Expression and Sequence Variants of Inflammatory Genes; Effects on Plasma Inflammation Biomarkers Following a 6-Week Supplementation with Fish Oil
Abstract
:1. Introduction
2. Results
3. Discussion
Strengths and Limitations
4. Experimental Section
4.1. Subjects
4.2. Study Design and Diets
4.3. Anthropometric Measurements
4.4. Biochemical Parameters
4.5. Measurement of FA Composition in Plasma Phospholipids
4.6. SNP Selection and Genotyping
4.7. Gene Expression
4.8. Statistical Analyses
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| n-3 | omega-3 |
| FAs | fatty acids |
| SNPs | single nucleotide polymorphisms |
| TNF-α | tumor necrosis factor alpha |
| IL6 | interleukin 6 |
| CRP | C-reactive protein |
| PBMCs | peripheral blood mononuclear cells |
| BMI | body mass index |
| EPA | eicosapentaenoic acid |
| DHA | docosahexaenoic acid |
| FFQ | food-frequency questionnaire |
| TG | triglycerides |
| MAF | minor allele frequency |
| HWE | Hardy-Weinberg equilibrium |
References
- Stryjecki, C.; Mutch, D.M. Fatty acid–gene interactions, adipokines and obesity. Eur. J. Clin. Nutr. 2011, 65, 285–297. [Google Scholar] [CrossRef] [PubMed]
- Calder, P.C. N-3 polyunsaturated fatty acids, inflammation, and inflammatory diseases. Am. J. Clin. Nutr. 2006, 83, 1505S–1519S. [Google Scholar]
- Serhan, C.N.; Chiang, N.; Van Dyke, T.E. Resolving inflammation: Dual anti-inflammatory and pro-resolution lipid mediators. Nat. Rev. Immunol. 2008, 8, 349–361. [Google Scholar] [CrossRef] [PubMed]
- Serhan, C.N. Novel omega-3-derived local mediators in anti-inflammation and resolution. Pharmacol. Ther. 2005, 105, 7–21. [Google Scholar] [CrossRef] [PubMed]
- Tjonahen, E.; Oh, S.F.; Siegelman, J.; Elangovan, S.; Percarpio, K.B.; Hong, S.; Arita, M.; Serhan, C.N. Resolvin E2: Identification and anti-inflammatory actions: Pivotal role of human 5-lipoxygenase in resolvin E series biosynthesis. Chem. Biol. 2006, 13, 1193–1202. [Google Scholar] [CrossRef] [PubMed]
- Joffe, Y.T.; Collins, M.; Goedecke, J.H. The relationship between dietary fatty acids and inflammatory genes on the obese phenotype and serum lipids. Nutrients 2013, 5, 1672–1705. [Google Scholar] [CrossRef] [PubMed]
- Grimble, R.F.; Howell, W.M.; O′Reilly, G.; Turner, S.J.; Markovic, O.; Hirrell, S.; East, J.M.; Calder, P.C. The ability of fish oil to suppress tumor necrosis factor alpha production by peripheral blood mononuclear cells in healthy men is associated with polymorphisms in genes that influence tumor necrosis factor α production. Am. J. Clin. Nutr. 2002, 76, 454–459. [Google Scholar] [PubMed]
- Calder, P.C. Polyunsaturated fatty acids, inflammatory processes and inflammatory bowel diseases. Mol. Nutr. Food Res. 2008, 52, 885–897. [Google Scholar] [CrossRef] [PubMed]
- Rudkowska, I.; Paradis, A.M.; Thifault, E.; Julien, P.; Tchernof, A.; Couture, P.; Lemieux, S.; Barbier, O.; Vohl, M.C. Transcriptomic and metabolomic signatures of an n-3 polyunsaturated fatty acids supplementation in a normolipidemic/normocholesterolemic caucasian population. J. Nutr Biochem. 2013, 24, 54–61. [Google Scholar] [CrossRef] [PubMed]
- Bouwens, M.; van de Rest, O.; Dellschaft, N.; Bromhaar, M.G.; de Groot, L.C.; Geleijnse, J.M.; Muller, M.; Afman, L.A. Fish–oil supplementation induces antiinflammatory gene expression profiles in human blood mononuclear cells. Am. J. Clin. Nutr. 2009, 90, 415–424. [Google Scholar] [CrossRef] [PubMed]
- Thifault, E.; Cormier, H.; Bouchard-Mercier, A.; Rudkowska, I.; Paradis, A.M.; Garneau, V.; Ouellette, C.; Lemieux, S.; Couture, P.; Vohl, M.C. Effects of age, sex, body mass index and APOE genotype on cardiovascular biomarker response to an n-3 polyunsaturated fatty acid supplementation. J. Nutrigenet. Nutrigenom. 2013, 6, 73–82. [Google Scholar] [CrossRef] [PubMed]
- Tremblay, B.L.; Rudkowska, I.; Couture, P.; Lemieux, S.; Julien, P.; Vohl, M.C. Modulation of C-reactive protein and plasma omega-6 fatty acid levels by phospholipase A2 gene polymorphisms following a 6-week supplementation with fish oil. Prostaglandins Leukot. Essent. Fat. Acids 2015, 102–103, 37–45. [Google Scholar] [CrossRef] [PubMed]
- Ferrucci, L.; Cherubini, A.; Bandinelli, S.; Bartali, B.; Corsi, A.; Lauretani, F.; Martin, A.; Andres-Lacueva, C.; Senin, U.; Guralnik, J.M. Relationship of plasma polyunsaturated fatty acids to circulating inflammatory markers. J. Clin. Endocrinol. Metab. 2006, 91, 439–446. [Google Scholar] [CrossRef] [PubMed]
- Trebble, T.; Arden, N.K.; Stroud, M.A.; Wootton, S.A.; Burdge, G.C.; Miles, E.A.; Ballinger, A.B.; Thompson, R.L.; Calder, P.C. Inhibition of tumour necrosis factor-α and interleukin 6 production by mononuclear cells following dietary fish-oil supplementation in healthy men and response to antioxidant co-supplementation. Br J. Nutr. 2003, 90, 405–412. [Google Scholar] [CrossRef] [PubMed]
- Rees, D.; Miles, E.A.; Banerjee, T.; Wells, S.J.; Roynette, C.E.; Wahle, K.W.; Calder, P.C. Dose-related effects of eicosapentaenoic acid on innate immune function in healthy humans: A comparison of young and older men. Am. J. Clin. Nutr. 2006, 83, 331–342. [Google Scholar] [PubMed]
- Mahajan, A.; Tabassum, R.; Chavali, S.; Dwivedi, O.P.; Chauhan, G.; Tandon, N.; Bharadwaj, D. Obesity-dependent association of TNF-LTA locus with type 2 diabetes in north indians. J. Mol. Med. 2010, 88, 515–522. [Google Scholar] [CrossRef] [PubMed]
- Teuffel, O.; Ethier, M.C.; Beyene, J.; Sung, L. Association between tumor necrosis factor-α promoter –308 A/G polymorphism and susceptibility to sepsis and sepsis mortality: A systematic review and meta-analysis. Crit. Care Med. 2010, 38, 276–282. [Google Scholar] [CrossRef] [PubMed]
- Franke, A.; McGovern, D.P.; Barrett, J.C.; Wang, K.; Radford-Smith, G.L.; Ahmad, T.; Lees, C.W.; Balschun, T.; Lee, J.; Roberts, R.; et al. Genome-wide meta-analysis increases to 71 the number of confirmed crohn's disease susceptibility loci. Nat. Genet 2010, 42, 1118–1125. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Yu, X.; Wang, L.; Zhou, S.; Sun, J.; Feng, N.; Nie, S.; Wu, J.; Gao, F.; Fei, B.; et al. Four genetic polymorphisms of lymphotoxin-α gene and cancer risk: A systematic review and meta-analysis. PLoS ONE 2013, 8, e82519. [Google Scholar] [CrossRef] [PubMed]
- Song, Z.; Song, Y.; Yin, J.; Shen, Y.; Yao, C.; Sun, Z.; Jiang, J.; Zhu, D.; Zhang, Y.; Shen, Q.; et al. Genetic variation in the TNF gene is associated with susceptibility to severe sepsis, but not with mortality. PLoS ONE 2012, 7, e46113. [Google Scholar] [CrossRef] [PubMed]
- Nieters, A.; Becker, N.; Linseisen, J. Polymorphisms in candidate obesity genes and their interaction with dietary intake of n-6 polyunsaturated fatty acids affect obesity risk in a sub-sample of the EPIC-heidelberg cohort. Eur. J. Nutr. 2002, 41, 210–221. [Google Scholar] [CrossRef] [PubMed]
- Joffe, Y.T.; van der Merwe, L.; Carstens, M.; Collins, M.; Jennings, C.; Levitt, N.S.; Lambert, E.V.; Goedecke, J.H. Tumor necrosis factor-α gene –308 G/A polymorphism modulates the relationship between dietary fat intake, serum lipids, and obesity risk in black south african women. J. Nutr. 2010, 140, 901–907. [Google Scholar] [CrossRef] [PubMed]
- Meydani, S.N.; Lichtenstein, A.H.; Cornwall, S.; Meydani, M.; Goldin, B.R.; Rasmussen, H.; Dinarello, C.A.; Schaefer, E.J. Immunologic effects of national cholesterol education panel step-2 diets with and without fish-derived n-3 fatty acid enrichment. J. Clin. Investig. 1993, 92, 105–113. [Google Scholar] [CrossRef] [PubMed]
- Kroeger, K.M.; Carville, K.S.; Abraham, L.J. The -308 tumor necrosis factor-α promoter polymorphism effects transcription. Mol. Immunol. 1997, 34, 391–399. [Google Scholar] [CrossRef]
- Wilson, A.G.; Symons, J.A.; McDowell, T.L.; McDevitt, H.O.; Duff, G.W. Effects of a polymorphism in the human tumor necrosis factor α promoter on transcriptional activation. Proc. Natl. Acad. Sci. USA 1997, 94, 3195–3199. [Google Scholar] [CrossRef] [PubMed]
- Antonicelli, R.; Olivieri, F.; Cavallone, L.; Spazzafumo, L.; Bonafe, M.; Marchegiani, F.; Cardelli, M.; Galeazzi, R.; Giovagnetti, S.; Perna, G.P.; et al. Tumor necrosis factor-α gene –308G>A polymorphism is associated with ST-elevation myocardial infarction and with high plasma levels of biochemical ischemia markers. Coron. Artery Dis. 2005, 16, 489–493. [Google Scholar] [CrossRef] [PubMed]
- Browning, L.M.; Krebs, J.D.; Moore, C.S.; Mishra, G.D.; O′Connell, M.A.; Jebb, S.A. The impact of long chain n-3 polyunsaturated fatty acid supplementation on inflammation, insulin sensitivity and CVD risk in a group of overweight women with an inflammatory phenotype. Diabetes Obes. Metab. 2007, 9, 70–80. [Google Scholar] [CrossRef] [PubMed]
- Ciubotaru, I.; Lee, Y.S.; Wander, R.C. Dietary fish oil decreases C-reactive protein, interleukin-6, and triacylglycerol to HDL-cholesterol ratio in postmenopausal women on HRT. J. Nutr. Biochem. 2003, 14, 513–521. [Google Scholar] [CrossRef]
- Rasic-Milutinovic, Z.; Perunicic, G.; Pljesa, S.; Gluvic, Z.; Sobajic, S.; Djuric, I.; Ristic, D. Effects of n-3 PUFAs supplementation on insulin resistance and inflammatory biomarkers in hemodialysis patients. Ren. Fail. 2007, 29, 321–329. [Google Scholar] [CrossRef] [PubMed]
- Poudel-Tandukar, K.; Sato, M.; Ejima, Y.; Nanri, A.; Matsushita, Y.; Imaizumi, K.; Mizoue, T. Relationship of serum fatty acid composition and desaturase activity to C-reactive protein in japanese men and women. Atherosclerosis 2012, 220, 520–524. [Google Scholar] [CrossRef] [PubMed]
- Eating well with canada's food guide. Available online: http://www.hc-sc.gc.ca/fn-an/food-guide-aliment/index-eng.php (accessed on 3 February 2016).
- Goulet, J.; Nadeau, G.; Lapointe, A.; Lamarche, B.; Lemieux, S. Validity and reproducibility of an interviewer-administered food frequency questionnaire for healthy french-canadian men and women. Nutr. J. 2004, 3, 13. [Google Scholar] [CrossRef] [PubMed]
- Callaway, C.W.; Chumlea, W.C.; Bouchard, C.; Himes, J.H.; Lohman, T.G.; Martin, A.D.; Mitchell, C.D.; Mueller, W.H.; Roche, A.F.; Seefeldt, V.D. The Airlie (VA) Consensus Conference; Human Kinetics Publishers: Champaign, IR, USA, 1988; pp. 39–80. [Google Scholar]
- Pirro, M.; Bergeron, J.; Dagenais, G.R.; Bernard, P.M.; Cantin, B.; Despres, J.P.; Lamarche, B. Age and duration of follow-up as modulators of the risk for ischemic heart disease associated with high plasma C-reactive protein levels in men. Arch. Intern. Med. 2001, 161, 2474–2480. [Google Scholar] [CrossRef] [PubMed]
- Shaikh, N.A.; Downar, E. Time course of changes in porcine myocardial phospholipid levels during ischemia. A reassessment of the lysolipid hypothesis. Circ. Res. 1981, 49, 316–325. [Google Scholar] [CrossRef] [PubMed]
- Fozzard, H.A.; Haber, E.; Jennings, R.B.; Katz, A.M. The Heart and Cardiovascular System; Raven Press: New York, NY, USA, 1986. [Google Scholar]
- Lepage, G.; Roy, C.C. Direct transesterification of all classes of lipids in a one-step reaction. J. Lip. Res. 1986, 27, 114–120. [Google Scholar]
- Livak, K.J. Allelic discrimination using fluorogenic probes and the 5′ nuclease assay. Genet. Anal. Biomol. Eng. 1999, 14, 143–149. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]

| Characteristics | All (n = 191) | Men (n = 95, 49.7%) | Women (n = 96, 50.3%) |
|---|---|---|---|
| Age | 30.8 ± 8.7 | 30.9 ± 8.1 | 30.8 ± 9.3 |
| Waist circumference, cm | 93.2 ± 10.6 | 94.8 ± 10.9 | 91.6 ± 10.1 |
| BMI, kg/m2 | 27.2 ± 3.6 | 27.5 ± 3.6 | 27.9 ± 3.5 |
| Biomarkers | Pre n-3 FA | Post n-3 FA | p-Value 1 |
|---|---|---|---|
| CRP levels (mg/L) 2 | 1.78 ± 2.09 | 1.81 ± 2.07 | 0.95 |
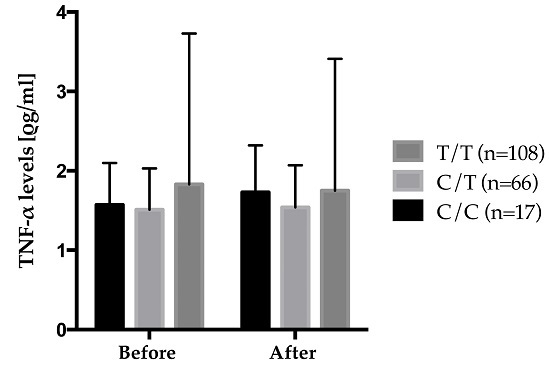
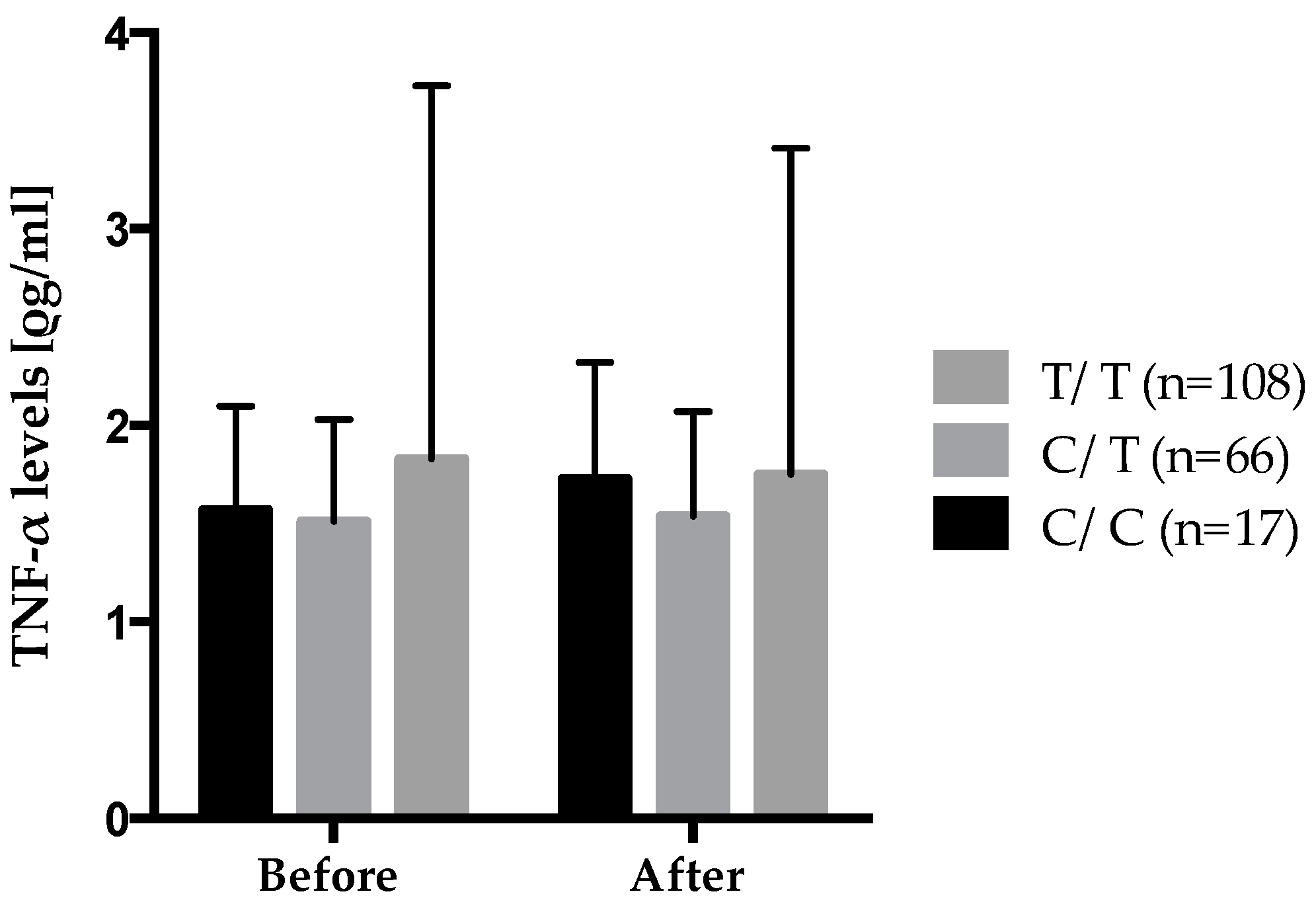
| TNF-α levels (ρg/mL) 3 | 1.70 ± 1.48 | 1.68 ± 1.30 | 0.69 |
| IL-6 levels, (ρg/mL) 2 | 1.34 ± 1.13 | 1.28 ± 0.98 | 0.54 |
| Dietary Intakes | Pre-Suppl. | Post-Suppl. | p-Value 1 |
|---|---|---|---|
| (Including n-3 FA Supplements) | |||
| Energy, (kcal) | 2290 ± 599 | 2196 ± 570 | 0.003 |
| Carbohydrate, (% of TEI) | 50.5 ± 7.2 | 48.5 ± 7.8 | 0.001 |
| Carbohydrate, (g/day) | 288.9 ± 79.8 | 264.1 ± 78.1 | <0.0001 |
| Protein, (% of TEI) | 17.4 ± 3.3 | 16.9 ± 3.1 | 0.15 |
| Protein, (g/day) | 98.6 ± 30.6 | 93.2 ± 30.1 | 0.002 |
| Total fat, (% of TEI) | 32.5 ± 6.0 | 35.2 ± 6.3 | <0.0001 |
| Total fat, (g/day) | 85.0 ± 29.7 | 86.8 ± 29.9 | 0.56 |
| SFA, (% of TEI) | 11.1 ± 3.1 | 10.3 ± 3.0 | 0.001 |
| SFA, (g/day) | 29.1 ± 12.1 | 25.4 ± 10.5 | <0.0001 |
| MUFA, (% of TEI) | 11.8 ± 2.8 | 12.0 ± 3.3 | 0.41 |
| MUFA, (g/day) | 30.8 ± 11.8 | 29.7 ± 12.5 | 0.17 |
| PUFA, (% of TEI) | 5.9 ± 2.1 | 6.9 ± 2.1 | <0.0001 |
| PUFA, (g/day) | 15.3 ± 6.7 | 16.9 ± 6.7 | 0.001 |
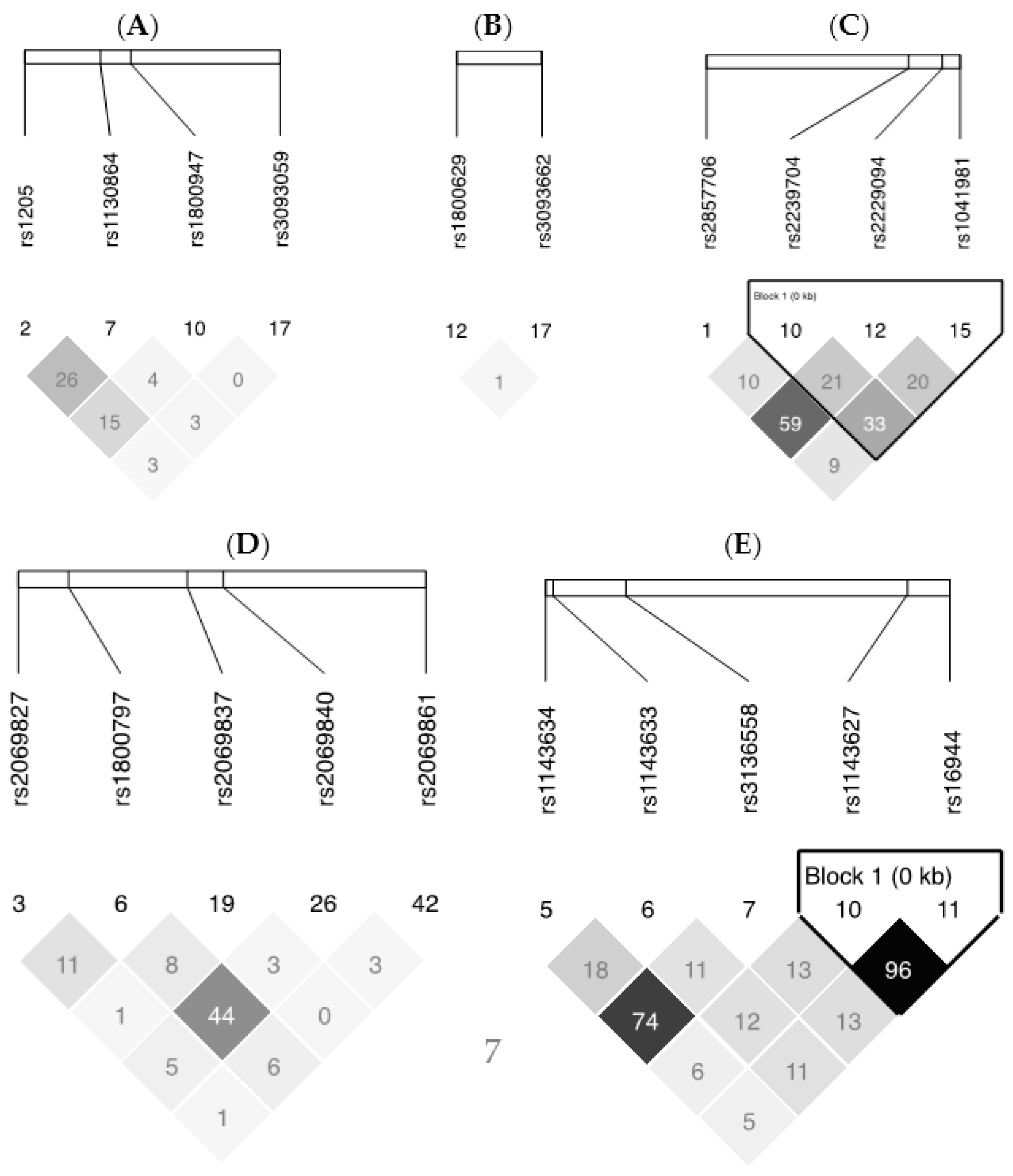
| Genes | dbSNP No. a | Sequence b | Position | AA | CC | CA | CG | CT | GA | GG | GT | AT | TT |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| n (%) | |||||||||||||
| TNF-LTA | rs1041981 | CA[A/C]CC | Missense [Thr]→[Asn] | 22 (10.5) | 97 (46.2) | 91 (43.3) | |||||||
| TNF-LTA | rs2857706 | TA[A/G]GT | Intron | 151 (7.2) | 52 (24.8) | 7 (3.3) | |||||||
| TNF-LTA | rs1800629 | TG[A/G]GG | nearGene-5 | 2 (1.0) | 45 (21.4) | 163 (77.6) | |||||||
| TNF-LTA | rs2239704 | GC[G/T]GG | UTR-5 | 49 (23.3) | 72 (34.3) | 89 (42.4) | |||||||
| TNF-LTA | rs3093662 | AC[A/G]GA | Intron | 178 (84.8) | 29 (13.8) | 3 (1.4) | |||||||
| TNF-LTA | rs2229094 | TG[C/T]GT | Missense [Cys]→[Arg] | 19 (9.1) | 74 (35.2) | 117 (55.7) | |||||||
| IL6 | rs2069861 | AA[C/T]AA | nearGene-3 | 177 (84.3) | 33 (15.7) | 0 (0.0) | |||||||
| IL6 | rs2069840 | AA[C/G]TT | Intron | 95 (45.2) | 92 (43.8) | 23 (11.0) | |||||||
| IL6 | rs2069837 | TA[A/G]AT | Intron | 181 (86.2) | 28 (13.3) | 1 (0.5) | |||||||
| IL6 | rs2069827 | TC[G/T]AT | nearGene-5 | 172 (81.9) | 38 (18.1) | 0 (0.0) | |||||||
| IL6 | rs1800797 | GG[A/G]TG | nearGene-5 | 73 (34.8) | 111 (52.9) | 26 (12.4) | |||||||
| IL1B | rs1143634 | TT[C/T]GA | Cds-synon [Phe]→[Phe] | 129 (61.4) | 70 (33.3) | 11 (5.2) | |||||||
| IL1B | rs1143633 | CC[A/G]CC | Intron | 32 (15.2) | 94 (44.8) | 84 (40.0) | |||||||
| IL1B | rs16944 | TC[A/G]GG | nearGene-5 | 28 (13.3) | 82 (39.1) | 100 (47.6) | |||||||
| IL1B | rs3136558 | GA[C/T]CT | Intron | 8 (3.8) | 77 (36.7) | 125 (59.5) | |||||||
| IL1B | rs1143627 | GC[C/T]AT | nearGene-5 | 29 (13.8) | 81 (38.6) | 100 (47.6) | |||||||
| CRP | rs1800947 | CT[C/G]TC | Cds-synon [Leu]→[Leu] | 190 (90.5) | 20 (9.5) | 0 (0.0) | |||||||
| CRP | rs3093059 | AT[C/T]GG | nearGene-5 | 2 (1.0) | 32 (15.2) | 176 (83.6) | |||||||
| CRP | rs1130864 | AA[C/T]GG | UTR-3 | 102 (48.6) | 84 (40.0) | 24 (11.4) | |||||||
| CRP | rs1205 | CA[C/T]AG | UTR-3 | 99 (47.1) | 89 (42.4) | 22 (10.5) | |||||||
| Gene | Biomarker | SNPs | Genotype by Suppl. | β ± SE | p-Value for the Genotype | p-Value for the n-3 Suppl. | p-Value for the Interaction |
|---|---|---|---|---|---|---|---|
| IL-1β | IL-6 levels (ρg/mL) | rs1143627 | CC × n-3 suppl. | −0.014 ± 0.025 | 0.91 | 0.78 | 0.02 |
| CT × n-3 suppl. | 0.037 ± 0.014 | ||||||
| TT × n-3 suppl. | −0.014 ± 0.013 | ||||||
| IL-6 levels (ρg/mL) | rs16944 | AA × n-3 suppl. | −0.018 ± 0.025 | 0.87 | 0.87 | 0.02 | |
| AG × n-3 suppl. | 0.038 ± 0.014 | ||||||
| GG × n-3 suppl. | −0.014 ± 0.013 | ||||||
| IL6 | IL-6 levels (ρg/mL) | rs1800797 | AA × n-3 suppl. | 0.025 ± 0.015 | 0.30 | 0.64 | 0.05 |
| AG × n-3 suppl. | −0.023 ± 0.013 | ||||||
| GG × n-3 suppl. | −0.016 ± 0.025 | ||||||
| IL-6 levels (ρg/mL) | rs2069840 | CC × n-3 suppl. | −0.010 ± 0.013 | 0.11 | 0.28 | 0.04 | |
| CG × n-3 suppl. | −0.016 ± 0.014 | ||||||
| GG × n-3 suppl. | 0.063 ± 0.029 | ||||||
| TNF-LTA gene family | TNF-α levels, (ρg/mL) | rs2229094 | CC × n-3 suppl. | 0.022 ± 0.012 | 0.85 | 0.20 | 0.03 |
| CT × n-3 suppl. | 0.005 ± 0.006 | ||||||
| TT × n-3 suppl. | −0.009 ± 0.005 | ||||||
| CRP levels (mg/L) | rs1800629 | GG × n-3 suppl. | −0.012 ± 0.016 | 0.37 | 0.17 | 0.04 | |
| [AA+GA] × n-3 suppl. | 0.058 ± 0.030 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cormier, H.; Rudkowska, I.; Lemieux, S.; Couture, P.; Vohl, M.-C. Expression and Sequence Variants of Inflammatory Genes; Effects on Plasma Inflammation Biomarkers Following a 6-Week Supplementation with Fish Oil. Int. J. Mol. Sci. 2016, 17, 375. https://doi.org/10.3390/ijms17030375
Cormier H, Rudkowska I, Lemieux S, Couture P, Vohl M-C. Expression and Sequence Variants of Inflammatory Genes; Effects on Plasma Inflammation Biomarkers Following a 6-Week Supplementation with Fish Oil. International Journal of Molecular Sciences. 2016; 17(3):375. https://doi.org/10.3390/ijms17030375
Chicago/Turabian StyleCormier, Hubert, Iwona Rudkowska, Simone Lemieux, Patrick Couture, and Marie-Claude Vohl. 2016. "Expression and Sequence Variants of Inflammatory Genes; Effects on Plasma Inflammation Biomarkers Following a 6-Week Supplementation with Fish Oil" International Journal of Molecular Sciences 17, no. 3: 375. https://doi.org/10.3390/ijms17030375
APA StyleCormier, H., Rudkowska, I., Lemieux, S., Couture, P., & Vohl, M.-C. (2016). Expression and Sequence Variants of Inflammatory Genes; Effects on Plasma Inflammation Biomarkers Following a 6-Week Supplementation with Fish Oil. International Journal of Molecular Sciences, 17(3), 375. https://doi.org/10.3390/ijms17030375