
- Cohen, S.P.; Liu, H.; Argueso, C.T.; Pereira, A.; Cruz, C.V.; Verdier, V.; Leach, J.E. RNA-Seq analysis reveals insight into enhanced rice xa7-mediated bacterial blight resistance at high temperature. PLoS ONE 2017, 12, e0187625. [Google Scholar] [CrossRef] [PubMed]
- Iwai, T.; Miyasaka, A.; Seo, S.; Ohashi, Y. Contribution of ethylene biosynthesis for resistance to blast fungus infection in young rice plants. Plant Physiol. 2006, 142, 1202–1215. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.-L.; Yang, Y.; He, Z. Roles of plant hormones and their interplay in rice immunity. Mol. Plant 2013, 6, 675–685. [Google Scholar] [CrossRef] [PubMed]
- Delaunois, B.; Jeandet, P.; Clément, C.; Baillieul, F.; Dorey, S.; Cordelier, S. Uncovering plant-pathogen crosstalk through apoplastic proteomic studies. Front. Plant Sci. 2014, 5, 249. [Google Scholar] [CrossRef] [PubMed]
- Figueiredo, A.; Monteiro, F.; Sebastiana, M. Subtilisin-like proteases in plant–pathogen recognition and immune priming: A perspective. Front. Plant Sci. 2014, 5, 739. [Google Scholar] [CrossRef] [PubMed]
- Cui, F.; Wu, S.; Sun, W.; Coaker, G.; Kunkel, B.; He, P.; Shan, L. The Pseudomonas syringae type III effector AvrPt2 promotes pathogen virulence via stimulating Arabidopsis auxin/indole acetic acid protein turnover. Plant Physiol. 2013, 162, 1018–1029. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Li, W.; Xiang, T.; Liu, Z.; Laluk, K.; Ding, X.; Zou, Y.; Gao, M.; Zhang, X.; Chen, S. Receptor-like cytoplasmic kinases integrate signaling from multiple plant immune receptors and are targeted by a Pseudomonas syringae effector. Cell Host Microbe 2010, 7, 290–301. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef] [PubMed]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Sakai, H.; Lee, S.S.; Tanaka, T.; Numa, H.; Kim, J.; Kawahara, Y.; Wakimoto, H.; Yang, C.C.; Iwamoto, M.; Abe, T.; et al. Rice annotation project database (rap-db): An integrative and interactive database for rice genomics. Plant Cell Physiol. 2013, 54, e6. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Pertea, G.; Trapnell, C.; Pimentel, H.; Kelley, R.; Salzberg, S.L. Tophat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013, 14, R36. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.; Kwan, G.; Van Baren, M.J.; Salzberg, S.L.; Wold, B.J.; Pachter, L. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar] [CrossRef] [PubMed]
- Zhou, D.; Xin, Z.; Yi, L.; Zhenhai, Z.; Zhen, S. agriGO: A GO Analysis Toolkit for the Agricultural Community. Available online: http://bioinfo.cau.edu.cn/agriGO/ (accessed on 15 September 2017).
- Minoru, K.; Susumu, G. KEGG: Kyoto Encyclopedia of Genes and Genomes. Available online: http://www.genome.jp/kegg/kegg2.html (accessed on 20 September 2017).
- Thimm, O.; Bläsing, O.; Gibon, Y.; Nagel, A.; Meyer, S.; Krüger, P.; Selbig, J.; Müller, L.A.; Rhee, S.Y.; Stitt, M. Mapman: A user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J. 2004, 37, 914–939. [Google Scholar] [CrossRef] [PubMed]
- Nicolas, J. Amplifx 1.5.4. Available online: http://www.brothersoft.com/amplifx-159421.html (accessed on 6 November 2017).
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using Real-Time quantitative Pcr and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]


| Samples | Raw Reads | Clean Reads | Total Mapped | Q Value (30%) | GC % |
|---|---|---|---|---|---|
| C0 | 46,594,244 | 44,372,944 | 36,489,861 (82.23%) | 91 | 54 |
| J0 | 51,085,434 | 48,289,010 | 39,611,914 (82.03%) | 91 | 54 |
| CBB23-12 hpi | 57,398,598 | 54,774,152 | 45,086,818 (82.31%) | 92 | 54 |
| CBB23-24 hpi | 52,122,832 | 49,294,374 | 41,275,771 (83.73%) | 91 | 55 |
| CBB23-36 hpi | 65,574,380 | 62,882,740 | 50,963,164 (81.04%) | 92 | 53 |
| CBB23-48 hpi | 41,893,788 | 40,539,554 | 33,922,512 (83.68%) | 92 | 54 |
| JG30-12 hpi | 49,890,010 | 48,749,132 | 40,250,051 (82.57%) | 93 | 55 |
| JG30-24 hpi | 54,891,574 | 52,116,660 | 43,303,751 (83.09%) | 91 | 55 |
| JG30-36 hpi | 45,507,160 | 44,119,566 | 35,292,193 (79.99%) | 92 | 53 |
| JG30-48 hpi | 57,418,698 | 54,514,722 | 45,769,331 (83.96%) | 91 | 55 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tariq, R.; Wang, C.; Qin, T.; Xu, F.; Tang, Y.; Gao, Y.; Ji, Z.; Zhao, K. Comparative Transcriptome Profiling of Rice Near-Isogenic Line Carrying Xa23 under Infection of Xanthomonas oryzae pv. oryzae. Int. J. Mol. Sci. 2018, 19, 717. https://doi.org/10.3390/ijms19030717
Tariq R, Wang C, Qin T, Xu F, Tang Y, Gao Y, Ji Z, Zhao K. Comparative Transcriptome Profiling of Rice Near-Isogenic Line Carrying Xa23 under Infection of Xanthomonas oryzae pv. oryzae. International Journal of Molecular Sciences. 2018; 19(3):717. https://doi.org/10.3390/ijms19030717
Chicago/Turabian StyleTariq, Rezwan, Chunlian Wang, Tengfei Qin, Feifei Xu, Yongchao Tang, Ying Gao, Zhiyuan Ji, and Kaijun Zhao. 2018. "Comparative Transcriptome Profiling of Rice Near-Isogenic Line Carrying Xa23 under Infection of Xanthomonas oryzae pv. oryzae" International Journal of Molecular Sciences 19, no. 3: 717. https://doi.org/10.3390/ijms19030717
APA StyleTariq, R., Wang, C., Qin, T., Xu, F., Tang, Y., Gao, Y., Ji, Z., & Zhao, K. (2018). Comparative Transcriptome Profiling of Rice Near-Isogenic Line Carrying Xa23 under Infection of Xanthomonas oryzae pv. oryzae. International Journal of Molecular Sciences, 19(3), 717. https://doi.org/10.3390/ijms19030717




